Abstract
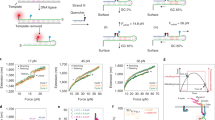
Understanding how cells sense and respond to mechanical forces is crucial for many biological processes, including adhesion, migration, differentiation and immune activation. In this protocol, we describe two advanced DNA-based tension probes, the reversible shearing DNA-based tension probe (RSDTP) and ForceChrono probe, which provide powerful tools for studying mechanotransduction in living cells. RSDTPs enable dynamic quantification of forces ranging from 4 to 60 pN, offering the advantage of reversibility without ligand depletion, making them ideal for ensemble force measurements across populations of cells. ForceChrono probes not only measure the magnitude of force but also capture its duration and loading rate, providing essential insights into the temporal dynamics of single-molecule force transmission. We detail the fundamental principles, design strategies and step-by-step procedures for synthesizing, purifying and applying these probes, including surface preparation, cell experiments, image acquisition and data analysis. In addition, we describe the computational tools for image analysis. Together, these probes enable a detailed analysis of cellular mechanobiology, with applications in integrin mechanobiology and cell adhesion biology. This protocol is suitable for researchers with a background in cell biology, molecular biology, surface chemistry, optical imaging and data analysis and can be completed by a graduate student in 3–4 days.
Key points
-
This protocol describes the use of two advanced DNA-based tension probes, the reversible shearing DNA-based tension probe (RSDTP) and ForceChrono probe, which provide powerful tools for studying mechanotransduction in living cells.
-
Compare to other probes, RSDTP enables reversible and repeatable measurements to be made across a wide range of forces, while ForceChrono probes capture force duration and loading rate, providing essential insights into the temporal dynamics of single-molecule force transmission.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The main images and analysis data in this protocol are available in the supporting primary research papers29,30. Additional source data have been uploaded to the online repository Figshare68 (https://doi.org/10.6084/m9.figshare.30016708.v1). Source data are provided with this paper.
Code availability
The data analysis tool (MATLAB code) used in this protocol can be accessed at https://doi.org/10.5281/zenodo.1489039365.
References
Orr, A. W., Helmke, B. P., Blackman, B. R. & Schwartz, M. A. Mechanisms of mechanotransduction. Dev. Cell 10, 11–20 (2006).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Shattil, S. J., Kim, C. & Ginsberg, M. H. The final steps of integrin activation: the end game. Nat. Rev. Mol. Cell Biol. 11, 288–300 (2010).
Kechagia, J. Z., Ivaska, J. & Roca-Cusachs, P. Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 20, 457–473 (2019).
Leckband, D. E., le Duc, Q., Wang, N. & de Rooij, J. Mechanotransduction at cadherin-mediated adhesions. Curr. Opin. Cell Biol. 23, 523–530 (2011).
Roca-Cusachs, P., Conte, V. & Trepat, X. Quantifying forces in cell biology. Nat. Cell Biol. 19, 742–751 (2017).
Polacheck, W. J. & Chen, C. S. Measuring cell-generated forces: a guide to the available tools. Nat. Methods 13, 415–423 (2016).
Lee, J. et al. Traction forces generated by locomoting keratocytes. J. Cell Biol. 127, 1957–1964 (1994).
Liu, Y., Galior, K., Ma, V. P. & Salaita, K. Molecular tension probes for imaging forces at the cell surface. Acc. Chem. Res. 50, 2915–2924 (2017).
Humphrey, J. D., Dufresne, E. R. & Schwartz, M. A. Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol. 15, 802–812 (2014).
Iskratsch, T., Wolfenson, H. & Sheetz, M. P. Appreciating force and shape—the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 15, 825–833 (2014).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
He, L. et al. Mechanical regulation of stem-cell differentiation by the stretch-activated Piezo channel. Nature 555, 103–106 (2018).
Huse, M. Mechanical forces in the immune system. Nat. Rev. Immunol. 17, 679–690 (2017).
Hu, K. H. & Butte, M. J. T cell activation requires force generation. J. Cell Biol. 213, 535–542 (2016).
Chakraborty, A. K. & Weiss, A. Insights into the initiation of TCR signaling. Nat. Immunol. 15, 798–807 (2014).
Vogel, V. & Sheetz, M. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 7, 265–275 (2006).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Plotnikov, S. V., Pasapera, A. M., Sabass, B. & Waterman, C. M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Zhao, Y., Wang, Y., Sarkar, A. & Wang, X. Keratocytes generate high integrin tension at the trailing edge to mediate rear de-adhesion during rapid cell migration. iScience 9, 502–512 (2018).
Stabley, D. R., Jurchenko, C., Marshall, S. S. & Salaita, K. S. Visualizing mechanical tension across membrane receptors with a fluorescent sensor. Nat. Methods 9, 64–67 (2011).
Zhang, Y., Ge, C., Zhu, C. & Salaita, K. DNA-based digital tension probes reveal integrin forces during early cell adhesion. Nat. Commun. 5, 5167 (2014).
Blakely, B. L. et al. A DNA-based molecular probe for optically reporting cellular traction forces. Nat. Methods 11, 1229–1232 (2014).
Liu, Y. et al. DNA-based nanoparticle tension sensors reveal that T-cell receptors transmit defined pN forces to their antigens for enhanced fidelity. Proc. Natl Acad. Sci. USA 113, 5610–5615 (2016).
Ma, R. et al. DNA probes that store mechanical information reveal transient piconewton forces applied by T cells. Proc. Natl Acad. Sci. USA 116, 16949–16954 (2019).
Bouchiat, C. et al. Estimating the persistence length of a worm-like chain molecule from force-extension measurements. Biophys. J. 76, 409–413 (1999).
Zhang, C. et al. The mechanical properties of RNA-DNA hybrid duplex stretched by magnetic tweezers. Biophys. J. 116, 196–204 (2019).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and notch signaling. Science 340, 991–994 (2013).
Li, H. et al. A reversible shearing DNA probe for visualizing mechanically strong receptors in living cells. Nat. Cell Biol. 23, 642–651 (2021).
Hu, Y. et al. DNA-based ForceChrono probes for deciphering single-molecule force dynamics in living cells. Cell 187, 3445–3459.e15 (2024).
Danilowicz, C. et al. Study of force induced melting of dsDNA as a function of length and conformation. J. Phys. Condens. Matter 22, 414106 (2010).
Mosayebi, M., Louis, A. A., Doye, J. P. & Ouldridge, T. E. Force-induced rupture of a DNA duplex: from fundamentals to force sensors. ACS Nano 9, 11993–12003 (2015).
Sun, F. et al. Exploring mechanical responses of cells to geometric information using micropatterned DNA-based molecular tension probes. ACS Nano 17, 18584–18595 (2023).
Wang, L. et al. Exploring integrin-mediated force transmission during confined cell migration by DNA-based tension probes. Anal. Chem. 94, 4570–4575 (2022).
Hu, Y. et al. Molecular force imaging reveals that integrin-dependent mechanical checkpoint regulates Fcγ-receptor-mediated phagocytosis in macrophages. Nano Lett. 23, 5562–5572 (2023).
Sethi, S. et al. Nuclease‐resistant L‐DNA tension probes enable long‐term force mapping of single cells and cell consortia. Angew. Chem. Int. Ed. Engl. 63, e202413983 (2024).
Zhao, Y., Sarkar, A. & Wang, X. Peptide nucleic acid based tension sensor for cellular force imaging with strong DNase resistance. Biosens. Bioelectron. 150, 111959 (2020).
Tu, Y., Pal, K., Austin, J. & Wang, X. Filopodial adhesive force in discrete nodes revealed by integrin molecular tension imaging. Curr. Biol. 32, 4386–4396.e3 (2022).
Roca-Cusachs, P., Gauthier, N. C., del Rio, A. & Sheetz, M. P. Clustering of α5β1 integrins determines adhesion strength whereas αvβ3 and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Galior, K. et al. Titin-based nanoparticle tension sensors map high-magnitude integrin forces within focal adhesions. Nano Lett. 16, 341–348 (2015).
Pawlak, M. R. et al. RAD-TGTs: high-throughput measurement of cellular mechanotype via rupture and delivery of DNA tension probes. Nat. Commun. 14, 2468 (2023).
Wang, X. et al. Constructing modular and universal single molecule tension sensor using protein G to study mechano-sensitive receptors. Sci. Rep. 6, 21584 (2016).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Gustafsson, M. G. L. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl Acad. Sci. USA 102, 13081–13086 (2005).
Brockman, J. M. et al. Live-cell super-resolved PAINT imaging of piconewton cellular traction forces. Nat. Methods 17, 1018–1024 (2020).
Schlichthaerle, T., Lindner, C. & Jungmann, R. Super-resolved visualization of single DNA-based tension sensors in cell adhesion. Nat. Commun. 12, 2510 (2021).
Curd, A. et al. Construction of an instant structured illumination microscope. Methods 88, 37–47 (2015).
Delattre, S. Igniting new confocal imaging potential—Nikon AX R Series with NSPARC. Micros. Today 31, 23–27 (2023).
Wang, W. et al. Hydrogel-based molecular tension fluorescence microscopy for investigating receptor-mediated rigidity sensing. Nat. Methods 20, 1780–1789 (2023).
Cheng, S. et al. Light sheet illumination in single-molecule localization microscopy for imaging of cellular architectures and molecular dynamics. Npj Imaging 2, 49 (2024).
Li, X. et al. Spatial redundancy transformer for self-supervised fluorescence image denoising. Nat. Comput. Sci. 3, 1067–1080 (2023).
Liu, J. & Yan, J. Unraveling the dual-stretch-mode impact on tension gauge tethers’ mechanical stability. J. Am. Chem. Soc. 146, 7266–7273 (2024).
Liu, J. et al. Tension gauge tethers as tension threshold and duration sensors. ACS Sens. 8, 704–711 (2023).
Guo, Y., Yan, J. & Goult, B. T. Mechanotransduction through protein stretching. Curr. Opin. Cell Biol. 87, 102327 (2024).
Zhou, Y. et al. A novel design for magnetic tweezers with wide-range temperature control. Biophys. J. 122, 3860–3868 (2023).
Woodside, M. T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl Acad. Sci. USA 103, 6190–6195 (2006).
Pfaff, M. et al. Selective recognition of cyclic RGD peptides of NMR defined conformation by αIIbβ3, αVβ3, and α5β1 integrins. J. Biol. Chem. 269, 20233–20238 (1994).
Vrancken, K. C. et al. Modelling of the reaction-phase interaction of γ-aminopropyltriethoxysilane with silica. J. Chem. Soc. Faraday Trans. 89, 2037–2040 (1993).
Zhang, F. & Srinivasan, M. P. Self-assembled molecular films of aminosilanes and their immobilization capacities. Langmuir 20, 2309–2314 (2004).
Jurchenko, C. et al. Integrin-generated forces lead to streptavidin-biotin unbinding in cellular adhesions. Biophys. J. 106, 1436–1446 (2014).
Liu, Y., Yehl, K., Narui, Y. & Salaita, K. Tension sensing nanoparticles for mechano-imaging at the living/nonliving interface. J. Am. Chem. Soc. 135, 5320–5323 (2013).
Piella, J., Bastús, N. G. & Puntes, V. Size-controlled synthesis of sub-10-nanometer citrate-stabilized gold nanoparticles and related optical properties. Chem. Mater. 28, 1066–1075 (2016).
Pan, H. et al. A simple procedure to improve the surface passivation for single molecule fluorescence studies. Phys. Biol. 12, 045006 (2015).
Tsunoyama, T. A. et al. Super-long single-molecule tracking reveals dynamic-anchorage-induced integrin function. Nat. Chem. Biol. 14, 497–506 (2018).
Wu, P. et al. Data analysis for single-molecule mechanical force dynamics. Zenodo https://doi.org/10.5281/zenodo.14890393 (2025).
Yao, M. et al. Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation. Sci. Rep. 4, 4610 (2014).
Wang, Y. et al. Force-dependent interactions between talin and full-length vinculin. J. Am. Chem. Soc. 143, 14726–14737 (2021).
Wu, P. Measure cellular force using DNA-based tension probes: from ensemble to single-molecule studies—source data. figshare https://doi.org/10.6084/m9.figshare.30016708 (2025).
Haiss, W., Thanh, N. T., Aveyard, J. & Fernig, D. G. Determination of size and concentration of gold nanoparticles from UV-vis spectra. Anal. Chem. 79, 4215–4221 (2007).
Acknowledgements
We sincerely thank the Jie Yan laboratory for their guidance and support in the theoretical calculations of DNA mechanics.
Author information
Authors and Affiliations
Contributions
Z.L. supervised the project. P.W. and Y.H. designed and carried out the experiments and performed data analysis. H.L. wrote the code for data-analysis tools for the ForceChrono probes. P.W., Y.H., W.C. and Z.L. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key references
Li, H. et al. Nat. Cell Biol. 23, 642–651 (2021): https://doi.org/10.1038/s41556-021-00691-0
Hu, Y. et al. Cell 187, 3445–3459.e15 (2024): https://doi.org/10.1016/j.cell.2024.05.008
Supplementary information
Supplementary Information (download PDF )
Supplementary Note 1, Supplementary Figures 1–5 and Supplementary Tables 1–6
Supplementary Data 1 (download XLSX )
Statistical source data of Supplementary Information figures
Source data
Source Data Figs. 2, 3 and 6 (download XLSX )
Statistical source data of Fig. 2f,g. Statistical source data of Fig. 3e–h. Statistical source data of Fig. 6a
Source Data Fig. 6 (download PDF )
Unprocessed gels of Fig. 6b,c
Source Data Fig. 6 (download PDF )
Unprocessed gels of Fig. 6d
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, P., Hu, Y., Li, H. et al. Measuring cellular force using DNA-based tension probes: from ensemble to single-molecule studies. Nat Protoc (2025). https://doi.org/10.1038/s41596-025-01277-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41596-025-01277-y