Abstract
Solemyidae, an ancient lineage of protobranch bivalves, are characterized by unique morphology and obligate symbiosis with sulfur-oxidizing bacteria, enabling survival in sulfide-rich sediments. However, limited genomic resources have hindered understanding of their evolutionary history, symbiotic interactions, and environmental adaptation. Here, we report a chromosome-level reference genome of Acharax haimaensis, assembled using PacBio, Illumina, and Hi-C sequencing. The 4.27 Gb genome, with a scaffold N50 of 195.52 Mb, was anchored to 22 chromosomes and achieved high completeness (98.2%) based on BUSCO. Transposable elements occupy 50.17% of the assembly, dominated by long interspersed nuclear elements (14.20%). We predicted 38,343 protein-coding genes, of which 87.25% were functionally annotated. Macrosynteny analysis revealed each chromosome comprises two to four segments of ancestral linkage groups, indicating extensive chromosomal breakage and fusion in early bivalve evolution. Phylogenetic inference suggested A. haimaensis diverged from the common ancestor of Autobranchia ~550 Mya. This first deep-sea protobranch genome provides an essential resource for exploring bivalve evolution and the genetic basis of symbiosis and adaptation to extreme environments.
Similar content being viewed by others
Background & Summary
Protobranchia constitute a relatively understudied yet phylogenetically primitive subclass of bivalve molluscs, offering valuable insights into early bivalve evolution and ecological diversification1,2. Within this group, the Solemyidae represent one of the most ancient lineages, with fossil evidence tracing their origins back to the Early Ordovician3. As among the earliest-diverging bivalve lineages, solemyids may have retained ancestral traits that were subsequently lost in other clades, thereby providing a critical window into the origin and evolutionary history of Bivalvia. However, genomic resources for Solemyidae remain extremely limited; to date, only the genome of the shallow-water solemyid Solemya velum has been recently published, severely constraining our understanding of this lineage’s evolutionary trajectory4.
Morphologically, solemyids are characterized by a distinctive thickened frill of radially pleated periostracum that extends beyond the calcified shell margins5. Their valves are equilateral, anteriorly elongate, and subcylindrical, with generally parallel dorsal and ventral margins and an edentate hinge, the latter representing an early apomorphic trait6. The family Solemyidae comprises approximately 30 described extant species belonging to the genera Solemya Lamarck, 1818 and Acharax Dall, 19085. Species of Solemya typically inhabit anaerobic muddy and sandy substrates on continental shelves and upper slopes at depths of 0–600 m7, whereas members of Acharax are primarily restricted to deep-sea chemosynthetic ecosystems, including cold seeps and hydrothermal vents, at depths of 400–5379 m8,9. Consequently, Acharax species are rarely collected alive, and knowledge of their phylogeny, diversity, and genomic architecture remains scarce.
The genus Acharax currently comprises approximately 20 fossil species and nine extant species10,11. Recent morphological examinations combined with molecular phylogenetic analyses led to the identification of Acharax haimaensis as a new species from the Haima cold seep in the South China Sea12. Over long evolutionary timescales, A. haimaensis has developed a suite of adaptive strategies that enable survival in extreme deep-sea habitats characterized by hypoxia, low temperatures, and high hydrostatic pressure. A key feature of this adaptation is its obligate symbiosis with chemoautotrophic, gill-associated sulfur-oxidizing bacteria, which provide an alternative carbon source in nutrient-limited environments12. These microbial partners fulfill dual ecological roles: they sustain primary production within the host and facilitate detoxification and metabolic flexibility in sulfide-rich, reducing conditions13. Due to this unique symbiotic association and its specialization to extreme environments, A. haimaensis represents a valuable model system for elucidating the ecological, physiological, and genomic mechanisms underlying adaptation to extreme environments in deep-sea organisms. Despite its recognized evolutionary and ecological significance, the paucity of genomic resources remains a major obstacle to elucidating the origin and evolution of A. haimaensis, its metabolic interactions with endosymbionts, and the molecular mechanisms underlying deep-sea adaptation.
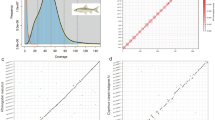
Here, we report a high-quality chromosome-level reference genome of A. haimaensis, generated using PacBio HiFi long reads, Illumina short reads, and Hi-C scaffolding (Table 1). The genome size of A. haimaensis was estimated at ~4.0 Gb using Jellyfish v2.3.1, with a heterozygosity of 1.47%. The initial assembly comprised 2,866 contigs with a contig N50 of 9.45 Mb (Table 2). Following Hi-C scaffolding, 91.57% of contigs were anchored to 22 chromosomes (Fig. 1; Table 3), yielding a final assembly of 4.27 Gb with a contig N50 of 10.53 Mb and a scaffold N50 of 195.52 Mb (Tables 2 and 3). The mapping rate of illumina short reads to the assembly was 99.34%. Genome completeness was further supported by the identification of 937 out of 954 metazoan Benchmarking Universal Single-Copy Orthologs (BUSCOs), corresponding to 98.2% completeness (Table 2). Repetitive element annotation revealed that transposable elements (TEs) account for 50.17% of the genome, with LINEs, SINEs, LTRs, and DNA transposons contributing 14.20%, 0.35%, 5.91%, and 8.56%, respectively (Table 4). We predicted 38,343 protein-coding genes, of which 87.25% could be functionally annotated against at least one public database (Table 5). In addition, 35,928 tRNAs, 18,341 rRNAs, 3,972 snRNAs, and 81 miRNAs were annotated (Table 5). Macrosynteny analysis showed that each chromosome of A. haimaensis comprises segments derived from two to four ancestral linkage groups (ALGs) of bilaterians, cnidarians, and sponges (BCnS), indicating extensive chromosomal breakage and fusion during its evolutionary history (Fig. 2). Phylogenetic analysis revealed that A. haimaensis diverged from the common ancestor of Autobranchia ~550 Mya, underscoring its basal position and primitive evolutionary status within Bivalvia (Fig. 3).
Genomic characteristics of A. haimaensis. (A) Hi-C contact heatmap of chromosome interactions. The scale bar indicates interaction intensity from yellow (low) to red (high). (B) Circos view of the assembled chromosomes showing marker distributions at 1-Mb sliding windows from outer to inner circle: (a) chromosome length, (b) GC content, (c) gene density, (d) long interspersed nuclear element (LINE) density, (e) short interspersed nuclear element (SINE) density, (f) long terminal repeat (LTR) density, and (g) DNA transposon density.
Chromosome-level macrosynteny between A. haimaensis and ancestral linkage groups (ALGs) of bilaterians, cnidarians, and sponges (BCnS). Distinct colors represent individual ALGs. Colored dots denote genes with statistically significant conserved synteny (P < 0.05), whereas black dots indicate non-significant synteny.
A phylogenetic tree was constructed using the single-copy orthologs identified from A. haimaensis and other 21 bivalve species.
Methods
Sampling and sequencing
Individuals of A. haimaensis were collected from the Haima cold seep in the South China Sea at a depth of 1,375 m (16°43′43″ N, 110°28′21″ E) using a TV grab. Adductor muscle tissue from one individual was aseptically dissected, immediately flash-frozen in liquid nitrogen, and stored at –80 °C for subsequent Illumina, PacBio, and Hi-C sequencing. High-molecular-weight genomic DNA was extracted using the SDS method and further purified with a QIAGEN Genomic Kit (QIAGEN, Germany). DNA integrity and contamination were assessed by 0.75% agarose gel electrophoresis, while DNA purity was evaluated using a NanoDrop spectrophotometer (Thermo Fisher Scientific, USA). High-quality DNA was then used for library preparation and sequencing. For Illumina sequencing, a short-insert library was first constructed to estimate genome complexity. Approximately 2 μg of genomic DNA was fragmented to an average size of 350 bp via ultrasonication, and the resulting library was sequenced on the Illumina NovaSeq 6000 platform (Illumina, USA), yielding 221.71 Gb of 150 bp paired-end reads with an coverage of ~52 × (Table 1). For PacBio HiFi long-read sequencing, 10 μg of genomic DNA was sheared into 10–20 kb fragments using a g-TUBE (Covaris, USA). The fragments were subjected to damage repair, end polishing, and ligation with stem–loop adapters. Unligated DNA was removed by exonuclease treatment, and target fragments were size-selected using a BluePippin system (Sage Science, USA). Sequencing on the PacBio Sequel II platform in CCS mode produced seven SMRT cells of data, generating 143.67 Gb of clean HiFi reads (~34 × coverage) (Table 1). For Hi-C library construction, muscle cells from another individual were crosslinked with 1% formaldehyde and quenched with 0.2 M glycine. Crosslinked chromatin was digested with MboI restriction enzyme, end-repaired, and biotin-labeled. Proximity-ligated chimeric DNA fragments were circularized with T4 DNA ligase, purified, and sheared into smaller fragments. Biotin-labeled fragments were enriched using streptavidin-coated magnetic beads (Invitrogen, USA), and the resulting Hi-C library was sequenced on the Illumina NovaSeq 6000 platform, producing 230.89 Gb of clean reads with ~54 × coverage (Table 1).
RNA-seq was performed to facilitate genome annotation. Total RNA was extracted from gill, foot, and mantle tissues using the TRIzol reagent (Invitrogen, USA). RNA integrity was evaluated by 1% agarose gel electrophoresis, while RNA purity and concentration were assessed with a NanoDrop spectrophotometer (Thermo Fisher Scientific, USA). Poly(A)+ mRNA was enriched using oligo(dT)-attached magnetic beads, and first-strand cDNA was synthesized with M-MuLV reverse transcriptase. Library construction was carried out using Phusion High-Fidelity DNA Polymerase. Sequencing was performed on the Illumina NovaSeq platform, generating a total of 37.20 Gb of paired-end clean reads with an average Q30 value of 93.25% (Table 1).
Genome assembly and Hi-C scaffolding
To estimate genome size, heterozygosity, and repeat content, K-mer analysis was performed using Jellyfish v2.3.1 with k = 2114. The resulting K-mer frequency distribution was analyzed using GenomeScope v2.0.1 with the parameters -m 17 -p 215. The genome size of A. haimaensis was estimated to be approximately 4.0 Gb, with a heterozygosity of 1.47%. Based on PacBio HiFi reads, a contig-level assembly was generated using Hifiasm v0.19.4 with default parameters16. HiFi reads were aligned to the draft assembly with Minimap2 v2.2417, and redundant sequences were removed using Purge_Dups v1.2.3, followed by haplotype purging with Purge Haplotigs v1.1.2 (https://github.com/dfguan/purge_dups). The initial assembly consisted of 2,866 contigs with a contig N50 of 9.45 Mb (Table 2). For chromosome-level scaffolding, raw Hi-C reads were first filtered using HiC-Pro v3.1.0 and processed with Juicer v1.618,19. Contigs were then assembled into pseudo-chromosomes using 3D-DNA v20100820. Contact maps were visualized and manually curated with Juicebox v1.11.08, during which misassemblies were corrected and chromosomal boundaries refined based on Hi-C interaction heatmaps19. A final chromosome-level assembly was produced by re-running 3D-DNA after manual correction. Following Hi-C scaffolding, 97.08% of the reads were anchored to 22 chromosomes (Fig. 1; Table 3), resulting in a 4.27 Gb assembly with a scaffold N50 of 195.52 Mb (Table 2). Assembly completeness was assessed using BUSCO v5.7.1 with the metazoan odb10 dataset, which showed a high assembly completeness of 98.2% (Table 2)21.
Repeat and gene annotation
Transposable elements (TEs) were identified using a combination of de novo and homology-based approaches. A de novo repeat library was constructed with MITE-Tracker and RepeatModeler v2.0.1 under default parameters22,23. Long terminal repeat (LTR) elements were detected with LTR_Finder v1.07 and LTR_retriever v2.9.024,25. The resulting de novo repeat library and LTR dataset were merged to generate a comprehensive repeat library. For homology-based detection, RepeatMasker v4.1 was applied to screen the genome against both the Dfam/Repbase databases and the custom repeat library26,27. Gene prediction was performed on the repeat-masked genome by integrating de novo, homology-based, and transcriptome-based strategies. For homology-based annotation, BLAST v2.16.0 was used to align protein sequences from representative bivalves (Archivesica marissinica, Mercenaria mercenaria, Sinonovacula constricta, Tegillarca granosa, Gigantidas platifrons, and Patinopecten yessoensis), and gene structures were subsequently inferred using GeneWise v2.4.128. RNA-seq reads were aligned to the genome using HISAT2 v2.2.129, and StringTie v2.2.130 was used to reconstruct transcripts to provide transcript-based evidence for genome annotation. Assembled transcripts were further integrated with PASA v2.5.231. De novo gene prediction was conducted with AUGUSTUS v3.5.0 and BRAKER2 v2.1.632,33. The final consensus gene set was generated using EvidenceModeler v2.1.034, and untranslated regions as well as alternative splicing events were annotated with GUSHR and GeMoMa based on RNA-seq alignments35. In total, 38,343 protein-coding genes were predicted from the A. haimaensis genome (Table 5). The gene set exhibited a BUSCO completeness score of 96.0% (Table 2). Functional annotation revealed that 33,458 (87.25%) of the predicted genes matched entries in at least one major public database, including Swiss-Prot, TrEMBL, NR, Pfam, KEGG, GO, and COG/KOG (Table 5). Additionally, non-coding RNAs were annotated: transfer RNAs (tRNAs) were identified using tRNAscan-SE v1.3.136, microRNAs (miRNAs) and small nuclear RNAs (snRNAs) were predicted using Infernal with the Rfam database37,38, and ribosomal RNAs (rRNAs) were detected using RNAmmer and Barrnap39.
Data Records
The raw Illumina, PacBio, Hi-C sequencing, and RNA-seq data of Acharax haimaensis were deposited in the NCBI SRA under the accession number SRP61790140. Specifically, Illumina sequencing data are available under accession number SRX30378101; PacBio sequencing data under accession numbers SRX30378102, SRX30378104, SRX30378094, SRX30378095, SRX30378096, and SRX30378097; Hi-C sequencing data under accession number SRX30378098; and RNA-seq data under accession numbers SRX30378099, SRX30378100, and SRX30378103. The chromosome-level genome assembly has been deposited in the NCBI GenBank database under accession number GCA_054131465.141. In addition, the assembled genome sequence and genome annotation files are publicly available at Figshare42.
Technical Validation
Evaluating genome assembly and annotation completeness
The DNA fragments used for PacBio sequencing were predominantly distributed around 16.82 Kb, with an N50 read length of 17.54 Kb (Table 1). The assembled genome size of A. haimaensis was 4.27 Gb, consistent with the estimate from Jellyfish v2.3.1 (~4.0 Gb)14. The assembly achieved a quality value of 62.28, as calculated with Merqury v1.34743, reflecting high assembly accuracy. The initial assembly comprised 2,866 contigs with a contig N50 of 9.45 Mb (Table 2). The 19.87% of Hi-C reads were deemed valid, and after scaffolding, 91.57% (239/261) of contigs were successfully anchored to 22 chromosomes (Table 3). The illumina short reads were mapped to the final assembly using BWA v0.7.1944, with mapping statistics calculated by samtools flagstat v1.22.145, yielding a mapping rate of 99.34%. RNA-seq reads and PacBio HiFi reads were further aligned to the assembly using HISAT2 and Minimap2, respectively17,29, with mapping rates of 81.47% and 99.91%. Assembly completeness was evaluated using BUSCO v5.7.1 with the metazoan odb10 dataset21, which recovered 937 of 954 single-copy orthologs, corresponding to 98.2% completeness. Gene set quality was also assessed with BUSCO, yielding a completeness score of 96.0%, consistent with that of the assembly (Table 2). Collectively, these results demonstrate that the A. haimaensis genome assembly and annotation are of high quality and completeness.
Macrosynteny analysis
Oxford Dot Plot (ODP) software (https://github.com/conchoecia/odp?tab=readme-ov-file) was employed to investigate chromosomal-level synteny between A. haimaensis and BCnS ALGs46. Macrosyntenic relationships were visualized in ODP, with each dot representing an orthologous gene pair. The analysis revealed relatively weak syntenic conservation between A. haimaensis and BCnS ALGs. Most ALGs were fragmented into two or more parts, each mapping to different chromosomes of A. haimaensis. Each A. haimaensis chromosome comprised segments derived from two to four ALGs, indicating extensive chromosomal breakage and fusion during its evolutionary history (Fig. 2).
Phylogenetic analysis
A phylogenetic tree was constructed using A. haimaensis together with 22 additional bivalve species, with the chiton Acanthopleura granulata designated as the outgroup. Coding gene and protein sequences were retrieved from the MolluscDB 2.0 database (http://mgbase.qnlm.ac/home). Orthologous genes were identified with OrthoFinder v2.5.5 under default parameters47, and the single-copy orthologous groups (OGs) were selected for phylogenetic reconstruction. Amino acid sequences were aligned using MUSCLE v3.8.1551, and conserved regions were extracted with Gblocks v0.91b under the parameters -b4 = 5 -b5 = h -t = p -e = .248. A maximum-likelihood phylogenetic tree was constructed using IQ-TREE v2.3.0 under the insect + I + R4 substitution model, with nodal support assessed using 1,000 bootstrap replicates49. Divergence times were estimated with the MCMCtree module in PAML v4.10.750, calibrated with fossil records at the following nodes: G. platifrons–Modiolus philippinarum (130–139 Mya), Venerida–Mytilida (230–524 Mya), and A. granulata–Bivalvia (480–559 Mya) (https://timetree.org/). As shown in Fig. 3, A. haimaensis was placed as a basal lineage within Protobranchia, diverging early from the ancestral node, suggesting its primitive evolutionary status among bivalve molluscs.
Data availability
The raw sequencing data of A. haimaensis were deposited into NCBI SRA database under the accession number SRP61790140. Illumina sequencing data are available under accession number SRX30378101. PacBio sequencing data are deposited under accession numbers SRX30378102, SRX30378104, SRX30378094, SRX30378095, SRX30378096, and SRX30378097. Hi-C sequencing data are available under accession number SRX30378098, and RNA-seq data are available under accession numbers SRX30378099, SRX30378100, and SRX30378103. The genome assembly was deposited in the NCBI Genbank under accession number GCA_054131465.141. Moreover, the assembled genome sequence and annotation files are accessible at Figshare42.
Code availability
No custom script was used in this work. Software that was used to analyse data was listed in methods in detail and commands were used based on the manuals.
References
Allen, J. & Sanders, H. J. P. i. O. The zoogeography, diversity and origin of the deep-sea protobranch bivalves of. the Atlantic: the epilogue 38, 95–153 (1996).
Sharma, P. P. et al. Into the deep: A phylogenetic approach to the bivalve. subclass Protobranchia. Molecular Phylogenetics and Evolution 69, 188–204, https://doi.org/10.1016/j.ympev.2013.05.018 (2013).
Cope, J. C. W. Early Ordovician (Arenig) bivalves from the Llangynog Inlier, south Wales. Palaeontology 39, 979–1025 (1996).
Chen, Z. et al. A genome-based phylogeny for Mollusca is concordant with fossils and morphology. Science 387, 1001–1007, https://doi.org/10.1126/science.ads0215 (2025).
Taylor, J. D., Glover, E. A. & Williams, S. T. J. M. O. T. Q. M. Ancient chemosynthetic bivalves: Systematics of Solemyidae from eastern and southern Australia (Mollusca: Bivalvia). Memoirs of the Queensland Museum, 54 (2008).
Perez-Barria, L. & Nielsen, S. N. Solemyidae (Bivalvia, Protobranchia) from the lower Miocene of south-central Chile, with description of a new species. Journal of Paleontology 94, 56–63, https://doi.org/10.1017/jpa.2019.60 (2020).
Conway, N. M., Howes, B. L., Capuzzo, J. E. M., Turner, R. D. & Cavanaugh, C. M. Characterization and site description of solemya-borealis (Bivalvia, Solemyidae), Another bivalve-bacteria symbiosis. Marine Biology 112, 601–613, https://doi.org/10.1007/bf00346178 (1992).
Walton, K. New Zealand living Solemyidae (Bivalvia: Protobranchia). Molluscan Research 35, 246–261, https://doi.org/10.1080/13235818.2015.1053168 (2015).
Bailey, J. B. The genera that never were: The impact of Janeia and Janacekia on phyletic and taxonomic relations within the Solemyidae (Bivalvia: Protobranchia). Palaeontologia Electronica 24, https://doi.org/10.26879/945 (2021).
Amano, K. & Ando, H. Giant fossil Acharax (Bivalvia: Solemyidae) from the Miocene of Japan. Nautilus 125, 207–212 (2011).
Saether, K. P., Sha, J., Little, C. T. S. & Campbell, K. A. New records and a new species of bivalve (Mollusca: Bivalvia) from Miocene hydrocarbon seep deposits, North Island, New Zealand. Zootaxa 4154, 1–26, https://doi.org/10.11646/zootaxa.4154.1.1 (2016).
Yang, M., Li, B., Gan, Z., Dong, D. & Li, X. A new chemosymbiotic bivalve species of the genus Acharax Dall, 1908 (Bivalvia, Solemyida, Solemyidae) from the Haima cold seep of the South China Sea. Zookeys, 185-192, https://doi.org/10.3897/zookeys.1198.112618 (2024).
Rodrigues, C. F., Webster, G., Cunha, M. R., Duperron, S. & Weightman, A. J. Chemosynthetic bacteria found in bivalve species from mud volcanoes of the Gulf of Cadiz. Fems Microbiology Ecology 73, 486–499, https://doi.org/10.1111/j.1574-6941.2010.00913.x (2010).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770, https://doi.org/10.1093/bioinformatics/btr011 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics 33, 2202–2204, https://doi.org/10.1093/bioinformatics/btx153 (2017).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nature Methods 18, 170–+, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100, https://doi.org/10.1093/bioinformatics/bty191 (2018).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biology 16, https://doi.org/10.1186/s13059-015-0831-x (2015).
Durand, N. C. et al. Juicer Provides a One-Click System for Analyzing Loop-Resolution Hi-C Experiments. Cell Systems 3, 95–98, https://doi.org/10.1016/j.cels.2016.07.002 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Seppey, M., Manni, M. & Zdobnov, E. M. in Methods in molecular biology 1962, 227–245 (2019).
Crescente, J. M., Zavallo, D., Helguera, M. & Vanzetti, L. S. MITE Tracker: an accurate approach to identify miniature inverted-repeat transposable elements in large genomes. Bmc Bioinformatics 19, https://doi.org/10.1186/s12859-018-2376-y (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proceedings of the National Academy of Sciences of the United States of America 117, 9451–9457, https://doi.org/10.1073/pnas.1921046117 (2020).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Research 35, W265–W268, https://doi.org/10.1093/nar/gkm286 (2007).
Ou, S. & Jiang, N. LTR_retriever: A Highly Accurate and Sensitive Program for Identification of Long Terminal Repeat Retrotransposons. Plant Physiology 176, 1410–1422, https://doi.org/10.1104/pp.17.01310 (2018).
Wheeler, T. J. et al. Dfam: a database of repetitive DNA based on profile hidden Markov models. Nucleic Acids Research 41, D70–D82, https://doi.org/10.1093/nar/gks1265 (2013).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA 6, https://doi.org/10.1186/s13100-015-0041-9 (2015).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Research 14, 988–995, https://doi.org/10.1101/gr.1865504 (2004).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nature Biotechnology 37, 907–+, https://doi.org/10.1038/s41587-019-0201-4 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature Biotechnology 33, 290–+, https://doi.org/10.1038/nbt.3122 (2015).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Research 31, 5654–5666, https://doi.org/10.1093/nar/gkg770 (2003).
Stanke, M. & Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Research 33, W465–W467, https://doi.org/10.1093/nar/gki458 (2005).
Bruna, T., Hoff, K. J., Lomsadze, A., Stanke, M. & Borodovsky, M. BRAKER2: automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR genomics and bioinformatics 3, lqaa108–lqaa108, https://doi.org/10.1093/nargab/lqaa108 (2021).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biology 9, https://doi.org/10.1186/gb-2008-9-1-r7 (2008).
Keilwagen, J., Hartung, F. & Grau, J. GeMoMa: Homology-Based Gene Prediction Utilizing Intron Position Conservation and RNA-seq Data. Methods in molecular biology (Clifton, N.J.) 1962, 161–177, https://doi.org/10.1007/978-1-4939-9173-0_9 (2019).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Research 25, 955–964, https://doi.org/10.1093/nar/25.5.955 (1997).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935, https://doi.org/10.1093/bioinformatics/btt509 (2013).
Kalvari, I. et al. Rfam 14: expanded coverage of metagenomic, viral and microRNA families. Nucleic Acids Research 49, D192–D200, https://doi.org/10.1093/nar/gkaa1047 (2021).
Lagesen, K. et al. RNAmmer:: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Research 35, 3100–3108, https://doi.org/10.1093/nar/gkm160 (2007).
Zhou, C. et al. NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP617901 (2025).
Zhou, C. et al. GenBank https://identifiers.org/ncbi/insdc.gca:GCA_054131465.1 (2025).
Zhou, C. et al. Acharax haimaensis genome and annotation files. figshare. Dataset. https://doi.org/10.6084/m9.figshare.30069271.v1 (2025).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biology 21, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Jung, Y. & Han, D. BWA-MEME: BWA-MEM emulated with a machine learning approach. Bioinformatics 38, 2404–2413, https://doi.org/10.1093/bioinformatics/btac137 (2022).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Schultz, D. T. et al. Ancient gene linkages support ctenophores as sister to other animals. Nature 618, 110–+, https://doi.org/10.1038/s41586-023-05936-6 (2023).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biology 20, https://doi.org/10.1186/s13059-019-1832-y (2019).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Systematic Biology 56, 564–577, https://doi.org/10.1080/10635150701472164 (2007).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular biology and evolution 32, 268–274, https://doi.org/10.1093/molbev/msu300 (2015).
Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Molecular Biology and Evolution 24, 1586–1591, https://doi.org/10.1093/molbev/msm088 (2007).
Acknowledgements
This work was supported by the the Marine S&T Fund of Shandong Province for Pilot National Laboratory for Marine Science and Technology (Qingdao) (2022QNLM030004), National Natural Science Foundation of China (Grant No. 42376058), the Science & Technology Fundamental Resources Investigation Program (Grant No. 2024FY101001), the National Key R&D Program of China (Grant No. 2024YFC2816000), the NSFC Innovative Group Grant (Grant No. 42221005 to WMX), the China Postdoctoral Science Foundation (Grant No. 2025M770892), and the Shandong Provincial Postdoctoral Innovation Program (Grant No. SDCX-ZG-202503097). The authors acknowledge the research vessel KEXUE for assistance with sample collection and the Oceanographic Data Center, IOCAS, for support in bioinformatics analyses.
Author information
Authors and Affiliations
Contributions
C.L. and M.W. conceived the idea. Z.Z. and Y.G. collected the sample. C.Z. and Z.Z. performed the experiments. C.Z., Y.Y. and J.W. performed the analysis. C.Z. wrote the manuscript. M.W. and C.L. supervised the study. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, C., Zhong, Z., Guo, Y. et al. Chromosome-level genome assembly of the deep-sea solemyid bivalve Acharax haimaensis. Sci Data 13, 559 (2026). https://doi.org/10.1038/s41597-026-06755-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06755-w