Abstract
Circulating tumor DNA (ctDNA) variants hold significant promise as cancer biomarkers in liquid biopsies, owing to their minimally invasive testing approach and capacity to capture the comprehensive tumor landscape. However, a comprehensive resource for studying ctDNA variants is still lacking. Here, we developed CTDdgv to systematically identify ctDNA variants and interpret their clinical relevance. We manually curated 1674 experimentally validated clinical interpretations for ctDNA variants, concerning prognosis, drug resistance, tumor characteristics, metastasis monitoring, therapy, prospect for detection, and others. Furthermore, we developed and integrated a pipeline that identifies tumor driver genes (TDGs) and variants (TDVs) and evaluates the prognostic significance of potential TDGs. Using publicly available ctDNA mutation spectra, we identified potential TDGs and TDVs from 38 datasets across 17 cancer types. Based on these datasets, we provide a multi-dimensional analysis of TDG gene sets in specific cancer types, providing insights into their driving effects. Collectively, CTDdgv has significant potential to serve as a valuable resource for molecular diagnostics and therapeutic decision-making in cancer via liquid biopsy.
Similar content being viewed by others
Background & Summary
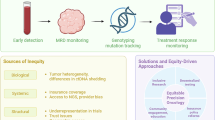
Cancer arises from genetic mutations, DNA damage responses, immune dysfunction, and chronic inflammation, among other factors1,2,3,4. The detection, identification, and interpretation of tumor driver genes (TDGs) and tumor driver variants (TDVs) are fundamental for devising personalized treatment strategies, predicting disease prognosis, and assessing therapeutic efficacy5,6,7. However, traditional approaches for detecting DNA variants rely on patient tissue sampling, which can be invasive and challenging. Liquid biopsy is a minimally invasive approach for circulating tumor DNA (ctDNA) detection, which overcomes many limitations of traditional tissue biopsy, enabling the identification of minimal residual disease, early detection of cancer recurrence, and dynamic monitoring of molecular changes in tumors8,9.
Currently, numerous bioinformatics methods have been developed to identify driver mutations in tissue-based multi-omics data, broadly classified into three main categories. The first category is frequency-based approaches, such as DriverML10, ActiveDriver11, and OncodriveFML12, which identify genes that are mutated more frequently than expected by background mutation rates. The second category is sub-network-based approaches, including tools like DriverNet13 and DawnRank14, which utilize prior knowledge of biological pathways and protein-protein interaction networks to identify driver genes. The third approach focuses on the hotspot mutations region that lie within functional domains or key residues in the three-dimensional structure of proteins, such as MSEA15 and e-Driver16. While these tools have been validated and benchmarked using tissue mutation data from consortia like The Cancer Genome Atlas (TCGA) project17, their efficacy in ctDNA mutation profiles, characterized by lower sensitivity and fewer detectable mutations, remains insufficiently explored.
In addition to the identification of driver genes and mutations, it is equally important to elucidate their biological functions and clinical significance. The concerted efforts of many scientists have advanced the standardization of mutation interpretation, enabling a comprehensive understanding of mutations from multiple perspectives (Table S1). For example, databases such as ClinVar18 and ClinGen19 comprehensively encompass currently reported human cancer and hereditary disease variants. Some drug-related databases like The Gene Drug Knowledge Database20 and PharmGKB21, mine existing resources to illustrate drug-gene/variant associations such as drug sensitivity/resistance, drug response, dosing guidelines, etc, while The Cancer Genome Interpreter22 integrates algorithms into an online platform to annotate the driver potential of alterations and their possible effect on treatment response. More comprehensive databases, like CIVIC23, not only contain therapeutic-related knowledge but also employ a strategy that combines expert review and community input to depict prognostic, diagnostic, and predisposing associations of all types of genetic and somatic variants. However, aside from part resources restricting public access, the main limitation of these resources is that the vast majority of variants were curated from tissue samples, which are not entirely applicable in clinical variant interpretation for patients who cannot undergo tissue sampling or tissue biopsy carries a high risk.
Here, we introduce CTDdgv, a comprehensive resource that provides researchers with an advanced platform for identifying TDGs and TDVs in ctDNA. CTDdgv integrates clinical functional information on ctDNA from thousands of research papers. By simply uploading a ctDNA mutation profile, researchers can efficiently identify genes and variants with cancer-driving potential and obtain clinical interpretations of the genes and variants of interest. Additionally, CTDdgv offers a suite of tools for visualizing driver genes and mutations, as well as exploring their prognostic and driver effects. In summary, CTDdgv is an invaluable resource for the in-depth study of ctDNA. A schematic overview of this study design is shown in Fig. 1.
Content and interface of CTDdgv. (A) Experimentally supported ctDNA variants. (B) Identified tumor driver genes and variants in ctDNA mutation profiles. (C) Searching experimentally supported clinical interpretation. (D) Genome browser for illustrating the genomic landscape of experimentally supported mutations and potential tumor driver variants (TDVs). (E) Downloading ctDNA mutation spectra and curated data. (F) A collection of tools for identifying tumor driver genes (TDGs) and TDVs for tissues or ctDNA mutation spectra, evaluating the prognostic value of driver genes, and explaining the effect of driver gene set from multiple aspects. (G) Searching potential TDGs and TDVs.
Methods
Collection and annotation of experimentally verified ctDNA variants
The associations between ctDNA variants and clinical interpretation were manually curated from the published literature by systematically screening in the PubMed database using the following keyword combinations: (i) cancer and ctDNA mutation/mutated; (ii) ctDNA mutation; (iii) tumor and ctDNA mutation; (iv) cancer and circulating tumor DNA mutation; (v) ctDNA mutation detection. In total, 4,797 pieces of literature were collected as of 8 May 2024. All articles underwent a rigorous review process by at least two researchers to ensure their relevance and accuracy. During the screening process, cfDNA variants that might result from clonal hematopoiesis, inflammatory damage, or release from blood cells were filtered out24. The following information was retrieved from the eligible studies: PubMed ID, gene symbol, variant type (e.g., missense, insertion, deletion, complex rearrangement), variant description (e.g., G1202R, G12C), disease type (e.g., breast cancer and metastatic breast cancer, with cancer subtypes specified in strict accordance with the original publications), body fluid used for ctDNA extraction (e.g., plasma, cerebrospinal fluid), sequencing method, experimental method, evidence of ctDNA variants with drug resistance, association with tumor characteristics (e.g., tumor staging, tumor diameter), prognostic value, marker for cancer metastasis, prospects of substituting tissue detection, impact on treatment (e.g., targeted therapy, immunotherapy), and other clinical interpretation. Here, experimentally supported clinical interpretation records of ctDNA driver genes and driver variants were categorized into two groups: Gene-Level and Variant-Level. The Gene-Level group provides clinical interpretation for overall gene variations while the Variant-Level group focuses on specific variations.
Data expansion of ctDNA variants
To facilitate a more comprehensive study and interpretation of driver genes and their variants, the following enhancements have been implemented: (i) Databases such as GeneCards25, HGNC26, Ensembl27, NCBI GenBank28, CIVIC23, COSMIC29, and Jax-Clinical Knowledgebase30 were linked to acquiring information from external resources, allowing users to retrieve a substantial amount of annotation and functional information. (ii) TransVar31 was used for the conversion of multi-omics mutation coordinates. Users can access detailed information about the DNA strand where the mutation occurs, along with the mutation coordinates across various omics levels, including gDNA, cDNA, and protein. This enables users to conduct a deeper investigation of the variants across multiple omics layers. (iii) CTDdgv offers links to the genome browser in UCSC32 and Ensembl databases27, allowing users to retrieve variant distribution within a 10,000 bp region upstream or downstream of the corresponding mutation site. (iv) For missense mutations, which make up the majority in the Variant-Level, AlphaFold33 and PyMOL (https://pymolwiki.org/) were utilized to predict the optimal protein structures before and after the mutation. Structure visualization and PDB format files were provided to assist users in researching protein macromolecular structure changes due to mutations, thereby aiding the development of targeted drugs and enhancing the understanding of cancer mechanisms.
Design of geMERlb pipeline for identifying TDGs and TDVs in ctDNA
We introduce geMERlb (Genomic Element Mutation Enrichment Research in Liquid Biopsy) to identify tumor driver genes (TDGs) and variants (TDVs) in ctDNA by integrating nonsynonymous somatic mutations of liquid biopsy and the sequence information of genomic elements.
The genomic element set containing the coding sequences (CDS) (n = 20,185), splice sites (n = 18,729), 3′ untranslated regions (3′ UTRs) (n = 19,369), 5′ untranslated regions (5′ UTRs) (n = 19,188), and promoters (n = 20,164) were generated by PCAWG and downloaded from DriverPower34. Their genome coordinates were converted from GENCODE v37 to GENCODE v38 by Liftover to align with somatic mutations annotated in different reference genomes.
geMERlb simulates a walker walking along the sequence of a given genomic element. Mutation Accumulation Score (MAS) represents the cumulative value recorded at each genome position, starting with 0, increasing if patients encounter mutations, and decreasing without a mutation. The increment (\({S}_{{inc}}\)) and decrement (\({S}_{{dec}}\)) of MAS are separately calculated by:
where Y represents a vector recording the number of mutations at each genome position, and L denotes the length of the genomic element sequence, that is, Y = (y1, …, yL). The sum of MAS increments and MAS decrements both amounting to 1. Thus, MAS at the \({i}^{{th}}\) position is calculated by:
where \({L}^{M}\) denotes a vector of positions where mutations occur, and the increment at a mutant position is calculated by \({y}_{j}\times {S}_{{inc}}\), where 1 ≤ j ≤ L and \(j\in {L}^{M}\). The decrease of positions without mutations is consistently \({S}_{{dec}}\). Therefore, MAS bridges of the genomic element sequence are expected to sharply increase over a short distance in a region with mutation enrichment. Mutation Enrichment Score (MES) is defined as the maximum deviation of MAS, representing the location of the mutation enrichment region (MER) and the abundance of mutation enrichment. Taking into account the positional order of the maximum MAS (maxMAS) and minimum MAS (minMAS) within the genomic sequence, we determine MER and MES as follows:
-
1)
MER is identified as the region between the genomic position of minMAS and maxMAS when the genomic position coordinate of maxMAS is greater than minMAS’s. MES is calculated by:
$${MES}={\max }{MAS}-{\min }{MAS}$$Since MAS begins and ends with a value of 0, both the positions of maxMAS and minMAS can’t simultaneously occur at the beginning and end of a genomic sequence. When the genomic position coordinate of maxMAS is less than that of minMAS,
-
2)
If the maxMAS, with a value of 0, is located at the beginning of the genomic sequence, a novel maximum MAS (nvmaxMAS) is always present to the right of the minMAS coordinate. MER is defined as the region between the genomic position of minMAS and nvmaxMAS. MES is calculated by:
$${MES}={nvmaxMAS}-{\min }{MAS}$$ -
3)
If the minMAS, with a value of 0, is located at the end of the genomic sequence, a novel minimum MAS (nvminMAS) will consistently be found to the left of the maxMAS coordinate. MER is defined as the region between the genomic positions of nvminMAS and maxMAS. MES is calculated by:MES is equal to maxMAS minus nvminMAS.
$${MES}={\max }{MAS}-{nvminMAS}$$ -
4)
If minMAS and maxMAS are not located at the beginning or end of the genomic sequence, nvminMAS is always present to the left of the maxMAS coordinate, while nvmaxMAS is always situated to the right of the minMAS coordinate. MES is compared between the maxMAS minus nvminMAS group and the nvmaxMAS minus minMAS group. The MER is defined as the region between the genomic position of minMAS and maxMAS of the group with greater MES. MES is calculated by:
An empirical p-value is employed to assess the MES significance by a randomization-based test. The randomization process is conducted 1000 times. For each randomization, the number of mutations is kept the same as the actual mutation records, while the sites where mutations occur are randomly selected across the sequence, allowing for replacement. The p-value is calculated as:
where \({MES}(\pi )\) is a MES vector of randomly selected mutations in the given genomic element. Benjamini-Hochberg procedure is applied to adjust the p-values. Genes with an adjusted p-value (adj.p) < 0.05 are considered as TDGs for the given element and the variants within MER are defined as TDVs.
Mutation dataset of ctDNA and tissue
We collected 38 ctDNA mutation profiles (5 of which included corresponding survival information) representing 21 cancer types from published literature35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67, 15 datasets containing over 500 mutations were selected to assess the performance of TDGs identification tools (Table S2). We identified 2,305 TDGs and 9,645 TDVs from 38 profiles and organized them into a Prediction-Level dataset. We reclassified and consolidated the 21 cancer types into 17 categories based on their tumor origins. For example, we merged “Non-Small Cell Lung Cancer” and “Lung Cancer” with the same tumor origin under a single Lung Cancer category, and removed “Gynecologic Cancers” due to uncertain tumor origin. In addition, we downloaded somatic mutation data for whole cancer genomes in MAF format across 33 cancer types from The Cancer Genome Atlas (TCGA, https://www.cancer.gov/tcga) and identified TDGs based on tissue mutation profiles. To facilitate comparison with the cancer types in ctDNA datasets, these 33 cancer types were reorganized into 28 categories based on the same classification criteria.
Other datasets
For benchmarking geMERlb, the Cancer Gene Census (CGC) gene list was downloaded from the COSMIC Cancer Gene Census project68, the HiConf gene list was obtained from Erina Takai et al.69, and the experimental supported clinical interpretation (ESCI) gene list comprises all ctDNA driver genes with clinical interpretation curated by us. The non-small cell lung cancer dataset used to demonstrate the application of external data was obtained from Qian Wang et al.62 The breast cancer from Belinda Kingston et al.56 was utilized to explore the driving impact of the driver gene set. Ten hallmark cancer gene sets were obtained from LncACTdb 3.0.70. We collected high-throughput expression profiles across 33 cancer types and corresponding clinical information from the TCGA (https://www.cancer.gov/tcga) to investigate the oncogenic expression patterns and prognostic value of driver genes.
Statistical analysis
We performed gene expression differential analysis using DESeq 2 (v1.38.3). Survival analysis results were calculated using the survival package. Enrichment analysis was performed using clusterProfiler (v4.6.2).
Data Records
All the data files are stored in the Figshare71 and are available under the terms of CC-BY 4.0. The dataset contains a total of 4 datasets, covering the underlying data for the data composition and visualization tools. The Variant.csv and Gene.csv contain the experimentally supported clinical interpretation recorded including PubMed ID, gene symbol, variant type, variant description, disease type, body fluid used for ctDNA extraction, sequencing method, experimental method, evidence of ctDNA variants with drug resistance, association with tumor characteristics, prognostic value, marker for cancer metastasis, prospects of substituting tissue detection, impact on treatment, and other clinical interpretation. The Prediction.csv contains all potential TDGs and TDVs identified from 38 datasets across 17 cancers. The data mainly include disease name, PubMed ID, gene symbol, prognostic value, genomic element, start and end positions of TDV, reference nucleotide, alteration nucleotide, and mutation type. The RawData.csv detailed annotation information for all mutation datasets, including data source, cancer type, number of samples, number of mutations, and survival information.
Technical Validation
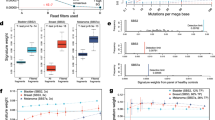
To confirm the accuracy of ctDNA TDGs and TDVs clinical interpretation supported by the experiment, data extraction was independently conducted by the authors and subsequently cross-checked. Discrepancies in the data extraction process were resolved through consensus. Through manual review of 4,797 publicly available studies, we curated a comprehensive set of 1,674 ctDNA-related records with clinically relevant interpretations, covering 238 TDGs and 577 TDVs. We categorized the dataset into 1,170 Variant-Level and 504 Gene-Level records (Table 1). The dataset encompasses 109 cancer types systematically ranked by Variant-Level record counts, as displayed on the database homepage. The top 15 most frequently reported diseases and associated genes are presented in Fig. 2A,B. At the Variant-Level, the majority of records are associated with disease prognosis (n = 439, Table 1) and drug resistance (n = 438, Table 1). At the Gene-Level, prognosis-related records are the most prevalent (n = 331, Table 1). Notably, at the Variant-Level, missense mutations predominate, accounting for 73.93% (n = 865) of all records, with the T790M mutation being the most frequently observed (Fig. 2C,D).
Statistics of clinical interpretation records and schematic of geMERlb pipeline. (A) Counts of clinical interpretation records for the top 15 disease types in the Variation-Level and Gene-Level datasets. (B) Counts of clinical interpretation records for the top 15 genes in the Variation-Level and Gene-Level datasets. (C) Statistics of clinical interpretation records for different variant types in the Variation-Level dataset. (D) Statistics of clinical interpretation records for different Missense mutations. (E) Schematic of the algorithm for identifying mutation enrichment region for genomic element.
We developed a pipeline, geMERlb, specifically designed to identify tumor driver genes (TDGs) and tumor driver variants (TDVs) in ctDNA by detecting mutation enrichment regions across genomic elements (see methods, Fig. 2E). To benchmark geMERlb, we performed a comprehensive performance evaluation by comparing it to two established tools, DriverPower34 and ActiveDriverWGS11, using 15 ctDNA mutation datasets. Following the user manuals of each tool, the same datasets were standardized to the required input formats for geMERlb, DriverPower, and ActiveDriverWGS. All tools were executed with default parameters for a consistent comparison. An adjusted p-value threshold of < 0.05 (i.e., the adjusted False Discovery Rate (FDR) < 0.05) was uniformly applied to identify candidate driver genes. Since the absence of a gold standard (i.e., true cancer driver genes) for evaluating performance with different tools, we employed two benchmarking measures as performance metrics (Table S3). The first benchmark reflects the capacity of tools for capturing well-known cancer-associated genes. The Cancer Gene Census (CGC) is a list of 748 driver genes whose mutations have been causally linked to cancer development72. A higher proportion of CGC genes in predictions is generally regarded as an indicator of better performance in numerous studies10,13. The second benchmark comprises a curated set of 99 high-confidence (HiConf) cancer genes identified by Kumar et al.69 through a comprehensive literature review. The overlaps between the potential driver genes and the CGC or HiConf gene sets were utilized as the gold standard for evaluating candidate ctDNA drivers. In most ctDNA datasets, Driverpower consistently predicted a larger number of driver genes, ActiveDriverWGS was more conservative in predictions, and geMERlb predicted an intermediate number of driver genes (Table S4). For all potential TDGs across 15 datasets, geMERlb identified the highest proportions of driver genes in the CGC gene set (11.23%) and the HiConf gene set (3.22%) (Fig. 3A). Moreover, geMERlb demonstrated superior performance when evaluated across individual datasets, achieving the highest proportions of potential driver genes in the CGC gene set and the HiConf gene set ranking second (Fig. 3B). Additionally, we evaluated the predictive performance of geMERlb on tissue mutation data from TCGA spanning 33 cancer types employing identical benchmarking criteria. The geMERlb demonstrated robust performance across 33 cancer types for both benchmarking metrics (CGC = 4.29%, HiConf = 0.58%; Fig. 3C) and consistently showed the highest proportions in both benchmark sets when assessed on individual datasets (Fig. 3D). Subsequently, we utilized these 238 genes with experimentally supported clinical interpretation (ESCI) as a novel benchmark to evaluate the performance of tools (Table S3). Among the three tools, geMERlb demonstrated the highest proportion of driver genes within the ESCI gene set (5.31%, Fig. 3E). Moreover, geMERlb consistently outperformed the other tools across different datasets (Fig. 3F). Then, we identified 2,305 TDGs and 9,645 distinct TDVs from publicly available publications, encompassing 17 cancer types (Fig. 3G). Retrieval examples of the identified results are provided in the supplementary materials (Method S2). Notably, several driver genes (115/2305) and driver variants (76/9645) have been previously demonstrated in independent studies to be associated with clinical outcomes such as treatment response and prognosis. Additionally, we also identified TDGs across 28 cancer types from TCGA data (Fig. 3H). In conclusion, the comparative analysis underscores the effectiveness of geMERlb in detecting driver genes in liquid biopsy and tissue samples relative to other established methodologies.
Performance evaluation of geMERlb and statistics of the Prediction-Level dataset. (A) The proportions of CGC and HiConf genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across all 15 ctDNA datasets. (B) The proportions of CGC and HiConf genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across 15 individual ctDNA datasets. (C) The proportions of CGC and HiConf genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across all 33 TCGA datasets. (D) The proportions of CGC and HiConf genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across 33 individual TCGA datasets. (E) The proportions of ESCI genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across all 15 ctDNA datasets. (F) The proportions of ESCI genes identified by the geMERlb, ActiveDriverWGS, and DriverPower tools across 15 individual ctDNA datasets. (G) Statistics of TDGs and TDVs identified from 38 ctDNA mutation profiles. (H) Statistics of TDGs identified from the TCGA dataset.
Usage Notes
To our knowledge, CTDdgv (http://bio-bigdata.hrbmu.edu.cn/CTDdgv/) is currently the most comprehensive resource on ctDNA variants. This platform offers researchers user-friendly modules to explore ctDNA TDGs and TDVs, including clinical annotations (Method S1, Fig. S1A–H, Table S5), driver potential predictions (Method S2, Fig. S1I,J), as well as online analysis (Method S3, Fig. S2, Table S6-7) and visualization tools (Method S4, Fig. S3, Table S8). In this study, our core algorithm, geMERlb, by focusing purely on the genomic positions of mutations and their counts in tumor samples, can detect rare mutations with greater sensitivity by calculating the MAS and MES, particularly in the underexplored non-coding regions of the genome. Nevertheless, the pipeline does not include a correction mechanism for undetected mutations in plasma that may fall below the sensitivity threshold. Moreover, while geMERlb has been successfully applied to identify driver genes in large-scale TCGA cohorts, most currently available plasma-derived mutation data originate from targeted sequencing, which typically captures fewer mutations. To address this limitation, we plan to incorporate larger-scale plasma mutation datasets in future research.
Code availability
The code generated during this study is available at https://github.com/yuncwang/CTDdgv.
References
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421, https://doi.org/10.1038/nature12477 (2013).
Pan, M. R., Li, K., Lin, S. Y. & Hung, W. C. Connecting the Dots: From DNA Damage and Repair to Aging. International journal of molecular sciences 17, https://doi.org/10.3390/ijms17050685 (2016).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science (New York, N.Y.) 331, 1565–1570, https://doi.org/10.1126/science.1203486 (2011).
Balkwill, F., Charles, K. A. & Mantovani, A. Smoldering and polarized inflammation in the initiation and promotion of malignant disease. Cancer cell 7, 211–217, https://doi.org/10.1016/j.ccr.2005.02.013 (2005).
Martínez-Jiménez, F. et al. A compendium of mutational cancer driver genes. Nature reviews. Cancer 20, 555–572, https://doi.org/10.1038/s41568-020-0290-x (2020).
Garraway, L. A. & Lander, E. S. Lessons from the cancer genome. Cell 153, 17–37, https://doi.org/10.1016/j.cell.2013.03.002 (2013).
Zhang, J. et al. Identifying driver mutations from sequencing data of heterogeneous tumors in the era of personalized genome sequencing. Briefings in bioinformatics 15, 244–255, https://doi.org/10.1093/bib/bbt042 (2014).
Levy, B. et al. Clinical Utility of Liquid Diagnostic Platforms in Non-Small Cell Lung Cancer. The oncologist 21, 1121–1130, https://doi.org/10.1634/theoncologist.2016-0082 (2016).
Diaz, L. A. Jr. & Bardelli, A. Liquid biopsies: genotyping circulating tumor DNA. Journal of clinical oncology: official journal of the American Society of Clinical Oncology 32, 579–586, https://doi.org/10.1200/jco.2012.45.2011 (2014).
Han, Y. et al. DriverML: a machine learning algorithm for identifying driver genes in cancer sequencing studies. Nucleic acids research 47, e45, https://doi.org/10.1093/nar/gkz096 (2019).
Zhu, H. et al. Candidate Cancer Driver Mutations in Distal Regulatory Elements and Long-Range Chromatin Interaction Networks. Molecular cell 77, 1307–1321.e1310, https://doi.org/10.1016/j.molcel.2019.12.027 (2020).
Mularoni, L., Sabarinathan, R., Deu-Pons, J., Gonzalez-Perez, A. & López-Bigas, N. OncodriveFML: a general framework to identify coding and non-coding regions with cancer driver mutations. Genome biology 17, 128, https://doi.org/10.1186/s13059-016-0994-0 (2016).
Bashashati, A. et al. DriverNet: uncovering the impact of somatic driver mutations on transcriptional networks in cancer. Genome biology 13, R124, https://doi.org/10.1186/gb-2012-13-12-r124 (2012).
Hou, J. P. & Ma, J. DawnRank: discovering personalized driver genes in cancer. Genome medicine 6, 56, https://doi.org/10.1186/s13073-014-0056-8 (2014).
Jia, P. et al. MSEA: detection and quantification of mutation hotspots through mutation set enrichment analysis. Genome biology 15, 489, https://doi.org/10.1186/s13059-014-0489-9 (2014).
Porta-Pardo, E. & Godzik, A. e-Driver: a novel method to identify protein regions driving cancer. Bioinformatics (Oxford, England) 30, 3109–3114, https://doi.org/10.1093/bioinformatics/btu499 (2014).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218, https://doi.org/10.1038/nature12213 (2013).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic acids research 44, D862–868, https://doi.org/10.1093/nar/gkv1222 (2016).
Rehm, H. L. et al. ClinGen–the Clinical Genome Resource. The New England journal of medicine 372, 2235–2242, https://doi.org/10.1056/NEJMsr1406261 (2015).
Dienstmann, R., Jang, I. S., Bot, B., Friend, S. & Guinney, J. Database of genomic biomarkers for cancer drugs and clinical targetability in solid tumors. Cancer discovery 5, 118–123, https://doi.org/10.1158/2159-8290.Cd-14-1118 (2015).
Whirl-Carrillo, M. et al. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clinical pharmacology and therapeutics 110, 563–572, https://doi.org/10.1002/cpt.2350 (2021).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome medicine 10, 25, https://doi.org/10.1186/s13073-018-0531-8 (2018).
Griffith, M. et al. CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer. Nature genetics 49, 170–174, https://doi.org/10.1038/ng.3774 (2017).
Stejskal, P. et al. Circulating tumor nucleic acids: biology, release mechanisms, and clinical relevance. Molecular cancer 22, 15, https://doi.org/10.1186/s12943-022-01710-w (2023).
Safran, M. et al. GeneCards Version 3: the human gene integrator. Database: the journal of biological databases and curation 2010, baq020, https://doi.org/10.1093/database/baq020 (2010).
Yates, B. et al. Genenames.org: the HGNC and VGNC resources in 2017. Nucleic acids research 45, D619–d625, https://doi.org/10.1093/nar/gkw1033 (2017).
Hubbard, T. et al. The Ensembl genome database project. Nucleic acids research 30, 38–41, https://doi.org/10.1093/nar/30.1.38 (2002).
Karsch-Mizrachi, I. & Ouellette, B. F. The GenBank sequence database. Methods of biochemical analysis 43, 45–63 (2001).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic acids research 43, D805–811, https://doi.org/10.1093/nar/gku1075 (2015).
Patterson, S. E. et al. The clinical trial landscape in oncology and connectivity of somatic mutational profiles to targeted therapies. Human genomics 10, 4, https://doi.org/10.1186/s40246-016-0061-7 (2016).
Zhou, W. et al. TransVar: a multilevel variant annotator for precision genomics. Nature methods 12, 1002–1003, https://doi.org/10.1038/nmeth.3622 (2015).
Raney, B. J. et al. The UCSC Genome Browser database: 2024 update. Nucleic acids research 52, D1082–d1088, https://doi.org/10.1093/nar/gkad987 (2024).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589, https://doi.org/10.1038/s41586-021-03819-2 (2021).
Shuai, S., Gallinger, S. & Stein, L. D. Combined burden and functional impact tests for cancer driver discovery using DriverPower. Nature communications 11, 734, https://doi.org/10.1038/s41467-019-13929-1 (2020).
Pereira, E. et al. Personalized Circulating Tumor DNA Biomarkers Dynamically Predict Treatment Response and Survival In Gynecologic Cancers. PloS one 10, e0145754, https://doi.org/10.1371/journal.pone.0145754 (2015).
Zhou, J. et al. Application of Circulating Tumor DNA as a Non-Invasive Tool for Monitoring the Progression of Colorectal Cancer. PloS one 11, e0159708, https://doi.org/10.1371/journal.pone.0159708 (2016).
Ma, C. X. et al. Neratinib Efficacy and Circulating Tumor DNA Detection of HER2 Mutations in HER2 Nonamplified Metastatic Breast Cancer. Clinical cancer research: an official journal of the American Association for Cancer Research 23, 5687–5695, https://doi.org/10.1158/1078-0432.Ccr-17-0900 (2017).
Phallen, J. et al. Direct detection of early-stage cancers using circulating tumor DNA. Science translational medicine 9, https://doi.org/10.1126/scitranslmed.aan2415 (2017).
Beije, N. et al. Somatic mutation detection using various targeted detection assays in paired samples of circulating tumor DNA, primary tumor and metastases from patients undergoing resection of colorectal liver metastases. Molecular oncology 10, 1575–1584, https://doi.org/10.1016/j.molonc.2016.10.001 (2016).
Ng, C. K. Y. et al. Genetic profiling using plasma-derived cell-free DNA in therapy-naïve hepatocellular carcinoma patients: a pilot study. Annals of oncology: official journal of the European Society for Medical Oncology 29, 1286–1291, https://doi.org/10.1093/annonc/mdy083 (2018).
Miller, A. M. et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 565, 654–658, https://doi.org/10.1038/s41586-019-0882-3 (2019).
Peng, M. et al. Resectable lung lesions malignancy assessment and cancer detection by ultra-deep sequencing of targeted gene mutations in plasma cell-free DNA. Journal of medical genetics 56, 647–653, https://doi.org/10.1136/jmedgenet-2018-105825 (2019).
Wang, Y. et al. Circulating tumor DNA analyses predict progressive disease and indicate trastuzumab-resistant mechanism in advanced gastric cancer. EBioMedicine 43, 261–269, https://doi.org/10.1016/j.ebiom.2019.04.003 (2019).
Cai, Z. et al. Comprehensive Liquid Profiling of Circulating Tumor DNA and Protein Biomarkers in Long-Term Follow-Up Patients with Hepatocellular Carcinoma. Clinical cancer research: an official journal of the American Association for Cancer Research 25, 5284–5294, https://doi.org/10.1158/1078-0432.Ccr-18-3477 (2019).
Wei, Q. et al. Clinicopathologic Characteristics of HER2-positive Metastatic Colorectal Cancer and Detection of HER2 in Plasma Circulating Tumor DNA. Clinical colorectal cancer 18, 175–182, https://doi.org/10.1016/j.clcc.2019.05.001 (2019).
Azad, T. D. et al. Circulating Tumor DNA Analysis for Detection of Minimal Residual Disease After Chemoradiotherapy for Localized Esophageal Cancer. Gastroenterology 158, 494–505.e496, https://doi.org/10.1053/j.gastro.2019.10.039 (2020).
Ritch, E. et al. Identification of Hypermutation and Defective Mismatch Repair in ctDNA from Metastatic Prostate Cancer. Clinical cancer research: an official journal of the American Association for Cancer Research 26, 1114–1125, https://doi.org/10.1158/1078-0432.Ccr-19-1623 (2020).
Bacon, J. V. W. et al. Plasma Circulating Tumor DNA and Clonal Hematopoiesis in Metastatic Renal Cell Carcinoma. Clinical genitourinary cancer 18, 322–331.e322, https://doi.org/10.1016/j.clgc.2019.12.018 (2020).
Chabon, J. J. et al. Integrating genomic features for non-invasive early lung cancer detection. Nature 580, 245–251, https://doi.org/10.1038/s41586-020-2140-0 (2020).
Shi, Y. et al. Circulating tumor DNA predicts response in Chinese patients with relapsed or refractory classical hodgkin lymphoma treated with sintilimab. EBioMedicine 54, 102731, https://doi.org/10.1016/j.ebiom.2020.102731 (2020).
Remon, J. et al. Outcomes in oncogenic-addicted advanced NSCLC patients with actionable mutations identified by liquid biopsy genomic profiling using a tagged amplicon-based NGS assay. PloS one 15, e0234302, https://doi.org/10.1371/journal.pone.0234302 (2020).
Rivas-Delgado, A. et al. Mutational Landscape and Tumor Burden Assessed by Cell-free DNA in Diffuse Large B-Cell Lymphoma in a Population-Based Study. Clinical cancer research: an official journal of the American Association for Cancer Research 27, 513–521, https://doi.org/10.1158/1078-0432.Ccr-20-2558 (2021).
Warner, E. et al. BRCA2, ATM, and CDK12 Defects Differentially Shape Prostate Tumor Driver Genomics and Clinical Aggression. Clinical cancer research: an official journal of the American Association for Cancer Research 27, 1650–1662, https://doi.org/10.1158/1078-0432.Ccr-20-3708 (2021).
Vandekerkhove, G. et al. Plasma ctDNA is a tumor tissue surrogate and enables clinical-genomic stratification of metastatic bladder cancer. Nature communications 12, 184, https://doi.org/10.1038/s41467-020-20493-6 (2021).
Zhu, G. et al. Tissue-specific cell-free DNA degradation quantifies circulating tumor DNA burden. Nature communications 12, 2229, https://doi.org/10.1038/s41467-021-22463-y (2021).
Kingston, B. et al. Genomic profile of advanced breast cancer in circulating tumour DNA. Nature communications 12, 2423, https://doi.org/10.1038/s41467-021-22605-2 (2021).
Pereira, B. et al. Cell-free DNA captures tumor heterogeneity and driver alterations in rapid autopsies with pre-treated metastatic cancer. Nature communications 12, 3199, https://doi.org/10.1038/s41467-021-23394-4 (2021).
Liu, T., Yao, Q. & Jin, H. Plasma Circulating Tumor DNA Sequencing Predicts Minimal Residual Disease in Resectable Esophageal Squamous Cell Carcinoma. Frontiers in oncology 11, 616209, https://doi.org/10.3389/fonc.2021.616209 (2021).
Fujii, Y. et al. Identification and monitoring of mutations in circulating cell-free tumor DNA in hepatocellular carcinoma treated with lenvatinib. Journal of experimental & clinical cancer research: CR 40, 215, https://doi.org/10.1186/s13046-021-02016-3 (2021).
Lim, Y. et al. Circulating tumor DNA sequencing in colorectal cancer patients treated with first-line chemotherapy with anti-EGFR. Scientific reports 11, 16333, https://doi.org/10.1038/s41598-021-95345-4 (2021).
Takai, E. et al. Clonal dynamics of circulating tumor DNA during immune checkpoint blockade therapy for melanoma. Cancer science 112, 4748–4757, https://doi.org/10.1111/cas.15088 (2021).
Qiu, B. et al. Dynamic recurrence risk and adjuvant chemotherapy benefit prediction by ctDNA in resected NSCLC. Nature communications 12, 6770, https://doi.org/10.1038/s41467-021-27022-z (2021).
Cabalag, C. S. et al. Potential Clinical Utility of a Targeted Circulating Tumor DNA Assay in Esophageal Adenocarcinoma. Annals of surgery 276, e120–e126, https://doi.org/10.1097/sla.0000000000005177 (2022).
Carvajal, R. D. et al. Clinical and molecular response to tebentafusp in previously treated patients with metastatic uveal melanoma: a phase 2 trial. Nature medicine 28, 2364–2373, https://doi.org/10.1038/s41591-022-02015-7 (2022).
Watanabe, K. et al. Tumor-Informed Approach Improved ctDNA Detection Rate in Resected Pancreatic Cancer. International journal of molecular sciences 23, https://doi.org/10.3390/ijms231911521 (2022).
Wang, Q. et al. RB1 aberrations predict outcomes of immune checkpoint inhibitor combination therapy in NSCLC. Frontiers in oncology 13, 1172728, https://doi.org/10.3389/fonc.2023.1172728 (2023).
Pascual, J. et al. Baseline Mutations and ctDNA Dynamics as Prognostic and Predictive Factors in ER-Positive/HER2-Negative Metastatic Breast Cancer Patients. Clinical cancer research: an official journal of the American Association for Cancer Research 29, 4166–4177, https://doi.org/10.1158/1078-0432.Ccr-23-0956 (2023).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nature reviews. Cancer 18, 696–705, https://doi.org/10.1038/s41568-018-0060-1 (2018).
Kumar, R. D., Searleman, A. C., Swamidass, S. J., Griffith, O. L. & Bose, R. Statistically identifying tumor suppressors and oncogenes from pan-cancer genome-sequencing data. Bioinformatics (Oxford, England) 31, 3561–3568, https://doi.org/10.1093/bioinformatics/btv430 (2015).
Wang, P. et al. LncACTdb 3.0: an updated database of experimentally supported ceRNA interactions and personalized networks contributing to precision medicine. Nucleic acids research 50, D183–d189, https://doi.org/10.1093/nar/gkab1092 (2022).
Wang, Y. CTDdgv: a comprehensive database for the identification and clinical interpretation of ctDNA driver genes and variants in cancer. figshare https://doi.org/10.6084/m9.figshare.28193990.v2 (2025).
Futreal, P. A. et al. A census of human cancer genes. Nature reviews. Cancer 4, 177–183, https://doi.org/10.1038/nrc1299 (2004).
Acknowledgements
This work was supported by the National Natural Science Foundation of China [32170674] and the Heilongjiang Provincial Natural Science Foundation (YQ2024C040).
Author information
Authors and Affiliations
Contributions
H.Z. conceived and led the study; Y.C.W. compiled the application, and performed data analyses; J.G. and Y.C.W. designed the geMERlb pipeline; H.Z., Y.C.W. and J.G. co-wrote the manuscript. H.Z., S.W.N. and P.W. supervised the students and supported funding acquisition. All authors participated in the investigation and data curation; All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Gan, J., Hu, H. et al. A comprehensive database for identifying and interpreting ctDNA driver genes and variants in cancer. Sci Data 12, 1244 (2025). https://doi.org/10.1038/s41597-025-05550-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05550-3