Abstract
The Homo sapiens Chromosomal Location Ontology (HSCLO) is designed to facilitate the integration of human genomic features into biomedical knowledge graphs from releases GRCh37 and GRCh38 at multiple resolutions. HSCLO comprises two distinct versions, HSCLO37 and HSCLO38, each tailored to its respective human genome release. This ontology supports the efficient integration and analysis of human genomic data across scales ranging from entire chromosomes to individual base pairs, thereby enhancing data retrieval and interoperability within large-scale biomedical datasets. Unlike existing ontologies that primarily focus on genomic feature identification or annotation, HSCLO is specifically engineered to optimize the interoperability and scalability of genomic data within biomedical knowledge graphs. The utility and performance of HSCLO are demonstrated through a case study involving the integration of high-resolution chromatin interaction data, which reveals significant improvements in query efficiency and data linkage. HSCLO represents a valuable resource for advancing research in disease genetics, personalized medicine, and other domains that require complex genomic data integration.
Similar content being viewed by others
Introduction
Knowledge graphs (KGs) have become important tools in integrating and analyzing heterogeneous biomedical data, enabling the linkage of diverse datasets in a structured and semantically enriched manner. However, integrating genomic feature data into KGs presents significant challenges due to the need for precise alignment across varying chromosomal resolutions, from entire chromosomes to individual base pairs. Current methods for identifying overlapping genomic features can involve computationally intensive searches, particularly when applied to large-scale biomedical knowledge graphs containing millions of data nodes. This highlights the need for an ontological framework that could readily be integrated and utilized in knowledge graphs (in the form of subject/predicate/object triples) that standardizes the representation of chromosomal locations and reduces computational complexity during data integration.
In biomedical research, analyzing chromosomal locations of genomic features and their positional relationships is critical for understanding disease mechanisms, guiding therapeutic development, and personalizing medical treatments. An ontology that systematically catalogs and categorizes chromosomal positions is essential for supporting these research endeavors. For example, a researcher may wish to rapidly interrogate a knowledge graph for all genomic features and annotations that are proximal to a chromosomal variant. Such a framework would enable these queries by linking variants to associated genes, regulatory elements, or chromatin interaction data across different resolution levels, thereby expediting complex analyses and supporting hypothesis generation and experimental validation.
Existing ontologies have contributed significantly to the standardization of genomic data, particularly in annotating specific genomic features or chromosomal structures1,2,3,4. However, these ontologies are not optimized for integrating positional information at multiple chromosomal resolutions within KGs, with a standing gap in the ability to link genomic data across different resolution scales efficiently.
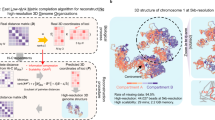
To address this gap, we introduce the Homo sapiens Chromosomal Location Ontology (HSCLO), developed for the GRCh375 (HSCLO37) and GRCh386 (HSCLO38) genome assemblies (Fig. 1). HSCLO is specifically designed to facilitate the integration of positional information into biomedical knowledge graphs, supporting rapid querying and efficient data linkage across diverse experimental datasets. In this study, we detail the construction of HSCLO, present its application in a knowledge graph environment, and demonstrate its utility through a case study involving high-resolution chromatin interaction data integration.
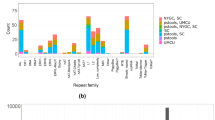
Graphical Schema of HSCLO. Entity X could be any chromosomal feature, including chromosomal bands, genes, exons, introns, regulatory elements, QTLs, variants, accessible chromatin regions, viral integration sites, human endogenous retroviruses, transposons, tandem repeats, chromosomal contact regions, TADs, telomere, centromeres, and any other type.
The HSCLO is currently utilized in two knowledge graph projects, Petagraph7 and the Common Fund Data Ecosystem’s Data Distillery Project8. In the Data Distillery project, HSCLO has been indispensable in integrating Common Fund genomic datasets across different resolution levels, for example, in connecting 4DN chromosomal loop data9 to individual genes or variants from sources such as GTEx10 or Kids First11.
Methods
Building HSCLO versions
To construct the HSCLO versions for GRCh37 and GRCh38, we downloaded each genomic coordinate file from the UCSC website (https://hgdownload.soe.ucsc.edu/goldenPath/hg19/bigZips/hg19.chrom.sizes and https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/hg38.chrom.sizes). We parsed them into binned location nodes that were organized by chromosome size and scale using custom R scripts (https://github.com/TaylorResearchLab/HSCLO/blob/main/Codes/HSCLO38.R). The nodes were defined at five resolution levels (1 kbp, 10 kbp, 100 kbp, 1 Mb, chromosome) and connected through hierarchical relationships reflecting their chromosomal positioning. Each level in the ontology was linked to both its parent level and neighboring nodes using specific edge names (e.g., “above_1Mbp_band,” “precedes_10kbp_band”). Nodes were named using a UCSC coordinate format-like structure12 as for example HSCLO38:chrN:nnnn-nnnn. The preprocessing scripts are available on GitHub (https://github.com/TaylorResearchLab/HSCLO/blob/main/Codes/HSCLO38.R).
Use-case methodology
For our use case, we show how we establish the intersection between the 4DN dataset and Gencode to profile the overlap between 4DN loops and human genes in GRCh38. Once ingested in a knowledge graph in the Neo4j database management environment (Neo4j Desktop Community Edition 1.5.7), we used the Cypher query language to query ENSEMBL nodes in Gencode12 and their corresponding HSCLO nodes at 1kbp resolution using the query in Fig. 2. The query utilizes the mapping of ENSEMBL genes and transcript onto the HSCLO38 through the GENCODE_HSCLO mapping dataset:
//Query1: GENCODE Overlap with HSCLO38 at 1kbp MATCH (o1:Code {SAB:'ENSEMBL'})<-[:CODE]-(c1:Concept)-[r1:us_5_prime]-> (c2:Concept)-[:CODE]->(o2:Code{SAB:’HSCLO’}),(c1:Concept)- [r2:ds_3_prime]->(c3:Concept)-[:CODE]->(o3:Code {SAB:’HSCLO’}), p = shortestPath((c2)-[:precedes_1kbp_band*]->(c3)) WHERE ((c2)<>(c3)) RETURN * LIMIT 1
Loop dotcall files of the following 4DN datasets were obtained from the 4D Nucleome project website data repository (https://data.4dnucleome.org). The dataset URLs and descriptions are provided in Table 3. The following query can identify 4DN loops and their anchor coordinates along HSCLO38 at 1kbp resolution.
//Query2: 4DN Overlap with HSCLO38 at 1kbp MATCH (o1:Code {SAB:'4DNL'})<-[a:CODE]-(c1:Concept)-[r1]-> (c2:Concept)-[b:CODE]->(o2:Code {SAB:'HSCLO'}),(c1:Concept)-[r2]-> (c3:Concept)-[c:CODE]->(o3:Code {SAB:'HSCLO'}),p=shortestPath((c2)- [:precedes_1kbp_band*]->(c3)) WHERE ((c2) <> (c3)) RETURN * LIMIT 1
Note that the query uses the Neo4j shortestPath procedure to extract HSCLO38 nodes connected using the “precedes_1kbp_band” relationship. Subsequently, genes that partially or entirely overlapped with human loops were identified through querying for shared links to HSCLO38:
//Query3: 4DN Overlap with GENCODE through HSCLO at 1 kbp resolution MATCH (o1:Code {SAB:'4DNL'})<-[a:CODE]-(c1:Concept)- [r1:loop_us_start]->(c2:Concept)-[b:CODE]->(o2:Code {SAB:'HSCLO'}) MATCH (c1:Concept)-[r2:loop_ds_end]->(c3:Concept)-[c:CODE]->(o3:Code {SAB:'HSCLO'}),p = shortestPath((c2)-[:precedes_1kbp_band*]->(c3)) WITH o1.CODE AS loop, nodes(p) AS P MATCH (o1:Code {SAB:'ENSEMBL'})<-[:CODE]-(c1:Concept)-[r1]->(c2:Concept) WHERE (c2 in P) RETURN loop, o1.CODE
After merging with the list of loops that overlap genes (reciprocal of Query 3), the complete list of human genes and overlapping loops was obtained. Analysis for density estimation was performed in R (RStudio 2022.12.0.35313,14 and R v4.2.214). Functional annotation of the gene list associated with the chromosomal loop with the highest number of genes per 100kbp loop length used Metascape15.
Results
HSCLO effectively maps chromosomal locations across five genomic resolution levels: whole chromosome, 1 Mbp, 100 kbp, 10 kbp, and 1 kbp. Over 3.4 million nodes are interconnected to enable detailed mapping and querying within the knowledge graph environment. The hierarchical relationships allow seamless transitions between different scales of genomic data, supporting complex queries such as those involving 1 kbp nodes and their positional relationships within larger chromosomal contexts (Table 1).
HSCLO38 defines chromosomal locations within the GRCh38 release for chr 1–22, X, Y, and mtDNA (Fig. 1). It provides hierarchical relationships across five genomic resolution levels: whole chromosome, 1 megabase pair (Mbp), 100 kilobase pairs (kbp), 10 kbp, and 1 kbp. Each node within these class levels is interconnected to its scale parent and the immediate neighbors on either side to support mapping and association between genomic datasets and features. For example, the 1kbp element HSCLO38:chr1.20517001–20518000 is connected to its “scale parent” at a lower resolution, HSCLO38:chr1.20510001–20520000 through the “below_1kbp_band” relationship as well as to its immediate neighbors: its 5′ neighbor HSCLO38:chr1.20516001–20517000 and 3′ neighbor HSCLO38:chr1.20518001–20519000 through “precedes_1kbp_band” relationships. Using human genome version GRCh38, the HSCLO38 schema results in 3,431,155 nodes and 6,862,195 relationships (Table 1). Similarly, the summary statistics of GRCh37 are provided in Table 2.
We provide a use case for linking biodata at different resolutions to demonstrate the practical application of HSCLO38 in knowledge organization and discovery. A researcher may be interested in identifying genes found within large-scale chromatin organization features, such as Hi-C data hosted by the 4DN project9. We began by importing HSCLO38 into Petagraph, our custom biomedical KG7, and then creating edges in the KG to link all gene nodes from GENCODE v4116 to their respective 1 kbp HSCLO38 nodes. We then created edges for the chromosomal loops from a set of files at the 4DN project9 to their respective 1 kbp locations in HSCLO38. Using a Cypher query in the Neo4j v5 environment, we retrieved the overlap in 1 kbp nodes between the spans of the GENCODE gene definitions and the start and end points of the 4DN loops. Figure 2 depicts the example query results outlined above. Figure 2A provides a sequence of HSCLO38 connecting the start and end of a 4DN loop upstream anchor. Figure 2B shows how HSCLO38 can extract the corresponding 1 kbp nodes, such as for a human transcript location. Figure 3A illustrates the frequency distribution of human genes overlapping the 4DN loops provided by the 4DN dot call dataset 4DNFIIQP46FO as a function of chromosome 1 coordinates as an example of how HSCLO38 could be utilized to bridge independent datasets in a knowledge graph context. Figure 3 shows the distribution of 4DN loop sizes (Fig. 3B) and the number of GENCODE-defined genes overlapping the 4DN dataset loops in 4DNFIIQP46FO (Fig. 3C,D, respectively). Further analysis of this data reveals ~36,000 loops (from 4DNFIIQP46FO) that overlap at least one gene. To explore the biological relevance of this analysis, we performed functional annotation of the gene list from the loop 4DNFIIQP46FO.chr12.3050000–3060000.chr12.10310000–10320000, which was identified as having the highest number of overlapping genes (173). The analysis provides the top 10 enriched pathways (Table 4), top 10 DisGenNet diseases (Table 5), and top 10 MSigDB cell types (Table 6), implying the disruption in the loop structure and, subsequently, the expression regulation of the overlapping genes could be associated with developmental disorders primarily related to muscular development.
Chromosomal features annotated with HSCLO. Example query results derived from Queries 1 and 2 outlined in the Methods section depict a 4DN loop upstream anchor (A) and a human gene transcript (B) as spanned along the HSCLO38.
Use case distributions. (A) Frequency distribution of genes overlapping the 4DN-computed loops in the 4DN dot call dataset 4DNFIIQP46FO ingested in Petagraph plotted for human chromosome 1 as a function of loop midpoints. (B) Loop size distribution according to 4DNFIIQP46FO dataset (in base-pairs), (C) gene count per loop distribution and (D) distribution of averaged gene count per 100 kbp of loop span derived from the intersection of 4DN loops recorded in 4DNFIIQP46FO and GENCODE genes identified through their connection to HSCLO38.
Discussion
The HSCLO is a structured ontological framework essential for organizing and categorizing information on the precise physical positions of genes, genetic markers, and other genomic elements along chromosomes in two human genome versions (GRCh37 and GRCh38). This ontology establishes a standardized vocabulary and hierarchical structure for accurately describing chromosomal positions in knowledge graphs, ensuring uniformity in data representation and sharing across diverse databases and research endeavors. Given the sizable and heterogeneous character of genomic data from multiple sources and studies, HSCLO functions as a unifying framework to enable the integration of datasets based on chromosomal coordinates.
HSCLO offers several advantages that enhance its utility for graphs in biomedical research. The design specifically addresses the challenge of integrating genomic data across multiple resolution levels within knowledge graphs, which supports accurate data alignment across scales, from entire chromosomes to 1kbp segments. This capability is particularly valuable when integrating large-scale studies where different datasets must be linked at varying feature resolutions. HSCLO’s hierarchical structure also enables rapid querying and efficient data retrieval, which are essential for handling the vast amounts of data typically involved in genomic studies.
Despite these strengths, HSCLO does have certain limitations. The only currently available versions are based on GRCh37 and GRCh38 genome assemblies, which may limit its applicability for other assemblies until an update is made to accommodate additional reference genomes. Another limitation inherent to large reference datasets in graphs is the computational demand associated with maintaining and querying the large number of nodes and relationships, especially at HSCLO’s finer resolution of 1 kbp. Large-scale analyses at a finer resolution can be resource-intensive and may pose challenges for researchers working in environments with limited computational infrastructure.
Another potential limitation of employing HSCLO is inherent to the complexity involved in its implementation and use. While HSCLO is designed to facilitate data integration, the initial setup process to annotate a new dataset —such as mapping and linking data coordinates to the ontology—can require data preparation and standardization efforts. To address these challenges, future developments will focus on creating more user-friendly tools and documentation and releasing prepared datasets that simplify the process of using HSCLO.
In clinical and biomedical research contexts, understanding the chromosomal locations of genes associated with diseases or genetic variants assumes critical importance. HSCLO38 and HSCLO37 facilitate the systematic cataloging and classification of such pertinent genetic information, thereby supporting investigations into disease genetics and personalized medicine applications. Furthermore, due to the reliance of computational tools and algorithms on structured data, HSCLO can be a foundational resource for developing robust computational methodologies for genomic analysis and interpretation. Thus, HSCLO can play a pivotal role in harmonizing, integrating, and standardizing genomic data, enhancing data interoperability, fostering interdisciplinary research collaborations, and catalyzing advancements in computational tools essential for fundamental research and applied biomedical applications.
HSCLO stands out from earlier ontologies by focusing on ontologized genomic coordinate binning, facilitating integration across various resolution levels in biomedical knowledge graphs. It addresses the challenge of handling genomic data with differing experimental resolutions while ensuring compatibility with the GRCh38 and GRCh37 genome assemblies. HSCLO can be a valuable tool for researchers and data scientists aiming to integrate and analyze genomic data in large-scale biomedical knowledge environments.
Data availability
Knowledge-graph-ready edgelists (triple format) can be found on the HSCLO project page at the OSF website: https://osf.io/pe8v7/, https://doi.org/10.17605/OSF.IO/PE8V7.
Code availability
The code used to generate and query HSCLO is available in a public repository: https://github.com/TaylorResearchLab/HSCLO/tree/main/HSCLO38.
References
Eilbeck, K. et al. The Sequence Ontology: a tool for the unification of genome annotations. Genome Biol. 6, R44 (2005).
Baran, J. et al. GFVO: the Genomic Feature and Variation Ontology. PeerJ 3, e933 (2015).
Mungall, C. J., Emmert, D. B. & FlyBase Consortium. A Chado case study: an ontology-based modular schema for representing genome-associated biological information. Bioinformatics 23, i337–46 (2007).
Feng, F. et al. GenomicKB: a knowledge graph for the human genome. Nucleic Acids Res. 51, D950–D956 (2023).
Church, D. M. et al. Modernizing reference genome assemblies. PLoS Biol 9, e1001091, https://doi.org/10.1371/journal.pbio.1001091 (2011).
Schneider, V. A. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Research 27, 849–864 (2017).
Stear, B. J. et al. Petagraph: A large-scale unifying knowledge graph framework for integrating biomolecular and biomedical data. Sci Data 11, 1338, https://doi.org/10.1038/s41597-024-04070-w (2024).
NIH Common Fund Data Ecosystem Data Distillery Partnership Repository. GitHub https://github.com/nih-cfde/data-distillery.
Dekker, J. et al. Spatial and temporal organization of the genome: Current state and future aims of the 4D nucleome project. Mol. Cell 83, 2624–2640 (2023).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Heath, A. P. et al. Abstract 2464: Gabriella Miller Kids First Data Resource Center: Harmonizing clinical and genomic data to support childhood cancer and structural birth defect research. vol. 79 2464–2464 (American Association for Cancer Research, 2019).
James, W. et al. The Human Genome Browser at UCSC. Genome Research 12(6), 996–1006, https://doi.org/10.1101/gr.229102 (2002).
Posit team. RStudio: Integrated Development Environment for R. (Posit Software, PBC, Boston, MA, 2022).
R Development Core Team, R. & Others. R: A language and environment for statistical computing (2011).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10 (2019).
Harrow, J. et al. GENCODE: the reference human genome annotation for the ENCODE Project. Genome Res. 22 (2012).
Acknowledgements
Funding for this project is acknowledged from the NIH Common Fund through the Office of Strategic Coordination/Office of the NIH Director under awards R03OD030600 and OT2OD030162 (DMT), and OT2OD026663, OT2OD026675 (JCS); The Department of Biomedical Informatics at The Children’s Hospital of Philadelphia (DMT).
Author information
Authors and Affiliations
Contributions
T.M.A. wrote the code, and provided analyses and figures. T.M.A., D.M.T. and B.J.S. wrote and edited the paper. T.M.A., D.M.T. and B.J.S. designed the HSCLO schema. T.M.A. and B.J.S. implemented HSCLO in the knowledge graphs. C.M.N. was the project manager. J.C.S. and J.A.S. were involved in the overall design of the knowledge graph environment around HSCLO and advised on its initial design. D.M.T. conceived of, guided, and funded work on HSCLO.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mohseni Ahooyi, T., Stear, B., Simmons, J.A. et al. Homo Sapiens Chromosomal Location Ontology: A Framework for Genomic Data in Biomedical Knowledge Graphs. Sci Data 12, 52 (2025). https://doi.org/10.1038/s41597-024-04358-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04358-x