Abstract
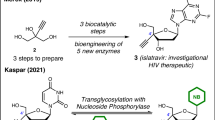
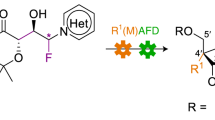
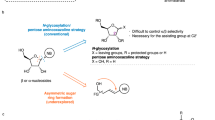
C4′-modified nucleoside analogs continue to attract global attention for treating infectious diseases and as components in oligonucleotide therapeutics. Current preparations mostly employ lengthy semi-synthetic approaches that do not allow for the efficient exploration of the chemical space associated with this valuable nucleoside subclass. Here we describe the pilot-scale (250 mg) and process-scale (85 g) preparation of 4′-methyl-ribothymidine (4′,5-dimethyluridine) using a de novo strategy. Both l- and d-nucleoside analogs are accessible, and 10 different C4′ modifications and 20 different nucleobases can be used interchangeably to create new analogs. This protocol involves the use of an enantioselective proline-catalyzed aldol between 2,2-dimethoxyacetaldehyde and a dioxanone. A 1,2-addition into the aldol product installs the C4′ modification. Subsequent cyclization via intramolecular trans-acetalization delivers the modified ribose core of the nucleoside analog. Peracetylation, followed by Vorbrüggen glycosylation, completes the route. The pilot- and process-scale protocols can be completed in ~5 and ~7 d, respectively, to deliver C4′-modified nucleoside analogs in good yields and excellent enantiopurity.
Key points
-
C4′-modified nucleoside analogs can be prepared via this five-step process. Pilot-scale (250 mg) and process-scale (85 g) procedures are described.
-
Although not limited to these, 10 different C4′ modifications and 20 different nucleobases have been shown to be compatible with this procedure.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The relevant data for this protocol can be found in the text and Supplementary Information of this paper and/or its supporting primary research paper. A preprint article containing the raw data for the additional C4′ modifications and nucleobases has been deposited at ChemRxiv https://doi.org/10.26434/chemrxiv.10001711/v1.
References
Shelton, J. et al. Metabolism, biochemical actions, and chemical synthesis of anticancer nucleosides, nucleotides, and base analogs. Chem. Rev. 116, 14379–14455 (2016).
Seley-Radtke, K. L. & Yates, M. K. The evolution of nucleoside analogue antivirals: a review for chemists and non-chemists. Part I: early structural modifications to the nucleoside scaffold. Antiviral Res. 154, 66–86 (2018).
Yates, M. K. & Seley-Radtke, K. L. The evolution of nucleoside analogue antivirals: a review for chemists and non-chemists. Part II: complex modifications to the nucleoside scaffold. Antiviral Res. 162, 5–21 (2019).
Galmarini, C. M., Mackey, J. R. & Dumontet, C. Nucleoside analogues and nucleobases in cancer treatment. Lancet Oncol. 3, 415–424 (2002).
Monroe, J. et al. N1-Methylpseudouridine and pseudouridine modifications modulate mRNA decoding during translation. Nat. Commun. 15, 8199 (2024).
Bege, M. & Borbás, A. The medicinal chemistry of artificial nucleic acids and therapeutic oligonucleotides. Pharmaceuticals 15, 909–948 (2022).
Chang, J. 4′-Modified nucleosides for antiviral drug discovery: achievements and perspectives. Acc. Chem. Res. 55, 565–578 (2022).
Betson, M., Allanson, N. & Wainwright, P. A review of methods to synthesize 4′-substituted nucleosides. Org. Biomol. Chem. 12, 9291–9306 (2014).
Ito, Y. et al. Generation of 4′-carbon radicals via 1,5-hydrogen atom transfer for the synthesis of bridged nucleosides. Org. Lett. 24, 7696–7700 (2022).
Abe, R. et al. Photoredox-catalyzed site-selective intermolecular C(sp3)-H alkylation of tetrahydrofurfuryl alcohol. Org. Lett. 27, 795–801 (2025).
Muir, G. et al. Unmasking the halide effect in diastereoselective Grignard reactions applied to C4′ modified nucleoside synthesis. Nat. Commun. 16, 1679 (2025).
Nuligonda, T. et al. An enantioselective and modular platform for C4′-modified nucleoside analogue synthesis enabled by intramolecular trans-acetalizations. Nat. Commun. 15, 7080 (2024).
Nuligonda, T., Kumar, G., Wang, J. W., Mahal, L. K. & Meanwell, M. W. Exploring scope and scale limitations of a modular route to C4′-modified nucleoside analogues. Preprint at https://doi.org/10.26434/chemrxiv.10001711/v1 (2026).
Gunic, E., Girardet, J.-L., Pietrzkowski, Z., Esler, C. & Wang, G. Synthesis and cytotoxicity of 4′-C- and 5-C-substituted toyocamycins. Bioorg. Med. Chem. 9, 163–170 (2001).
Lee, S., Uttamapinant, C. & Verdine, G. L. A concise synthesis of 4′-fluoro nucleosides. Org. Lett. 9, 5007–5009 (2007).
Perrone, P. et al. First example of phosphoramidate approach applied to a 4′-substituted purine nucleoside (4′-azidoadenosine): conversion of an inactive nucleoside to a submicromolar compound versus hepatitis C virus. J. Med. Chem. 50, 5463–5470 (2007).
Meanwell, M. et al. A short de novo synthesis of nucleoside analogs. Science 369, 725–730 (2020).
Huffman, M. A. et al. Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366, 1255–1259 (2019).
McLaughlin, M. et al. Enantioselective synthesis of 4ʹ-ethynyl-2-fluoro-2ʹ-deoxyadenosine (EFdA) via enzymatic desymmetrization. Org. Lett. 19, 926–929 (2017).
Nawrat, C. C. et al. Nine-step stereoselective synthesis of islatravir from deoxyribose. Org. Lett. 22, 2167–2172 (2020).
Patel, N. R. et al. Five-step enantioselective synthesis of islatravir via asymmetric ketone alkynylation and an ozonolysis cascade. Chem. Eur. J. 26, 14118–14123 (2020).
Patel, N. R. et al. Synthesis of islatravir enabled by a catalytic enantioselective alkynylation of a ketone. Org. Lett. 22, 4659–4664 (2020).
Kaspar, F. et al. Diversification of 4ʹ-methylated nucleosides by nucleoside phosphorylases. ACS Catal. 11, 10830–10835 (2021).
Weber, E., Cordingley, M. G., Burnie, A. J., Darapaneni, C. M. & Paladino, M. Nucleosides for treating cancer. PrimeFour Therapeutics WO2023192499A1 (2023).
Sommadossi, J.-P., Gosselin, G. & Imbach, J.-L. Methods and compositions for treating flaviviruses and pestiviruses using 4′-modified nucleoside. US Patent 2004/0006002 A1 (2004).
Devos, R. et al. 4′-Substituted nucleoside derivatives as inhibitors of HCV RNA replication. US Patent 2003/0236216 A1 (2003).
Schinazi, R., Amlard, F., Gavegnano, C., Cox, B. & Mengshetti, S. Combined modalities for nucleosides and/or NADPH oxidase (NOX) inhibitors as myeloid-specific antiviral agents. WO2019133712A1 (2019).
Weber, E. et al. Nucleoside line-1 inhibitors. Transposon Therapeutics Inc. WO2023192491A2 (2023).
Grondal, C. & Enders, D. Direct asymmetric organocatalytic de novo synthesis of carbohydrates. Tetrahedron 62, 329–337 (2006).
Ishido, Y., Sakairi, N., Okazaki, K. & Nakazaki, N. Partial protection of carbohydrate derivatives. Part 4. Regioselective 2′-O-deacylation of fully acylated purine and pyrimidine ribonucleosides with hydroxylaminium acetate. J. Chem. Soc. Perkin Trans. I 0, 563–573 (1980).
Xia, R. & Chen, L.-S. Efficient synthesis of cladribine via the metal-free deoxygenation. Nucleosides Nucleotides Nucleic Acids 34, 729–735 (2015).
Prakash, G. K. S., Krishnamurti, R. & Olah, G. A. Fluoride-induced trifluoromethylation of carbonyl compounds with trifluoromethyltrimethylsilane (TMS-CF3). A trifluoromethide equivalent. J. Am. Chem. Soc. 111, 393–395 (1989).
Carr, G. & Whittaker, D. Lactone formation in superacidic media. J. Chem. Soc. Perkin Trans. II 1987, 1877–1880 (1987).
Li, Y. et al. Chemoselective and diastereoselective synthesis of C-aryl nucleoside analogues by nickel-catalyzed cross-coupoling of furanosyl acetates with aryl iodides. Angew. Chem. Int. Ed. 61, e202114619 (2022).
Thiesen, L. J. H., Cabral, N., Silva, M. J., Bezerra, G. & Doboszewski, B. larger laboratory scale synthesis of 5-methyluridine and formal synthesis of its l-enantiomer. Arkivoc 4, 249–264 (2017).
Lawhorn, B. G., Mehl, R. A. & Begley, T. P. Biosynthesis of the thiamin pyrimidine: the reconstruction of a remarkable rearrangement reaction. Org. Biomol. Chem. 2, 2538–2546 (2004).
Gurjar, M. K., Reddy, L. K. & Hotha, S. Synthesis of ethyl 5-O-(α-d-arabinofuranosyl)-6-O-(β-d-galactofuranosyl)-β-d-galactofuranoside present in motif E of the Mycobacterium tuberculosis cell well. Org. Lett. 3, 321–323 (2001).
Anderson, B. G., Bauta, W., Cantrell, W. R. Jr., Engles, T. & Lovett, D. P. Isolation, synthesis, and characterization of impurities and degradants from the clofarabine process. Org. Process Res. Dev. 12, 1229–1237 (2008).
Desai, S. P., Yatzoglou, G., Turner, J. A. & Taylor, M. S. Boronic acid-catalyzed regio- and stereoselective N-glycosylations of purines and other azole heterocycles: access to nucleoside analogues. J. Am. Chem. Soc. 146, 4973–4984 (2024).
Acknowledgements
We thank funding from the Alberta Ministry of Technology and Innovation through SPP–ARC (Striving for Pandemic Preparedness–The Alberta Research Consortium). M.W.M. is thankful for the support from the Manley and Marian Johnston Professorship in Chemistry. G.K. and L.K.M. acknowledge the Canada Excellence Research Chairs Program (CERC in Glycomics). We thank M. Ferguson for his work on structural confirmation with X-ray crystallography. We also thank E. Fu (Hall Lab) for his work on %ee determination.
Author information
Authors and Affiliations
Contributions
T.N. optimized the process-scale protocol, wrote the pilot- and process-scale procedures and the Supplementary Information. T.N. and G.K. expanded the substrate scope of the protocol. J.W.W. helped optimize the process-scale protocol. L.K.M. edited the manuscript and the pilot- and process-scale procedures. M.W.M. wrote the manuscript and supervised the project.
Corresponding author
Ethics declarations
Competing interests
M.W.M. and T.N. declare the following competing interests: the University of Alberta holds a patent (WO 2025/199647 A1) describing the synthesis of C4ʹ-modified NAs via the process presented in this manuscript. The other authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Stanislaw Wnuk, who co-reviewed with Mukesh Mudgal; Noriko Saito-Tarashima; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key reference
Nuligonda, T. et al. Nat. Commun. 15, 7080 (2024): https://doi.org/10.1038/s41467-024-51520-5
Extended data
Extended Data Fig. 1
A) Aldol reaction: synthesis of aldol product. B) Purification of process scale aldol reaction. The aldol reaction needs to be maintained at between 0 - 4oC, which can be done in a refrigerator as shown in panel A. Purification on process scale requires the use of column with an internal diameter of 10 cm.
Extended Data Fig. 2
A) Cannulation setup for Grignard reaction. B) Setup for the filtration through celite for quenched Grignard reaction. On process scale, the Grignard should be slowly cannulated into the reaction mixture (panel A). The quenched reaction mixture is filtered through a pad of celite to remove insoluble salts (panel B).
Extended Data Fig. 3
A) pilot scale setup of glycosylation reactions; B) Process setup of glycosylation reaction. On pilot scale, it is possible to set up multiple glycosylation reactions in the same heating bath (panel A). A 1 L round-bottom flask is required for process scale glycosylation (panel B).
Extended Data Fig. 4
Concentration of crude 4′-methylribothymidine (19) (process scale); B) Blow drying pilot scale sample of 19. Rotary evaporators are used to remove solvents from crude mixtures and purified samples (panel A). In order to remove the acetamide by-product from the glycosylation, blow drying the sample with a steady stream of air for several hours is required (panel B).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Rights and permissions
About this article
Cite this article
Nuligonda, T., Kumar, G., Wang, J.W. et al. Preparation of C4′-modified nucleoside analogs. Nat Protoc (2026). https://doi.org/10.1038/s41596-026-01353-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41596-026-01353-x