Abstract
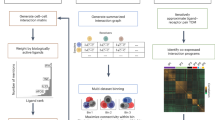
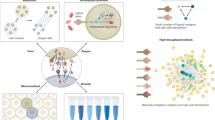
Ligand–receptor interactions constitute a fundamental mechanism of cell–cell communication and signaling. NicheNet is a well-established computational tool that infers ligand–receptor interactions that potentially regulate gene expression changes in receiver cell populations. Whereas the original publication delves into the algorithm and validation, this paper describes a best practices workflow cultivated over four years of experience and user feedback. Starting from the input single-cell expression matrix, we describe a ‘sender-agnostic’ approach that considers ligands from the entire microenvironment and a ‘sender-focused’ approach that considers ligands only from cell populations of interest. As output, users will obtain a list of prioritized ligands and their potential target genes, along with multiple visualizations. We include further developments made in NicheNet v2, in which we have updated the data sources and implemented a downstream procedure for prioritizing cell type–specific ligand–receptor pairs. Although a standard NicheNet analysis takes <10 min to run, users often invest additional time in making decisions about the approach and parameters that best suit their biological question. This paper serves to aid in this decision-making process by describing the most appropriate workflow for common experimental designs like case-control and cell-differentiation studies. Finally, in addition to the step-by-step description of the code, we also provide wrapper functions that enable the analysis to be run in one line of code, thus tailoring the workflow to users at all levels of computational proficiency.
Key points
-
This protocol describes NicheNet, which is a computational tool to infer the ligand–receptor interactions that potentially regulate gene expression changes in receiver cell populations. This protocol produces a list of prioritized ligands and their potential target genes and includes multiple visualizations.
-
Unlike many other cell–cell communication tools, which include information only about the expression of ligands and receptors, NicheNet incorporates information about the transcriptional responses triggered by these interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The original NICHE-seq data can be accessed at the Gene Expression Omnibus with accession number GSE104054. The processed Seurat object can be downloaded on Zenodo41 (https://zenodo.org/record/3531889). NicheNet networks can be downloaded at https://zenodo.org/record/7074291/ (ref. 42).
Code availability
NicheNet is publicly available on https://github.com/saeyslab/nichenetr/ as an R package. The code shown in this paper and the code for reproducing the figures can be found on https://github.com/saeyslab/nichenet_protocol. The code in this protocol has been peer reviewed.
References
Armingol, E., Baghdassarian, H. M. & Lewis, N. E. The diversification of methods for studying cell–cell interactions and communication. Nat. Rev. Genet. 25, 381–400 (2024).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Kim, N., Kang, H., Jo, A., Yoo, S.-A. & Lee, H.-O. Perspectives on single-nucleus RNA sequencing in different cell types and tissues. J. Pathol. Transl. Med. 57, 52–59 (2023).
Liu, Y.-J. IPC: professional type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annu. Rev. Immunol. 23, 275–306 (2005).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396.e38 (2022).
Wijesooriya, K., Jadaan, S. A., Perera, K. L., Kaur, T. & Ziemann, M. Urgent need for consistent standards in functional enrichment analysis. PLoS Comput. Biol. 18, e1009935 (2022).
Browaeys, R. et al. MultiNicheNet: a flexible framework for differential cell-cell communication analysis from multi-sample multi-condition single-cell transcriptomics data. Preprint at https://www.biorxiv.org/content/10.1101/2023.06.13.544751v1 (2023).
Bonnardel, J. et al. Stellate cells, hepatocytes, and endothelial cells imprint the Kupffer cell identity on monocytes colonizing the liver macrophage niche. Immunity 51, 638–654.e9 (2019).
Rodriguez, E. et al. Sialic acids in pancreatic cancer cells drive tumour-associated macrophage differentiation via the Siglec receptors Siglec-7 and Siglec-9. Nat. Commun. 12, 1270 (2021).
Karras, P. et al. A cellular hierarchy in melanoma uncouples growth and metastasis. Nature 610, 190–198 (2022).
Loft, A. et al. A macrophage-hepatocyte glucocorticoid receptor axis coordinates fasting ketogenesis. Cell Metab. 34, 473–486.e9 (2022).
Liu, X. et al. Distinct human Langerhans cell subsets orchestrate reciprocal functions and require different developmental regulation. Immunity 54, 2305–2320.e11 (2021).
Mourcin, F. et al. Follicular lymphoma triggers phenotypic and functional remodeling of the human lymphoid stromal cell landscape. Immunity 54, 1788–1806.e7 (2021).
Voss, A. J. et al. Identification of ligand–receptor pairs that drive human astrocyte development. Nat. Neurosci. 26, 1339–1351 (2023).
Chan, J. M. et al. Lineage plasticity in prostate cancer depends on JAK/STAT inflammatory signaling. Science 377, 1180–1191 (2022).
Hoste, E. et al. OTULIN maintains skin homeostasis by controlling keratinocyte death and stem cell identity. Nat. Commun. 12, 5913 (2021).
Reyes, N. S. et al. Sentinel p16INK4a+ cells in the basement membrane form a reparative niche in the lung. Science 378, 192–201 (2022).
Mair, F. et al. Extricating human tumour immune alterations from tissue inflammation. Nature 605, 728–735 (2022).
Wilk, A. J., Shalek, A. K., Holmes, S. & Blish, C. A. Comparative analysis of cell–cell communication at single-cell resolution. Nat. Biotechnol. 42, 470–483 (2024).
He, C., Zhou, P. & Nie, Q. exFINDER: identify external communication signals using single-cell transcriptomics data. Nucleic Acids Res. 51, e58 (2023).
Xin, Y. et al. LRLoop: a method to predict feedback loops in cell–cell communication. Bioinformatics 38, 4117–4126 (2022).
Rao, N. et al. Charting spatial ligand-target activity using Renoir. Preprint at bioRxiv https://doi.org/10.1101/2023.04.14.536833 (2024).
Yuan, Y. et al. CINS: Cell Interaction Network inference from single cell expression data. PLoS Comput. Biol. 18, e1010468 (2022).
Kyaw, W. et al. ENTRAIN: integrating trajectory inference and gene regulatory networks with spatial data to co-localize the receptor–ligand interactions that specify cell fate. Bioinformatics 39, btad765 (2023).
Mason, K. et al. Niche-DE: niche-differential gene expression analysis in spatial transcriptomics data identifies context-dependent cell-cell interactions. Genome Biol. 25, 14 (2024).
Armingol, E., Officer, A., Harismendy, O. & Lewis, N. E. Deciphering cell–cell interactions and communication from gene expression. Nat. Rev. Genet. 22, 71–88 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Hu, Y., Peng, T., Gao, L. & Tan, K. CytoTalk: de novo construction of signal transduction networks using single-cell transcriptomic data. Sci. Adv. 7, eabf1356 (2021).
Baruzzo, G., Cesaro, G. & Di Camillo, B. Identify, quantify and characterize cellular communication from single-cell RNA sequencing data with scSeqComm. Bioinformatics 38, 1920–1929 (2022).
Cheng, J., Zhang, J., Wu, Z. & Sun, X. Inferring microenvironmental regulation of gene expression from single-cell RNA sequencing data using scMLnet with an application to COVID-19. Brief. Bioinform. 22, 988–1005 (2021).
Jiang, P. et al. Systematic investigation of cytokine signaling activity at the tissue and single-cell levels. Nat. Methods 18, 1181–1191 (2021).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat. Rev. Genet. 11, 733–739 (2010).
Luecken, M. D. & Theis, F. J. Current best practices in single‐cell RNA‐seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Ching, T., Huang, S. & Garmire, L. X. Power analysis and sample size estimation for RNA-Seq differential expression. RNA 20, 1684–1696 (2014).
Vieth, B., Ziegenhain, C., Parekh, S., Enard, W. & Hellmann, I. powsimR: power analysis for bulk and single cell RNA-seq experiments. Bioinformatics 33, 3486–3488 (2017).
Schmid, K. T. et al. scPower accelerates and optimizes the design of multi-sample single cell transcriptomic studies. Nat. Commun. 12, 6625 (2021).
Medaglia, C. et al. Spatial reconstruction of immune niches by combining photoactivatable reporters and scRNA-seq. Science 358, 1622–1626 (2017).
Browaeys, R., Saelens, W. & Saeys, Y. Dataset to demonstrate the use of NicheNet on a Seurat object. Zenodo https://zenodo.org/records/3531889 (2019).
Browaeys, R. NicheNet v2: final networks and ligand-target matrice. Zenodo https://zenodo.org/records/7074291 (2022).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-Seq data. Nat. Commun. 13, 3224 (2022).
Acknowledgements
C.S.-a. is funded by the Ghent University Special Research Fund (grant number BOF21-DOC-105), R.S. is funded by the Flemish Government under the Flanders AI Research Program and Y.S. is funded by Ghent University Special Research Fund (grant number BOF18-GOA-024), the Belgian Excellence of Science (EOS) program, FWO SBO (grant number S001121N) and the Leducq project “Cellular and Molecular Drivers of Acute Aortic Dissections.”
Author information
Authors and Affiliations
Contributions
R.B. conceptualized the NicheNet algorithm. R.B. and C.S.-a. wrote the code. R.S. and Y.S. supervised the work. C.S. wrote the manuscript. All authors edited, read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key references
Browaeys, R., Saelens, W. & Saeys, Y. Nat. Methods 17, 159–162 (2020): https://doi.org/10.1038/s41592-019-0667-5
Bonnardel, J. et al. Immunity 51, 638–654.e9 (2019): https://doi.org/10.1016/j.immuni.2019.08.017
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sang-aram, C., Browaeys, R., Seurinck, R. et al. Unraveling cell–cell communication with NicheNet by inferring active ligands from transcriptomics data. Nat Protoc 20, 1439–1467 (2025). https://doi.org/10.1038/s41596-024-01121-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41596-024-01121-9