Abstract
Fungi are closely associated with various diseases, among which Candida albicans (C. albicans) is recognized as an important pathogen in inflammatory bowel disease (IBD). Fungal pathogenicity is primarily mediated by virulence factors (VFs); therefore, comprehensive identification of fungal virulence factors is critical for targeted drug development and disease treatment. However, current databases contain limited numbers of fungal VFs, lack effective predictive algorithms, and do not directly provide protein structural information relevant for drug discovery. In this study, we constructed a positive dataset comprising 18,072 fungal VFs. Utilizing machine learning approaches, we further predicted and identified 390 potential VFs from 8,081 representative protein sequences across the proteomes of 99 C. albicans strains, generating a dedicated C. albicans VF dataset. Additionally, five IBD-associated pathogenic VFs were identified, and their protein structural data included in the dataset were leveraged to facilitate small-molecule compound screening. Collectively, this study provides a comprehensive data resource and theoretical foundation for the identification of fungal VFs and the development of related therapeutics.
Similar content being viewed by others
Background & Summary
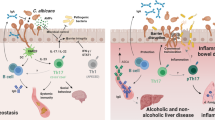
Fungi, as ubiquitous eukaryotic organisms, encompass an estimated five million species1. While the majority of fungi are non-pathogenic and some can colonize the gut of healthy individuals2, nearly three hundred species are known to pose threats to human health3. Fungal infections are associated with considerable morbidity and mortality, particularly among immunocompromised populations. Among pathogenic fungi, Candida albicans (C. albicans) stands out as a common opportunistic pathogen intricately involved in the onset and progression of diverse diseases, with an especially noteworthy role in inflammatory bowel disease (IBD). Emerging evidence suggests that C. albicans is a key pathogenic microorganism in IBD, characterized by increased abundance in affected individuals and a strong association with impaired intestinal barrier function and exacerbated inflammation.
The pathogenicity of fungi principally relies on a repertoire of virulence factors (VFs) that facilitate adhesion, invasion, immune evasion, stress adaptation, and morphological transitions. These properties enable fungi to survive within host environments, circumvent immune responses, and precipitate tissue damage or systemic inflammatory events4,5. Fungal VFs are therefore not only central to understanding disease mechanisms, but also represent promising targets for the development of novel antifungal therapeutics, especially in light of the significant clinical challenges posed by the limited arsenal of antifungal drugs and the rising prevalence of drug resistance6. This underscores the urgent need for new molecular targets and therapeutic agents.
Conventional methodologies for identifying VFs rely on protein isolation, purification, and characterization, which are often time-consuming and labor-intensive. The advent of artificial intelligence has revolutionized this domain, with machine learning approaches facilitating the discovery of VFs across microbial proteomes. Previous efforts have yielded a variety of predictive models tailored to bacteria and viruses, such as VirulentPred7, MP38, DeepVF9, HyperVR10, VFNet11, PreVFs-RG12, ViPal13, yet dedicated models for fungal VF prediction remain scarce.
The successful application of machine learning to fungal VF prediction is contingent on access to accurate and comprehensive datasets. Existing resources, such as Victors14, PHI-base15, UniProt16, and DFVF17, offer valuable collections but exhibit limitations in terms of coverage, sequence annotation standardization, and the availability of protein structure information. The construction of a more inclusive, fungus-specific VF dataset, enriched with protein structural data, is imperative for advancing both predictive modeling and structure-based drug discovery that targets fungal VFs.
In this study, we present a comprehensive dataset encompassing fungal VFs, including a dedicated collection for C. albicans generated via machine learning approaches and a subset of IBD-associated VFs. This resource establishes a robust foundation for accurate identification of fungal VFs and provides a theoretical basis for targeted therapy and antifungal drug development for IBD.
Methods
Construction of the positive VF dataset
Fungal VF sequences were downloaded from the DFVF17 (http://sysbio.unl.edu/DFVF), PHI-base15 (http://www.phi-base.org), Victors14 (http://www.phidias.us/victors) databases. Simultaneously, sequences were retrieved from UniProt16 (https://www.uniprot.org) using the keywords “fungi AND virulence” retaining only manually reviewed entries. All sequences acquired from these four databases were merged based on UniProt IDs and complete sequence identity, resulting in 5,812 non-redundant fungal VF protein sequences. To minimize the influence of sequence length variability, sequences were filtered to retain only those between 200 and 900 amino acids, yielding 4,426 sequences.
To further enhance the VF dataset, non-redundant fungal VF sequences from the combined databases were compared with unreviewed sequences in UniProt16, applying a threshold of e-value < 1e-5 and identity > 62% to identify putative VFs. Ultimately, a positive dataset consisting of 18,072 VF sequences was established for model training and testing.
Construction of the negative dataset
The primary source for the negative dataset was the UniProt16 database. Reviewed fungal protein sequences were collected, and fungal VF sequences were excluded based on UniProt ID, resulting in a pool of non-virulence fungal protein sequences. To enhance the model’s ability to identify features beyond sequence similarity, the non-virulence sequences were aligned with the positive dataset and any matches were removed, producing a negative dataset of 18,131 non-VF sequences, matching the scale of the positive dataset.
Additionally, to better simulate real-world predictive scenarios, a validation set was constructed by randomly sampling fungal non-virulence proteins, resulting in 18,173 negative sequences, comparable in number to the positive dataset.
Integration of structural information
For each VF in the dataset, predicted or experimentally resolved three-dimensional protein structure data were collected from the publicly available AlphaFold.
Protein Structure Database18. Predicted AlphaFold models were linked via UniProt accession mapping.
Construction of the C. albicans VF prediction dataset
Genomic, proteomic, and annotation data of C. albicans were obtained from GenBank19, Ensembl20, and the NCBI database. Orthofinder was used to generate orthologous protein groups among C. albicans strains. Based on their presence across strains, these proteins were classified as core proteins (present in over 90% of strains), unique proteins (present in fewer than 5% of strains), or accessory proteins (present in 5–95% of strains). The longest sequence within each orthologous group was extracted as its representative. These representative sequences were then subjected to machine learning-based prediction to identify potential VFs.
Functional annotation was performed using emapper with eggNOG as the reference database. KEGG and GO enrichment analysis of core VFs was conducted using the R package clusterProfiler, with all proteins as the background, and FDR-corrected p-value < 0.05 set as the threshold for statistical significance.
Prediction of IBD-associated C. albicans VFs
Metagenomic data (project number PRJNA38928021), comprising samples from 282 IBD patients and 66 healthy individuals, were obtained from the NCBI SRA database. After quality control and removal of low-quality reads, paired-end sequence files were merged and assembled with SOAPdenovo2. Fungal sequences were identified using EukRep, and GeneMark was used to predict ORFs within contigs, from which corresponding gene and protein sequences were extracted.
Within each sample, BLAT (BLAST-like Alignment Tool) was employed to identify clusters of highly similar gene sequences (identity > 95%), thus generating non-redundant gene sets. These non-redundant genes were compared with the 390 predicted C. albicans VFs and their abundance was calculated. Relative abundance differences of these VFs were evaluated between healthy individuals and IBD patients.
Molecular Docking
The structures of 9,637 small molecules in SDF format were downloaded from the DrugBank22 database (https://go.drugbank.com/). The protein structure of GeneID_003944-T1 was predicted using AlphaFold2. Each small molecule was individually docked to this protein structure using Autodock4. The parameters were set as ga_num_evals = 25,000,000, ga_run = 100, ga_pop_size = 300, and sw_max_its = 3,000. The result with the lowest binding energy was selected.
Data Record
The datasets are available in the Figshare online repository under an open-access license23.
Fungal VF positive dataset
This dataset includes sequences collected directly from VF databases, as well as an expanded set of 18,072 non-redundant fungal VF protein sequences (in FASTA format) for use in machine learning applications. Each sequence is annotated with the corresponding source database: DFVF17, Victors14, PHI-base15, and reviewed entries from UniProt16, the database-specific ID, the converted UniProt ID, and the corresponding AlphaFold Protein Structure Database protein structure ID. Of these, 16,464 VFs have either available or predicted protein structures retrievable from public databases.
The folder ‘Database_raw’ contains the 5,812 unprocessed VF sequences collected from the original databases and 4,426 sequences that have been filtered by length.
The file ‘Reference_data/positive-18072.fa’ contains the 18,072 protein sequences from the positive dataset of fungal VFs.
Metadata for the VFs is recorded in ‘Reference_data/positive_VF_meta.xlsx’.
Negative dataset
Negative dataset 1 contains 18,131 non-VF protein sequences selected to ensure no overlap with the positive set, available in ‘Reference_data/negative-18131.fa’.
Negative dataset 2 includes 18,173 randomly selected non-VF sequences, used for model validation, and is stored in ‘Reference_data/negative-18173.fa’.
C. albicans VF dataset
The predicted C. albicans VF dataset comprises 390 protein sequences, available in ‘Candida albicans/vf_family-390.fa’.
VF metadata for C. albicans is provided in ‘Candida albicans/candida_VF_meta.xlsx’.
Five protein sequences, along with corresponding structural information significantly associated with IBD, are provided in the folder ‘Candida albicans/IBD’.
Technical Validation
Sequence features of the fungal VF datasets
An analysis of VF sequences collected from DFVF17, Victors14, PHI-base15, and UniProt16 via UniProt ID mapping revealed that most VFs appear in only one database, with only a small subset shared among two or more databases (Fig. 1a). Additionally, sequence length statistics showed significant differences in maximum, minimum, and average lengths across the four databases (Fig. 2b). These findings underscore the necessity of integrating and jointly analyzing fungal VFs from diverse databases to construct a comprehensive resource.
Source and sequence characteristics of the datasets. (a) Overlapping and unique fungal VFs among four databases. (b) Sequence length statistics of entries from each database. (c) Sequence length distributions of raw VF and non-VF entries collected from databases. (d) Sequence length distributions of VFs and non-VFs used for model training and prediction.
Construction and performance evaluation of machine learning models. (a) Overview of the prediction workflow, including feature extraction from protein sequences, selection of individual models, development of ensemble models, and prediction of VFs. (b) F1 scores of baseline models. (c) F1 scores of ensemble models. (d) Comprehensive model evaluation using precision, recall, F1 score, accuracy, area under the curve (AUC), and Matthews correlation coefficient (MCC).
Given the limited number of VFs available solely from databases, we employed a homology-based expansion strategy, selecting homologous proteins from UniProt to augment the positive set, thereby balancing the number of positive and negative sequences in the machine learning datasets.
We further evaluated the sequence length distribution for the final positive and two negative datasets used for model training and testing. The range for all three datasets was 200–900 bp, consistent with the length distribution observed in the original database-derived VF sequences, which ensures length parity between positive and negative samples (Fig. 1c,d). This strategy reduces potential sequence length bias, providing a robust foundation for subsequent model training and evaluation.
Machine learning-based prediction of fungal VFs
Multiple sequence-derived features and machine learning algorithms were employed to create benchmark models for VF identification within fungal proteomes (Fig. 2a). The extracted features included amino acid composition (AAC), dipeptide composition (DPC), and dipeptide deviation (DDE), which are sequence-based features; quasi-sequence order (QSO), representing physicochemical properties; and evolutionary information-based features such as PSSM-composition, S-FPSSM, and RPM-PSSM. These features were integrated into vector representations serving as model inputs.
Seven machine learning algorithms were tested: Random Forest (RF), Support Vector Machine (SVM), k-Nearest Neighbors (KNN), XGBoost, AdaBoost, Gradient Boosting Decision Tree (GBDT), and Multilayer Perceptron (MLP), which resulted in 49 baseline models.
Comparison of algorithms revealed that RF, KNN, and XGBoost consistently performed well across various feature types, while SVM and MLP delivered outstanding performance with dipeptide composition features (DPC, DDE). AdaBoost and GBDT performed comparably less well. While predictive performance differences among feature sets were generally minimal, dipeptide composition features frequently produced higher accuracy across multiple algorithms. To ensure robust ensemble performance, models with F1 scores exceeding 0.8 were selected, resulting in 25 baseline models used in subsequent ensemble model construction (Fig. 2b).
The prediction results from these 25 selected models were used as input features for three types of ensemble approaches: mode, mean, and XGBoost. On the validation set, all three ensemble methods achieved F1 scores above 0.9. Notably, the XGBoost-based ensemble model outperformed individual models and BLAST, and exhibited performance comparable to existing published models (Fig. 2c,d; Table 1).
Prediction and validation of C. albicans VFs
C. albicans, a common opportunistic pathogen, is associated with a variety of diseases24,25. To predict candidate VFs and further validate the reliability of the machine learning strategy, a pan-genome analysis was performed on 99 C. albicans strains, leading to the identification of 8,081 representative gene sequences for subsequent VF prediction.
Among these, 390 sequences were predicted as VFs (Fig. 3a). Of these, 280 (71.79%) were classified as core VFs present in most strains, while 63 (16.15%) were specific to a minority of strains. BLAST comparative analysis of the predicted VFs against the four published databases showed that 193 proteins (49.49%) shared high sequence identity (≥80%) with database entries, the majority (172, 89%) being core factors. Twenty-two proteins (5.64%) had low similarity (<80%) to database records, and 175 (44.87%) did not match any database entries, comprising 93 core, 23 accessory, and 59 unique proteins (Fig. 3b,c). These findings confirm that the model is capable of predicting novel candidate VFs not captured by database or sequence similarity alone.
Comparison and functional analysis of predicted C. albicans VFs with database entries. (a) Pan-genome classification of 390 predicted C. albicans VFs. (b) Distribution of sequence similarity between predicted C. albicans VFs and entries from existing databases. (c) Similarity distribution in different pan-genome categories of C. albicans VFs compared to database entries. (d) Pathway enrichment of core C. albicans VFs in KEGG.
KEGG pathway enrichment of the core VFs indicated significant association with 24 pathways, spanning organismal systems (n = 7), metabolic pathways (n = 2), human diseases (n = 7), environmental information processing (n = 3), and cellular processes (n = 5; Fig. 3d).
Structural modeling of VFs for IBD-specific antifungal drug screening
Of the predicted C. albicans VFs, 93 were detected in gut metagenomes, with five factors exhibiting significantly increased abundance in IBD patient samples: GeneID_003944-T1, RLP66883.1, KHC88809.1, RLP65258.1, and GeneID_002228-T1 (Fig. 4a). Annotation was performed based on the NCBI NR database. GeneID_003944-T1 encodes glucan 1,3-beta-glucosidase, an enzyme involved in the degradation and remodeling of fungal cell wall glucans and reported to be associated with immune evasion26. RLP66883.1 is a mitogen-activated protein kinase, and KHC88809.1 is part of an osmolarity two-component system; these two enzymes may be involved in the fungal response to hyperosmotic stress caused by intestinal barrier damage during gut inflammation27,28. RLP65258.1 is an inositol phosphorylceramide synthase, participating in sphingolipid synthesis, and dysregulation of sphingolipid metabolism is closely linked to intestinal inflammation29. GeneID_002228-T1 belongs to hydrolase family 3, which may disrupt the host mucus layer and degrade important immune defense proteins30.
IBD-associated C. albicans VFs. (a) Changes in relative abundance of predicted C. albicans VFs among IBD individuals. The five significantly associated factors are highlighted in red. (b) Comparison of the relative abundance of the five C. albicans VFs between IBD patients and healthy controls. P values were calculated using Wilcoxon rank-sum test.
Among these, GeneID_003944-T1 showed the greatest and most significant fold enrichment (Fig. 4a,b). To assess the potential of the structural model of GeneID_003944-T1 for IBD-specific antifungal drug screening, molecular docking was performed. A total of 9,637 small molecules from DrugBank were docked to the predicted protein structure (Fig. 5a). Based on binding energy, the top 100 compounds were selected for clustering, resulting in two distinct groups (Fig. 5b,c). The five compounds with the best binding energies were distributed across the clusters: Trilaciclib (−9.23 kcal/mol), Beta carotene (−9.19 kcal/mol), DB02559 (−9.01 kcal/mol), Gentamicin (−8.96 kcal/mol), and DB08683 (−8.81 kcal/mol). Trilaciclib is an approved cyclin-dependent kinase 4/6 inhibitor. Clinical trials have demonstrated its safety and efficacy31. Interaction analysis of Trilaciclib revealed hydrogen bonds with Glu300, Glu230, Asp183, and aromatic stacking involving Phe296 (Fig. 5d). These results highlight the potential of VF structural models in providing new therapeutic targets for drug development.
Virtual screening of small molecules targeting the IBD-associated VF GeneID_003944-T1. (a) Protein structure of GeneID_003944-T1. (b) Classification of physicochemical properties of the top 100 compounds with the lowest binding energies. (c) Distribution of binding energies between compounds and GeneID_003944-T1. (d) Interaction pattern between Trilaciclib and GeneID_003944-T1.
Usage Notes
We anticipate that the comprehensive fungal VF dataset generated in this study—including both the VFs identified from the C. albicans genome and those associated with IBD—will serve as a valuable resource for researchers engaged in antimicrobial target identification. In the future, this dataset can be applied in virtual screening and drug development targeting fungal virulence factors by leveraging comprehensive peptide databases (e.g., DRAMP32) or small-molecule compound databases (e.g., PubChem33), thereby facilitating the discovery of novel therapeutic agents.
Limitations of the dataset
Firstly, all fungal VFs included in our dataset are derived from publicly available VF databases; newly incorporated sequences are homologous to already known VFs. Thus, novel VFs identified in future publications may not be immediately reflected in our dataset. Secondly, although we have endeavored to collect protein structural information for all entries, certain VFs remain without experimentally determined structures; these structures may be predicted in the future through computational or AI-based approaches. Finally, we provide only a single, well-performing prediction strategy in this study. As this work presents a dataset, we hope it will encourage the development of more innovative algorithms for fungal target identification and drug discovery.
The uniqueness of our dataset is summarized as follows: (1) It comprehensively collects and expands the repertoire of fungal VFs; (2) it provides supplementary protein structural information; and (3) it contains two machine learning-derived sub-datasets, including predicted VFs of C. albicans and those associated specifically with IBD.
Data availability
The datasets are publicly available in the Figshare online repository23.
Code availability
The code related to model construction, application, and data analysis is publicly available on GitHub: https://github.com/fengshuo112/Candida-albicans-Virulence-Factors-Predicted-Using-Machine-Learning.git.
References
Köhler, J. R., Hube, B., Puccia, R., Casadevall, A. & Perfect, J. R. Fungi that Infect Humans. in The Fungal Kingdom 811–843 (ASM Press, Washington, DC, USA, 2017).
Hallen-Adams, H. E. & Suhr, M. J. Fungi in the healthy human gastrointestinal tract. Virulence 8, 352–358 (2017).
Brunke, S., Mogavero, S., Kasper, L. & Hube, B. Virulence factors in fungal pathogens of man. Curr. Opin. Microbiol. 32, 89–95 (2016).
Saputo, S., Kumar, A. & Krysan, D. J. Efg1 Directly Regulates ACE2 Expression To Mediate Cross Talk between the cAMP/PKA and RAM Pathways during Candida albicans Morphogenesis. Eukaryot. Cell 13, 1169–1180 (2014).
Phan, Q. T. et al. Als3 Is a Candida albicans Invasin That Binds to Cadherins and Induces Endocytosis by Host Cells. PLoS Biol. 5, e64 (2007).
Fisher, M. C. et al. Tackling the emerging threat of antifungal resistance to human health. Nat. Rev. Microbiol. 20, 557–571 (2022).
Garg, A. & Gupta, D. VirulentPred: a SVM based prediction method for virulent proteins in bacterial pathogens. BMC Bioinformatics 9, 62 (2008).
Gupta, A., Kapil, R., Dhakan, D. B. & Sharma, V. K. MP3: A Software Tool for the Prediction of Pathogenic Proteins in Genomic and Metagenomic Data. PLoS One 9, e93907 (2014).
Xie, R. et al. DeepVF: a deep learning-based hybrid framework for identifying virulence factors using the stacking strategy. Brief. Bioinform. 22 (2021).
Ji, B. et al. HyperVR: a hybrid deep ensemble learning approach for simultaneously predicting virulence factors and antibiotic resistance genes. NAR Genomics Bioinforma. 5 (2023).
Zheng, D., Pang, G., Liu, B., Chen, L. & Yang, J. Learning transferable deep convolutional neural networks for the classification of bacterial virulence factors. Bioinformatics 36, 3693–3702 (2020).
Zhang, S. & Jing, Y. PreVFs-RG: A Deep Hybrid Model for Identifying Virulence Factors Based on Residual Block and Gated Recurrent Unit. IEEE/ACM Trans. Comput. Biol. Bioinforma. 20, 1926–1934 (2023).
Yin, R. et al. ViPal: A framework for virulence prediction of influenza viruses with prior viral knowledge using genomic sequences. J. Biomed. Inform. 142, 104388 (2023).
Sayers, S. et al. Victors: a web-based knowledge base of virulence factors in human and animal pathogens. Nucleic Acids Res. 47, D693–D700, https://doi.org/10.1093/nar/gky999 (2019).
Urban, M. et al. PHI-base: a new interface and further additions for the multi-species pathogen–host interactions database. Nucleic Acids Res. 45, D604–D610, https://doi.org/10.1093/nar/gkw1089 (2017).
Bateman, A. et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49 (2021).
Lu, T., Yao, B. & Zhang, C. DFVF: database of fungal virulence factors. Database 2012, bas032–bas032, https://doi.org/10.1093/database/bas032 (2012).
Varadi, M. et al. AlphaFold Protein Structure Database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2012).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res. 50, D988–D995 (2022).
Schirmer, M. et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 3, 337–346 (2018).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082, https://doi.org/10.1093/nar/gkx1037 (2018).
Feng, S. et al. Fungal Virulence Factors Datasets for IBD-Specific Antifungal Drug Discovery. Figshare https://doi.org/10.6084/m9.figshare.30112165.v2 (2025).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Mukherjee, P. K. et al. Mycobiota in gastrointestinal diseases. Nat. Rev. Gastroenterol. Hepatol. 12, 77–87 (2015).
Liu, H. et al. Plant immunity suppression by an exo-β-1,3-glucanase and an elongation factor 1α of the rice blast fungus. Nat. Commun. 14, 5491 (2023).
Liao, B. et al. The two-component signal transduction system and its regulation in Candida albicans. Virulence 12, 1884–1899 (2021).
Román, E., Correia, I., Prieto, D., Alonso, R. & Pla, J. The HOG MAPK pathway in Candida albicans: more than an osmosensing pathway. Int. Microbiol. 23, 23–29 (2020).
Mota Fernandes, C. & Del Poeta, M. Fungal sphingolipids: role in the regulation of virulence and potential as targets for future antifungal therapies. Expert Rev. Anti. Infect. Ther. 18, 1083–1092 (2020).
Pichová, I. et al. Secreted aspartic proteases of Candida albicans, Candida tropicalis, Candida parapsilosis and Candida lusitaniae. Eur. J. Biochem. 268, 2669–2677 (2001).
Weiss, J. M. et al. Myelopreservation with the CDK4/6 inhibitor trilaciclib in patients with small-cell lung cancer receiving first-line chemotherapy: a phase Ib/randomized phase II trial. Ann. Oncol. 30, 1613–1621 (2019).
Ma, T. et al. DRAMP 4.0: an open-access data repository dedicated to the clinical translation of antimicrobial peptides. Nucleic Acids Res. 53, D403–D410 (2025).
Kim, S. et al. PubChem 2025 update. Nucleic Acids Res. 53, D1516–D1525 (2025).
Acknowledgements
The present study is supported by the National Natural Science Foundation of China Grants (32170062 and 82273834).
Author information
Authors and Affiliations
Contributions
X.-Z.L. and J.L. conceived and designed the study. S.F. and Z.-T.W. performed the data analysis. Z.-T.W., Y.-J.H., A.-B.Z., M.G. help to collect the data. S.F. wrote the manuscript. Z.-L.S. and X.Z. contributed to text revision. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, S., Hou, Yj., Zhang, Ab. et al. Fungal virulence factors datasets for inflammatory bowel disease-specific antifungal drug discovery. Sci Data 12, 1796 (2025). https://doi.org/10.1038/s41597-025-06087-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06087-1