Abstract
This study introduces the first hybrid brain-computer interface dataset specifically designed for research on intracerebral hemorrhage (ICH) rehabilitation. It offers a novel data source through the synchronized acquisition of electroencephalogram (EEG) and functional near-infrared spectroscopy (fNIRS) signals. The dataset innovatively incorporated neural recordings from 17 normal subjects and 20 patients with ICH under standardized left-right hand motor imagery (MI) paradigms, featuring systematically collected and preprocessed dual-modality neural data. Beyond raw neural signals, the resource provides feature-engineered data optimized for classification algorithms and multidimensional signal decoding. The public availability of this dataset can facilitate the validation and optimization of MI decoding algorithms and advance the development of precision rehabilitation systems based on multimodal neural feedback.
Similar content being viewed by others
Background & Summary
Stroke remains a principal contributor to chronic disability worldwide1, with 55%–75% of stroke survivors developing motor deficits that critically impair functional independence and socioeconomic function2,3,4. ICH needs particular attention as a stroke subtype, accounting for 6.5%–19.6% of all stroke cases, contributing disproportionately to more than 40% of stroke-related mortality and posing exceptional neurorehabilitation challenges5,6. Upper limb motor impairment, a major sequela among ICH survivors, necessitates innovative neurorehabilitation strategies to address the limitations of conventional therapies, which often results in suboptimal recovery due to disrupted corticospinal pathways. In this context, motor imagery (MI)-based brain-computer interfaces (MI-BCIs) have emerged as a transformative approach, leveraging neuroplasticity to facilitate motor network reorganization through closed-loop feedback mechanisms7,8. However, the clinical translation of MI-BCIs for ICH rehabilitation remains limited due to technological and physiological challenges. Particularly, there is a lack of robust multimodal neural correlates specific to this population of patients.
Although electroencephalography (EEG) has been widely adopted for MI-BCI systems due to its millisecond temporal resolution9, it suffers from inherent limitations in spatial specificity and susceptibility to motion artifacts10,11. These shortcomings are critical drawbacks when monitoring hemodynamically compromised brains subjected to ICH12. Functional near-infrared spectroscopy (fNIRS) complements EEG by offering hemodynamic response measurements with superior spatial localization (5–10 mm resolution) and resistance to motion artifact, enabling precise detection of fluctuations in oxygenated hemoglobin (Oxy-Hb) in the sensorimotor cortex13. The multimodal integration of EEG and fNIRS, also known as hybrid BCIs, capitalizes on their spatiotemporal synergy. Specifically, EEG captures rapid neuronal activation patterns during MI tasks, while fNIRS tracks slower changes in hemodynamics associated with cortical reorganization14. This hybrid approach has enhanced classification accuracy in normal subjects, offering 5%-10% improvement compared to unimodal systems15. However, the multimodal integration of EEG and fNIRS remains underexplored in ICH populations where neurovascular uncoupling may alter signal dynamics.
Contemporary hybrid BCI repositories, characterized by Shin’s dataset, predominantly aggregate neurotypical cohorts performing standardized paradigms, such as MI and mental arithmetic16. Subsequent studies introduced protocols, like n-back working memory tasks, discrimination/selection response, and lexical generation paradigms, but still mainly focused on normal populations17. Crucially, BCI signals from patients with ICH exhibit fundamental divergences from normative baselines due to neurostructural compromise, neurocognitive reorganization, and age-related cerebrovascular alterations. Although emerging stroke-related datasets address certain neurological conditions, a major gap persists in ICH-specific repositories. Liu et al. pioneered an open-access dataset for upper limb MI tasks in acute stroke rehabilitation; however, it exclusively comprises unimodal EEG recordings from 50 patients with cerebral ischemia and does not include patients with ICH18. A similar study conducted by Isaev et al. released the first public fNIRS dataset including 15 stroke survivors with unilateral cortical lesions performing MI tasks under various cognitive loads, but still excluding patients with ICH19. Notably, both resources suffer from modality confinement (EEG or fNIRS alone) and etiological exclusivity, failing to address the unique pathophysiological signatures of ICH. The two limitations, including the absence of both ICH-specific and multimodal datasets, not only severely impede neurorehabilitation research but also perpetuate algorithmic biases in feature extraction and classifier design, as machine learning models trained on non-ICH data fail to capture the unique neuroelectrophysiological and hemodynamic signatures of ICH.
To address these limitations, we present the Hybrid EEG-fNIRS MI Dataset for Intracerebral Hemorrhage (HEFMI-ICH), a comprehensive dataset providing data from 17 normal subjects and 20 patients with ICH through hybrid EEG-fNIRS acquisition. The dataset comprises three components: raw data, pre-processed trial-ready data (directly applicable for classification tasks), and clinical characteristics of participants. This study offers academic contributions across three critical dimensions. First, it provided the hybrid EEG-fNIRS MI dataset encompassing both healthy individuals and patients with ICH, filling a crucial gap in neurorehabilitation research. Second, we developed a unified deep-learning framework capable of effective neural signal classification in both healthy subjects and patients, addressing the critical need for developing generalizable models in clinical BCI applications. Third, this study introduces an innovative dual-modal integration methodology that combines feature-level fusion with adaptive weighting mechanisms, thereby enhancing multimodal data synergy. The HEFMI-ICH represents a pivotal resource for advancing personalized rehabilitation BCI development, offering substantial potential for the development of more robust and accurate hybrid systems tailored for MI applications in ICH.
Methods
Participants
HEFMI-ICH comprised 17 normal subjects (right-handed; 12 males and 5 females; mean age 23.6 ± 1.8 years) and 20 patients with ICH (right-handed; 17 males and 3 females; mean age 50.8 ± 10.3 years; time since onset ranging from 2 days to 2 months). The participants were recruited from Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology (Table 1). Clinical assessments were conducted based on the Fugl-Meyer Assessment for Upper Extremities (FMA-UE), Modified Barthel Index (MBI; mean score 85.8 ± 28.7), and modified Rankin Scale (mRS; mean score 2.0 ± 1.7). All participants and their families provided written informed consent after being fully informed of the procedures and objectives of this study. And all participants provided written informed consent for participation in this study and explicitly consented to the public sharing of their anonymized data for research purposes. The study protocol was approved by the Ethics Committee of Tongji Hospital (Approval No. TJ-IRB202412123) and registered with the National Medical Research Registration and Filing Information System of China (Registration No. MR-42-25-008403).
Experimental paradigm
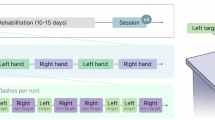
The experimental paradigm (Fig. 1) comprised at least two consecutive sessions for each subject, and each session included 15 trials of either left-hand and right-hand MI. Sufficient inter-session rest intervals were implemented to mitigate fatigue effects and ensure data quality.
Overview of the experimental procedures. The experimental protocol began with a grip-strength calibration phase using a dynamometer and stress ball to enhance the kinesthetic sensation and consistency of motor imagery. The main paradigm employed a block design, consisting of multiple sessions, each containing multiple trials. Throughout the task, subjects performed kinesthetic motor imagery, focusing on the sensation of the movement rather than its visual representation. Data was acquired by synchronously recording 32-channel EEG (256 Hz) and 90-channel fNIRS (11 Hz) signals.
In our conversations with participants revealed that some were unable to grasp the concept of “MI”. In MI, the brain issues a command to the left (or right) hand without any overt movement—this is not picturing the act of motion, but rather simulating its kinesthetic sensation. To strengthen participants’ perception of the imagined motion, we introduced a grip strength calibration procedure before data acquisition, using both a dynamometer and a stress ball. This preparatory phase comprised: 1) repeated 5 kg maximal force exertions (or voluntary maximum efforts) using a dynamometer; 2) equivalent force applications using a stress ball; 3) grip training at a rate of one contraction per second. By reinforcing the tactile and force-related aspects of the grasping movement and standardizing its temporal rhythm, this calibration enhanced participants’ MI vividness and improved consistency across trials.
Participants were seated 25 cm away from the display monitor in an ergonomic position. The experimental sequence initiated with baseline signal recording: 1-minute eyes-closed followed by 1-minute eyes-open states, demarcated by an auditory cue (200 ms beep). The MI paradigm then started with 30 trials per session, structured as follows (Fig. 2):
-
(1)
Visual cue presentation (2 s): A yellow directional arrow (left/right) in a blue background suggesting the required MI;
-
(2)
Execution phase (10 s): Central yellow fixation cross display following auditory cue (200 ms beep), during which the participants conducted kinesthetic MI, specifically, according to the directional cue (left or right), they were instructed to imagine a grasping movement with the corresponding hand at a rate of one imagined grasp per second;
-
(3)
Inter-trial interval (15 s): Blank screen suggesting the rest period.
The block design of motor imagery. A single trial was structured as follows: a visual cue (2 s, directional arrow) was followed by an execution phase (10 s, kinesthetic motor imagery of left or right hand grasping), and ended with a rest interval (15 s).
Data were automatically archived after completing the trial. The protocol was repeated across at least two sessions (total 60 trials: 30 left/right each or more), with intersession breaks adjusted based on participants’ readiness. This standardized approach enhanced signal consistency while maintaining ecological validity through phased task administration and physiological preparation.
Data acquisition
In the experimental setup, we integrated a g.HIamp amplifier (g.tec medical engineering GmbH, Graz, Austria) and a continuous-wave multifunctional fNIRS system (NirScan, Danyang Huichuang Medical Equipment Co. Ltd., China) for synchronized neurophysiological recording. The acquisition system incorporated dual sampling rates of 256 Hz for EEG signals and 11 Hz for fNIRS hemodynamic measurements. We developed a custom-designed hybrid EEG-fNIRS cap (Model M, designed for head circumferences ranging from 54 to 58 cm), with 32 EEG electrodes (32-channel configuration), 32 optical sources, and 30 photodetectors arranged in an optimized topography to achieve 90 fNIRS measurement channels through source-detector pairing (Fig. 3). The neurophysiological recording configuration employed systematic expansion of the international 10–20 system for EEG electrode placement, strategically positioning 32 channels to ensure comprehensive coverage of major functional areas in the cortex. Complementing the electrophysiological array, the fNIRS optode arrangement followed an anatomically-guided configuration aligned with functional neuroanatomical parcellations, deploying 32 laser sources and 30 photodetectors in an optimized geometric matrix. This topological design provided 90 distinct measurement channels through systematic source-detector pairing at controlled separation distances (3 cm), enabling hemodynamic monitoring across prefrontal, motor, and association cortices. The channel-specific optode coupling relationships and corresponding signal pathways are detailed in Table 2, recording the photon migration paths underlying each fNIRS measurement channel. Table 320 record the 3D coordinates of the EEG electrodes, and Tables 4, 5 record the 3D coordinates of the fNIRS optodes under the MNI template, respectively. Temporal synchronization between modalities was conducted using event markers transmitted from E-Prime 3.0 (Psychology Software Tools Inc., USA), which simultaneously triggered both recording systems during experimental paradigms.
Layout configurations of EEG electrodes and fNIRS optodes. (a) Arrangement of the hybrid system: Blue circles denote EEG electrodes, yellow squares indicate fNIRS detectors, and red squares represent fNIRS light sources; (b) Placement of EEG electrodes; (c) Arrangement of fNIRS optodes.
Clinical measures
The fugl-meyer assessment for upper extremities (FMA-UE)
The FMA-UE is a standardized, performance-based quantitative measure widely employed to assess motor function, coordination, and reflex activity in patients suffering from hemiplegia or other neurological impairments after stroke21. Developed as part of the broader Fugl-Meyer Assessment (FMA) scale, this domain-specific tool systematically assesses voluntary movement patterns across proximal-to-distal joints of the upper limb and grades tasks based on a 3-point ordinal scale (0: cannot perform; 1: partial performance; and 2: full performance). The FMA-UE comprises 33 items (total score: 66 points), encompassing shoulder/elbow/forearm, wrist, hand, and coordination/speed subscales, with higher scores suggesting better motor recovery. High inter-rater reliability, sensitivity to longitudinal changes, and alignment with neurophysiological recovery stages make it a gold standard for clinical trials and rehabilitation studies.
Modified barthel index (MBI)
The MBI is a validated, clinician-rated scale designed to assess functional independence in activities of daily living (ADLs) among patients with impaired neuromuscular function or mobility, particularly among stroke survivors22. It assesses 10 main ADL domains, including feeding, grooming, bathing, dressing, bowel/bladder control, toileting, chair-bed transfers, ambulation, and stair climbing, with weighted scoring (0–100 points) assigned based on the level of required assistance (0: complete dependence; 5–15: partial dependence; and 10–15: independence).
Modified rankin scale (mRS)
The mRS is a global disability assessment tool adopted to assess functional outcomes among patients with neurological disorders, particularly among patients with stroke23. This 6-point ordinal scale (0: no symptoms; and 6: death) emphasizes independence rather than specific impairments, categorizing patients based on their ability to perform daily activities (e.g., 2: slight disability [independent but restricted]; and 5: severe disability [bedridden, requiring constant care]). The mRS is highly applied in clinical settings since it is brief and can predict long-term morbidity/mortality.
Data Records
All BCI data are publicly accessible via Figshare24, with data from 17 healthy participants and 20 patients with ICH stored in separate repositories (Fig. 4). The raw electrophysiological data include EEG recordings in HDF5 format and fNIRS data in NIRS and HCX formats. A standardized preprocessing pipeline was conducted using MATLAB to extract critical information from raw files, including metadata, event markers (types and timestamps), channel configurations, and experimental events.
-
(1)
Raw Data Handling: EEG raw data were collected in a common HDF5 format, while fNIRS data were recorded in HCX format (a proprietary format by Huichuang Near-Infrared) and the standard NIRS format. In MATLAB, we directly read these files using the appropriate -mat file reading methods. The source files contain all necessary information, from which we extracted EEG electrode signals, fNIRS optical signals, event markers and their labels for both EEG and fNIRS, as well as sampling frequencies (fs) and other relevant metadata.
-
(2)
Data Processing: Our event coding was primarily performed in the MATLAB environment. It is important to note that during data acquisition, the original labels in EEG files exhibited occasional omissions. To ensure data integrity and accuracy, we created an EEG label template based on the experimental paradigm settings and complete fNIRS labels to replace the incomplete original labels. Through this label template, we aligned critical time points during the data acquisition process, enabling accurate extraction of neural signals during motor imagery tasks. For fNIRS data, which were originally recorded as light intensity, we applied the Beer-Lambert law to convert these signals into concentrations of oxy- and deoxy-Hb. In our desynchronization/synchronization (ERD/ERS) analysis example, we employed the following preprocessing pipeline: band-pass filtering (0.5Hz-30Hz) and baseline correction methods, with all preprocessing steps implemented using Python’s MNE library. The detailed description of preprocessing pipeline is in the section on ERD/ERS analysis.
-
(3)
Data Segmentation and Alignment: Both EEG and fNIRS data were segmented separately based on the event labels and their time points. Then, data from both modalities were aligned according to identical labels to ensure temporal synchronization of valid data across the two modalities.
Detailed structure of HEFMI-ICH.
It is noted that at this stage, we did not perform filtering or correction operations because different filtering methods can affect the data differently. We intend to provide the raw data in full to enable future researchers to analyze it without bias introduced by prior preprocessing choices. Regarding the shared code, it includes the scripts related to these preprocessing steps for both EEG and fNIRS data, enabling reproducibility of the data preparation pipeline.
Processed data were restructured into MAT files (x_epo.mat), where “x” denotes the number of sessions the participants made. Each MAT file contains the following structured components:
-
(1)
fs: Sampling rates (EEG: 256 Hz; fNIRS: 11 Hz);
-
(2)
x: A 3D tensor (trials × channels × samples) comprising 90 trials of left/right hand MI data. The sample dimension corresponds to 37-second epochs (12 s pre-task, 10 s task execution, 15 s post-task), ensuring compatibility with diverse temporal analysis frameworks;
-
(3)
y: Trial labels encoded as a 90-element array;
-
(4)
channelsName: Channel nomenclature;
-
(5)
EO/EC: Resting-state data obtained in eyes-open and eyes-closed conditions.
For fNIRS data, hemodynamic activity was quantified based on changes in the concentration of Oxy-Hb, deoxygenated hemoglobin (Deoxy-Hb), and total hemoglobin (HbT), which were derived from raw optical intensity signals through the Beer-Lambert law25.
Technical Validation
Qualitative analysis
Data quality analysis
To ensure the reliability and validity of our dataset, we conducted comprehensive data quality assessments for both EEG and fNIRS modalities. Quality evaluation is essential for multimodal neuroimaging studies, as it directly impacts the interpretability and reproducibility of research findings.
For EEG data quality assessment, we calculated signal-to-noise ratio (SNR) values across all channels. SNR provides a quantitative measure of signal quality by comparing the amplitude of the neural signal to background noise levels. Higher SNR values indicate better data quality with less contamination from artifacts such as muscle activity, eye movements, or electrical interference. We computed SNR using the ratio of signal power during task-relevant periods to baseline noise power, following established protocols in the field.
For fNIRS data quality evaluation, we employed the scalp coupling index (SCI) as the primary metric. SCI quantifies the quality of optode-scalp contact by measuring the coupling between near-infrared light sources and detectors through scalp tissue. This metric is particularly important for fNIRS recordings as poor optode contact can significantly degrade signal quality and lead to unreliable hemodynamic measurements.
All analytical methods were implemented using the MNE-Python library. For EEG data, the raw datasets were directly used to construct the corresponding data structures required by MNE. For fNIRS data, the original NIRS format files were converted to SNIRF format using MATLAB’s Homer3 toolbox, which enabled the construction of appropriate raw data structures compatible with MNE’s analytical functions. This standardized approach facilitated seamless integration of both modalities within the MNE framework for subsequent data processing and analysis. The relevant analysis files are included in the code folder.
Both SNR and SCI metrics serve as objective indicators for data screening and channel selection. Researchers can utilize these quality metrics to filter datasets according to their specific analytical requirements and quality thresholds. By providing these quantitative assessments alongside our raw data, we enable users to make informed decisions about data inclusion while maintaining the flexibility to apply their own quality control standards and preprocessing methodologies.
Time-frequency analysis
The time-domain features of time series signals consist of mean, root mean square, standard deviation, extremum, skewness, and kurtosis. For a discrete time-domain signal, these features were calculated using the Eq. (1).
where N represents the signal length, and \(x(n)\) signifies the discrete-time signal, TD denotes the characteristics in the time domain, \({{TD}}_{{Mean}}\) represents the mean value of the signal \(x(n)\), \({{TD}}_{S{td}}\) represents the standard deviation of the signal \(x(n)\), \({{TD}}_{{Var}}\) represents the variance of the signal \(x(n)\), \({{TD}}_{M{ax}}\) and \({{TD}}_{M{in}}\) represents the maximum and minimum of the signal \(x(n)\), \({{TD}}_{{Skewness}}\) represents the skewness of the signal \(x(n)\), \({{TD}}_{{Kurtosis}}\) represents the Kurtosis of the signal \(x(n)\).
In time series signal analysis, time-domain features effectively capture the characteristics of fundamental signals, such as periodicity and amplitude, with high sensitivity to anomalies and abrupt transitions. However, inherent limitations emerge due to challenging signal acquisition in non-invasive systems: Environmental noise and non-neural signal fluctuations significantly compromise the reliability of feature extraction. Furthermore, inter-individual variations in age, gender, and physiological conditions limit feature generalizability. Hybrid analytical frameworks integrating Fourier transform and wavelet analysis have been developed to enhance feature robustness and address these challenges.
As a fundamental frequency-domain analysis method, the Fourier transform processes global signal characteristics. EEG signals exhibit five principal frequency bands with distinct neurophysiological characteristics26. Delta waves (0.5–4 Hz) are associated with deep sleep and unconscious physiological regulation. Theta oscillations (4–8 Hz) manifest in hypnagogic states and exhibit pathological increase in depressive disorders. Alpha rhythms (8–14 Hz) dominate relaxed wakefulness and meditative states. Beta waves (14–30 Hz) are linked to active cognition and decision-making processes. Gamma activity (>30 Hz) suggests facilitated higher-order neural integration.
Time-frequency analysis techniques offer particular advantages for non-stationary signals with time-varying statistical properties. Short-Time Fourier Transform (STFT) employs sliding window mechanisms to decompose non-stationary signals into sufficiently small segments that can be treated as stationary signals for localized spectral estimation. However, the window size exerts a profound influence on the time-frequency resolution trade-off, where wider windows provide enhanced frequency resolution at the expense of temporal precision, while narrower windows offer superior time localization but impose limitations on frequency resolution due to insufficient signal duration for accurate spectral characterization. Wavelet Transform offers adaptive time-frequency resolution through Eq. (2):
where j represents the scaling parameter controlling frequency resolution, and k represents the temporal shift parameter. The mother wavelet function \(\varPsi \left. (x\right)\) generates wavelet basis \({\varPsi }_{j,k}(t)\) through scaling j and translation operations k. When the scaling parameter j increases, the wavelet function becomes progressively narrower in the time domain while expanding in the frequency domain, thereby enhancing its capacity to capture fine-grained high-frequency details and transient phenomena within the signal. Conversely, when j decreases, the wavelet function exhibits broader temporal support with enhanced frequency localization, primarily reflecting the low-frequency structural characteristics and sustained oscillatory patterns of the signal. The temporal shift parameter k enables systematic translation of wavelets at identical scales across different temporal segments of the signal, facilitating comprehensive examination of localized features at each temporal position. This orthogonal wavelet basis set constitutes a mathematically robust framework that enables simultaneous capture of transient high-frequency components and sustained low-frequency characteristics through its inherent multi-resolution decomposition capabilities.
The hierarchical nature of this wavelet decomposition scheme provides an elegant solution to the fundamental time-frequency trade-off inherent in traditional Fourier analysis, where the uncertainty principle dictates that improved frequency resolution necessarily compromises temporal precision, and vice versa. It is particularly well-suited for analyzing non-stationary neurophysiological signals that exhibit complex temporal dynamics across multiple frequency scales.
Figure 5 illustrates characteristic electrophysiological patterns in C3/C4 EEG channels during bilateral hand motor imagery trials. This multi-resolution analytical framework can better decode complex temporal patterns within biological signals (Subject 8 as the example).
Time-frequency representations of C3 and C4 channels during left-hand and right-hand motor imagery (Subject 8 as the example).
fNIRS topographic mapping
Optimal sensor placement for neurophysiological monitoring necessitates precise alignment with cerebral functional topography. The cerebral cortex is anatomically divided into four principal regions. The frontal lobe controls motor function, executive function, and verbal expression; the parietal lobe mediates somatosensory integration and spatial navigation; the occipital lobe is the primary center processing visual inputs; and the temporal lobe is responsible for auditory perception, semantic comprehension, and mnemonic-affective processes. fNIRS optode arrays are strategically configured based on these neuroanatomical landmarks.
Spatial normalization via the Montreal Neurological Institute (MNI) template enables cross-modal neural signal co-registration. This stereotaxic framework projects individual EEG/fNIRS source signals into a standardized 3D coordinate system, thereby effectively mitigating craniometric variability in signal localization. As depicted in Fig. 6, fNIRS-based hemodynamic mapping visualizes task-specific oxyhemoglobin dynamics during hand motor imagery. In particular, it visualizes: 1) mean concentration profiles during bilateral limb imagination; 2) task-rest differential patterns of hemoglobin fluctuations; and 3) spatiotemporal gradient evolution featuring cortical activation propagation during motor cognitive processes (Subject 8 as the example).
Cortical activation maps during left-hand and right-hand motor imagery under fNIRS signals. Top row: The concentration of oxygenated hemoglobin; Middle row: The concentration of deoxygenated hemoglobin; Bottom row: The concentration of total hemoglobin. Left to right columns: Mean hemoglobin concentration during left-hand motor imagery (LH-MI); mean difference between LH-MI and resting state; hemoglobin gradient during LH-MI; Mean hemoglobin concentration during right-hand motor imagery (RH-MI); mean difference between RH-MI and resting state; hemoglobin gradient during RH-MI (Subject 8 as the example).
Event-related desynchronization/synchronization (ERD/ERS)
Neurophysiological analysis reveals that rhythmic potential components in EEG signals, which are associated with neuronal oscillations, can serve as critical biomarkers for differentiating stimulus/behavior patterns.
Three characteristic EEG responses are observed during functional activation: event-related potential (ERP), ERD, and ERS27. MI suppresses α-band and β-band amplitudes (ERD phenomenon) in the contralateral sensorimotor cortex while enhancing α-band and β-band amplitudes (ERS phenomenon) in the ipsilateral hemisphere.
Prior to computing ERD/ERS, we performed several preprocessing steps on the EEG data. First, to reduce the impact of noise, we applied a bandpass filter to the raw EEG data, retaining the frequency range of 0.5 Hz to 30 Hz. Second, to enhance the stability of the data, we conducted baseline correction using the mean value of the 5 s to 2 s prior to the instruction onset. All the preprocessing operations were implemented using the MNE library in Python. ERD/ERS for C3/C4 channels (primary motor areas) can be calculated following three steps:
-
(1)
Extracting all trial-based EEG recordings from C3/C4 electrodes;
-
(2)
Computing trial-averaged spectral power within α/β frequency bands;
-
(3)
Calculating ERD/ERS indices based on Eq. (3).
where \(X\) denotes spectral power during MI, and \(M\) represents baseline power averaged over pre-stimulus interval (−2~0 s).
The resulting ERD/ERS topographic distributions are presented in Fig. 7 (Subject 8 as the example).
Event-related desynchronization/synchronization maps and topographic distributions of alpha and beta oscillations in left-hand and right-hand motor imagery. Topographic map labels: R: Resting state; I: Cue period (motor imagery instruction); T: Task execution (left-hand/right-hand motor imagery) (Subject 8 as the example).
Quantitative analysis
Neurophysiological analysis revealed that although MI tasks can be identified based on ERD/ERS patterns, interference from environmental noise and physiological artifacts undermines the classification stability of conventional manual feature extraction. We implemented machine learning frameworks with preprocessed data (signal calibration and bandpass filtering) across all subjects to enhance robustness. Technical implementations of these approaches are detailed below (Fig. 8):
Comparison of the performance of different classification methods. (a) Time-frequency feature-based classification; (b) CSP-SVM multimodal framework; (c) FBCSP-SVM enhanced EEG classification; (d) Depth-wise separable convolution networks; (e) Accuracy-weighted linear hybrid model.
Time-frequency feature-based classification
This approach adopted manual feature engineering using time-domain and spectral characteristics described in Section 4.1. The constructed feature vectors were classified through SVM, establishing a baseline model according to raw signal attributes.
CSP-SVM multimodal framework
Common spatial patterns (CSP), a spatial filtering technique for binary classification, was employed to optimize spatial projection matrices through matrix diagonalization and maximize inter-class variance differences28. The derived discriminative features were processed using SVM classifiers to construct maximum-margin hyperplanes, thereby enabling modality-specific classification for EEG and fNIRS.
FBCSP-SVM enhanced EEG classification
Building upon standard CSP, this improved method adopts frequency band segmentation (4 Hz bandwidth for EEG)29. Subsequent CSP feature extraction and optimal feature selection per sub-band were conducted to generate composite feature sets for final SVM-based classification.
EEGBaseNet/fNIRSBaseNet
A dual-modal foundation network architecture based on depthwise separable convolution (EEGBase/fNIRSBase) is proposed that is specifically adapted to the signal characteristics of EEG and fNIRS modalities. The feature extractor employs a three-stage convolutional architecture: the first two stages utilize one-dimensional convolutional layers to capture multi-scale spatiotemporal features, while the third stage implements depthwise separable convolution through a cascaded structure of depthwise grouped convolution and pointwise convolution, significantly reducing computational complexity while maintaining feature representation capability. This structure draws inspiration from MobileNet’s optimization principles, achieving modular decoupling of feature channels through grouped convolution to enhance network cohesion.
EEGBaseNet and fNIRSBaseNet share identical network architectures but employ different kernel sizes due to the distinct sampling frequencies of the two modalities (EEG at 256 Hz, fNIRS at 11 Hz). Specifically, EEGBaseNet configures kernel lengths of 64, 30, and 15 for the first, second, and third convolutional layers, respectively. fNIRSBaseNet maintains the same kernel configuration with lengths of 64, 30, and 15 for the corresponding layers. The network incorporates a cascaded structure of Batch Normalization and average pooling: each convolutional layer is followed by a Bath Normalization (BN) layer to accelerate convergence and mitigate gradient vanishing, while global average pooling at the terminal stage generates compact modal feature vectors. The classifier comprises a feature flattening layer and fully connected layer, achieving end-to-end signal classification through the Sigmoid function. Detailed network specifications and parameters are provided in the Table 6. During network training, cross-entropy loss serves as the loss function, with the AdamW optimizer chosen for weight optimization and the learning rate is 0.001 for parameter updating.
Accuracy-weighted linear hybrid model
We developed an accuracy-driven linear hybrid mechanism to address the limitation of conventional feature fusion methods in overlooking modality performance disparities. Training-set accuracy differentials determine feature combination coefficients, while a learnable linear layer enables dynamic multimodal fusion, thereby improving collaborative classification performance.
where, \(F\) is the fused feature, and \({F}_{{eeg}}\) and \({F}_{{nirs}}\) are the two kinds of features. \({A}_{{eeg}}\) and \({A}_{{nirs}}\) were the average accuracies of EEG and fNIRS in the training set, \(\alpha \) and \(\beta \) were the ratio of mixing features.
Comparative analysis of the aforementioned methodologies supported the accuracy distributions of subject-specific classification across algorithmic frameworks, with distinct decoding efficacy patterns emerging between EEG and fNIRS modalities (Figs. 9, 10). The implementation of Time-Frequency Feature-based Classification, CSP-SVM Multimodal Framework, and FBCSP-SVM Enhanced EEG Classification was conducted using MATLAB, while EEGBaseNet/fNIRSBaseNet and the Accuracy-Weighted Linear Hybrid Model were implemented using the PyTorch library in Python. During network training, we employed the Cross-Entropy loss function with the AdamW optimizer for weight optimization. The learning rate was set to 0.001 for parameter updating, with a batch size of 64. Each model was trained for 200 epochs to ensure convergence. All computational results were validated using 5-fold cross-validation. Specifically, the samples were randomly partitioned into five equal subsets, with each subset serving as the test set in turn while the remaining four subsets constituted the training set. This process was repeated five times, and the final classification performance was calculated as the average across all folds. The classification accuracy represents the average performance across all folds of the cross-validation process, providing a robust and unbiased estimate of the model’s generalization capability within each subject.
Classification accuracy of different methods among normal subjects and patients with ICH.
Classification accuracy across all subjects. Radar chart legend: (a) Time-frequency feature-based EEG classification; (b) Time-frequency feature-based fNIRS classification; (c) Common spatial pattern (CSP)-based EEG classification; (d) Filter bank CSP (FBCSP)-based EEG classification; (e) CSP-based fNIRS classification; (f) Deep neural network for EEG classification; (g) Deep neural network for fNIRS classification; (h) Deep neural network for hybrid EEG-fNIRS classification. N denotes subject ID.
Usage Notes
This dataset is licensed under the Creative Commons Attribution (CC-BY) terms. Users should recognize the dual functionality of the multimodal database as a foundational training substrate for developing deep learning-based decoding networks for MI tasks and a pivotal resource for the construction of personalized prognostic prediction systems in the field of ICH. Nevertheless, as a limitation, there is a lack of structured therapeutic intervention data (particularly upper limb rehabilitation protocols) that could elucidate neuroplastic changes associated with motor imagery after neurological interventions.
Data availability
The dataset has been deposited to the Figshare data deposition service (https://doi.org/10.6084/m9.figshare.28955456.v4).
Code availability
The complete experimental data processing code is systematically organized in the “code.zip” archive, structured with three dedicated directories. The “EpoDataMaking” folder contains MATLAB routines for raw data preprocessing. Specifically, it comprises marker calibration modules that address partial label absence in raw EEG recordings through manual marker correction, along with data integration algorithms. The “DLClassification” directory encompasses Python scripts with deep learning architectures for unimodal (EEG/fNIRS) and multimodal classification tasks, developed using the PyTorch framework. The “CLFResult” folder comprises traditional machine learning implementations, leveraging CSP and FBCSP algorithms for EEG pattern recognition and employing MATLAB visualization to generate diagrams for performance metrics and comparative results.
References
Feigin, V. L. et al. World Stroke Organization: Global Stroke Fact Sheet 2025. International Journal of Stroke, 17474930241308142 (2025).
Khan, M. A., Das, R., Iversen, H. K. & Puthusserypady, S. Review on motor imagery based BCI systems for upper limb post-stroke neurorehabilitation: From designing to application. Computers in biology and medicine 123, 103843 (2020).
Lee, S.-H., Kim, S. S. & Lee, B.-H. Action observation training and brain-computer interface controlled functional electrical stimulation enhance upper extremity performance and cortical activation in patients with stroke: a randomized controlled trial. Physiotherapy theory and practice 38, 1126–1134 (2022).
Gareev, I., Beylerli, O., Ilyasova, T., Mashkin, A. & Shi, H. The use of bioinformatic analysis to study intracerebral hemorrhage. Brain Hemorrhages (2024).
Chen, D. et al. Evolving therapeutic landscape of intracerebral hemorrhage: Emerging cutting-edge advancements in surgical robots, regenerative medicine, and neurorehabilitation techniques. Translational Stroke Research, 1–15 (2024).
Magid-Bernstein, J. et al. Cerebral hemorrhage: pathophysiology, treatment, and future directions. Circulation research 130, 1204–1229 (2022).
Wang, H. et al. Motor network reorganization after motor imagery training in stroke patients with moderate to severe upper limb impairment. CNS neuroscience & therapeutics 29, 619–632 (2023).
Yan, W. et al. Enhancing Neuroplasticity for Post-Stroke Motor Recovery: Mechanisms, Models, and Neurotechnology. IEEE Transactions on Neural Systems and Rehabilitation Engineering (2025).
Mishuhina, V. Feature extraction from EEG signals and regularization for brain-computer interface (2019).
Michel, C. M. et al. EEG source imaging. Clinical neurophysiology 115, 2195–2222 (2004).
Li, R. et al. Concurrent fNIRS and EEG for brain function investigation: a systematic, methodology-focused review. Sensors 22, 5865 (2022).
Yun, S. Advances, challenges, and prospects of electroencephalography-based biomarkers for psychiatric disorders: a narrative review. Journal of Yeungnam Medical Science 41, 261–268 (2024).
Almajidy, R. K., Mankodiya, K., Abtahi, M. & Hofmann, U. G. A newcomer’s guide to functional near infrared spectroscopy experiments. IEEE reviews in biomedical engineering 13, 292–308 (2019).
Liu, Z. et al. A systematic review on hybrid EEG/fNIRS in brain-computer interface. Biomedical Signal Processing and Control 68, 102595 (2021).
Xu, T. et al. Motor imagery decoding enhancement based on hybrid EEG-fNIRS signals. IEEE Access 11, 65277–65288 (2023).
Shin, J. et al. Open access dataset for EEG+ NIRS single-trial classification. IEEE Transactions on Neural Systems and Rehabilitation Engineering 25, 1735–1745 (2016).
Shin, J. et al. Simultaneous acquisition of EEG and NIRS during cognitive tasks for an open access dataset. Scientific data 5, 1–16 (2018).
Liu, H. et al. An EEG motor imagery dataset for brain computer interface in acute stroke patients. Scientific Data 11, 131 (2024).
Isaev, M. R. et al. A multiple session dataset of NIRS recordings from stroke patients controlling brain–computer interface. Scientific Data 11, 1168 (2024).
Okamoto, M. et al. Three-dimensional probabilistic anatomical cranio-cerebral correlation via the international 10–20 system oriented for transcranial functional brain mapping. Neuroimage. 21(1):99-111 (2004 Jan 1).
Hernández, E. D. et al. Intra-and inter-rater reliability of Fugl-Meyer Assessment of Lower Extremity early after stroke. Brazilian journal of physical therapy 25, 709–718 (2021).
Ohura, T., Hase, K., Nakajima, Y. & Nakayama, T. Validity and reliability of a performance evaluation tool based on the modified Barthel Index for stroke patients. BMC medical research methodology 17, 1–8 (2017).
Broderick, J. P., Adeoye, O. & Elm, J. Evolution of the modified Rankin scale and its use in future stroke trials. Stroke 48, 2007–2012 (2017).
Jian, S. et al. HEFMI-ICH: A Hybrid EEG-fNIRS Motor Imagery Dataset for Brain-Computer Interface in Intracerebral Hemorrhage. figshare. Dataset. https://doi.org/10.6084/m9.figshare.28955456.v4 (2025).
Swinehart, D. F. The beer-lambert law. Journal of chemical education 39, 333 (1962).
Jensen, M. P., Adachi, T. & Hakimian, S. Brain oscillations, hypnosis, and hypnotizability. American Journal of Clinical Hypnosis 57, 230–253 (2015).
Peter, J. et al. Movement-related beta ERD and ERS abnormalities in neuropsychiatric disorders. Frontiers in neuroscience 16, 1045715 (2022).
Saranya, S. & Menaka, R. A Quantum-Based Machine Learning Approach for Autism Detection using Common Spatial Patterns of EEG Signals. IEEE Access 2169–3536 (2025).
Ang, K. K., Chin, Z. Y., Zhang, H. & Guan, C. Filter bank common spatial pattern (FBCSP) in brain-computer interface. IEEE international joint conference on neural networks (IEEE world congress on computational intelligence) 2390–2397 (2008).
Acknowledgements
This work was supported by the Major Program (JD) of Hubei Province (2023BAA005), the Fundamental Research Funds for the Central Universities (YCJJ20251401), the National Natural Science Foundation of China (92148206), the Key Research and Development Program of Wuhan (2024020702030123), Huazhong University of Science and Technology (2024JCYJ044), and the Research Fund of Tongji Hospital (2022ZHFY01, AI2024B03).
Author information
Authors and Affiliations
Contributions
Pi.Z., B.T. and ZP.T. contributed to the conception and design of the work. J.S., DY.C., XW.Z., ZX.Z., SJ.L., YG.X., T.D., Z.Z., Pe.Z., Q.Y. and YX.T. collected the data, organized the data and prepared parts of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
Zheng Zhu is employed by GigaAI. All other authors disclose no relevant relationships.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, J., Chen, D., Zhao, X. et al. HEFMI-ICH: a hybrid EEG-fNIRS motor imagery dataset for brain-computer interface in intracerebral hemorrhage. Sci Data 12, 1816 (2025). https://doi.org/10.1038/s41597-025-06100-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06100-7