Abstract
Brain metastases occur in approximately 30–40% of lung cancer patients, and outcomes are heterogeneous, dependent on a combination of clinical, radiologic, pathologic, and molecular characteristics. Though artificial intelligence-based prognostic models show promise, their use is limited by a lack of high-quality training data. Thus, there is a critical need for a large, heterogeneous, annotated, open-access brain metastasis dataset with matched radiologic and histopathologic imaging. We present such a dataset composed of 111 cases of magnetic resonance (MR) and histopathologic imaging from patients with brain metastasis from primary lung cancer. We provide pre-operative T1-weighted contrast-enhanced (T1CE) and fluid-attenuated inversion recovery (FLAIR) MR images with matched whole slide images of formalin-fixed, paraffin-embedded (FFPE) brain metastasis biopsies. The dataset also includes segmentations of contrast enhancement and FLAIR hyperintensity, radiomic features, and clinical information. A Kaplan-Meier analysis validated that the dataset’s patients are consistent with prior large retrospective studies of patients with brain metastasis from lung cancer. The provided dataset will facilitate the development of multimodal models for improved management of lung cancer patients.
Similar content being viewed by others
Background & Summary
Brain metastases occur in approximately 30–40% of lung cancer patients1 and represent about 40% of all brain metastasis cases2. They remain a clinical challenge for oncologists and are associated with substantial morbidity and mortality, with reported median survival times ranging from 7 to 46 months, depending on histologic, clinical, and molecular characteristics3.
Current treatment modalities for brain metastases include surgical resection, whole brain radiotherapy (WBRT), stereotactic radiosurgery (SRS), and systemic therapies, such as targeted agents and immune checkpoint inhibitors4,5,6. While these approaches offer varying degrees of efficacy, factors including patient age, baseline cognitive function, the number and size of metastatic lesions, and molecular profile heavily influence outcomes7. Integrating the various elements which influence outcomes to provide accurate risk stratification remains an ongoing clinical challenge.
Advanced imaging techniques, particularly T1-weighted contrast-enhanced (T1CE) and T2-weighted fluid-attenuated inversion recovery (FLAIR) MRI, play a central role in brain metastasis diagnosis, monitoring, and treatment planning8,9,10,11,12,13,14. To date, however, MR imaging has not played a large role in prognosis estimation. Image-based predictive modeling has shown promise by leveraging both deep learning and radiomic approaches15,16,17; however, these methods fail to integrate clinical, pathologic, and molecular data, which have been shown to influence outcomes. The clinical applicability of these models is often constrained by small, homogeneous datasets that lack multimodal data to enhance granularity18,19,20.
To address these challenges, this dataset provides multimodal data for 111 cases of brain metastasis from lung cancer, including pre-treatment T1CE and FLAIR MR images, comprehensive segmentations of peritumoral edema and enhancing tumor regions, and matched histopathological whole slide images derived from surgical biopsy specimens. Segmentations were performed within a research-grade Picture Archiving and Communication System (PACS), ensuring high accuracy and reproducibility. Radiomic features were then extracted from these segmentations and are provided for further downstream analysis and machine learning model training.
The inclusion of histopathologic images establishes a direct link between radiologic findings with tissue morphology, tumor microenvironment, and cellular composition. This integration bridges the gap between non-invasive imaging and definitive tissue analysis, enabling more comprehensive tumor characterization21,22. The provided dataset of matched radiologic and tissue imaging will allow for true multimodal risk stratification studies.
By offering this high-quality, annotated dataset, we aim to catalyze the development and validation of multimodal artificial intelligence algorithms tailored for brain metastasis management.
Methods
Subject characteristics
Prospectively collected data from within the Yale Central Nervous System (CNS) Metastasis Biorepository was queried for patients who had a diagnosis of brain metastasis consistent with primary lung cancer. Of the 129 cases meeting this criterion, 6 cases were excluded for lacking any available pre-operative brain MRI with T1CE enhancement and/or FLAIR hyperintensity. Another 4 cases were excluded due to extreme motion artifact degrading image quality. Finally, 8 cases were excluded for lacking pathology images. Thus, 111 cases with matched histologic-radiologic imaging are provided in this dataset. This dataset focuses on preoperative imaging and resected tissue samples; treatment data beyond surgery (e.g., radiation, chemotherapy, targeted therapy) was not collected and is not included.
Breakdown of the cohort by histologic subtype is as follows: 12 (11%) small-cell lung cancer (SCLC) and 99 (89%) non-small cell lung cancer (NSCLC). Of the NSCLC cases, 79 (71%) are adenocarcinoma and 20 (18%) are non-adenocarcinoma. A summary of demographic and clinical characteristics is provided in Table 1. Details are provided in the dataset’s accompanying Excel file.
Resections of brain metastasis recurrence
There were 8 instances of archived brain metastasis recurrence, each with entries from brain tissue resection at two different time points, so 103 unique patients were included in this dataset. For each of the 8 patients with archived recurrence of brain metastasis, the diagnosed histologic subtype remained consistent across both time points. The provided dataset includes unique case identifiers and shared patient identifiers for each of the 8 pairs.
Tissue sample processing and histopathologic image acquisition
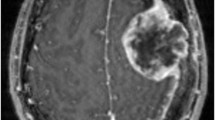
Fresh brain metastasis tissue was collected at the time of surgery by the Yale CNS Biorepository and processed within one hour of collection. Tissues were fixed with 10% neutral buffered formalin for 12–24 hours and were paraffin-embedded by standard procedures. Histology slides were also obtained in this study from the Yale Pathology Tissue Archive after receiving Institutional Review Board approval. FFPE tissue processing by Yale Pathology followed standard procedures. Sections of 5 µm thickness were stained with hematoxylin and eosin (HE) for histologic evaluation. HE-stained slides were reviewed by a pathologist to assess tumor cellularity and tissue staining quality. A representative slide from each case was scanned at 20x magnification using the MoticEasyScan Infinity whole slide imaging system (San Francisco, CA) and MoticEasyScan software. Quality assurance of whole slide digital images was performed using QuPath software.
Radiologic image acquisition
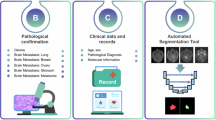
This dataset is hosted at The Cancer Imaging Archive (TCIA), providing open access to MRI sequences, radiomic features, clinical metadata, and pathology images23. Fig. 1 outlines the dataset’s composition and curation process.
Dataset Overview. Workflow figure highlights the patient cohort as those with resection of brain metastasis from primary lung cancer. It also outlines the three matched data sources: (1) pre-operative radiologic imaging and its derivatives, including T1CE and FLAIR MRI, core enhancement and whole peritumoral edema segmentations, and radiomic features extracted from these segmentations; (2) clinical data from the EMR system (2) whole slide histopathologic images of brain metastasis tissue; and (3).
For each included brain tissue sample, a matched pre-operative MR imaging study was identified through a search of the electronic medical record (EMR) system. DICOM metadata from the associated T1CE and FLAIR MRI sequences were obtained via the Visage (Visage 7, Visage Imaging, Inc., San Diego, CA) Python application programming interface (API). These metadata were, in turn, used to read acquisition parameters for the included imaging studies. The images were obtained on 0.7 T, 1.5 T, and 3 T MRI scanners. Scanner vendors included Siemens, General Electric, Philips, and Hitachi. Image acquisition parameters for each scan are available in the accompanying Excel file. A summary of these parameters is provided in Table 2.
Segmentation procedure
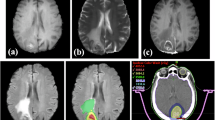
A copy of the DICOM files of the included MR imaging studies on the clinical production PACS was de-identified and transferred to a research instance of the PACS (AI Accelerator). Prior to segmentation, the location of the excised lesion was confirmed via a review of the EMR. Then, within the research PACS instance, tumor boundaries were manually annotated. Annotators were instructed to segment the core tumor as the area of contrast enhancement on T1CE image sequences, corresponding to the actively proliferating and vascularized portion of the lesion. The whole tumor was defined as the combination of the core tumor and the surrounding peritumoral edema, as identified by hyperintensity on FLAIR images.
Of the 111 imaging studies included in this dataset, 100 studies (90%) had both core and whole tumor segmentations, 4 studies (4%) had only whole tumor segmentations, and 7 studies (6%) had only core tumor segmentations. Segmentations were not performed if the relevant imaging sequence (T1CE or FLAIR) was not obtained, or contrast enhancement or peritumoral edema were absent on the respective sequences. Segmentations were saved as annotations on each imaging study and batch exported as NIfTI files via the Visage Python API.
Radiomic feature extraction
PyRadiomics24 version 3.1.0 was used to extract radiomic features from each three-dimensional (3D) segmentation volume. The feature extraction was performed in compliance with the Image Biomarker Standardisation Initiative25. Prior to feature extraction, images underwent N4 bias field correction to mitigate inter-scanner and inter-protocol inhomogeneities. All images were then resampled to an isotropic voxel size of 1 × 1 × 1 mm³ using B-spline interpolation, while segmentation masks were resampled using nearest-neighbor interpolation. Image intensities were z-score normalized to further reduce scanner and protocol-related variability.
Texture features were computed after discretizing image intensities using a fixed bin width of 25 intensity units, ensuring consistent grey-level quantization across cases. Texture matrices were calculated in 3D and aggregated across directions and slices using the mean of all directional matrix-based features, following PyRadiomics defaults.
A total of 107 features were extracted: 14 shape-based, 18 first-order, and 75 textural features. The full extraction script is available on GitHub (https://github.com/Aneja-Lab-Yale/BM-Pathology-Dataset), ensuring reproducibility.
These radiomic features were selected to capture a wide range of tumor properties, including shape (e.g., elongation, sphericity), intensity distribution (e.g., entropy, kurtosis), and texture (e.g., gray-level co-occurrence matrix features like contrast and homogeneity). Such features are commonly used in radiomics studies of brain tumors and have been shown to correlate with tumor grade, molecular subtype, and prognosis15,26,27,28. Including these features enables users of the dataset to explore associations between quantitative imaging biomarkers and clinical or molecular outcomes.
Clinical data acquisition
Clinical data for each patient was collected from the EMR. They include the following: sex, pack-year smoking history, race, age at resection, size of dominant lesion, location of lesion, and histologic subtype. Additional factors were collected to calculate the Graded Prognostic Assessment (GPA)3 score for each patient, including age, Karnofsky Performance Status (KPS), number of brain metastases, and presence of extracranial metastasis (ECM) at time of brain metastasis diagnosis, as well as EGFR, ALK, and PD-L1 mutational status. These data were collected from the EMR as of December 2024.
Anonymization
A copy of the radiologic data for our study was de-identified on the clinical production PACS and sent to the research PACS instance. The de-identification process removed or modified all identifiable metadata in accordance with the DICOM standard PS3.15 2018b Appendix E “Attribute Confidentiality Profiles.”29 Clinical medical record numbers and case accession numbers were replaced with anonymized patient and case identifiers. MR images and 3D segmentation masks were exported from the research server as NIfTI files using the Visage Python API. To further ensure patient anonymity, all radiologic images underwent brain extraction using HD-BET30 prior to the publication of the dataset.
Histopathologic images were scanned without patient identifiers on slides. SVS files were renamed with the anonymous case identifiers, aligning names between imaging modalities.
Ethical approval
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Institutional Review Board (or Ethics Committee) of Yale University, protocol 2000027592, approved on 04/20/2020. The Institutional Review Board determined that informed consent can be waived for this study.
Data Records
The data records have been published at TCIA collections23. MR images with tumor segmentations are provided alongside histopathologic imaging and an accompanying Excel file with separate sheets for clinical data, radiologic image acquisition parameters, and extracted radiomic features. For each brain metastasis resection, there are up to four files with the T1CE (caseID_t1ce_img) and/or FLAIR (caseID_flair_img) pre-operative images, as well as the core enhancement (caseID_core_seg) and/or whole hyperintensity (caseID_whole_seg) tumor segmentations. All sequences and segmentations were exported from the research PACS in the NIfTI format and brain-extracted. For each brain metastasis resection, there is also one associated whole slide image in SVS format (caseID_wsi.svs). All filenames include an anonymous case identifier and a short description of the contents, as indicated above.
As an example, the T1CE image YG_0CBM148C1MFN_t1ce_img.nii.gz corresponds to the core segmentation file named YG_0CBM148C1MFN_core_seg.nii.gz and the matched histopathology slide YG_0CBM148C1MFN_wsi.svs. These identifiers allow for cross-referencing across imaging modalities. Additional details on file structure are available through the repository hosted at TCIA23. Moreover, lesion location information based on biopsy labels is provided in the accompanying clinical data sheet to assist users in matching histopathology samples to specific MR lesions.
Technical Validation
Pathological confirmation of brain metastasis and the primary tumor of origin was obtained for all patients using data from pathology reports. The quality of whole slide pathology images was verified by a double board-certified cyto/molecular pathologist (D.D.). While the MRI scanner vendors and field strengths varied, image quality was assessed during dataset curation to ensure sufficient diagnostic clarity. Only high-quality T1CE and/or FLAIR MR imaging free from extreme motion artifacts was included in the final dataset. Differences in scanner type and field strength can influence image contrast and resolution; however, this heterogeneity reflects real-world clinical variation and enhances the generalizability of the dataset for algorithm development.
To validate the representativeness of the patient population in this study, prognoses were compared with those reported in the GPA studies by Sperduto et al.3,7 GPA, a widely-used prognostic index, stratifies patients with brain metastases based on clinical and molecular characteristics, as detailed in Methods. These factors were collected, and GPA scores were calculated for each patient.
GPA scores are provided as a validated surrogate for prognosis. In Fig. 2, pie charts are shown to present the distribution of patients in this dataset by GPA risk group and histologic subtype. Kaplan-Meier survival analyses were performed for each histologic subtype, and the Mantel-Cox log-rank test was used to evaluate statistical significance between GPA risk groups. Figure 3 plots the results from this analysis. Statistically significant differences in survival between GPA risk groups were observed among NSCLC non-adenocarcinoma patients (p = 0.020) and NSCLC adenocarcinoma patients (p = 0.002). However, no statistically significant differences were identified for SCLC patients (p = 0.397) in the setting of small sample size (n = 11). Based on these findings, the patient population in this study is considered consistent with previous large population studies of patients with brain metastasis from lung cancer, supporting the validity of the provided dataset.
Breakdown of patients by GPA risk group. Pie charts display the distribution of patients stratified by GPA risk groups in three brain metastasis histologic subtypes: (A) NSCLC non-adenocarcinoma, (B) NSCLC adenocarcinoma, (C) SCLC.
Kaplan-Meier survival curves stratified by GPA risk group. Kaplan-Meier survival curves illustrate differences in survival outcomes across GPA risk groups for (A) NSCLC non-adenocarcinoma, (B) NSCLC adenocarcinoma, and (C) SCLC. P-values were calculated using the Mantel-Cox log-rank test to evaluate statistical differences between GPA risk groups within each histologic subtype.
Usage Notes
The NIfTI files can be downloaded from TCIA public collections (https://www.cancerimagingarchive.net/) and opened on segmentation platforms that support NIfTI format23. SVS files can also be downloaded from TCIA and can be opened on most platforms that support viewing whole slide images acquired by medical microscope scanners.
Data availability
The dataset is available at TCIA collections (https://doi.org/10.7937/k0sm-y874)23.
Code availability
Code used for radiomic feature extraction, as well as brain extraction, is freely available on the Aneja Lab GitHub page (https://github.com/Aneja-Lab-Yale/BM-Pathology-Dataset).
References
Sacks, P. & Rahman, M. Epidemiology of Brain Metastases. Neurosurgery Clinics of North America 31, 481–488 (2020).
Tabouret, E. et al. Recent Trends in Epidemiology of Brain Metastases: An Overview. Anticancer Research 32, 4655–4662 (2012).
Sperduto, P. W. et al. Graded Prognostic Assessment (GPA) for Patients With Lung Cancer and Brain Metastases: Initial Report of the Small Cell Lung Cancer GPA and Update of the Non-Small Cell Lung Cancer GPA Including the Effect of Programmed Death Ligand 1 and Other Prognostic Factors. International Journal of Radiation Oncology*Biology*Physics 114, 60–74 (2022).
Gondi, V. et al. Radiation Therapy for Brain Metastases: An ASTRO Clinical Practice Guideline. Pract Radiat Oncol 12, 265–282 (2022).
Aizer, A. A. et al. Brain metastases: A Society for Neuro-Oncology (SNO) consensus review on current management and future directions. Neuro Oncol 24, 1613–1646 (2022).
Vogelbaum, M. A. et al. Treatment for Brain Metastases: ASCO-SNO-ASTRO Guideline. JCO 40, 492–516 (2022).
Sperduto, P. W. et al. Survival in Patients With Brain Metastases: Summary Report on the Updated Diagnosis-Specific Graded Prognostic Assessment and Definition of the Eligibility Quotient. JCO 38, 3773–3784 (2020).
Brenner, A. W. & Patel, A. J. Review of Current Principles of the Diagnosis and Management of Brain Metastases. Front Oncol 12, 857622 (2022).
Fink, K. R. & Fink, J. R. Imaging of brain metastases. Surg Neurol Int 4, S209–S219 (2013).
Lignelli, A. & Khandji, A. G. Review of Imaging Techniques in the Diagnosis and Management of Brain Metastases. Neurosurgery Clinics 22, 15–25 (2011).
Kotecha, R. & Aneja, S. Opportunities for integration of artificial intelligence into stereotactic radiosurgery practice. Neuro Oncol 23, 1629–1630 (2021).
Familiar, A. M. et al. Towards consistency in pediatric brain tumor measurements: Challenges, solutions, and the role of artificial intelligence-based segmentation. Neuro-Oncology 26, 1557–1571 (2024).
Ramakrishnan, D. et al. Comparison of Volumetric and 2D Measurements and Longitudinal Trajectories in the Response Assessment of BRAF V600E-Mutant Pediatric Gliomas in the Pacific Pediatric Neuro-Oncology Consortium Clinical Trial. American Journal of Neuroradiology 45, 475–482 (2024).
von Reppert, M. et al. Comparison of volumetric and 2D-based response methods in the PNOC-001 pediatric low-grade glioma clinical trial. Neuro-Oncology Advances 6, vdad172 (2024).
Chang, E. et al. Comparison of radiomic feature aggregation methods for patients with multiple tumors. Sci Rep 11, 9758 (2021).
Aneja, S. & Omuro, A. Imaging biomarkers for brain metastases: more than meets the eye. Neuro Oncol 21, 1493–1494 (2019).
Bhatia, A. et al. MRI radiomic features are associated with survival in melanoma brain metastases treated with immune checkpoint inhibitors. Neuro Oncol 21, 1578–1586 (2019).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 278, 563–577 (2016).
Chen, B. T. et al. Predicting Survival Duration With MRI Radiomics of Brain Metastases From Non-small Cell Lung Cancer. Front Oncol 11, 621088 (2021).
Kniep, H. C. et al. Radiomics of Brain MRI: Utility in Prediction of Metastatic Tumor Type. Radiology 290, 479–487 (2019).
Lipkova, J. et al. Artificial intelligence for multimodal data integration in oncology. Cancer Cell 40, 1095–1110 (2022).
Waqas, A. et al. Digital pathology and multimodal learning on oncology data. BJR|Artificial Intelligence 1, ubae014 (2024).
Chadha, S. et al. MR Imaging and Segmentations with Matched Brain Biopsy Pathology Slides from Patients with Brain Metastases from Primary Lung Cancer (Brain-Mets-Lung-MRI-Path-Segs). The Cancer Imaging Archive https://doi.org/10.7937/k0sm-y874 (2025).
van Griethuysen, J. J. M. et al. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Research 77, e104–e107 (2017).
Zwanenburg, A. et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-based Phenotyping. Radiology 295, 328–338 (2020).
Aldawsari, A. M. et al. The role and potential of using quantitative MRI biomarkers for imaging guidance in brain cancer radiotherapy treatment planning: A systematic review. Phys Imaging Radiat Oncol 27, 100476 (2023).
Avery, E., Sanelli, P. C., Aboian, M. & Payabvash, S. Radiomics: A Primer on Processing Workflow and Analysis. Seminars in Ultrasound, CT and MRI 43, 142–146 (2022).
Aboian, M. et al. Development of a workflow efficient PACS based automated brain tumor segmentation and radiomic feature extraction for clinical implementation (N2.003). Neurology 98, (2022).
NEMA PS3 / ISO 12052. Digital Imaging and Communications in Medicine (DICOM) Standard.
Isensee, F. et al. Automated brain extraction of multisequence MRI using artificial neural networks. Hum Brain Mapp 40, 4952–4964 (2019).
Acknowledgements
The authors would like to thank the Yale School of Medicine Center for Outcomes Research and Evaluation, Department of Pathology, Department of Therapeutic Radiology, and Department of Radiology and Biomedical Imaging for providing the images and helping to make the data publicly available. Research reported in this publication was supported by the Richard K. Gershon Endowed Medical Student Research Fellowship and Yale School of Medicine Fellowship for Medical Student Research. The content is solely the responsibility of the authors and does not necessarily represent the official views of the Richard K. Gershon Endowed Medical Student Research Fellowship and Yale School of Medicine Fellowship for Medical Student Research.
Author information
Authors and Affiliations
Contributions
D.X.N. and S.A. conceived of the project, provided administrative support, supervision, resources, and acquired funding. S.C. curated the radiologic imaging, segmentations, and extracted radiomic features. D.D. and S.C. curated the pathologic imaging. D.D. and D.V.S. conducted the chart review. M.S.A., K.B. and M.L. provided technical resources. S.C. drafted the original manuscript. V.C. procured and provided access to brain tissue samples. All authors contributed to and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
D.X.N. has received research funding from AstraZeneca Inc. and honoraria from Daiichi Sankyo unrelated to this work. M.L. and K.B. are employees of Visage Imaging, and M.L. is a stockholder of Visage Imaging. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chadha, S., Sritharan, D.V., Dolezal, D. et al. Matched MRI, Segmentations, and Histopathologic Images of Brain Metastases from Primary Lung Cancer. Sci Data 13, 40 (2026). https://doi.org/10.1038/s41597-025-06353-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06353-2