Abstract
Studying the genetic diversity of non-human great apes is important for research questions in evolution as well as human diversity and disease. Genomic data of the three great ape clades (Pan, Gorilla, Pongo) has been published across multiple studies over more than one decade. However, unlike in humans, no comprehensive dataset on great ape diversity is available, due to different scopes of the original studies. Here, we present a curated dataset of 332 high coverage (≥12-fold) whole genomes, including 198 chimpanzee, 16 bonobo, 77 gorilla and 41 orangutan individuals sequenced on the Illumina platform. By integrating data from captive individuals, we contextualize them with data from wild individuals. We discuss issues with previously published data leading to removal of individuals due to low sequencing depth, missing data, or occurrence of duplicate individuals. This resource of files in CRAM and gVCF format, as well as segregating sites per clade, will allow researchers to address questions related to human and great ape evolution and diversity in a comparative manner.
Similar content being viewed by others
Background & Summary
Great apes have been of long-standing interest as the closest living relatives of humans. Studying their phylogenetic relationships to humans and amongst each other has been an important field of genetic and genomic research over the past few decades1,2,3,4, fostering investigations into human uniqueness and understanding evolution and disease5,6,7. An assembly of the chimpanzee reference genome was published soon after the human reference genome8, and by now high-quality reference genomes of all great ape species are available9,10,11. However, beyond the insights from single genome assemblies, the diversity within a clade is an important aspect to understand their evolution and their species-specific traits12,13.
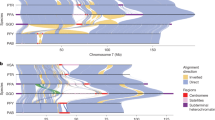
While the advent of high-throughput sequencing technologies allowed characterizing the genetic makeup of many individuals, in humans on a large scale and with high quality14, this is not possible in great apes. Great apes are all endangered or critically endangered15, with rapidly shrinking habitats and a high risk of extinction in the wild in the near future, and small captive populations from a limited pool of founder individuals. Hence, a limited number of individuals is available for genomic studies, and often, access to genetic diversity is only possible through non-invasive sampling16, or from other degraded sources such as historical collections17. However, several key publications generated diversity data from all present-day great ape species (Fig. 1): chimpanzees (Pan troglodytes)18,19, bonobos (Pan paniscus)18, western gorillas (Gorilla gorilla)18,20 and eastern gorillas (Gorilla beringei)18,21,22, as well as Bornean, Sumatran and Tapanuli orangutans (Pongo pygmaeus, Pongo abelii, Pongo tapanuliensis)18,23. This is complemented by a number of genomes from mostly captive individuals across these clades, published in the context of genome assemblies9,10,11,24, trio sequencing for the estimation of mutation rates25,26,27, functional studies28, or large-scale studies of primate diversity29.
Phylogeny and overview of individuals from each of the clades. Orangutans (Pongo): Sumatran = P. abelii; Tapanuli = P. tapanuliensis; Bornean = P. pygmaeus. Gorillas (Gorilla): Western lowland gorilla (WLG) = G. gorilla gorilla; Cross-river gorilla (CRG) = G. gorilla diehli; Mountain gorilla (MG) = G. beringei beringei; Eastern lowland gorilla (ELG) = G. beringei graueri. Chimpanzees and bonobos (Pan): Bonobo = P. paniscus; Central chimpanzee = P. troglodytes troglodytes; Eastern chimpanzee = P. troglodytes schweinfurthii; Nigeria-Cameroon chimpanzee (NC) = P. troglodytes ellioti; Western chimpanzee = P. troglodytes verus. Divergence times in million years ago (Mya) from previous studies. Numbers of individuals as reference panel plus (+) captive panel.
In many studies of human and hominin evolution, great apes are used for comparison, either as outgroup when calculating statistics30 or for demographic modelling31, or with the purpose to put diversity or variation into context5,32,33,34,35,36. However, in many cases the great ape reference genomes are used for such purposes rather than diversity data, even though it is often important to put variation within species into context. For example, examining regions of homozygosity37 or inbreeding patterns in archaic hominins are better understood by variation data from great ape species38. Furthermore, information on great ape and primate variation guides studies on human disease39 and can be used to predict the effects of mutations in present-day people40.
Given the importance of such data for human-related research questions, we suggest that it is necessary to provide a comprehensive, curated panel of published great ape genomes. Importantly, this should contain information on non-variant sites within populations, to ascertain the status of each position in the genome. A considerable number of individuals is available as short-read sequencing data (i.e. using the Illumina platform), while long-read sequencing data was primarily used for reference genome assemblies. Hence, here we focus on the single-nucleotide variant (SNV) diversity, providing a coherent dataset for 332 high-coverage genomes from all extant great ape species41. Given that great apes are very closely related to humans and each other, all data was mapped to the human reference genome (GrCH38)19. We provide both the mappings (in CRAM format) and intermediate raw gVCF files, as well as sets of called segregating sites across clades, all of which we hope will be a useful resource for numerous studies on human and primate genomics.
Methods
Samples
We used publicly available great ape genomes published in different studies (Table 1). These entail 23 chimpanzees, 12 bonobos, 27 gorillas and 10 orangutans from a landmark study on great ape diversity18; 32 chimpanzees19, 21 gorillas20,21,22 and 15 orangutans23 from subsequent population-scale studies on different wild-born individuals; and a captive panel of in total 143 chimpanzees, 4 bonobos, 29 gorillas and 16 orangutans from multiple studies with different focus10,11,24,25,26,27,28,29,42,43,44,45,46,47. All sequencing data was publicly available on the Sequencing Read Archive (SRA), and obtained through the European Nucleotide Archive branch for this study (Table S1).
We did not consider individuals from studies reporting partial genomic data (such as chromosome 2116 or the exome48) or with insufficient sequencing coverage17,49 (below 12-fold, on average, across the genome). In some of the studies considered here, sequencing data was reported for additional individuals18,19,20,23,46, which we excluded due to low average coverage, or reported evidence of cross-contamination18. In the case of one individual (SAMEA10436153923), no data was available for one sequencing run accession (ERX2240355), leading to insufficient coverage. Finally, we merged data for identical individuals published using different identifiers or in different studies in order to reach sufficient coverage (see section “Captive panel” in Technical Validation). We only considered data generated through Illumina short-read sequencing, in an effort of building a coherent dataset. We note that both long-read and short-read sequencing data was generated for some of the individuals from which the most recent genome assemblies were generated – in such cases the short-read data are included here10,11.
Bioinformatic processing
Raw fastq files were downloaded using sratoolkit (https://hpc.nih.gov/apps/sratoolkit.html, version 3.0.6), and fastQC50 was applied for initial quality assessment. Adapter trimming was performed with trimmomatic (version 0.39)51. Reads were then mapped to the human genome version GRCh38 (GCA_000001405.15_GRCh38_no_alt_analysis_set from the UCSC genome browser) with bwa mem (version 0.7.16a)52, sorted with samtools (version 1.14)53 and unmapped reads removed. Read groups were assigned with picardtools (version 2.21.4) (http://broadinstitute.github.io/picard/), and duplicated reads were marked with GATK (version 4.1.4.0)54 MarkDuplicatesSpark. Finally, reads from all sequencing libraries for each individual were merged with samtools merge into a single CRAM file. These files are reported in the associated dataset. For two individuals, sequencing depth was more than 200-fold27, resulting in excessive spurious heterozygous calls. For coherence of the dataset, we restricted the analysis to a subset of the raw sequencing data (only one out of two run accessions each, as reported in Table S1).
Using this merged CRAM file per individual, genotypes were called per chromosome with GATK HaplotypeCaller, using the flag ‘-ERC GVCF’ to generate genome-wide VCF files. These files are reported in the associated dataset. For haploid sex chromosomes in male individuals, we performed haploid genotype calling.
For downstream analyses, we created joint callsets with GATK GenomicsDBImport, and GATK GenotypeGVCFs. After creating the callset for the wild-born individuals, we added the captive panels. We report sets of segregating sites per clade (Pan, Gorilla, Pongo) with and without the captive panel within the PHAIDRA repository41 (see Data Records section). Both sets are available as VCF files and in PLINK2 format after conversion using plink255 (version 2.00a5) with the parameters ‘–max-alleles 2–snps-only–make-pgen–maf 0.00’. A permissive set contains all information on segregating sites per individual. We also obtained a more stringent set after filtering using bcftools (version 1.21)56, retaining only bi-allelic SNPs passing a 36-basepair mappability filter57, excluding sites outside the central 98% of the coverage distribution per individual with a minimum of 5-fold coverage per site, and removing heterozygous positions with less than 15% of reads supporting one allele. We also report a joint set of segregating sites across all species41.
Data analysis
We estimated depth of coverage using mosdepth58 (version 0.3.3) on the CRAM files, and used bcftools56 (version 1.16) to generate summary statistics on the VCF files. Genetic sexing was performed using ratios of the mean coverage per chromosome, with chrX:chr1 smaller than 0.75 and chrY:chrX larger than 0.1 to determine male sex.
We performed Principal Component Analysis (PCA) using VCF2PCACluster (version 1.41)59 on VCF files before and after filtering. For ADMIXTURE analyses, we subsampled 1,000,000 random autosomal loci from the VCF files and ran ADMIXTURE (version 1.3.0)60. Relatedness estimates were calculated using ngsRelate61 (version 2.0). Runs of Homozygosity were detected using bcftools roh62 (version 1.21) per chromosome per individual. Human contamination on captive individuals was estimated using HuConTest63. Subspecies assignment with f3-statistics was performed using admixtools264. As outgroup, ancestral alleles were approximated by liftover of genomic coordinates to the macaque reference genome (rheMac10)65 using rtracklayer66 (version 1.58.0 in R version 4.2.2) and bedtools getfasta67 (version 2.31.1). Geolocalization of captive chimpanzees was performed using rareCAGA16 after liftover of genotypes to the human genome version hg19 with bcftools liftover68.
We used R69 versions 4.2.3 and 4.2.2 for plotting, with packages ggplot270 (versions 3.4.4 and 3.5.1), gridExtra (version 2.3; 10.32614/CRAN.package.gridExtra), dplyr71 (version 1.1.4), tidyverse72 (version 2.0.0), ggh4x73 (version 0.2.8).
Data Records
The full dataset is available through PHAIDRA with the University of Vienna under the following link: https://doi.org/10.25365/phaidra.51441. This dataset contains the CRAM files (mapped reads) for 332 individuals, as well as gVCF files (genotype calls) for all 332 individuals for autosomes and X chromosomes, as well as Y chromosomes for the male individuals. Note that gVCF files are in the intermediate format provided by GATK HaplotypeCaller, which can be used for joint or individual genotype calling. For all files, md5sums are provided in Table S7. Furthermore, joint genotype calls in VCF format are available for the three species complexes Pan, Gorilla and Pongo, for the full set of genotype calls as well as a filtered set. A set of joint genotype call files across all 332 individuals are available on PHAIDRA41, as well as the EVA platform74 under the accession PRJEB9732475.
Technical Validation
Sequencing data
We report a curated dataset of previously published genomic data for 138 wild-born great ape individuals18,19,20,21,22,23,25, which constitute a reference panel for population genetic studies41. We also included 194 captive individuals from multiple studies10,11,18,24,25,26,27,28,29,42,43,45,46,47, resulting in a total dataset of 332 individuals. Only individuals with at least 12-fold average coverage across the genome were included, with a median of 23-fold and a maximum of 141-fold coverage (as obtained by mosdepth). Since in some cases the coverage of called genotypes was below this threshold, we also excluded three such individuals (SAMN02736775, SAMN01920524 and SAMEA104361528). Using the average coverage for the sex chromosomes, we report 208 (63%) of individuals as female, and 124 (37%) as male. We provide this information, alongside other summary metrics, in Table S2, S3, as well as Fig. S1.
We obtained several quality control measures to ensure completeness of the data: average coverage per chromosome, the last position per chromosome, the numbers of non-reference records and heterozygous positions per chromosome, and the ratio of transitions to transversions per individual (Figs. S2–7). We present the genome-wide average coverage and heterozygosity in Fig. 2. Heterozygosity values recapitulate findings from previous studies when stratified by the different subspecies3,18,19,21,22,23. Furthermore, we estimated potential human contamination63 in captive individuals. We set a threshold of 1% in order to include individuals (Fig. 4a, Table S5), leading to exclusion of some individuals (SAMN29543728, SAMN29543727, SAMN29543724, SAMN29543729)46 with values above 1%.
Distributions of coverage and heterozygosity. (a) Average coverage per individual across subspecies. Note: captive Pan is cut at 70-fold for three samples with more than 100-fold coverage. (b) Genome-wide heterozygosity per 1,000 base pairs (bp) per individual across subspecies.
Combined genotype calls of segregating sites per clade contain 117,472,161 sites for Pan (23,468,769 high quality sites after filtering), 80,907,619 sites for Gorilla (28,214,170 high quality sites after filtering), and 139,874,238 sites for Pongo (42,620,590 high quality sites after filtering). For downstream analyses, usually a coverage-based filtering is recommended. We report the central 98% of the coverage distribution for each individual, separately for autosomes and chromosome X in Table S4, with a lower cutoff of 5-fold coverage in cases where this value was below five. We advise the user to carefully consider additional filtering depending on their intended use of this dataset.
Population genetic validation
We performed basic population genetic characterisation of the individuals in this dataset, which allows to assess the quality of the data in the context of previous findings. First, we performed a PCA, showing the expected population clustering of all individuals within the respective clades Pan, Gorilla and Pongo (Fig. 3a-c; Figs. S8-12). Captive individuals are shown in grey. Notably, a PCA on the unfiltered data shows outliers for the three orangutan individuals with the highest amount of human contamination in the sequencing data (see section below; Fig. S12). We also performed clustering with ADMIXTURE (Fig. 3d; Figs. S13-15), which recapitulates known patterns of subspecies stratification in these great ape species3,18,19,21,22,23.
Basic population genetic characterisation of great ape individuals in this dataset. (a) PCA for the Pan clade (chimpanzees and bonobos), (b) PCA for the Gorilla clade, and (c) PCA for the Pongo clade, all calculated on a filtered SNV callset. (d) ADMIXTURE clustering for the Pan, Gorilla and Pongo clades (based on 1,000,000 random SNVs per clade). PPA, Pan paniscus; PTE, Pan troglodytes ellioti; PTS, Pan troglodytes schweinfurthii; PTT, Pan troglodytes troglodytes; PTV, Pan troglodytes verus; GBB, Gorilla beringei beringei; GBG, Gorilla beringei graueri; GGD, Gorilla gorilla diehli; GGG, Gorilla gorilla gorilla; PA, Pongo abelii; PP, Pongo pygmaeus; PT, Pongo tapanuliensis; CP, captive population.
Since we initially included all data reported in previous studies, we found several identical individuals based on relatedness estimates61 (KING relatedness larger than 0.4). No indication of identity was given in these respective publications. This affects the orangutan individual PD_0262/ORAN2328,29, as well as the gorilla individuals Banjo18,26, Mimi18,26, Mawenzi/PD_026426,29 and PD_0189/PD_02629, each of which have two unique SRA biosample IDs. Furthermore, a total of 17 identical chimpanzee individuals sequenced in different studies were identified. Remarkably, the individual Donald18 appears to have been sequenced in three independent studies (as 4x051947 and NS0760246). In most duplicate cases, the more recent study yielded high-coverage genomes (>30-fold), which we used for building this dataset. In some cases, in order to increase coverage we merged data after the additional step of inspecting heterozygosity. We report heatmaps of relatedness estimates including these identical individuals in the Supplementary Materials (Fig. S22-23), and provide a table of the biosample IDs for duplicated individuals (Table S5). We conclude that our data is comprehensive and reflects the original data published through these studies.
Usage Notes
Beyond the well-characterized datasets of wild-born great apes presented here as a reference dataset, we included 198 captive individuals from different studies. As described above, we assessed human contamination, retaining only individuals with less than 1% contamination. However, for three orangutan individuals, values close to 1% apparently still lead to false genotype calls and a shift in the PCA (Fig. S12). We conclude that quality filtering is recommended for subsequent analyses.
Furthermore, we provide an accurate assignment on the subspecies level based on f3-statistics (Table S6, Fig. S20)76, since 24 gorilla and 131 chimpanzee individuals did not have subspecies-level information in their SRA record, as well as two orangutans which were only labelled as Pongo. We assigned these two orangutans as Pongo pygmaeus, as reported in supplementary materials of a corresponding study77, though not in the SRA database. Most gorilla individuals are Gorilla gorilla gorilla, with the exception of PD_0179, a Gorilla beringei graueri, also reported as such only in the supplementary of the corresponding publication29 (Fig. 4b). Among chimpanzees, we identify PD_0259 and Rogger as Pan troglodytes schweinfurthii, CH114 as Pan troglodytes troglodytes, and 88A020 as Pan troglodytes ellioti, while all other chimpanzees are Pan troglodytes verus. The individual Donald/4x051946/NS0760245 is a known subspecies hybrid18. Furthermore, we performed geolocalisation of the captive chimpanzees (Fig. S21), finding, for example, an approximate origin of PD_0259 in northern Democratic Republic of Congo (Fig. 4c). We also identify 16 further individuals as likely subspecies hybrids in captivity (Fig. S21, Table S6). These analyses give a meaningful context for the genomes of these individuals, as they can complement diversity datasets of their respective subspecies or local population groups.
Characterizing the captive panel. (a) Human contamination estimates in captive individuals. (b) Subspecies assignment using f3-statistics for PD_0179. GBG, Gorilla beringei beringei; GBG, Gorilla beringei graueri; GGD, Gorilla gorilla diehlii; GGG, Gorilla gorilla gorilla. C) Geolocalization of a captive chimpanzee individual (PD_0259).
We also estimated pairwise relatedness between individuals61, recapitulating different degrees of background relatedness in some of the groups (Fig. S16-18), e.g. among bonobos (Fig. 5a) or Mountain gorillas (Fig. S17). Individuals from studies aimed at mutation rate estimation through trio sequencing24,25,26,27 were clearly identifiable by their first-degree relationships. Furthermore, multiple first-degree relationships were determined in captive chimpanzees. Known relationships are provided in Table S2, as well as inferred first-degree relationships (based on KING relatedness larger than 0.2), allowing to exclude such individuals from downstream analyses. Finally, we estimated runs of homozygosity62 (RoHs), a measure informative on long-term small effective population sizes, bottlenecks, and recent inbreeding78. We largely recapitulate previous findings18, e.g. more such RoHs in bonobos than chimpanzees or more in eastern gorillas than western lowland gorillas, while the captive individuals do not seem to show a systematic increase in RoHs (Fig. 5b; Fig. S19). Metadata are presented in Supplementary Tables and Figures.
Kinship and runs of homozygosity. (a) Relatedness among bonobos (Pan paniscus) as estimated by ngsRelate (KING method)61, as an example of individual relationships. Individuals from Trio sequencing (Mhudiblu, Loretta, PR00251) are distinguishable. Plots for all clades are found in Supplementary Materials. (b) Runs of homozygosity as estimated by bcftools roh62 across all species and subspecies. For abbreviations see Table S2.
Data availability
The dataset is available at https://doi.org/10.25365/phaidra.514, and has been deposited to EVA [PRJEB97324].
Code availability
The code used is available under https://github.com/admixVIE/Great_Ape_genomes.
References
Kaessmann, H. & Pääbo, S. The genetical history of humans and the great apes. J. Intern. Med. 251, 1–18 (2002).
Wall, J. D. Great ape genomics. ILAR J. 54, 82–90 (2013).
Kuhlwilm, M. et al. Evolution and demography of the great apes. Curr. Opin. Genet. Dev. https://doi.org/10.1016/j.gde.2016.09.005 (2016).
Yousaf, A., Liu, J., Ye, S. & Chen, H. Current Progress in Evolutionary Comparative Genomics of Great Apes. Front. Genet. 12 (2021).
Pollen, A. A., Kilik, U., Lowe, C. B. & Camp, J. G. Human-specific genetics: new tools to explore the molecular and cellular basis of human evolution. Nat. Rev. Genet. 24, 687–711 (2023).
Varki, A. & Altheide, T. K. Comparing the human and chimpanzee genomes: Searching for needles in a haystack. Genome Res. 15, 1746–1758 (2005).
Juan, D., Santpere, G., Kelley, J. L., Cornejo, O. E. & Marques-Bonet, T. Current advances in primate genomics: novel approaches for understanding evolution and disease. Nat. Rev. Genet. 24, 314–331 (2023).
Consortium, T. C. S. and A. & The Chimpanzee Sequencing and Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437, 69–87 (2005).
Gordon, D. et al. Long-read sequence assembly of the gorilla genome. Science (80-.). 352, aae0344–aae0344 (2016).
Mao, Y. et al. A high-quality bonobo genome refines the analysis of hominid evolution. Nature 594, 77–81 (2021).
Kronenberg, Z. N. et al. High-resolution comparative analysis of great ape genomes. Science (80-.). 360, eaar6343 (2018).
Zeberg, H., Jakobsson, M. & Pääbo, S. The genetic changes that shaped Neandertals, Denisovans, and modern humans. Cell 187, 1047–1058 (2024).
Han, S., Andrés, A. M., Marques-Bonet, T. & Kuhlwilm, M. Genetic variation in Pan species is shaped by demographic history and harbors lineage-specific functions. Genome Biol. Evol. evz047 https://doi.org/10.1093/gbe/evz047 (2019).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Caldecott, J. & Miles, L. World Atlas of Great Apes and their Conservation. in Environmental Conservation 33, 456 (University of California Press, 2005).
Fontsere, C. et al. Population dynamics and genetic connectivity in recent chimpanzee history. Cell Genomics 2 (2022).
van der Valk, T., Díez-del-Molino, D., Marques-Bonet, T., Guschanski, K. & Dalén, L. Historical Genomes Reveal the Genomic Consequences of Recent Population Decline in Eastern Gorillas. Curr. Biol. 29, 165–170.e6 (2019).
Prado-Martinez, J. et al. Great ape genetic diversity and population history. Nature 499, 471–5 (2013).
De Manuel, M. et al. Chimpanzee genomic diversity reveals ancient admixture with bonobos. Science (80-.). 354 (2016).
Alvarez-Estape, M. et al. Past Connectivity but Recent Inbreeding in Cross River Gorillas Determined Using Whole Genomes from Single Hairs. Genes vol. 14 (2023).
Xue, Y. et al. Mountain gorilla genomes reveal the impact of long-term population decline and inbreeding. Science (80-.). 348, 242–245 (2015).
Pawar, H. et al. Ghost admixture in eastern gorillas. Nat. Ecol. Evol. https://doi.org/10.1038/s41559-023-02145-2 (2023).
Nater, A. et al. Morphometric, Behavioral, and Genomic Evidence for a New Orangutan Species. Curr. Biol. 27, 3487–3498.e10 (2017).
Makova, K. D. et al. The complete sequence and comparative analysis of ape sex chromosomes. Nature https://doi.org/10.1038/s41586-024-07473-2 (2024).
Venn, O. et al. Strong male bias drives germline mutation in chimpanzees. Science (80-.). 344, 1272–1275 (2014).
Besenbacher, S., Hvilsom, C., Marques-Bonet, T., Mailund, T. & Schierup, M. H. Direct estimation of mutations in great apes reconciles phylogenetic dating. Nat. Ecol. Evol. 3, 286–292 (2019).
Tatsumoto, S. et al. Direct estimation of de novo mutation rates in a chimpanzee parent-offspring trio by ultra-deep whole genome sequencing. Sci. Rep. 7, 13561 (2017).
García-Pérez, R. et al. Epigenomic profiling of primate lymphoblastoid cell lines reveals the evolutionary patterns of epigenetic activities in gene regulatory architectures. Nat. Commun. 12, 3116 (2021).
Kuderna, L. F. K. et al. A global catalog of whole-genome diversity from 233 primate species. Science (80-.). 380, 906–913 (2023).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–9 (2014).
Gronau, I., Hubisz, M. J., Gulko, B., Danko, C. G. & Siepel, A. Bayesian inference of ancient human demography from individual genome sequences. Nat. Genet. 43, 1031–1034 (2011).
Staes, N. et al. FOXP2 variation in great ape populations offers insight into the evolution of communication skills. Sci. Rep. 7, 16866 (2017).
Dennis, M. Y. et al. Evolution of Human-Specific Neural SRGAP2 Genes by Incomplete Segmental Duplication. Cell 149, 912–922 (2012).
Mangan, R. J. et al. Adaptive sequence divergence forged new neurodevelopmental enhancers in humans. Cell 185, 4587–4603.e23 (2022).
Weiss, C. V. et al. The \textit{cis}-regulatory effects of modern human-specific variants. Elife 10, e63713 (2021).
Huang, X., Kruisz, P. & Kuhlwilm, M. sstar: A Python package for detecting archaic introgression from population genetic data with S *. Mol. Biol. Evol. 39, 1–6 (2022).
Kuhlwilm, M. et al. Ancient gene flow from early modern humans into Eastern Neanderthals. Nature 530, 429–433 (2016).
Skov, L. et al. Genetic insights into the social organization of Neanderthals. Nature 610, 519–525 (2022).
Richard, D. et al. Evolutionary Selection and Constraint on Human Knee Chondrocyte Regulation Impacts Osteoarthritis Risk. Cell 181, 362–381.e28 (2020).
Gao, H. et al. The landscape of tolerated genetic variation in humans and primates. Science (80-.). 380, eabn8153 (2023).
Kuhlwilm, M. Great ape genome diversity panel. https://doi.org/10.25365/phaidra.514 (2025).
Shao, Y. et al. Phylogenomic analyses provide insights into primate evolution. Science (80-.). 380, 913–924 (2023).
Solis-Moruno, M. et al. Potential damaging mutation in LRP5 from genome sequencing of the first reported chimpanzee with the Chiari malformation. Sci. Rep. 7, 15224 (2017).
Shukla, N., Shaban, B. & Gallego Romero, I. Genetic Diversity in Chimpanzee Transcriptomics Does Not Represent Wild Populations. Genome Biol. Evol. 13, evab247 (2021).
Gokcumen, O. et al. Primate genome architecture influences structural variation mechanisms and functional consequences. Proc. Natl. Acad. Sci. 110, 15764–15769 (2013).
Bracci, A. N. et al. The evolution of the human DNA replication timing program. Proc. Natl. Acad. Sci. 120, e2213896120 (2023).
Fair, B. J. et al. Gene expression variability in human and chimpanzee populations share common determinants. Elife 9, e59929 (2020).
Teixeira, J. C. et al. Long-Term Balancing Selection in LAD1 Maintains a Missense Trans-Species Polymorphism in Humans, Chimpanzees, and Bonobos. Mol. Biol. Evol. 32, 1186–1196 (2015).
Locke, D. P. et al. Comparative and demographic analysis of orang-utan genomes. Nature 469, 529–33 (2011).
Andrews, S. FASTQC. A quality control tool for high throughput sequence data. (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, s13742-015–0047–8 (2015).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Karimzadeh, M., Ernst, C., Kundaje, A. & Hoffman, M. M. Umap and Bismap: quantifying genome and methylome mappability. Nucleic Acids Res. 46, e120–e120 (2018).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics 34, 867–868 (2018).
He, W. et al. VCF2PCACluster: a simple, fast and memory-efficient tool for principal component analysis of tens of millions of SNPs. BMC Bioinformatics 25, 173 (2024).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Korneliussen, T. S. & Moltke, I. NgsRelate: A software tool for estimating pairwise relatedness from next-generation sequencing data. Bioinformatics 31, 4009–4011 (2015).
Narasimhan, V. et al. BCFtools/RoH: a hidden Markov model approach for detecting autozygosity from next-generation sequencing data. Bioinformatics 32, 1749–1751 (2016).
Kuhlwilm, M., Fontsere, C., Han, S., Alvarez-Estape, M. & Marques-Bonet, T. HuConTest: Testing Human Contamination in Great Ape Samples. Genome Biol. Evol. 13, 2021.03.30.437753 (2021).
Maier, R. et al. On the limits of fitting complex models of population history to \textit{f}-statistics. Elife 12, e85492 (2023).
Warren, W. C. et al. Sequence diversity analyses of an improved rhesus macaque genome enhance its biomedical utility. Science (80-.). 370, eabc6617 (2020).
Lawrence, M., Gentleman, R. & Carey, V. rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Genovese, G. et al. BCFtools/liftover: an accurate and comprehensive tool to convert genetic variants across genome assemblies. Bioinformatics 40, btae038 (2024).
R Core Team. R: A Language and Environment for Statistical Computing. (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. (Springer-Verlag, New York, 2009).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: A Grammar of Data Manipulation (2023).
Wickham, H. et al. Welcome to the Tidyverse. J. Open Source Softw. 4, 1686 (2019).
van den Brand, T. ggh4x: Hacks for ‘ggplot2’. (2024).
Cezard, T. et al. The European Variation Archive: a FAIR resource of genomic variation for all species. Nucleic Acids Res. 50, D1216–D1220 (2022).
European Variation Archive. http://identifiers.org/ena.embl:PRJEB97324 (2025).
Maier, R. & Patterson, N. admixtools: Inferring demographic history from genetic data. (2024).
Ferrández-Peral, L. et al. Transcriptome innovations in primates revealed by single-molecule long-read sequencing. Genome Res. 32, 1448–1462 (2022).
Kirin, M. et al. Genomic runs of homozygosity record population history and consanguinity. PLoS One 5, 1–7 (2010).
Acknowledgements
We thank N. Schulmeister for revising parts of the data. This project has been funded by the Vienna Science and Technology Fund (WWTF) [10.47379/VRG20001] to M.K. S.H. was supported by the Austrian Science Fund (FWF) [10.55776/ESP546]. The computational results of this work have been achieved using supercomputer resources provided by the Vienna Scientific Cluster (VSC) and the Life Science Compute Cluster (LiSC) of the University of Vienna.
Author information
Authors and Affiliations
Contributions
M.K. conceived the study, analysed data, and wrote the manuscript with feedback from all coauthors. S.H., S.R. and X.H. analysed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, S., Riyahi, S., Huang, X. et al. A curated dataset of great ape genome diversity. Sci Data 12, 1835 (2025). https://doi.org/10.1038/s41597-025-06124-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06124-z