Abstract
Accurate and early differentiation between bacterial and nonbacterial pharyngitis is crucial for optimizing treatment and minimizing unnecessary antibiotic use. The similar clinical presentation of sore throat in bacterial and nonbacterial infections poses significant diagnostic challenges, even for experienced clinicians. To address this, we developed a publicly available dataset consisting of high-resolution throat images captured using smartphone cameras. These images were analyzed through deep neural networks to distinguish between bacterial and nonbacterial infections based on visual features and symptoms. The dataset is the largest publicly available dataset in this field, which includes images from 742 patients experiencing common cold symptoms. For each patient, it also records the presence or absence of 20 symptoms, age, gender, and between 4 to 9 diagnoses by different physicians. Furthermore, three baseline models were established to differentiate bacterial from nonbacterial infections. Our goal is to enhance the field of non-invasive and accurate pharyngitis diagnosis, drive the development of AI-driven diagnostic tools, promote remote healthcare solutions, and inspire future innovations in medical image analysis.
Similar content being viewed by others
Background & Summary
Pharyngitis, commonly referred to as sore throat, is an inflammation of the pharynx that primarily affects the ear, nose, and throat (ENT) system, impacting millions of individuals annually1. This condition can manifest as nonbacterial pharyngitis, often caused by common viruses such as adenoviruses and rhinoviruses, or as bacterial pharyngitis, most commonly attributed to Group A Streptococcus bacteria2. Accurate differentiation between these two forms is critical, as bacterial infections typically require antibiotic therapy, whereas nonbacterial infections do not3.
The standard diagnostic process for pharyngitis relies on a combination of patient history, physical examination, laboratory tests including rapid antigen detection tests (RADTS), and throat cultures4. However, these methods can be time-consuming and may not provide immediate results5, delaying appropriate treatment. Accurate identification of bacterial versus nonbacterial infections is essential for determining whether antibiotics are necessary. In cases of bacterial infection, timely antibiotic administration is crucial to avoid complications5,6. Conversely, the misdiagnosis of nonbacterial infections and the resulting unnecessary use of antibiotics contribute significantly to the global issue of antibiotic resistance7, a growing public health crisis that threatens the effectiveness of these life-saving drugs8.
In response to the need for more accurate and rapid pharyngitis diagnoses, we developed the PGUPharyngitis dataset, comprising 742 high-resolution images of patients’ throats. Each case in the dataset was meticulously annotated with insights from four to nine physicians who reviewed and labeled the records based on the images and symptoms to ensure the highest diagnostic accuracy. The images were captured using two smartphones, the Samsung Galaxy S21 Ultra and the Xiaomi Redmi 8, selected for their ability to produce clear and detailed visuals. Figure 1 presents two samples from the dataset, one being bacterial and the other nonbacterial.
(a) Example of a nonbacterial throat image from the dataset (b) Example of a bacterial throat image from the dataset.
Data collection was conducted in two Iranian cities with distinct climatic conditions—one with a colder, mountainous climate and the other with a warm, humid, and coastal environment—to ensure the dataset represents a diverse range of patients from varied geographic and environmental backgrounds. The data was collected from October 2023 to May 2024. This dataset aims to facilitate the training of advanced deep learning models designed to improve the accuracy and reliability of pharyngitis diagnosis.
By providing a rich and diverse collection of throat images, the PGUPharyngitis dataset is the largest publicly available dataset in this field, and addresses a critical gap in existing medical data resources, offering researchers and healthcare professionals the opportunity to develop and refine intelligent diagnostic tools. These tools can operate effectively in various clinical settings and support the integration of machine learning techniques into healthcare.
The availability of this dataset is a crucial step toward advancing the field of automated diagnostics and tackling the challenges posed by misdiagnosis and unnecessary antibiotic use. Accurately diagnosing a condition can be difficult, even for experienced physicians. In fact, within our dataset, there were cases where physicians disagreed on the diagnosis for a single patient. Our goal is to substantially enhance healthcare outcomes, particularly in regions with limited access to specialized medical facilities. Since the images are captured using smartphones, the dataset can support the integration of deep learning models into smartphone applications. This enables rapid and accurate diagnostics, empowering patients to actively manage their health and reducing the risk of complications caused by misdiagnoses or the overuse of antibiotics9.
The work by Yoo et al.10 shares a similar goal with our study, aiming to leverage smartphone-captured throat images and deep learning for pharyngitis diagnosis. Their dataset included 131 pharyngitis images and 208 normal throat images, collected from online sources and augmented using CycleGAN. However, the reliance on web-sourced images raises concerns regarding the dataset’s representativeness and clinical validation. Additionally, synthetic augmentation may not fully reflect real-world variability in throat presentations. In contrast, our dataset provides significant advantages. It consists of 742 high-resolution images gathered directly from diverse geographical and clinical settings, accompanied by detailed demographic and symptom data. These attributes enhance its diversity, applicability, and reliability for real-world clinical applications, supporting the development of robust AI-driven diagnostic models designed for practical healthcare scenarios.
Methods
Data Collection
The data for this study were meticulously gathered during patient visits to general practitioners across two geographically distinct regions of Iran. One region, located in the mountainous areas, experiences a cold climate, while the other, situated near the coast of Persian Gulf, is characterized by a hot and humid environment. These contrasting climatic conditions provided a unique opportunity to collect data from a diverse range of environmental contexts, enriching the dataset and enhancing its applicability across various population segments. Data collection occurred from October 2023 to May 2024, focusing on patients with common cold related symptoms. Participation was entirely voluntary, with patients being fully informed about the research objectives and potential outcomes. Patients provided informed written consent, after being informed by the attending physician of the study’s purpose, voluntary nature, and confidentiality measures, including permission for anonymized clinical data and images to be used in current and future research and publications internationally. Ethical approval was obtained prior to the study (research ethics committee Approval ID: IR.BPUMS.REC.1403.282), and informed consent was secured from all participants.
All collected data, including throat images and associated demographic information, were anonymized and used solely for research purposes to advance the development of deep learning diagnostics. The risk of participant identification was carefully considered. All personally identifiable information, including names, dates of birth, contact details, and any other direct identifiers, was removed from the dataset. Each participant was assigned a unique code to replace identifying information. Additionally, facial features in medical images (if present) were obscured or cropped where applicable.
Image Processing
Throat images were captured using two smartphone models—Samsung Galaxy S21 Ultra and Xiaomi Redmi 8 Pro—selected for their high-quality cameras. Using two smartphone models introduced natural variations in image quality, enriching the dataset. Additionally, the differing lighting conditions in the two cities further contributed to the robustness of the dataset. Each throat image was taken in a well-lit room, utilizing the smartphones’ flashlight function to ensure clear visibility of the throat area. The camera was positioned directly in front of the patient’s open mouth, focusing on the back of the throat to capture the most relevant region for pharyngitis diagnosis.
Alongside the images, key demographic and clinical data were recorded, including the patient’s age, gender, and symptoms. These additional data points allowed for a more comprehensive analysis and facilitated the exploration of potential correlations between demographic characteristics and the different types of pharyngitis.
Following the collection, the images underwent a rigorous quality control process. Each image was carefully reviewed for clarity, with blurred or poorly lit images excluded from the dataset. Misaligned images were manually corrected through rotation and cropping to ensure uniformity. The throat area was emphasized to focus on the regions most indicative of pharyngitis. The final images in the repository are cropped sections of the original images and were not resized. Therefore, they each have varying resolutions. After this initial review, a second round of quality control was performed to ensure the dataset’s reliability and uniformity. Ultimately, images that at least 3 physicians agreed were not suitable for diagnosis have been excluded from the dataset, and 742 high-quality images were selected from the original 860 collected.
Diagnostic Process
Upon finalizing the dataset, a standardized diagnostic process was used to classify each image based on the type of pharyngitis. This classification was performed by a team of experienced physicians. On average, each image was reviewed by six physicians, although the number of reviewers varied, with some images being assessed by as few as four and others by as many as nine experts. The goal was to ensure a reliable and accurate classification by leveraging the collective expertise of the physician team. One doctor made the diagnosis by examining the patient’s throat in person, while the others based their diagnoses solely on images.
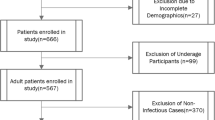
Pharyngitis is typically classified into two main categories: bacterial and nonbacterial (e.g., viral, allergic, fungal, and normal). Similarly, our records are divided into these two categories: bacterial and nonbacterial. Since there have been no cases of fungal pharyngitis in this dataset, our nonbacterial category includes only viral, allergic, and normal cases. In general, the treatment for all these cases is supportive care. A majority vote among the physicians determined the final classification for each image. In cases where significant discrepancies in diagnosis were observed, the image was reassessed by an additional physician who provided an independent evaluation. This step helped resolve any inconsistencies and ensured a high degree of accuracy in the final dataset. The overall workflow and summary of the dataset creation process are presented in Fig. 2 and Table 1.
The overall data acquisition and quality assessment workflow used in this work.
Data Records
The PGUPharyngitis dataset is a publicly accessible collection containing clinical and demographic data alongside diagnostic results from 742 patients who presented with pharyngitis symptoms. This dataset is instrumental for training and evaluating machine learning models aimed at differentiating between bacterial and nonbacterial pharyngitis. Each entry comprises comprehensive symptom indicators, demographic details, and verified diagnostic outcomes.
Data Format
The dataset is organized as a structured Excel file, where each row represents an individual patient. The columns are arranged to include patients’ age, gender, symptoms, and diagnoses. Table 2 describes fields in the dataset.
Symptom Indicators and Diagnostic Data
This dataset includes 20 binary symptom indicators that capture common clinical features of pharyngitis, such as sore throat, headache, and fever, outlined in Table 2. Each symptom is represented by a binary value, where 1 indicates the presence and 0 is the absence of the symptom. Additionally, 9 columns in each patient record contain the diagnostic results, where each patient might have between 4 to 9 diagnoses by different physicians.
Quality Control and Data Validation
To ensure data integrity, the PGUPharyngitis dataset underwent rigorous quality control and validation processes, and records missing essential information were excluded from the final dataset. Diagnostic results were confirmed via reviews conducted by multiple physicians, resulting in high reliability of the diagnostic data.
Technical Validation
The dataset’s statistical breakdown is detailed in Figs. 3 and 4. As illustrated in Fig. 3a, the most common symptoms among patients are sore throat, cough, and rhinorrhea, affecting 19.8%, 14.7%, and 12.1% of the sample, respectively. Figure 3b,d validate that the dataset is balanced in terms of age and gender distribution, as it includes a broad range of age groups, with nearly equal representation of male and female participants (51.6% and 48.4%, respectively).
Statistical analysis of the dataset (a) Distribution and prevalence of symptoms, each patient can have up to 20 symptoms (b) Distribution of age groups in the dataset and the number of patients in each age group (c) Percentage and total number of bacterial and nonbacterial diagnoses (d) Percentage and number of male and female participants. (e) Distribution of the number of diagnoses per patient.
Symptoms co-occurrence heatmap.
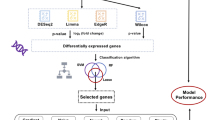
Moreover, to validate and evaluate the dataset, as well as to demonstrate a sample use case, we conducted experiments using four different models: DenseNet12111 Swin Tiny12, MobileNet V3 Small13, and ConvNeXt Small14. These models were used to develop a binary image classifier (i.e., classifying the images into bacterial or nonbacterial). The inputs of the models are images of size 224*224, and each image in the dataset has a different size, they are resized in the training process to 224*224 as well. We trained each model on four different subsets of the dataset: the full dataset, and individual subsets corresponding to the warm city, cold city, Redmi phone, and Samsung phone images.
The results of these experiments are shown in Table 3. To mitigate any potential bias due to data splitting, we applied 3-fold cross-validation. K-Fold cross-validation (K = 3) involves dividing the dataset into K equal subsets (folds). The model is trained K times, each time using K-1 folds for training and the remaining fold for validation. This process ensures that every data point is used for both training and validation exactly once. It reduces the risk of performance variance due to a particular train and validation split and provides a more generalized estimate of the model’s accuracy.
The experiments used a batch size of 20, the Adam optimizer15, and data augmentation techniques including random rotation, horizontal flip, and random affine transformations. The results are presented across five metrics: accuracy, precision, recall, F1-score, and AUC. While using all of the dataset to train the model, MobileNet outperforms the other models in all metrics except for AUC, achieving an accuracy of 80.50% ± 12.79%, a precision of 69.30% ± 21.99%, a recall of 55.26% ± 1.75%, an F1-score of 54.26% ± 1.81%, and an AUC of 0.554 ± 0.073. Although these models show satisfactory performance, incorporating the symptoms, exploring other state-of-the-art models, and developing new architectures to improve metrics remains an ongoing challenge.
Usage Notes
The dataset is openly available on Figshare (https://doi.org/10.6084/m9.figshare.28163513)16. The root directory contains a folder with all the images, each labeled with a unique identifier. Additionally, an Excel file is included, listing each patient’s symptoms, age, gender, and diagnosis.
Images in this dataset were obtained using smartphone cameras under carefully managed yet varied lighting conditions to enhance the visibility of the throat area. Variations in smartphone models, handling, lighting, and patient positioning may affect image color, brightness, and clarity. For instance, blood pressure and oxygenation can affect the color of the mucous membranes. To mitigate the potential effects of the aforementioned factors on the visibility of redness or inflammation, histogram matching during the preprocessing stage is recommended to standardize brightness and color. Additionally, data augmentation techniques, like minor rotations or lighting adjustments, are recommended to adapt the model to real-world conditions.
In some images, the focus is on the lips rather on the throat. As one of the main goals of this dataset is to support the development of remote healthcare solutions and mobile applications, particularly for use in underdeveloped regions. Therefore, in such scenarios, users are likely to capture throat images themselves using mobile devices, which can often result in suboptimal focus. Including these images ensures that AI models trained on the dataset are robust and better equipped to handle real-world usage conditions. Future researchers are free to exclude these images if their specific use cases require only high-quality, well-focused data.
The collected dataset reflects the actual distribution of symptoms and patients in the cities where the data was gathered. Since the real-world distribution of symptoms among patients is imbalanced, this imbalance is also present in the dataset. This point has been added to the manuscript. Therefore, future researchers using this dataset are encouraged to apply methods that address the effects of imbalanced data.
To complement the images, essential contextual information is included, such as the patient’s age, gender, and presence or absence of 20 types of symptoms. This supplementary data provides insights that can significantly enhance diagnostic accuracy and model performance. Incorporating this information allows researchers to gain an understanding of the diagnostic results and refine models to better address real-life scenarios.
Future users of the data are encouraged to use methods that address label uncertainty such as stochastic and probabilistic methods. Among the samples, 182 images had full consensus among all doctors, with identical diagnostic opinions. This subset can serve as a “gold standard” for model evaluation. Future researchers can use these 182 samples as test data, while using the remaining data (potentially with probabilistic or self-supervised methods) for training the model.
While the dataset includes a wide range of bacterial and nonbacterial pharyngitis cases, some limitations exist. Rare forms of pharyngitis, such as those due to fungal infections, are underrepresented due to their rarity. Additionally, cases of chronic pharyngitis, which may present distinct clinical characteristics compared to acute cases, are not included.
Code availability
The Python code for the three models for the classification of pharyngitis into bacterial or nonbacterial categories is available at: https://github.com/habibrostami/basepharyngitis.
References
Mustafa, Z. & Ghaffari, M. Diagnostic methods, clinical guidelines, and antibiotic treatment for group a streptococcal pharyngitis. Frontiers in Cellular and Infection Microbiology 10, 563627, https://doi.org/10.3389/fcimb.2020.563627 (2020).
Gerber, M. A. et al. Prevention of rheumatic fever and diagnosis and treatment of acute streptococcal pharyngitis: a scientific statement from the american heart association rheumatic fever, endocarditis, and kawasaki disease committee. Circulation 119, 1541–1551, https://doi.org/10.1161/CIRCULATIONAHA.109.191959 (2009).
Gerber, M. A. Diagnosis and treatment of pharyngitis in children. Pediatric Clinics of North America 52, 729–747, https://doi.org/10.1016/j.pcl.2005.02.004 (2005).
Ebell, M. H., Smith, M. A., Barry, H. C., Ives, K. & Carey, M. The rational clinical examination: Does this patient have strep throat? JAMA 284, 2912–2918, https://doi.org/10.1001/jama.284.22.2912 (2000).
Cohen, J.-F. et al. Accuracy of rapid antigen detection tests for group a streptococcal pharyngitis: a systematic review and meta-analysis. Clinical Infectious Diseases 63, 487–496, https://doi.org/10.1093/cid/ciw418 (2016).
NICE Evidence Review. Suspected sepsis: recognition, diagnosis and early management: Evidence review b https://www.ncbi.nlm.nih.gov/books/NBK602484/bin/niceng51er2-appf-et2.pdf (2024).
Centor, R. et al. Antibiotic stewardship for patients with pharyngitis: Implementing the centor criteria. The Journal of Infectious Diseases 220, 12–17, https://doi.org/10.1093/infdis/jiz051 (2019).
Murray, C. J. et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet 399, 629–655, https://doi.org/10.1016/S0140-6736(21)02724-0 (2022).
Jiang, F. et al. Deep learning for mobile health applications: A systematic review. Journal of Healthcare Engineering 2017, 1–8, https://doi.org/10.1155/2017/1464158 (2017).
Yoo, T. K., Choi, J. Y., Jang, Y., Oh, E. & Ryu, I. H. Toward automated severe pharyngitis detection with smartphone camera using deep learning networks. Computers in Biology and Medicine 125, 103980 https://doi.org/10.1016/j.compbiomed.2020.103980 (2020).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2261–2269, https://doi.org/10.1109/CVPR.2017.243 (2017).
Liu, Z. et al. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. 2021 IEEE/CVF International Conference on Computer Vision (ICCV), Montreal, QC, Canada, pp. 9992-10002, https://doi.org/10.1109/ICCV48922.2021.00986 (2021).
Howard, A. et al. Searching for MobileNetV3. 2019 IEEE/CVF International Conference on Computer Vision (ICCV). Seoul, Korea (South), pp. 1314-1324, https://doi.org/10.1109/ICCV.2019.00140 (2019)
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. In International Conference on Learning Representations (ICLR 2015), Poster Session. Retrieved from http://arxiv.org/abs/1412.6980 (2015).
Liu, Z. et al. A ConvNet for the 2020s. 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). New Orleans, LA, USA, pp. 11966-11976, https://doi.org/10.1109/CVPR52688.2022.01167 (2022)
Shojaei, N. et al. A publicly available pharyngitis dataset and baseline evaluations for bacterial or nonbacterial classification Figshare https://doi.org/10.6084/m9.figshare.28163513 (2025).
Acknowledgements
The authors would like to express their gratitude to the following physicians for their invaluable efforts in diagnoses and labeling the data: Mina Mirzad, Yasaman Sharifian, Bardiya Hosseini, Zeinab Najar, Mahsa Farhadiyan, Fatemeh Hosseini, Arefeh Ahmadi, Nastaran Shaban, and Mehdi Zandavi.
Author information
Authors and Affiliations
Contributions
Negar Shojaei performed data collection, investigation, preprocessing, and wrote the original draft; Habib Rostami conceptualized, designed, supervised, evaluated, and administrated the project, implemented the deep learning algorithms, and provided suggestions for manuscript improvement; Mohammad Barzegar conducted technical validation, wrote the respective section in the paper, and reviewed and edited the final manuscript; Shokooh Saadat Farzaneh, Majid Alimohammadi, and Jahanbakhsh Keyvani provided medical diagnoses and provisioned access to relevant patients; Zohreh Farrar reviewed and verified the diagnoses and labels; Mehdi Mirzad collaborated in writing the initial draft; Leila Gonbadi performed visualizations.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shojaei, N., Rostami, H., Barzegar, M. et al. A publicly available pharyngitis dataset and baseline evaluations for bacterial or nonbacterial classification. Sci Data 12, 1418 (2025). https://doi.org/10.1038/s41597-025-05780-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05780-5