Abstract
The dataset (Elsafty_Reports_of_Myeloid_Neoplasms_2024) consists of 14,441 comprehensive clinicopathologic correlations (CPCs) for myeloid neoplasms (MN) with corresponding laboratory results. Specialized online platform utilizing these laboratory results automatically generated the diagnostic and prognostic CPCs. These results include complete blood counts (CBC), peripheral blood smear (PBS) findings, blast/promyelocyte count with dysplasia screening by flow cytometry, and molecular results by NGS/PCR. Two Hematopathologists, who performed the CBC and PBS review, collected their 10,794 real-world reports of cases served in Egypt with molecular studies performed in the USA and Europe. These newly-diagnosed cases included 243 chronic myeloid leukemia (CML) cases with 257 mutations/aberrations; 4,567 non-CML MN cases with 7,883 Tiers I/II mutations/aberrations, amino acid changes, and allele frequencies; and 5,984 benign/inconclusive cases with negative NGS for myeloid mutations/aberrations in 66 DNA/RNA genes. Additionally, there are 3,647 CPCs with synthetic genomics simulating all new/follow-up CML cases and complex/rare non-CML MN cases. Stringent validation by 51 international professors/consultants/specialists from multiple medical centres confirmed 100% medical and clerical accuracy, referencing the WHO 5th edition and relevant esteemed publications.
Similar content being viewed by others
Background & Summary
Hematologic malignancies are broadly classified into myeloid and lymphoid neoplasms. The myeloid neoplasms (MN) include acute myeloid leukemia (AML), myelodysplastic neoplasms (MDS), myeloproliferative neoplasms (MPN) which include chronic myeloid leukemia (CML), overlapping MDS/MPN, and clonal haematopoiesis (CH). In 2024, approximately 500 million complete blood counts (CBC) were conducted in the USA alone, while globally, the total could reach up to 3.6 billion1. In the same year, over 50,000 new cases of AML, MDS, MPN, CML, and MDS/MPN were diagnosed in the USA2,3,4,5,6. Diagnosing and treating these neoplasms requires comprehensive clinicopathologic correlation (CPC), which involves integrating clinical findings, suspicions, and concerns with the interpretation of both current and previous laboratory/pathology results to create a comprehensive accurate diagnosis, prognosis/risk assessment, and follow-up evaluation. Essential tests for building a CPC include CBC with peripheral blood smear (PBS) review, flow cytometry for precise blast cell counting and screening for dysplastic changes, cytogenetics and molecular analysis for diagnosis and prognosis, and histopathologic examination of aspirates and biopsies when bone marrow specimens are available7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24. In specialized Hematopathology reference laboratories, 10–50% of samples are from bone marrow, while the remaining 50–90% consist of peripheral blood specimens. Nevertheless, the CBC and PBS review for qualitative findings, such as dysplastic changes and teardrop-shaped red blood cells (RBCs), remain fundamental to any CPC25,26.
The interpretation of CBC and PBS review has seen minimal evolution over the decades, remaining a time-consuming and resource-intensive process, even with the automation of descriptive comments for the CBC quantitative parameters27. In contrast, the CPC requires considerable up-to-date knowledge, mental effort, time, and experience28,29. There is a projected shortage of 5,000 to 5,700 pathologists - out of approximately 21,000 in the USA - by 203030,31. Examples of the CBC contribution to the diagnosis and the updates in the CPC since 2022: Example 1, the JAK2 V617F, CALR, and MPL mutations are highly specific to the essential thrombocythemia (ET) and primary myelofibrosis (PMF) diseases which are within the MPN category. However, serving as evidence of clonality, their presence alone may confirm the diagnosis of chronic neutrophilic leukemia (CNL) or MDS, for example, if the other diagnostic criteria are met4,32,33. Example 2, polycythaemia vera (Pv), one of the MPNs, is diagnosed in the presence of erythrocytosis and a mutation in the JAK2 gene specifically32,33. Erythrocytosis is defined as an elevated haemoglobin level above 16.0 g/dL in females and 16.5 g/dL in males, or (not and) when there is no anemia and the haematocrit level exceeds 48.0% in females and 49.0% in males. In certain situations, iron deficiency may mask erythrocytosis; therefore, Pv with superimposed iron deficiency cannot be excluded. In such cases, re-assessment after replenishing iron stores should clarify the differential diagnosis. Example 3, there are AML-defining aberrations such as CBFB::MYH11, GATA2::MECOM, KAT6A::CREBBP, MLLT3::KMT2A, PML::RARA, and RUNX1::RUNX1T1 RNA gene fusions, along with NPM1 mutations32,33. The detection of one of these AML-defining aberrations/mutations confirms the diagnosis of AML when blasts/blast-equivalents are at 10.0% or more, instead of the standard 20.0% cutoff in their absence33, and suspects progression to AML when the blasts/blast-equivalents are below 10%, including cases with as few as 0.0%. Example 4, an SF3B1 mutation is necessary to confirm the diagnosis of MDS with low blasts and SF3B1 mutation (MDS-SF3B1) and for MDS/MPN with SF3B1 mutation and thrombocytosis (MDS/MPN-T-SF3B1)32,33. Example 5, a multi-hit TP53 mutation is essential for confirming the diagnosis of MDS with biallelic TP53 inactivation (MDS-biTP53)32,33. Example 6, the CSF3R “T618I” mutation lowers the total leukocyte count required for diagnosing CNL from 25.0 K/µL to 13.0 K/µL33. Example 7, the absence of major “p210 M-bcr” BCR::ABL1 fusion transcripts excludes the CML category. According to the WHO 5th edition, the BCR::ABL1-negative atypical CML subcategory was eliminated and replaced with MDS/MPN with neutrophilia32.
Prior art
In 2011, a synoptic reporting system for PBS interpretation was introduced34. This semi-automated system features 150 titles of abnormal CBC and PBS findings, organized into checklists and drop-down menus. Experienced users, familiar with the abnormalities, select options/titles to retrieve corresponding pre-defined templates for editing. This method, where professionals compile templates for each case, remains the simplest and most basic practice in labs today. In 2018, a total of 862 CBC results were collected to verify the output of an automated draft report generator for PBS examinations35. This system, developed using the Java programming language, generates CBC and PBS draft/preliminary reports. It reads quantitative parameter results according to predefined reference ranges and produces separate, bullet-point, keyword-like descriptive (not diagnostic) comments and recommendations in a fixed structure. This technology is currently an available solution in large laboratories. In 2023, a study demonstrated the collection of 189 CBC results to verify the output of a software system, developed by US government employees and released into the public domain27. This software, named “PROSER”, combined concepts from the two previous efforts. It introduced a more agile, web-based platform featuring customizable reference ranges and generating full sentences instead of bullet points. Additionally, PROSER could be integrated with electronic health records (EHR), automating data entry from external sources into the system, where applicable. On the other hand, in contrast to the previously mentioned rule-based or code-based systems, there were state-of-technology learning-based/data-driven/AI-based systems for supporting diagnosis36,37,38,39,40,41,42. Such learning-based systems can assist in distinguishing between a few, but not all, differential diagnoses, with an overall accuracy below 95%, by utilizing the results of some flow cytometry markers or by integrating a few CBC and PBS parameters along with a limited number of mutations/aberrations. The major drawback in all these previous systems is their inability to provide specific and comprehensive diagnostic narratives, detailed prognosis, follow-up evaluation/recommendation, and CPCs for malignant/neoplastic or benign/non-neoplastic cases. They are unable to produce concise and complex sentences that prioritize significant findings, as a senior Hematopathologist would. A key novelty of this study is that none of the mentioned rule-based systems, AI-based systems or any other related enterprise AI systems have made their valuable comprehensive datasets publicly available.
Utility
The aim of the collected dataset in this work, (Elsafty_Reports_of_Myeloid_Neoplasms_2024) dataset43, which is freely accessible on the Figshare data repository, is to provide specialized, world-class, and comprehensive templates for myeloid neoplasms that serve purposes in Hematology/Hematopathology clinical practice, research, teaching/training/examination, and automation, including verification and validation of relevant multimodal automated systems. This validated, up-to-date, 100% accurate, machine-readable, and novel/unprecedented dataset has been meticulously reviewed word-by-word by both Hematopathologists and attending physicians from various specialties who detect or diagnose such cases at multiple international medical centers. Such meticulous review, referencing the WHO 5th edition32, the international consensus classification of myeloid neoplasms and acute leukemias (ICC)33, along with other relevant esteemed publications7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,28,29, justifies the complexity and diversity of the dataset while maintaining benchmark scientific quality. Including attending physicians in the evaluation and validation process was crucial for assessing the added value of this dataset from clinical perspectives. The nonspecific signs and symptoms of MN diseases make MN one of the differential diagnoses that is considered or suspected across a wide range of medical and surgical specialties. For instance, in the USA, the incidence of venous thromboembolism including the deep vein thrombosis (DVT) in cases of MN is over 5%, translating to more than 2,500 cases in MN, whereas the incidence of venous thromboembolism among various medical conditions can reach up to 900,000 cases annually44. Despite the relatively low incidence of DVT in MN compared to other conditions, recurrent DVT strongly raises suspicion of MN. While treatment for DVT may be temporary in other contexts, it is often long-term in MN cases. Therefore, a Hematopathology report that includes a robust CPC is essential in these instances.
Methods
This retrospective study received approval number ZU-IRB#: 458-12-06-2024 from the independent Research Ethics Committee at the Faculty of Medicine in Zagazig University, which operates independently from the authors. No data from any medical facilities affiliated with this university were utilized.
10,794 real-world reports collection
The original CPCs in Hematopathology reports were generated for the referred patients at specialized clinical facility in Egypt by two Hematopathologists since 2020. The patients were from all over Egypt with some residents/visitors from Libia, Sudan, Syria, Iraq, and Yemen. These reports were encrypted and stored on secure local healthcare systems with controlled access. During triage, each patient signed a clearly written legal consent form (in both Arabic and English) granting these two Hematopathologists full ownership of all collected specimens and related non-personal data, with unlimited rights for use and share in any research. Then, these two Hematopathologists collected the blood samples, conducted in-house CBC and PBS review, and sent the samples out to CAP accredited reference labs in Egypt, via a send-out lab-to-lab protocol. These reference labs performed in-house 10-color flow cytometry analyses on minimum 20,000 cells and average 40,000 cells and in turn sent out the samples to labs in the USA and Europe for genomics testing. The genomics testing included NGS for 66 driver myeloid DNA/RNA aberrations and PCR for analysis and/or confirmation of mutations/aberrations in JAK2 “V617F”, JAK2 exon 12, CALR, and FLT3. Such genomics testing was performed even with positive findings in other tests (e.g., presence of ring-sideroblasts, which is highly associated with SF3B1 mutations, did not preclude the genomics testing). As diagnostic and prognostic criteria have recently changed, the old CPCs built by these two Hematopathologists were excluded from this dataset. Instead, automated correlations for the same cases were generated using the online platform, discussed later.
The CBC analyses in these reports were performed using fully automated Haematology analyzers, including Sysmex XN-Series (Sysmex Corporation, Kobe, Japan) and Beckman Coulter DxH Series (Beckman Coulter Inc., Brea, CA, USA). The PBS reviews were performed by the two experienced Hematopathologists according to standard laboratory operating procedures to check and review for the abnormal findings. The 10-color flow cytometry gating was conducted using Kaluza software (Beckman Coulter Life Sciences, Brea, CA, USA). The NGS studies were performed using Illumina MiSeq platform (Illumina Inc., San Diego, CA, USA) and Ion S5 system (Thermo Fisher Scientific, Waltham, MA, USA). The PCR tests were performed using kits from Bio-Rad Laboratories (Hercules, CA, USA) and Thermo Fisher Scientific (Waltham, MA, USA).
The search process was limited to the two Hematopathologists mentioned earlier, the selected reports based on the inclusion and exclusion criteria were collected with their written informed consents, as patients/participants agreed to the open publication of the medical data only. The collected reports were then anonymized to protect privacy and ensure zero harm. This included assigning new case numbers after complete de-identification and removing all real identifiers, such as IDs, contact details, request numbers, names, dates of birth, gender types, clinical notes, referring physicians, testing dates, departments, and other investigations. Only medical data was retained after removing all patient and physician information with no link files or re-identification keys.
A total of 10,794 real-world reports were collected. Please refer to the Data Records section for the names and details of the collected parameters and findings. The inclusion criteria included all available final reports with complete results of the following four tests:
-
1.
Molecular analysis for mutations/aberrations by NGS in 66 myeloid DNA/RNA genes (with confirmation by PCR when necessary, especially for JAK2 Exon 12). For each mutation/aberration, the amino acid changes and the corresponding allele frequencies were collected. From total 8,140 mutations/aberrations, there were only 37 records with no data about the amino acid changes (34 FLT3 and three JAK2 Exon 12 aberrations confirmed by PCR), and there were 524 records with no data about the allele frequency (429 JAK2 “V617F”, 58 CALR type-1 and type-2, 34 FLT3, and three JAK2 Exon 12 aberrations confirmed by PCR).
-
2.
Flow cytometry count for blast/blast-equivalents/promyelocyte and screening for dysgranulopoiesis/dysmyelomonopoiesis.
-
3.
Peripheral blood smear (PBS) review for abnormal qualitative and quantitative findings.
-
4.
CBC.
To ensure data integrity, any report with incomplete results or unqualified preliminary draft was excluded. Only Tier I and Tier II mutations/aberrations, those with strong or potential clinical significance, were included. Variants with unknown clinical significance (VUS), Tier III, were not documented. Suspected germline variants, with variant allele frequency (VAF) close to 50% (45–55%) with no confirmation to be somatic mutations/aberrations were excluded by the labs and not recorded in the reports.
The real-world dataset included three groups as follows:
-
4,567 real non-CML MN cases with total 7,883 mutations/aberrations (co-mutations in up to seven DNA/RNA genes; approximately 1.7 aberrations per case).
-
243 real CML cases with 243 major “p210 M-bcr” BCR::ABL1 gene fusion transcripts results, six ABL1 kinase domain mutations, and other eight mutations/aberrations detected by myeloid NGS.
-
5,984 real myeloid NGS negative cases including 4,801 benign cases with secondary/reactive disorders and 1,183 inconclusive cases with recommended bone marrow evaluation and cytogenetics study.
The total real-world 8,140 myeloid driver mutations/aberrations were detected in different 42 genes including ABL1 kinase domain, ASXL1, BCOR, major BCR:ABL1 “p210 M-bcr”, CALR, CBFB::MYH11, CBL, CEBPA, CSF3R, DEK::NUP214, DNMT3A “R882”, DNMT3A-Others, ETV6, EZH2, FLT3, GATA2, IDH1, IDH2, IKZF1, JAK2 Exon 12, JAK2 “V617F”, JAK3, KIT, KRAS, MLLT3::KMT2A, MPL, NF1, NPM1, NRAS, PHF6, PML::RARA, PTPN11, RUNX1, RUNX1::RUNX1T1, SETBP1, SF3B1, SRSF2, STAG2, TET2, TP53-Multi-Hit, TP53-Single-VAF-Less-Than-50%, U2AF1, WT1, and ZRSR2. The synthetic dataset included CPCs with mutations/aberrations in nine more genes: BCORL1, CUX1, DDX41, GATA2::MECOM, GNAS, KAT6A::CREBBP, RAD21, SMC1A, and SMC3.
Synthetic genomics-positive dataset creation
To gather CPCs for atypical and complex cases, a dataset of total 3,647 CBC and PBS reports, different from the reports of the real-world dataset, and containing all possible degrees of normal and abnormal quantitative and qualitative results/findings, was collected. Then, at first, the seven myeloid mutations/aberrations with diagnostic specificity, discussed earlier in the background section, were assigned to numerous, but not all, CBC and PBS review reports of the corresponding suspected MN subcategories. Then using a uniform randomization, the genomic findings (including also the ones with diagnostic specificity) were assigned to the remaining reports. This was done by the following Excel formula = INDEX(A:A, RANDBETWEEN(1, COUNTA(A:A))), where column A contained a curated list of genomics. Such curated list included either CML or (not and) non-CML genomic findings. The CML list included baseline/previous and current major “p210 M-bcr” BCR::ABL1 gene fusion transcripts, 29 BCR::ABL1-kinase domain mutations, and 14 CML-associated secondary chromosomal abnormalities. While the non-CML list included 49 myeloid driver mutations/aberrations. Such conditionally-governed constrained uniform random assignment of genomic findings (total 4,181 assigned genomics/aberrations) created examples for all cases. There are examples of the entire possibilities for CML molecular response assessment (768 examples) and for non-CML MN cases (2,879 examples) including the complex and atypical ones.
The synthetic dataset included CPCs for new CML cases, where major “p210 M-bcr” BCR::ABL1 fusion transcripts were consistently detected, as well as CPCs for assessments of molecular response following CML treatment. In the latter, current results were compared with available previous results for major “p210 M-bcr” BCR::ABL1 fusion transcripts, which may be detected, not detected, or detected but non-quantifiable. Among these cases, 29 BCR::ABL1-kinase domain mutations and 14 CML-associated secondary chromosomal abnormalities, which have prognostic rather than diagnostic significance, were randomly distributed.
Generation of comprehensive clinicopathologic correlation (CPC)
The operable RESTful API “https://cbctst.com/PathOlOgics-Hematopath”45 was integrated with the lab information systems (LIS) to transfer the collected quantitative and qualitative results/findings of CBC, PBS review, flow cytometry, and molecular from the electronic health records. This software automatically generates narratives/comments/interpretations and recommendations for CBC and PBS reviews, as well as comprehensive Hematopathology CPCs, which include diagnosis and prognosis, when genomic data are incorporated. This automated tool was adopted to ensure consistent quality, eliminate clerical errors, and save valuable time allowing to focus on review, evaluation, and validation46,47. The output automated CPCs, along with the inputted CBC results, PBS findings, and genomic data, were collected and curated into an Excel workbook (.xlsx) file from the software database ensured that the same data used for generating the CPC were gathered. Critical manual data verification, for all cases, confirmed 100% correspondence between the generated CPCs and the inputted CBC results, PBS findings, and genomics data. This was performed directly through the online platform (https://cbctst.com)45, by the data verification team and the specialists in Hematology/Hematopathology and in other medical fields, who also participated in the initial phase of validation and evaluation (please see the Technical Validation section). As no discrepancies were identified, corrective procedures were not implemented.
Data Records
The dataset (Elsafty_Reports_of_Myeloid_Neoplasms_2024)43 is openly accessible through the Figshare data repository. It is organized within a single Excel workbook in .xlsx format, to enhance user searchability and facilitate machine readability. There are four sheets; “Real Non-CML MN”, “Real CML”, “Real NGS Negative”, and “Synthetic Genomics”.
Comprehensive clinicopathologic correlation (CPC) is detailed in column “A” across all sheets. Column “B” represents the “Myeloid Neoplasm Sub-Category” in both the “Real Non-CML MN” and “Synthetic Genomics” sheets. In the “Real Non-CML MN” sheet, this column includes: “Clonal Hematopoiesis (CH)”, “MDS”, “MPN”, “MDS/MPN”, and “Acute Leukemia”. In the “Synthetic Genomics” sheet, it additionally includes “CML Follow-up” and “CML New Case”. Real CML cases are listed separately in the “Real CML” sheet.
Then, there are two columns for “Age” and “Gender”, except in the “Synthetic Genomics” sheet, where all cases were assigned to adult female.
Then, the following columns are for the quantitative parameters of the CBC: Hgb, Hct, MCV, MCH, RDW%, PLT, WBC, NEUTRO%, LYMPH%, MONO%, EOSINO%, BASO%, Bands%, Meta%, Myelo%, Immature Mono Nuclear Cells (IMNC-%), Prolymphocytes%, Promyelocyte by Flow Cytometry%, BLAST by Flow Cytometry%, and nRBCs/100 WBCs. Empty cells indicate no significant presence.
Then, the following columns are for the qualitative PBS review findings: Dysplasia, Teardrop-RBCs, Atypical Lymph, Toxic Granulations, Plt Clumps, and Giant Plt. The qualitative findings are graded as (borderline, mild, moderate, severe) or (occasional, few, increased, numerous). Empty cells indicate no significant presence.
In the “Real Non-CML MN”, the columns “V”,”W”, and “X” are for “Number of genes with mutations/aberrations”, “Number of mutations/aberrations”, and “Number of AML-defining genes with mutations/aberrations”, respectively.
Then, the next columns are for genetic mutations/aberrations. The genes are sorted alphabetically starting with ASXL1 and ends with ZRSR2. Each RNA gene fusion has a column for its “Variant ID” (e.g., CBFB::MYH11.C5M33, DEK::NUP214.D9N18, KMT2A::MLLT3.K7M9, etc.), while each DNA mutation/aberration has two columns for its “Amino Acid Changes” (e.g., E929*, K700E, K385fs, etc.) and “VAF %”. In case of multiple mutations/aberrations in the same gene, the columns are repeated (e.g., “2nd ASXL1 VAF %” and “2nd ASXL1 Amino Acid Changes”, etc.).
In the “Synthetic Genomics”, there are columns for: CML history; current and previous major BCR::ABL1 fusion transcripts status (i.e., Not Detected, Detected but Non-Quantifiable, or Detected); current and previous major BCR::ABL1 fusion transcripts results; current and previous ABL1 Copies (i.e., ≥100K, 32 K-100 K, or <32K); BCR::ABL1 Kinase Domain Mutation; and CML-associated chromosomal abnormalities. Empty cells indicate that no testing was performed.
Data Overview
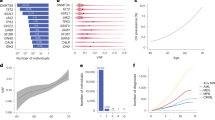
The curation of the collected CPCs identified comprehensive differential diagnoses, prognoses, and follow-up evaluations, each encompassing additional different details in the real and synthetic cases. For the counts of these differential diagnoses and prognoses across each MN subcategory in real and synthetic cases, please consult Table 1.
In MDS, the frequency of MPL mutations was noted to be threefold higher than that of JAK2 “V617F” or CALR mutations. The counts of real cases and mutations/aberrations in different genes across the five MN sub-categories (real AML, real MDS, real MDS/MPN, real MPN, and real CH) are displayed in Table 2.
There are examples for rare non-CML cases in the synthetic cases. For example, the SF3B1 mutations were present in 20 differential diagnoses beyond the two SF3B1-specific ones; (1) MDS with low blasts and SF3B1 mutation (MDS-SF3B1) and (2) MDS/MPN with SF3B1 mutation and thrombocytosis (MDS/MPN-T-SF3B1). These 20 differential diagnoses are: acute leukemia11,16,48, acute leukemic transformation of non-CML MN48, MDS with increased blasts (MDS-IB1)48, MDS with increased blasts (MDS-IB2)48, clonal haematopoiesis of indeterminate potential (CHIP)49, myelodysplastic chronic myelomonocytic leukaemia (MD-CMML-1)48,50, MD-CMML-248,50, accelerated phase of MDS/MPN48,50, MDS/MPN, not otherwise specified (MDS/MPN, NOS)48, chronic eosinophilic leukaemia (CEL)51, chronic neutrophilic leukaemia (CNL)52, ET48, MPN with impending progression to accelerated phase48, accelerated phase of MPN48, Pv48, Pv with leukocytosis48, pre-PMF48, PMF48, and triple negative essential thrombocythemia (TN-ET)53.
The CML subcategory included diagnoses and comprehensive assessments for post-treatment molecular response: covering achievement, continuation, or loss of deep molecular response (DMR), major molecular response (MMR), no MMR, and molecular relapse, with interval changes determined from the log fold calculations of the major BCR:ABL1 “p210 M-bcr” transcripts.
The cases in the real NGS negative group are either with benign changes or inconclusive. In the inconclusive cases (e.g., early low-grade MDS cannot be excluded), the CPCs comprehensively explain the case and recommend bone marrow evaluation with cytogenetics. While in the benign cases, the CPCs mention specific, personalized secondary/reactive causes for the detected disorders.
Technical Validation
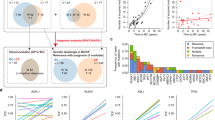
The validation/evaluation team consisted of 51 professors, consultants, and specialists from various international medical centers. This team included 16 professors/consultants and 13 specialists in Hematology/Hematopathology, as well as 11 professors/consultants and 11 specialists who are attending physicians in different medical fields, responsible for clinically diagnosing or identifying suspected MN cases. To ensure an impartial and rigorous third-party assessment, none of the medical staff involved in the development of the used platform (https://cbctst.com)45 participates in the validation/evaluation process. For a visual flowchart illustrating the steps of the technical validation process, please refer to Fig. 1.
Flowchart of the technical validation process: three-phase validation by 51 international professors/consultants and specialists.
The validators/evaluators independently conducted the validation/evaluation based on the following criteria:
-
1.
the accuracy, completeness, and comprehensiveness of the sentences summarizing the CBC and PBS review findings, and whether the narrative is self-sufficient and diagnosis-focused rather than merely descriptive.
-
2.
the correctness and usefulness of the intercalated numerical values for positive CBC quantitative results and whether they eliminate the need to refer to the raw results table.
-
3.
the relevance of highlighting the absence of certain negative qualitative or quantitative findings in supporting the diagnosis.
-
4.
the prioritization of findings based on their clinical significance.
-
5.
the accuracy, completeness, and comprehensiveness of the sentences summarizing genomic findings - both positive and negative - regarding their results, clinical significance, and implications.
-
6.
the accuracy, completeness, and comprehensiveness of the diagnostic contexts.
-
7.
the accuracy, completeness, and comprehensiveness of the prognostic contexts.
-
8.
the accuracy, completeness, and comprehensiveness of the follow-up evaluation contexts.
-
9.
the relevance of the recommendations and their consistency with medical necessity.
-
10.
the absence of clerical errors and typographical mistakes.
For each aspect, the validators/evaluators either approved or raised an objection, accompanied by explanation or uncertainty note. The validation/evaluation process was organized in three sequential phases. In the initial phase, each CPC, along with the corresponding CBC, PBS, and genomic data, was reviewed by both a Hematology/Hematopathology specialist and a specialist from a non-Hematopathology field. Each specialist in this initial phase independently reviewed and evaluated 1200–1300 different cases. If both validators/evaluators provided no objections or uncertainty notes, the case advanced to the second phase, where two professors/consultants in Hematology/Hematopathology independently reviewed that case, ensuring no errors were overlooked. Each Hematology/Hematopathology professor/consultant in this second phase reviewed and evaluated 1800–1900 different cases. While, if any objections or uncertainties were raised in the initial phase, the case was escalated to the third phase. In this third phase, each case was assigned to one professor/consultant in Hematology/Hematopathology and one professor/consultant from another clinical specialty. Each professor/consultant in this phase independently reviewed and evaluated 400–500 different cases. So, each CPC, along with its corresponding CBC, PBS, and genomic data, was reviewed by four different evaluators/validators, who included at least one specialist and one professor/consultant in Hematology/Hematopathology. The evaluation in all phases involved assessing the cases against the medical content referenced in esteemed published papers7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,28,29,32,33.
The approval rates in the initial phase conducted by the junior specialists were as follows: 88.2% for prognostic contexts, 95.0% for diagnostic contexts, 95.2% for CBC and PBS review narratives, 95.8% for follow-up evaluation contexts, 97.2% for genomic implications, 97.3% for relevance of highlighting negative findings, and 98.9% for relevance of recommendations. A perfect score of 100% was achieved for prioritization of findings, intercalated numerical values, and absence of clerical errors. The overall approval rate in the initial phase, after excluding all cases with notes, even minor ones, was 67.6%. In contrast, the second and third phases, performed by professors/consultants, with extensive experience, achieved 100% approval for all cases, with references supporting all medical content in this dataset. No cases were excluded, and the original dataset has been reviewed extensively. For a breakdown of objection types and their frequencies during the initial evaluation phase conducted by the junior specialists, please refer to Table 3.
The following 10 examples illustrate the responses from the third phase addressing the objections or uncertainties raised during the initial phase:
-
1.
Relevance of the recommendations: peripheral blood testing could be sufficient for confirming the diagnosis of CML. However, marrow evaluation is recommended when there is a need to test for secondary chromosomal abnormalities, which necessitate karyotyping testing, thereby requiring a bone marrow sample5,15.
-
2.
Relevance of highlighting negative quantitative findings: specifying the absence of leukoerythroblastosis or normoblastemia in the context of pancytopenia is essential for supporting the diagnosis of MDS over PMF, provided that all necessary diagnostic criteria are met4,33.
-
3.
Genomic implications: the statement (The BCR::ABL1 kinase domain “L384M” mutation confers resistance to Imatinib) is accurate17,18,19.
-
4.
Follow-up evaluation: the statement (The lack of detectable major “p210 M-bcr” BCR::ABL1 fusion transcripts with ≥100,000 ABL1 copies indicates deep molecular response at the 5.0 log fold level “DMR⁵“. No evidence of interval change is noted since the prior study (previous “p210 M-bcr” BCR::ABL1 fusion transcripts were not detected: ≥ MR⁴).) is accurate. The previously undetected “p210 M-bcr” BCR::ABL1 fusion transcripts, representing molecular response at or above the 4.0 log fold level, indicate that the laboratory was unable to collect more than 32 K ABL1 copies, and among them, no fusion transcripts were detected. It is inappropriate to conclude the previous level as exactly 4.0 log fold, compare it with the current 5.0 log fold level, and endorse a favorable interval decrease. There is a possibility that collecting more ABL1 copies would have still revealed no fusion transcripts. Therefore, attributing the partial lab uncertainty to significant clinical outcomes is not valid. Hence, the term “No evidence of interval change” is accurate19,21.
-
5.
Follow-up evaluation: the statement (deep molecular response at or above the 4.5 log fold level) is accurate. The lack of detectable major “p210 M-bcr” BCR::ABL1 fusion transcripts with ≥32,000 and <100,000 ABL1 copies indicates deep molecular response at or above the 4.5 log fold level. Identifying the level as exactly 4.5 log fold is incorrect and may be misleading when calculating interval changes19,21.
-
6.
Follow-up evaluation: by definition, MMR is achieved when there is more than a 3-log fold reduction, or (not and) when the results are below 0.1% on the International Scale (IS). It is not appropriate to describe a drop from a baseline of 55.0% IS (for example) to 0.06% IS as a lack of MMR simply because there is no 3-log fold reduction, despite the current result being below the 0.1% IS cutoff. The cutoffs for MMR and DMR are fundamental and cannot be further lowered. The concept of “MMR with more than a 3-log fold reduction” applies in cases where baseline results exceed 100% IS. In such rare cases, the MMR and DMR cutoffs could be elevated, making the condition of “MMR with more than 3-log fold reduction” perfectly applicable. Please refer to the method for calculating the IS for more details19,21.
-
7.
CBC and PBS review narrative interpretation: when the relative percentage of neutrophils, lymphocytes, monocytes, eosinophils, or basophils is abnormal but the absolute count is within normal range, the relative percentage should be disregarded and not considered in the comment. However, if the absolute count is increased and the relative percentage is normal or decreased, describe this as mild cytosis, depending on the cell type. If both absolute and relative counts are elevated, use the lower grade/degree among them to describe the elevation. For example, in one cell type, if there is severe absolute cytosis with mild relative cytosis, or vice versa, the description as mild cytosis is accurate. If the absolute count is decreased while the relative percentage is elevated or normal, focus on commenting on the absolute count and disregard the relative value23,24.
-
8.
CBC and PBS review narrative interpretation: when the absolute counts of neutrophils, lymphocytes, monocytes, eosinophils, and basophils are all elevated, do not specify each cell type in the comment, as this could be clinically misleading. Instead, simply refer to the condition as leukocytosis, if applicable23,24.
-
9.
Diagnostic contexts: no diagnosis of Pv can be established when all criteria are met, except for the detection of JAK2 mutations (specifically, V617F in over 95% of cases, Exon 12 in around 5%, and Exons 13 or 15 in extremely rare instances), even if other myeloid driver mutations are present. In such cases, the erythrocytosis may be secondary4,32,33.
-
10.
Prognostic contexts: the statement (DNMT3A “R882” mutations portend poor prognosis in myeloid neoplasms (MN), while up to 23% of similar cases with DNMT3A “R882” mutations are observed in Acute Leukemia (risk of post-MN AML)) is accurate11,16.
Usage Notes
To enhance the searchability of cases within the Excel workbook containing the dataset, please make use of the “Filter” and/or “Custom Sort” features in Excel. Column filters enable efficient navigation and retrieval of specific cases based on chosen criteria, providing a more streamlined and user-friendly experience for reviewing the dataset and finding the required cases. This could be whether by specifying a range of results in one or more parameters/columns, using the different options under the “Filter > Number Filters”, or by searching for specific words within the CPCs, using the different options under the “Filter > Text Filters”. Applying filters will swiftly narrow down the data, while the “Custom Sort” feature, using one or more sorting levels, allows for organizing cases in a preferred order.
Data availability
The dataset (Elsafty_Reports_of_Myeloid_Neoplasms_2024) is available at https://doi.org/10.6084/m9.figshare.29586065 on the Figshare data repository.
Code availability
No specialized code was employed. The data selection, collection, curation, check, and validation were all performed manually. Standard Excel functions within the workbook were used, including table generation and summarization. To facilitate the search for specific cases, users are encouraged to utilize the text filters or custom sort features in Excel.
References
Cohan, P. Complete blood count testers say their products can monitor covid-19 patients safely. Forbes. https://www.forbes.com/sites/petercohan/2020/03/18/complete-blood-count-testers-say-their-products-can-monitor-covid-19-patients-safely/?sh=1d0e64b34a06 (2020).
National Cancer Institute. Acute Myeloid Leukemia Treatment (PDQ®)–Health Professional Version. National Institutes of Health. https://www.cancer.gov/types/leukemia/hp/adult-aml-treatment-pdq#_359_toc (2024).
National Cancer Institute. Myelodysplastic Syndromes Treatment (PDQ®)–Health Professional Version. National Institutes of Health. https://www.cancer.gov/types/myeloproliferative/hp/myelodysplastic-treatment-pdq#_291_toc (2024).
Thapa, B., Fazal, S., Parsi, M. & Rogers, H. J. Myeloproliferative Neoplasms (StatPearls Publishing, 2024).
National Cancer Institute. Chronic Myeloid Leukemia Treatment (PDQ®)–Health Professional Version. National Institutes of Health. https://www.cancer.gov/types/leukemia/hp/cml-treatment-pdq#_224_toc (2024).
National Cancer Institute. Myelodysplastic/Myeloproliferative Neoplasms Treatment (PDQ®)–Health Professional Version. National Institutes of Health. https://www.cancer.gov/types/myeloproliferative/hp/mds-mpd-treatment-pdq#_127_toc (2024).
Tashakori, M. et al. Clinicopathologic spectrum of myeloid neoplasms with concurrent myeloproliferative neoplasm driver mutations and SRSF2 mutations. Mod Pathol. 35, 1677–1683 (2022).
Della Porta, M. et al. Minimal morphological criteria for defining bone marrow dysplasia: a basis for clinical implementation of WHO classification of myelodysplastic syndromes. Leukemia. 29, 66–75 (2015).
Yang, H. et al. High-resolution structural variant profiling of myelodysplastic syndromes by optical genome mapping uncovers cryptic aberrations of prognostic and therapeutic significance. Leukemia. 36, 2306–2316 (2022).
He, R. et al. Molecular markers demonstrate diagnostic and prognostic value in the evaluation of myelodysplastic syndromes in cytopenia patients. Blood Cancer J. 12, 12 (2022).
Gu, M. et al. Multiparameter prediction of myeloid neoplasia risk. Nat Genet. 55, 1523–1530 (2023).
Lee, W. H. et al. Validation of the molecular international prognostic scoring system in patients with myelodysplastic syndromes defined by international consensus classification. Blood Cancer J. 13, 120 (2023).
Bao, E. L. et al. Inherited myeloproliferative neoplasm risk affects haematopoietic stem cells. Nature. 586, 769–775 (2020).
Yang, Y. et al. SRSF2 mutation reduces polycythemia and impairs hematopoietic progenitor functions in JAK2V617F-driven myeloproliferative neoplasm. Blood Cancer J. 13, 171 (2023).
Hehlmann, R. et al. High-risk additional chromosomal abnormalities at low blast counts herald death by CML. Leukemia. 34, 2074–2086 (2020).
Kewan, T. et al. Molecular patterns identify distinct subclasses of myeloid neoplasia. Nat Commun. 14, 3136 (2023).
Soverini, S. et al. Next-generation sequencing for BCR-ABL1 kinase domain mutation testing in patients with chronic myeloid leukemia: a position paper. J Hematol Oncol. 12, 131 (2019).
Alves, R. et al. Resistance to Tyrosine Kinase Inhibitors in Chronic Myeloid Leukemia—From Molecular Mechanisms to Clinical Relevance. Cancers 13, 4820 (2021).
Deininger, M. W. et al. Chronic myeloid leukemia, version 2.2021. J Natl Compr Canc Netw. 18, 1385–1415 (2020).
Sanz, M. A. et al. Management of acute promyelocytic leukemia: updated recommendations from an expert panel of the European LeukemiaNet. Blood. 133, 1630–1643 (2019).
Cross, N. et al. Laboratory recommendations for scoring deep molecular responses following treatment for chronic myeloid leukemia. Leukemia. 29, 999–1003 (2015).
Landolfi, R. et al. Leukocytosis as a major thrombotic risk factor in patients with polycythemia vera. Blood 109, 2446–52 (2007).
Doig, K. & Thompson, L. A. A Methodical Approach to Interpreting the White Blood Cell Parameters of the Complete Blood Count. ASCLS. 30, 186–193 (2017).
Blumenreich, M. S. The White Blood Cell and Differential Count (Clinical Methods: The History, Physical, and Laboratory Examinations. 3rd edition. Boston: Butterworths. Chapter 153, 1990)
Sinno, S. A. J., Chakhachiro, Z. I. & Nassif, S. R. Review of Hematopathology Consult Cases: A Two-Year Experience in a Tertiary Referral Center in Lebanon. BioMed Research International. 2018, 3028625 (2018).
Elsafty, A., Soliman, A. & Ahmed, Y. 1 Million Segmented Red Blood Cells With 240 K Classified in 9 Shapes and 47 K Patches of 25 Manual Blood Smears. Sci Data. 11, 722 (2024).
Iscoe, M. S. et al. PROSER: A Web-Based Peripheral Blood Smear Interpretation Support Tool Utilizing Electronic Health Record Data. American Journal of Clinical Pathology. 160, 98–105 (2023).
Zhang, Y. et al. Comparison of the revised 4th (2016) and 5th (2022) editions of the World Health Organization classification of myelodysplastic neoplasms. Leukemia. 36, 2875–2882 (2022).
Lee, W. H. et al. Comparison of the 2022 world health organization classification and international consensus classification in myelodysplastic syndromes/neoplasms. Blood Cancer J. 14, 57 (2024).
Fu, L. et al. Virtual Pathology Elective Provides Uninterrupted Medical Education and Impactful Pathology Education During the COVID-19 Pandemic. Academic Pathology. 8, 23742895211010275 (2021).
Metter, D. M. et al. Trends in the US and Canadian Pathologist Workforces From 2007 to 2017. JAMA Netw Open. 2, e194337 (2019).
Khoury, J. D. et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia. 36, 1703–1719 (2022).
Arber, D. A. et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data. Blood. 140, 1200–1228 (2022).
Jaso, J., Nguyen, A. & Nguyen, A. N. D. A Synoptic Reporting System for Peripheral Blood Smear Interpretation. American Journal of Clinical Pathology. 135, 358–364 (2011).
Kim, Y. et al. An Automated Draft Report Generator for Peripheral Blood Smear Examinations Based on Complete Blood Count Parameters. Ann Lab Med. 38, 512–517 (2018).
Kimura, K. et al. Automated diagnostic support system with deep learning algorithms for distinction of Philadelphia chromosome-negative myeloproliferative neoplasms using peripheral blood specimen. Sci Rep. 11, 3367 (2021).
Zaccaria, G. M. et al. Electronic case report forms generation from pathology reports by ARGO, automatic record generator for onco-hematology. Sci Rep. 11, 23823 (2021).
Walter, W. et al. How artificial intelligence might disrupt diagnostics in hematology in the near future. Oncogene. 40, 4271–4280 (2021).
Lhermitte, L. et al. Automated database-guided expert-supervised orientation for immunophenotypic diagnosis and classification of acute leukemia. Leukemia. 32, 874–881 (2018).
Lhermitte, L. et al. Automated identification of leukocyte subsets improves standardization of database-guided expert-supervised diagnostic orientation in acute leukemia: a EuroFlow study. Mod Pathol. 34, 59–69 (2021).
Brasen, C. L. et al. Machine learning in diagnostic support in medical emergency departments. Sci Rep. 14, 17889 (2024).
Böttcher, S. et al. Expert-independent classification of mature B-cell neoplasms using standardized flow cytometry: a multicentric study. Blood Adv. 6, 976–992 (2022).
Elsafty, A. Elsafty_Reports_of_Myeloid_Neoplasms_2024. Figshare. https://doi.org/10.6084/m9.figshare.29586065 (2025).
Centers for Disease Control and Prevention. Data and Statistics on Venous Thromboembolism. Centers for Disease Control and Prevention. https://www.cdc.gov/blood-clots/data-research/facts-stats/index.html (2024).
CBC Test Support Team (CBCTST). Online Electronic Hematology Diagnosis, Prognosis, and Clinicopathologic Correlation. PathOlOgics, LLC. https://cbctst.com (2020).
Elsafty, A. et al. A Novel Automated Hematopathologist-Styled Commenter for Complete Blood Count and Smear Review. American Journal of Clinical Pathology. 156, S100 (2021).
Elsafty, A. & Ahmed, Y. Automated Clinicopathologic Correlation Integrating Clinical Findings, Suspicions, Hematopathology Tests and Previous Records. American Journal of Clinical Pathology. 158, S101 (2022).
Cilloni, D., Itri, F., Bonuomo, V. & Petiti, J. SF3B1 Mutations in Hematological Malignancies. Cancers 14, 4927 (2022).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Wudhikarn, K. et al. SF3B1-mutant CMML defines a predominantly dysplastic CMML subtype with a superior acute leukemia–free survival. Blood Adv 4, 5716–5721 (2020).
Shomali, W. & Gotlib, J. World Health Organization-defined eosinophilic disorders: 2019 update on diagnosis, risk stratification, and management. American Journal of Hematology 94, 1149–1167 (2019).
Szuber, N., Elliott, M. & Tefferi, A. Chronic neutrophilic leukemia: 2022 update on diagnosis, genomic landscape, prognosis, and management. American Journal of Hematology 97, 491–505 (2022).
Cattaneo, D. et al. Triple-Negative Essential Thrombocythemia: Clinical-Pathological and Molecular Features. A Single-Center Cohort Study. Front Oncol. 11, 637116 (2021).
Author information
Authors and Affiliations
Contributions
Conceptualization and study/method design: Ahmed Elsafty, Yomna, Yara, Mohamed Montaser, Ahmed Soliman, Mohamed Mahmoud Moustafa & Taher. Real-world data collection and synthetic data creation: Ahmed Elsafty & Yomna. Exporting the dataset from the online software database: Ahmed Soliman & Deyaa Elnady. Data verification: Howida & Hazem. Senior evaluators; Professors/Consultants in Hematology/Hematopathology: Shaoguang, Dongguang, Taher, Hatem, Ahmed Barefah, Hany, Samir, Nermeen, Naglaa, Maha, Manal, Ola Soliman, Heba-Allah, Iman, Ahmed S. Abdelhafiz & Walaa. Senior evaluators; Professors/Consultants in other medical fields: Yasmin, Amera, Osama, Amira, Alaa, Mohamed Saad Hamed Mahmoud, Haitham, Soha, Khaled, Ibrahim & Yara. Junior evaluators; Specialists in Hematology/Hematopathology: Samah, Nourhan, Elshaimaa, Menatalla, Lamis, Heldez, Aya, Fatma-Elzahraa, Marwa, Sara, Shady, May & Mohamed A. Saada. Junior evaluators; Specialists in other medical fields: Mahmoud, Ola M. S. Mahmoud, Abdel-Rahman, Ahmed K. Talaat, Mones, Mohamed K. Talaat, Hadeel, Sherif, Ashraf, Nora & Ramy. Statistics and review: Omar Elnady. Study management, follow-up, and manuscript editing: Howida, Elena, Maryana & Mohamed M. Moustafa. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Five of the co-authors are co-founders of PathOlOgics, LLC, a company providing AI-powered automated analytical and post-analytical pathology services. Ahmed Elsafty serves as the CEO, Yomna as the Chief Medical Officer (CMO), Ahmed Soliman as the Technical Officer, Howida as the Quality Officer, and Mohamed Montaser as the board member. The remaining co-authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Elsafty, A., Ahmed, Y., Halawa, T.F. et al. 14,441 Genomics-Based Validated Automated Comprehensive Clinicopathologic Correlations for Myeloid Neoplasms. Sci Data 12, 1819 (2025). https://doi.org/10.1038/s41597-025-06284-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06284-y