Abstract
Craigia yunnanensis, endemic to East Asia, is an endangered species with important economic and scientific research values. However, the absence of a reference genome has hindered studies on genetic variation and conservation management of C. yunnanensis. To address this gap, we present a high-quality chromosome-level genome sequence of C. yunnanensis by using PacBio HiFi sequencing and Hi-C scaffolding. The genome has a total length of 1,618.96 Mb with scaffold N50 of 39.39 Mb and 98.00% of the genome assigned to 41 chromosomes. BUSCO assessment yielded a completeness score of 99.40%. Furthermore, we predicted 58,969 proteincoding genes, and 94.09% of them was functionally annotated. Assembly of the C. yunnanensis genome facilitates a deeper understanding of adaptive evolution in Craigia, knowledge that is fundamental to promoting the conservation and enabling evidence-based management of this endangered plant.
Similar content being viewed by others
Background & Summary
Craigia yunnanensis W. W. Sm. & W. E. Evans, a deciduous tree belonging to the family Malvaceae, is a species endemic to East Asia1. The species is primarily distributed in the limestone mountain forests of southern China and northern Vietnam, where it exists in small, fragmented wild populations2. Due to extensive deforestation, its natural habitat has been severely degraded and fragmented. C. yunnanensis is currently listed as a protected species and classified as “Vulnerable” on the IUCN Red List (https://www.iucnredlist.org), reflecting its high conservation value3.
The core objective of endangered species conservation is to preserve genetic variation, such as genetic diversity and population structure, which serves as a foundation for formulating effective conservation strategies4. Previous studies have employed molecular markers to investigate the population structure of C. yunnanensis5,6. With the advancement of high-throughput sequencing technologies, the assessment of genetic variation has entered a new era. Whole-genome sequencing not only yields more robust data on phylogenetic relationships, genetic diversity, and population structure, but also provides valuable insights into species origin, differentiation, historical effective population sizes, and adaptive evolution.
However, the lack of a high-quality reference genome for C. yunnanensis impedes research on its genetic variation and conservation management. Without a chromosome-level assembly, key genomic features (e.g., nuclear genes and structural variants) remain inaccessible, undermining phylogenetic resolution and evidence-based conservation strategies. Recently, the chloroplast genome of C. yunnanensis was sequenced to investigate its phylogenetic placement7. However, chloroplast genome sequence represents only a small fraction of the total genetic content, with the majority residing in the nuclear genome. Therefore, a high-quality, chromosome-level genome assembly is crucial for comprehensively elucidating the phylogenetic relationships within the Craigia genus and for advancing conservation efforts for the endangered C. yunnanensis.
In this study, we present a high-quality, chromosome-scale genome assembly of C. yunnanensis, generated by integrating PacBio HiFi sequencing (51.44 Gb), Illumina short-read sequencing (94.54 Gb), and Hi-C sequencing (176.00 Gb). The assembled genome spans approximately 1,618.96 Mb, with a contig N50 length of 34.28 Mb. Of the assembled sequences, 1,586.60 Mb (98.00%) were anchored to 41 pseudo-chromosomes, with a scaffold N50 length of 39.39 Mb. Repetitive elements constitute 71.58% of the genome, with long terminal repeats (LTRs) being the most abundant, accounting for 54.48%. A total of 58,969 protein-coding genes were annotated. This high-quality genome provides an essential genetic resource for understanding the adaptive strategies of C. yunnanensis.
Methods
Plant materials collection and genomic DNA extraction
The plant materials of C. yunnanensis used in this study were collected from a naturally growing wild population in Mangbang Township, Tengchong City, Yunnan Province, China (98.73°E, 25.03°N) (Fig. 1a,b). Fresh tissue samples, including roots, stems, and leaves, were harvested, immediately frozen in liquid nitrogen, and stored at −80 °C for subsequent DNA and RNA extraction. In parallel, young leaves and root tips were collected for karyotype analysis using imaging techniques (Fig. 2a). Genomic DNA was isolated from leaf tissue using the CTAB method. The concentration and purity of the extracted DNA were measured with a Qubit fluorometer (Thermo Fisher Scientific), and the integrity of the DNA was assessed through agarose gel electrophoresis.
Plant morphology and sampling locations of C. yunnanensis (a). Morphological characteristics of C. yunnanensis, (b) Location where the samples were collected.
Karyotype analysis and genome survey of C. yunnanensis. (a) Karyotype analysis of C. yunnanensis. (b) Estimation of genome size based on 21 K-mer distribution. (c) Ploidy inference using the Smudgeplot tool.
Illumina sequencing and genome survey analysis
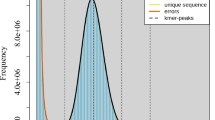
For Illumina sequencing, paired-end reads of 150 bp were generated on the NovaSeq platform. For RNA sequencing (RNA-seq), total RNA was isolated from all tissue samples using the NEBNext® Ultra™ II Directional RNA Library Prep Kit for Illumina® (New England Biolabs, MA, USA), after which paired-end 150-bp reads were also produced on the Illumina NovaSeq platform. To estimate the genome size of C. yunnanensis, a DNA library with a 150-bp insert size was constructed and sequenced by Annoroad Gene Technology Co., Ltd. (Beijing, China). After filtering low-quality reads, 94.54 Gb of clean data were used for the genome survey. The 21-mer frequency distribution was calculated using Jellyfish v2.2.98, and the genome size, along with the heterozygosity rate, was estimated using GenomeScope v2.09. Based on k-mer analysis, the genome size of C. yunnanensis was estimated to be 1,619.95 Mb, and the heterozygous ratio is 0.78% (Fig. 2b). Ploidy level was estimated using FastK v1.1 (https://github.com/thegenemyers/FASTK) to analyze k-mer distributions, followed by Smudgeplots v0.5.0 (https://github.com/KamilSJaron/smudgeplot). Based on the k-mer profile patterns, C. yunnanensis was confirmed to be diploid (Fig. 2c).
HiFi sequencing and draft genome assembly
For PacBio HiFi sequencing, a SMRTbell library was prepared with the SMRTbell Express Template Prep Kit 2.0 according to the manufacturer’s protocol (Pacific Biosciences, CA, USA). Sequencing was subsequently carried out on the PacBio Sequel II platform. After filtering low-quality reads, 51.44 Gb of clean data were used for genome assembly. De novo assembly of the HiFi reads was performed using Hifiasm v0.15.4 with the parameter ‘-l 3’10. The draft genome assembly resulted in a total size of 1.61 Gb, comprising 474 contigs with a contig N50 of 34.28 Mb (Table 1). This assembly length is consistent with the estimated genome size of 1,619.95 Mb obtained from k-mer analysis.
Hi-C sequencing and chromosome-scale assembly
For Hi-C sequencing, genomic DNA was first cross-linked with formaldehyde, followed by extraction and library construction. Sequencing was performed on the Illumina NovaSeq platform (Illumina, CA, USA) using a paired-end strategy. All Hi-C experiments were carried out by Annoroad Gene Technology Co., Ltd. (Beijing, China). To achieve chromosome-level genome assembly, clean Hi-C reads were initially mapped to the C. yunnanensis contig assembly using Juicer v1.6.11. The preliminary positioning and orientation of contigs were inferred using 3D-DNA v20100812. To minimize assembly errors, the initial 3D-DNA output was refined with Juicerbox v2.20.0013, which corrected misjoins and optimized contig clustering. The corrected contigs were then anchored into pseudo-chromosomes using 3D-DNA, and the Hi-C contact map was visualized using HiCExplorer v3.7.414. As a result, Hi-C data enabled the anchoring of 1,586.60 Mb (98.00%) of the assembled sequences onto 41 pseudo-chromosomes, which ranged in size from 23.53 Mb to 58.67 Mb (Table 1 and Fig. 3a,b).
Hi-C interaction, and genome assembly features of C. yunnanensis. (a) the Hi-C interaction heatmap for 41 pseudochromosomes in the C. yunnanensis genome. (b) the correlation between genome assembly and physical chromosome length. (c) genomic features of C. yunnanensis. Circus plot from the outer to the inner circle represents chromosome-scale pseudochromosomes from the inner to the outer circles, the order are: gene collinearity (connected by curved lines), GC content, gene density, TE density and LTR density.
Genome annotation
A combination of homology-based and de novo approaches was employed to identify transposable elements (TEs) and annotate genes in the C. yunnanensis genome. For TE identification, a de novo repeat library was constructed using RepeatModeler v2.0.615 with the ‘-LTRStruct’ option. This library was then integrated with homology-based repeat libraries from the Repbase database v2018102616 database and the Dfam database v3.717, and the combined dataset was used to identify repetitive sequences with RepeatMasker v4.0.915. By integrating both homology-based and de novo predictions, approximately 1,135.63 Mb (71.58%) of the genome was identified as repetitive sequences (Tables 1, 2). Among all repeat elements, long terminal repeat (LTR) retrotransposons were the most prevalent, representing 54.48% of the genome, followed by DNA transposons, which constituted 13.82%.
Protein-coding gene annotation combined homology-based evidence, RNA-seq data, and ab initio predictions. Homology-based gene models were generated using GeMoMa v1.918 with Arabidopsis thaliana (https://www.arabidopsis.org), Bombax ceiba19, Theobroma cacao20 and Durio zibethinus21 protein sequences. These models trained three ab initio gene predictors: Augustus v3.5.022 and GENEMARK-ES v4.7223. Simultaneously, a de novo transcriptome assembly was generated using Hisat2 v2.2.124, StringTie v2.2.325 and TransDecoder v5.7.1 (https://github.com/TransDecoder/TransDecoder). The trained predictors, homology-based evidence, and transcriptome assembly were then integrated using the EvidenceModeler v2.1.026 pipeline. This workflow ensured the accurate identification of transposable elements and comprehensive annotation of protein-coding genes in the C. yunnanensis genome. Gene identification and annotation in C. yunnanensis was performed using a combination of de novo, homology-based, and RNA-seq-based predictions, resulting in a total of 58,969 protein-coding genes being predicted (Table 3). Of the predicted genes, a total of 55,485 genes (94.09%) were simultaneously annotated in either of these databases, including nonredundant protein sequence (NR, https://www.ncbi.nlm.nih.gov/refseq/about/nonredundantproteins), Uniprot27, Pfam28, Eukaryotic orthologous groups of proteins (KOG)29, Gene Ontology (GO)30, and the Kyoto Encyclopedia of Genes and Genomes (KEGG)31 (Fig. 4a).
Evaluation of the genome assembly and annotation of C. yunnanensis. (a) the annotation of proteome of C. yunnanensis. (b) the BUSCO of contigs level and chromosome level of genome and proteome of C. yunnanensis.
Non-coding RNAs, including miRNA, rRNA, and snRNA genes, were detected using infernal v1.1.532 by searching the Rfam database, while tRNA genes were predicted using tRNAscan-SE v2.0.933. In addition to protein-coding genes, non-coding RNAs were also identified, including 372 miRNAs, 999 tRNAs, 359 rRNAs, and 6,330 snRNAs (Table 4).
Data Records
All raw sequencing data generated in this study were deposited in the Sequence Read Archive (SRA) database of the National Center for Biotechnology Information (NCBI), with the corresponding BioProject accession number PRJNA1327616. The HiFi, Hi-C, Illumina and RNA-seq sequencing datasets were archived under accession number (SRR3538602434, SRR3621621835, SRR3537130836, SRR3636973337). The genome assembly was submitted to GenBank with the accession number GCA_054051545.138. Both the genome assembly and its annotation are publicly accessible on Figshare via the following link: https://doi.org/10.6084/m9.figshare.3007572739.
Technical Validation
The somatic cells of C. yunnanensis are diploid and contain 41 pairs of chromosomes, including 16 metacentric, 44 short submetacentric, and 22 subtelocentric chromosomes. To verify the assembly, we compared the physical lengths of the 41 chromosome pairs in somatic cells with the lengths from the assembly. The results showed a high level of agreement, with a correlation coefficient of 0.99 and a p-value of 7.55e-35, suggesting a very strong positive correlation (Fig. 3b).
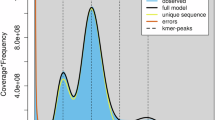
The quality of the C. yunnanensis genome assembly was evaluated to assess its continuity, completeness, and accuracy. To verify the accuracy of the assembly, both HiFi and Illumina sequencing reads were utilized. The HiFi reads were aligned to the assembly using Minimap2 v2.2840, while Illumina reads were mapped with BWA v0.7.1841. The mapping rate was calculated using Samtools v1.1742. The assembly showed mapping coverages of 99.64% for NGS and 99.91% for PacBio HiFi reads (Table 5). These sequencing reads were effectively mapped to the C. yunnanensis genome, further confirming the assembly’s accuracy and completeness. The completeness was assessed using BUSCO v5.4.343 with the ‘embryophyta_odb10’ database, revealing that 99.4% of core genes were complete, with only 0.3% missing (Fig. 4b). Additionally, the assembly’s accuracy was evaluated using Merqury v1.344 with a k-mer size of 20, resulting in an average quality value (QV) of 54.22. These findings collectively indicate that the C. yunnanensis genome assembly possesses high continuity, completeness, and accuracy.
In comparison with the 57,219 genes reported in Zhang’s assembly45, our version identified 58,969 protein-coding genes. Assessment with BUSCO v5.4.3 using the ‘embryophyta_odb10’ database indicated that 98.5% of conserved orthologous genes were present, including 43.4% single-copy and 55.1% duplicated genes (Fig. 4b). Furthermore, 55,485 genes were successfully annotated across six databases: NR, SwissProt, KEGG, KOG, GO, and Pfam (Fig. 4a). Relative to the recently published monoploid genome of C. yunnanensis, our assembly displayed comparable quality, with a contig N50 of about 34 Mb and a scaffold N50 of approximately 40 Mb (Table 6).
BLASTP46 was adopted to identify homologous genes between and within two genomes (diploid genome assembly and the autotetraploid haplotype), while syntenic blocks were delineated by JCVI47 based on these homologs. The synonymous substitution rates (Ks) of both paralogous and orthologous gene pairs were calculated using WGDI48, with JCVI further deployed to generate syntenic depth ratios and synteny dot plots. Our data revealed that the Ks peak of paralogous gene pairs reached approximately 1.5 in both genome assemblies (Fig. 5a,b). Notably, the Ks value of orthologous gene pairs also approximated 1.5 (Fig. 5c), showing no significant difference between the two genomes. This result indicated a high level of genomic homology between the diploid genome assembly and the haplotype of the autotetraploid genome, which belong to one species. Additionally, a 1:1 syntenic depth ratio was detected between the two assemblies (Fig. 5d), consistent with the observation of strict one-to-one chromosomal homology across all 41 chromosomes of both assemblies (Fig. 5e). Collectively, these syntenic analyses provided compelling evidence that the autotetraploid haplotype is highly congruent with the diploid genome assembly.
Comparison of C. yunnanensis neoautotetraploid and C. yunnanensis diploid (this study). (a) Ks peak of paralogous gene pairs in diploid genome assembly (this study). (b) Ks peak of paralogous gene pairs in autotetraploid haplotype genome assembly. (c) Ks peak of orthologous gene pairs between diploid genome assembly and the autotetraploid haplotype genome assembly. (d) syntenic depth ratio between diploid genome assembly and the autotetraploid haplotype genome assembly. (e) Syntenic dotplot illustrating the comparative analysis of the diploid genome assembly and the autotetraploid haplotype genome assembly.
Data availability
The raw sequencing data of C. yunnanensis have been deposited in the National Center for Biotechnology Information (NCBI) under the BioProject accession number PRJNA1327616 (SRR3538602434, SRR3621621835, SRR3537130836, SRR3636973337). The genome assembly was submitted to GenBank with the accession number GCA_054051545.138. Additionally, the genome assembly data of this species have been archived in Figshare and are accessible via the following persistent link: https://doi.org/10.6084/m9.figshare.3007572739.
Code availability
This study does not involve custom scripts or code. The software and code used are publicly accessible.
References
Wang, B. et al. A new occurrence of Craigia (Malvaceae) from the Miocene of Yunnan and its biogeographic significance. Historical Biology 33, 3402–3412, https://doi.org/10.1080/08912963.2020.1867980 (2021).
Gao, Z., Zhang, C. & Milne, R. I. Size-class structure and variation in seed and seedling traits in relation to population size of an endangered species Craigia yunnanensis (Tiliaceae). Australian Journal of Botany 58, 214–223 (2010).
de Kok, R. Craigia yunnanensis. The IUCN Red List of Threatened Species 2024, e.T32335A2815412 (2024).
Frankham, R. Challenges and opportunities of genetic approaches to biological conservation. Biological Conservation 143, 1919–1927, https://doi.org/10.1016/j.biocon.2010.05.011 (2010).
Yang, J., Gao, Z., Sun, W. & Zhang, C. High regional genetic differentiation of an endangered relict plant Craigia yunnanensis and implications for its conservation. Plant Diversity 38, 221–226, https://doi.org/10.1016/j.pld.2016.07.002 (2016).
Chen, Y. L., Yang, J. & Sun, W. B. Development of 14 microsatellite markers in the endangered relict plant Craigia yunnanensis (Tiliaceae). Russian Journal of Genetics 56, 123–127, https://doi.org/10.1134/S1022795420010032 (2020).
Wariss, H. M., Yaling, C. & Yang, J. The complete chloroplast genome of Craigia yunnanensis, an endangered plant species with extremely small populations (PSESP) from South China. Mitochondrial DNA Part B 4, 2740–2741, https://doi.org/10.1080/23802359.2019.1644228 (2019).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204, https://doi.org/10.1093/bioinformatics/btx153 (2017).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nature Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Systems 3, 95–98, https://doi.org/10.1016/j.cels.2016.07.002 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Systems 3, 99–101, https://doi.org/10.1016/j.cels.2015.07.012 (2016).
Wolff, J. et al. Galaxy HiCExplorer 3: a web server for reproducible Hi-C, capture Hi-C and single-cell Hi-C data analysis, quality control and visualization. Nucleic acids research 48, W177–w184, https://doi.org/10.1093/nar/gkaa220 (2020).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Current Protocols in Bioinformatics Chapter 4, 4.10.11–14.10.14, https://doi.org/10.1002/0471250953.bi0410s25 (2009).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA 6, 11, https://doi.org/10.1186/s13100-015-0041-9 (2015).
Wheeler, T. J. et al. Dfam: a database of repetitive DNA based on profile hidden Markov models. Nucleic acids research 41, D70–82, https://doi.org/10.1093/nar/gks1265 (2013).
Keilwagen, J., Hartung, F. & Grau, J. GeMoMa: Homology-based gene prediction utilizing intron position conservation and RNA-seq data. Methods Molecular Biology 1962, 161–177, https://doi.org/10.1007/978-1-4939-9173-0_9 (2019).
Shao, L. et al. High-quality genomes of Bombax ceiba and Ceiba pentandra provide insights into the evolution of Malvaceae species and differences in their natural fiber development. Plant Communications 5, 100832, https://doi.org/10.1016/j.xplc.2024.100832 (2024).
Argout, X. et al. The genome of Theobroma cacao. Nature Genetics 43, 101–108, https://doi.org/10.1038/ng.736 (2011).
Li, W., Chen, X., Yu, J. & Zhu, Y. Upgraded durian genome reveals the role of chromosome reshuffling during ancestral karyotype evolution, lignin biosynthesis regulation, and stress tolerance. Science China Life Sciences 67, 1266–1279, https://doi.org/10.1007/s11427-024-2580-3 (2024).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic acids research 32, W309–312, https://doi.org/10.1093/nar/gkh379 (2004).
Borodovsky, M. & Lomsadze, A. Eukaryotic gene prediction using GeneMark.hmm-E and GeneMark-ES. Current Protocols in Bioinformatics Chapter 4, 4.6.1–4.6.10, https://doi.org/10.1002/0471250953.bi0406s35 (2011).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nature Biotechnology 37, 907–915, https://doi.org/10.1038/s41587-019-0201-4 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature Biotechnology 33, 290–295, https://doi.org/10.1038/nbt.3122 (2015).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biology 9, R7, https://doi.org/10.1186/gb-2008-9-1-r7 (2008).
UniProt. the universal protein knowledgebase. Nucleic Acids Research 45, D158–169, https://doi.org/10.1093/nar/gkw1099 (2017).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Research 40, D290–301, https://doi.org/10.1093/nar/gkr1065 (2012).
Tatusov, R. L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4, 41, https://doi.org/10.1186/1471-2105-4-41 (2003).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genetics 25, 25–29, https://doi.org/10.1038/75556 (2000).
Tanabe, M. & Kanehisa, M. Using the KEGG database resource. Current Protocols in Bioinformatics Chapter 1, 1.12.11–11.12.43, https://doi.org/10.1002/0471250953.bi0112s38 (2012).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935, https://doi.org/10.1093/bioinformatics/btt509 (2013).
Chan, P. P., Lin, B. Y., Mak, A. J. & Lowe, T. M. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Research 49, 9077–9096, https://doi.org/10.1093/nar/gkab688 (2021).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR35386024 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36216218 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR35371308 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR36369733 (2025).
NCBI GenBank https://identifiers.org/ncbi/insdc.gca:GCA_054051545.1 (2025).
Cheng, Z. & Xing, Y. Y. A chromosome-level reference genome of an endangered plant Craigia yunnanensis. Figshare https://doi.org/10.6084/m9.figshare.30075727 (2025).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760, https://doi.org/10.1093/bioinformatics/btp324 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, https://doi.org/10.1093/gigascience/giab008 (2021).
Seppey, M., Manni, M. & Zdobnov, E. M. BUSCO: Assessing genome assembly and annotation completeness. Methods Molecular Biology 1962, 227–245, https://doi.org/10.1007/978-1-4939-9173-0_14 (2019).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biology 21, 245, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Zhang, R. G. et al. Reticulate allopolyploidy and subsequent dysploidy drive evolution and diversification in the cotton family. Nature Communications 16, 7480, https://doi.org/10.1038/s41467-025-62644-7 (2025).
Al-Fatlawi, A., Menzel, M. & Schroeder, M. Is Protein BLAST a thing of the past? Nat Commun 14, 8195, https://doi.org/10.1038/s41467-023-44082-5 (2023).
Tang, H. et al. JCVI: A versatile toolkit for comparative genomics analysis. Imeta. 3(4), e211 (2024).
Sun, P. et al. WGDI: A user-friendly toolkit for evolutionary analyses of whole-genome duplications and ancestral karyotypes. Mol Plant. 15(12), 1841–1851, https://doi.org/10.1016/j.molp.2022.10.018 (2022).
Acknowledgements
This research was supported by grants from the Yunnan Provincial Baoshan Administration of Gaoligongshan National Nature Reserve (202305AF150121 & GBP-2022-01), the National Natural Science Foundation of China (32370407, 31761143001 & 31870316).
Author information
Authors and Affiliations
Contributions
C.L.L., Z.L. and F.F.X. conceived the project. J.H.L. and C.L.X. collected the samples and coordinated the sequencing. Z.C., Y.Y.X., Y.M.P., J.W., X.X. W. and R.A.X. carried out the analysis. Z.C. and Y.Y.X. wrote and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, Z., Xing, Y., Pan, Y. et al. A chromosome-level reference genome of an endangered plant Craigia yunnanensis. Sci Data 13, 567 (2026). https://doi.org/10.1038/s41597-026-06746-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06746-x