Abstract
Phosphorus (P) and potassium (K) are essential fertilizer elements whose recommendations require soil analysis tailored to crop-specific needs. However, in developing countries like Morocco, access to open databases on nutrient bioavailability remains limited, hindering data-driven agricultural strategies. This paper presents the first national reference maps of available P and exchangeable K at 250 m resolution over Morocco’s croplands using digital soil mapping with machine learning algorithms and environmental covariates. Unlike previous efforts employing traditional interpolation methods, these maps were developed using Random Forest by integrating 5,276 soil samples for P and 6,978 for K with 76 environmental covariates representing climate, topography, vegetation, and parent material. Model validation using independent test datasets demonstrated strong performance, with R2 values of 0.78 for P and 0.80 for K. Uncertainty assessment through bootstrapping confirmed prediction stability across diverse agricultural landscapes. These baseline maps enhance fertilizer recommendations, promote precision farming, and support agricultural sustainability. The maps are freely available through open-access repository, enabling evidence-based decision-making for researchers, practitioners, and policymakers to improve nutrient management efficiency.
Similar content being viewed by others
Background & Summary
Spatially explicit information on available P and exchangeable K is crucial for understanding and managing agricultural soil fertility1,2. These nutrients are fundamental determinants of crop productivity, with phosphorus essential for root development and early crop growth, while potassium plays vital roles in photosynthesis, enzyme activation and plant stress resistance3,4. P and K’s availability and spatial distribution significantly influence fertilizer use efficiency and agricultural sustainability5, especially in semi-arid countries like Morocco where nutrient limitations in soil often constrain crop yields by restricting plant growth and development6.
Despite the recognized importance of P and K in agricultural systems, research efforts worldwide, specifically in Morocco, have primarily focused on soil organic carbon mapping7,8, leaving these essential nutrients comparatively understudied. While global and continental-scale soil mapping initiatives like iSDAsoil9 (https://isda-africa.com/isdasoil, last access Dec-23-2024) provide broad coverage of soil properties across Africa at 30-meter resolution, their predictions for Morocco were generated from a small number of samples, which introduces considerable uncertainty in their local applicability. Previous studies of soil nutrients in Morocco have been restricted mainly to local scales or specific agricultural regions10,11,12,13,14, limiting their utility for national-level planning and decision-making regarding fertilizer recommendations for strategic crops.
The only available cartographic information on soil fertility in Morocco was produced as part of the Fertimap project, which aimed to develop maps of key soil properties including organic matter, pH, phosphorus and potassium, along with fertility recommendation tools. Although these data are accessible via the project platform, their data cannot be downloaded or repurposed for further applications. Additionally, while the maps were generated using standard interpolation techniques, the lack of documented methodologies and model performance metrics hinders the clarity and reproducibility of the findings.
To address these limitations, this study aims to develop the first comprehensive national baseline maps of plant-available phosphorus and exchangeable potassium for Morocco’s croplands using advanced digital soil mapping techniques with machine learning algorithms and environmental covariates. We introduce a new gridded database of available-P (ppm) and exchangeable-K (ppm) at 250-meter spatial resolution covering Morocco’s cropland areas, developed using digital soil mapping techniques and soil databases. Our approach integrates Random Forest machine learning with a comprehensive set of environmental covariates, following specifications aligned with international soil mapping standards and the SCORPAN framework15.
The mapping methodology incorporated a set of environmental covariates, including climate data, terrain attributes, remote sensing indices, and soil parent material, enabling robust predictions across Morocco’s diverse agricultural landscapes. Unlike previous efforts through the Fertimap project that employed traditional interpolation methods16, this study represents the first application of machine learning-based digital soil mapping for comprehensive nutrient assessment at the national scale in Morocco. Particular attention was paid to assessing map accuracy using independent validation data and quantifying prediction uncertainty through ensemble modeling approaches.
These baseline maps represent a significant advance in understanding soil nutrient status across Moroccan cropland areas, providing spatially explicit nutrient information previously unavailable at the national scale. The maps are freely available, facilitating their use by researchers, practitioners, and policymakers for evidence-based decision-making and targeted interventions to improve agricultural productivity and sustainability.
Methods
Soil phosphorus and potassium data
Soil data used in this study were compiled from various sources collected between 2010 and 2022, with approximately 50% of the samples derived from the Fertimap project database and the remaining 50% from complementary datasets compiled through various Moroccan organizations including the Ministry of Agriculture, the National Institute for Agricultural Research, and the Regional Offices for Agricultural Development. In total, we used 5,276 and 6,978 georeferenced topsoil samples (0–30 cm) for available-P and exchangeable-K prediction, respectively (Fig. 1).
Spatial distribution of soil samples used for the prediction of the soil available phosphorus and exchangeable potassium.
For the Fertimap data, we utilized only a subset of the total dataset, knowing that the full dataset comprises approximately 32,000 samples collected under the project. The foundational Fertimap dataset is not publicly accessible, and while soil property maps are published on the Fertimap website (www.fertimap.ma) for visualization purposes, the underlying raster files (such as TIFF format) are not shared, limiting their reusability for further research applications. For this study, we obtained access to a subset of the Fertimap data through formal request from the responsible parties at the National Institute of Agricultural Research. Additional information about the Fertimap project can be found in Bouabid et al.17.
Environmental covariates
The spatial prediction of soil phosphorus and potassium was supported by a set of environmental covariates representing the major soil-forming factors according to the SCORPAN approach15. For the Soil property covariate, we used the soil organic carbon map from (https://github.com/abdelkrim-bsr/SOC_Morocco), details about this data can be found at Bouasria et al.8, with a 250 m resolution. Additionally, long-term Landsat bare earth spectral reflectance (2002–2022) was incorporated for six spectral bands (Blue, Green, Red, NIR, SWIR1, and SWIR2), generated following the methodology described in Demattê et al.18. Climate variables: temperature parameters (annual mean, maximum of warmest month, minimum of coldest month), annual precipitation were obtained from WorldClim version 2.0 database19 (https://www.worldclim.org), reference evapotranspiration (both mean and standard deviation) and the global aridity index were obtained from (https://doi.org/10.6084/m9.figshare.7504448.v5; Zomer et al.20). Hydrological variables included the height above the nearest drainage derived at multiple resolutions (30 m with 100 and 1000 river head threshold cells and 90 m with 1000 threshold cells) from (https://gee-community-catalog.org/projects/hand/; Donchyts et al.21). The global hydrologic curve number dataset (GCN250) provided additional information on runoff potential under dry, average, and wet antecedent conditions were extracted from Jaafar and Ahmad22. Vegetation and land use indicators were derived from multiple remote sensing sources. These included monthly median values of vegetation indices (NDVI, EVI) based on MODIS data (2000–2023) and extracted from (https://doi.org/10.5067/MODIS/MCD43A4.006; Schaaf and Wang23), Sentinel-1 radar backscatter measurements in VV and VH polarization (2016–2017) are available vis its DOI (https://doi.org/10.48436/n2d1v-gqb91; Bauer-Marschallinger et al.24), long-term cropping intensity (2001–2019) (https://doi.org/10.6084/m9.figshare.14099402; Liu et al.25), and the Biodiversity Intactness Index (2017–2020) (https://data.nhm.ac.uk/dataset/bii-bte; Hudson et al.26). Anthropogenic influences were represented through global human modification indices covering 1990–2015, including agricultural stress indicators and human intrusion metrics (https://doi.org/10.5281/zenodo.3963013; Theobald et al.27). Terrain attributes were derived from the MERIT Digital Elevation Model (https://hydro.iis.u-tokyo.ac.jp/~yamadai/MERIT_DEM/; Yamazaki et al.28), comprising slope, aspect, curvature, topographic wetness index, Valley Bottom Flatness, various roughness measures, and geomorphological classifications. Parent material information was incorporated through lithological classes from the Global Lithological Map database v1.1 (https://doi.pangaea.de/10.1594/PANGAEA.788537; Hartmann and Moosdorf29). All the 76 environmental covariates were preprocessed to ensure consistent spatial resolution (250 m) and projection. Continuous variables were resampled using bilinear interpolation, while categorical variables were processed using nearest neighbor resampling. All processing was performed using Google Earth Engine30, ensuring computational efficiency and reproducibility. All covariates were masked using a long-term mean of annual global cropping intensity layer to exclude non-agricultural areas.
Predictive modeling
Figure 2 outlines the methodological flowchart adopted in this study. Available-P and exchangeable-K prediction datasets were partitioned into training (70%) and testing (30%) subsets using a stratified random sampling approach. The stratification was performed by first sorting the data according to available-P and exchangeable-K prediction values and dividing it into 10 equal groups, ensuring that the full range of nutrient values was proportionally represented in both subsets. This approach maintains the statistical distribution of the target variables and minimizes potential sampling bias in model development and validation.
Methodological flowchart.
The 76 environmental covariates were considered potential predictors. Random Forest (RF) was selected as the primary modeling algorithm due to its demonstrated capability in providing high performance across diverse soil science applications, as highlighted by Lamichhane et al.31. Furthermore, Bouasria et al.8 found that RF models outperformed other state-of-the-art algorithms including XGBoost and LightGBM when applied to predict soil organic carbon across Morocco’s diverse landscapes. For feature selection, a Recursive Feature Elimination (RFE) approach was integrated with Random Forest modeling to determine the most influential covariates for each nutrient. The RFE-RF combination has demonstrated superior performance in various soil mapping studies in Morocco, proving effective for nutrient prediction applications8,32,33,34,35.
Random Forest Regression models (RF) were fitted separately for phosphorus and potassium using the ‘ranger’ package in R using the selected covariates. The grid search was performed to optimize four key hyperparameters. The optimization process evaluated combinations of different numbers of trees (100, 500, 1000), numbers of variables per split (2, 3, 4, 5), minimum node sizes (5, 10), and maximum tree depths (5, 10, 15, 20). Model performance under different parameter combinations was assessed using 10-fold cross-validation on the training dataset. The optimal parameter set was selected to minimize the Root Mean Square Error (RMSE). These optimized parameters were then used to develop the final prediction models for both nutrients. Model fitting and prediction were performed under R programming language.
Accuracy assessment and uncertainty analysis
The models for available-P and exchangeable-K were validated using independent test datasets, representing 30% of the total samples. Model performance and predictive accuracy were evaluated based on multiple statistical metrics, including the coefficient of determination (R2), Lin’s concordance correlation coefficient (LCCC), Root Mean Square Error (RMSE), Mean Error (ME), and the Ratio of Performance to Interquartile Range (RPIQ).
Spatial prediction uncertainty was quantified through an ensemble modeling approach using the bootstrapping method. The data splitting process was repeated 25 times, with each iteration using stratified random sampling approach for splitting the dataset into 70% training and 30% testing. The RF model was trained on the respective training subset for each bootstrapping iteration and applied to predict phosphorus and potassium values across all cropland areas. This generated 25 independent prediction maps for each nutrient. The ensemble of predictions was used to calculate based-pixel Prediction Interval Ratio (PIR), defined as the width of the 90% prediction interval (difference between the 95th and 5th percentiles of predictions) divided by the predicted median value (50th percentile)
Generating available-P and exchangeable-K prediction maps
The selected covariates for each parameter and the best RF models were used to generate spatial predictions of available-P (ppm) and exchangeable-K prediction (ppm) at 250 m resolution across Morocco’s cropland areas.
All spatial predictions were processed and stored as GeoTIFF files to maintain the spatial reference system and resolution of the input environmental covariates.
Data Records
Generated maps with the lower and upper interval can be downloaded from the Zenodo repository at https://doi.org/10.5281/zenodo.1507325736. The data was organized as follows. ‘available-P_250m_croplands.tif’: The zip file contains the generated map for the available phosphorus (P2O5 ppm) with a spatial resolution of 250 m and a coordinate system WGS84 (EPSG:4326). ‘exchangeable-K_250m_croplands.tif’: The zip file contains the generated map for the exchangeable potassium (K2O ppm) with a spatial resolution of 250 m and a coordinate system WGS84 (EPSG:4326).
Technical Validation
Prediction accuracy
Using the RFE method, 15 and 50 covariates were selected as important covariates for the prediction of available-P (ppm) and exchangeable-K (ppm), respectively. The variable importance of all selected covariates is presented in Fig. 3 with the spatial distribution of the top 3 features for each parameter, with an important contribution of climatic and topographic data. The predictive accuracy performance of the models for both parameters is given in Table 1 and Fig. 4.
Maps of the top three covariates and the importance of variables for predicting phosphorus and potassium.
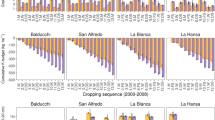
Random Forest model performance for available-P (ppm) and exchangeable-K (ppm) prediction for training and validation using bootstrapping method.
For exchangeable-K, the initial model achieved a high R2 of 0.95 and LCCC of 0.97 on the training dataset, with an RMSE of 34 ppm and MAE of 23 ppm. The model maintained strong performance using the validation dataset, with R2 of 0.80, LCCC of 0.88, and RMSE of 67 ppm. Similarly, the available-P model demonstrated robust performance on both training data (R2 = 0.95, LCCC = 0.97, RMSE = 17 ppm) and test dataset (R2 = 0.78, LCCC = 0.87, RMSE = 35 ppm). Box plots (Fig. 4) also show that model performance metrics remained stable under bootstrapping (25 times repeated resampling), particularly for LCCC and R2, indicating robust model behavior. In addition, the RPIQ values were consistently higher for exchangeable-K compared to available-P across bootstrap iterations, suggesting better relative prediction accuracy for potassium regardless of training data composition.
Spatial mapping and accuracy analysis
Figures 5 and 6 present the spatial distribution of available-P (ppm) and exchangeable-K (ppm) concentrations across Moroccan croplands, and their associated uncertainty. The nutrient concentration maps are classified according to Moroccan agricultural standards13,37, ranging from very low to very high fertility levels. For phosphorus, the classification ranges from very low (<15 ppm) to very high (>100 ppm), while potassium is categorized from very low (<60 ppm) to very high (>300 ppm). The spatial patterns reveal distinct regional variations in nutrient status across Morocco’s agricultural landscapes. The corresponding uncertainty maps display the Prediction Interval Ratio (PIR90%), with consistently low values (generally <0.8 for phosphorus and <0.5 for potassium) across most cropland areas. These low uncertainty values corroborate the robust performance metrics observed during model validation, demonstrating the models’ stability and reliability in predicting both soil nutrients (especially high R2 values). Globally, areas with higher uncertainty values typically correspond to regions with limited sampling density.
Spatial distribution of available-P (ppm) and associated uncertainty (expressed by Prediction Interval Ratio (PIR)).
Spatial distribution of exchangeable-K (ppm) and associated uncertainty (expressed by Prediction Interval Ratio (PIR)).
Limitations and future work
This study presents several limitations that should be acknowledged. The spatial distribution of soil samples, while comprehensive at the national scale, remains uneven across Morocco’s diverse agricultural landscapes, with some regions having lower sampling density than others. This sampling heterogeneity may affect prediction accuracy in undersampled areas, as reflected in the uncertainty maps. Additionally, the temporal inconsistency of input data, spanning collection periods from 2010 to 2022, introduces potential variability that may not fully capture recent changes in soil nutrient status. The reliance on remote sensing covariates of variable quality and temporal resolution further contributes to prediction uncertainty, particularly in areas with complex topography or frequent cloud cover.
Another important limitation is the focus on topsoil layers (0–30 cm), which provides limited information about nutrient distribution in deeper soil horizons that may be relevant for certain crops and management practices. The 250-meter spatial resolution, while appropriate for national-scale applications, may not capture fine-scale variability needed for precision agriculture at the field level.
Future research efforts should address these limitations through expanded sampling campaigns targeting undersampled regions to improve spatial coverage and prediction accuracy. Priority should be given to collecting new samples across diverse agroecological zones and establishing collaborations with other Moroccan research institutes to compile a comprehensive national soil database. The long-term objective is to develop an integrated platform that provides both raw soil data and derived raster products for the research community, subject to data sharing agreements.
Future work will also explore higher spatial resolution products, potentially at 30-meter resolution, to better support precision agriculture applications. This study represents a first attempt to demonstrate the advantages of integrating machine learning with environmental covariates for soil mapping in Morocco, establishing a methodological framework that can be recommended for future soil mapping projects as an alternative to traditional interpolation methods employed in previous studies.
Usage Notes
The generated maps of available phosphorus (ppm) and exchangeable potassium (ppm) across Moroccan croplands, developed at a 250 m resolution, provide an essential foundation for comprehending soil nutrient distribution nation-wide. Being the first maps of their kind, they offer insightful information to promote sustainable agricultural practices, allowing policymakers to make better decisions and farmers to manage fertilizer more precisely. In other terms, these maps can help optimize crop productivity, reduce nutrient imbalances and guide targeted interventions where soil amendments are most needed.
Researchers can also exploit these maps to explore nutrient-crop interactions, assess the impact of soil management practices on long-term soil health, and use them as baseline data for modeling studies in agronomy, soil, hydrology, and environmental sciences. Additionally, these maps represent essential inputs for a wide range of applications, including land degradation assessments, crop yield prediction models, and studies on the impacts of climate change, where soil nutrient data is a crucial factor.
Their utility extends beyond farm-level decision-making to broader agricultural and environmental planning. Specifically, these maps facilitate the design of efficient soil monitoring networks, inform evidence-based agricultural policies, and support sustainable land management strategies. Furthermore, they contribute to national and regional initiatives aimed at enhancing food security, optimizing fertilizer use, and promoting environmental sustainability by enabling data-driven interventions for soil conservation and ecosystem resilience. Furthermore, these maps bridge data gaps and support existing global and African efforts to generate and model soil information7,38.
Code availability
Codes associated with features selection, model development, uncertainty analysis and mapping are freely available from https://doi.org/10.5281/zenodo.1507325736. All analyses were performed in the R programming language (version 4.1.0).
References
Dharumarajan, S., Lalitha, M., Niranjana, K. V. & Hegde, R. Evaluation of digital soil mapping approach for predicting soil fertility parameters—a case study from Karnataka Plateau, India. Arabian Journal of Geosciences 15(5), 386, https://doi.org/10.1007/s12517-022-09629-8 (2022).
Mandal, H. R. et al. Assessment of Soil Fertility Status in Rupani Rural Municipality, Saptari, Nepal. AgroEnvironmental Sustainability 1(2), 111–121, https://doi.org/10.59983/s2023010204 (2023).
Mostofa, M. G. et al. Potassium in plant physiological adaptation to abiotic stresses. Plant Physiology and Biochemistry 186, 279–289, https://doi.org/10.1016/j.plaphy.2022.07.011 (2022).
Zhao, B. et al. Microbe‐dependent and independent nitrogen and phosphate acquisition and regulation in plants. New Phytologist 242(4), 1507–1522, https://doi.org/10.1111/nph.19263 (2024).
Bouwman, A. F. et al. Lessons from temporal and spatial patterns in global use of N and P fertilizer on cropland. Scientific reports 7(1), 40366, https://doi.org/10.1038/srep40366 (2017).
Devkota, K. P., Bouasria, A., Devkota, M. & Nangia, V. Predicting wheat yield gap and its determinants combining remote sensing, machine learning, and survey approaches in rainfed Mediterranean regions of Morocco. Eur. J. Agron. 158, 127195, https://doi.org/10.1016/j.eja.2024.127195 (2024).
Nenkam, A. M. et al. Applications and challenges of digital soil mapping in Africa. Geoderma 449, 117007, https://doi.org/10.1016/j.geoderma.2024.117007 (2024).
Bouasria, A., Bouslihim, Y., Mrabet, R. & Devkota, K. National baseline high-resolution mapping of soil organic carbon in Moroccan cropland areas. Geoderma Regional. https://doi.org/10.1016/j.geodrs.2025.e00941 (2025).
Hengl, T. et al. African soil properties and nutrients mapped at 30 m spatial resolution using two-scale ensemble machine learning. Scientific reports 11(1), 6130, https://doi.org/10.1038/s41598-021-85639-y (2021).
Al Masmoudi, Y., Bouslihim, Y., Doumali, K., Hssaini, L. & Ibno Namr, K. Use of machine learning in Moroccan soil fertility prediction as an alternative to laborious analyses. Modeling Earth Systems and Environment 8(3), 3707–3717, https://doi.org/10.1007/s40808-021-01329-8 (2022).
Eljebri, S., Mounir, M., Faroukh, A. T., Zouahri, A. & Tellal, R. Application of geostatistical methods for the spatial distribution of soils in the irrigated plain of Doukkala, Morocco. Modeling earth systems and environment 5, 669–687, https://doi.org/10.1007/s40808-018-0558-2 (2019).
Ismaili, S., Douik, A. & Moughli, L. Soil fertility mapping: comparison of three spatial interpolation techniques. International Journal of Engineering Research and Technology 3(11), 134–143, https://doi.org/10.17577/IJERTV3IS111043 (2014).
Lahmar, M. et al. GIS and statistical evaluation of soil quality of Sidi Yahya, Gharbplain (Morocco). Journal of Experimental Biology and Agricultural Sciences 9(6), 881–893, https://doi.org/10.18006/2021.9(6).881.893 (2021).
John, K. et al. Mapping soil nutrients via different covariates combinations: theory and an example from Morocco. Ecological Processes 11(1), 23, https://doi.org/10.1186/s13717-022-00368-y (2022).
McBratney, A. B., Santos, M. M. & Minasny, B. On digital soil mapping. Geoderma 117(1-2), 3–52, https://doi.org/10.1016/S0016-7061(03)00223-4 (2003).
Mamassi, A. et al. Relevance of soil fertility spatial databases for parameterizing APSIM-wheat crop model in Moroccan rainfed areas. Agronomy for Sustainable Development 42(5), 83, https://doi.org/10.1007/s13593-022-00813-4 (2022).
Bouabid, R., Badraoui, M., Balaghi, R., Lahlou, M. Soil Resource Information System and its Implications for Land Management in Morocco. Land issues in the Mediterranean Countries. Paris: CIHEAM. April 2014. Watch Letter N° 28 (2014).
Demattê, J. A. M. et al. Bare Earth’s Surface Spectra as a Proxy for Soil Resource Monitoring. Sci Rep 10, 4461, https://doi.org/10.1038/s41598-020-61408-1 (2020).
Fick, S. E. & Hijmans, R. J. WorldClim 2: new 1km spatial resolution climate surfaces for global land areas. International Journal of Climatology 37(12), 4302–4315, https://doi.org/10.1002/joc.5086 (2017).
Zomer, R. J., Xu, J. & Trabuco, A. Version 3 of the Global Aridity Index and Potential Evapotranspiration Database. Scientific Data 9, 409 https://www.nature.com/articles/s41597-022-01493-1 (2022).
Donchyts, G. et al. Global 30m Height Above the Nearest Drainage (HAND). Geophysical Research Abstracts, Vol. 18, EGU2016-17445-3, 2016, EGU General Assembly. https://doi.org/10.13140/RG.2.1.3956.8880 (2016).
Jaafar, H. H. & Ahmad, F. Jaafar, Hadi; Ahmad, Farah GCN250, global curve number datasets for hydrologic modeling and design. figshare. Dataset. https://doi.org/10.6084/m9.figshare.7756202.v1 (2019).
Schaaf, C. & Wang, Z. MCD43A4 MODIS/Terra+ aqua BRDF/albedo nadir BRDF adjusted RefDaily L3 global 500 m V006. NASA EOSDIS Land Processes DAAC, 670, https://doi.org/10.5067/MODIS/MCD43A4.006 Date Accessed: 2024-07-16 (2015).
Bauer-Marschallinger, B. et al. The normalised Sentinel-1 Global Backscatter Model, mapping Earth’s land surface with C-band microwaves. Scientific Data 8(no. 1), 1–18, https://doi.org/10.1038/s41597-021-01059-7 (2021).
Liu, X. et al. Annual dynamic dataset of global cropping intensity from 2001 to 2019. Scientific Data 8(1), 283, https://doi.org/10.1038/s41597-021-01065-9 (2021).
Hudson, L. N. et al. The database of the PREDICTS (projecting responses of ecological diversity in changing terrestrial systems) project. Ecology and evolution 7(1), 145–188, https://doi.org/10.1002/ece3.2579 (2017).
Theobald, D. M. et al. Earth transformed: detailed mapping of global human modification from 1990 to 2017. Earth System Science Data 12(no. 3), 1953–1972, https://doi.org/10.5194/essd-12-1953-2020 (2020).
Yamazaki, D. et al. A high‐accuracy map of global terrain elevations. Geophysical Research Letters 44(11), 5844–5853, https://doi.org/10.1002/2017GL072874 (2017).
Hartmann, J., & Moosdorf, N. The new global lithological map database GLiM: A representation of rock properties at the Earth surface. Geochemistry, Geophysics, Geosystems, 13(12). https://doi.org/10.1029/2012GC004370 (2012).
Gorelick, N. et al. Google Earth Engine: Planetary-scale geospatial analysis for everyone. Remote Sensing of Environment, https://doi.org/10.1016/j.rse.2017.06.031 (2017).
Lamichhane, S., Kumar, L. & Wilson, B. Digital soil mapping algorithms and covariates for soil organic carbon mapping and their implications: A review. Geoderma 352, 395–413, https://doi.org/10.1016/j.geoderma.2019.05.031 (2019).
Bouslihim, Y. et al. Soil Organic Carbon Prediction and Mapping in Morocco Using PRISMA Hyperspectral Imagery and Meta-Learner Model. Remote Sensing 17(8), 1363, https://doi.org/10.3390/rs17081363 (2025a).
Dai, L. et al. Prediction of soil organic carbon fractions in tropical cropland using a regional visible and near-infrared spectral library and machine learning. Soil and Tillage Research 245, 106297, https://doi.org/10.1016/j.still.2024.106297 (2025).
Dahhani, S., Raji, M. & Bouslihim, Y. Synergistic Use of Multi-Temporal Radar and Optical Remote Sensing for Soil Organic Carbon Prediction. Remote Sensing 16(11), 1871, https://doi.org/10.3390/rs16111871 (2024).
Hu, B. et al. A high-resolution map of soil organic carbon in cropland of Southern China. Catena 237, 107813, https://doi.org/10.1016/j.catena.2024.107813 (2024).
Bouslihim, Y. et al. Baseline high-resolution maps of soil nutrients in Morocco to support sustainable agriculture. Data and Scripts https://doi.org/10.5281/zenodo.15073257 (2025b).
DIAEA/DRHA/SEEN Direction de l’irrigation et de l’aménagement de l’espace Agricole, Service des Expérimentations, des Essais et de la Normalisation Rabat (2008)
Arrouays, D. et al. Impressions of digital soil maps: The good, the not so good, and making them ever better. Geoderma Regional 20, e00255, https://doi.org/10.1016/j.geodrs.2020.e00255 (2020).
Acknowledgements
The authors thank the Fertimap project and all institutions that shared the soil data used in this paper. We also thank MCGP, CGIAR/SFP-AoW3 for supporting this publication.
Author information
Authors and Affiliations
Contributions
Y.B.: writing, validation, methodology, formal analysis, data curation, conceptualization. A.B.: writing, software, formal analysis. A.J.: data curation, formal analysis, writing, review, editing. L.K.: writing, review, editing. S.D.: writing, review, editing. R.MR.: data curation, conceptualization, writing, review, editing. R.MO.: data curation, conceptualization, writing, review, editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bouslihim, Y., Bouasria, A., Jelloul, A. et al. Baseline high-resolution maps of soil nutrients in Morocco to support sustainable agriculture. Sci Data 12, 1389 (2025). https://doi.org/10.1038/s41597-025-05699-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05699-x