Abstract
Transcriptomic profiling of Traditional Chinese Medicine (TCM) perturbations is essential for elucidating the molecular mechanisms of therapeutic interventions. Although data from TCM treatment experiments are scattered across public repositories, a comprehensive, harmonized dataset remains unavailable due to heterogeneous experimental designs and inconsistent metadata. Here, we present a curated, harmonized resource comprising 362 human gene expression profiles derived from 27 TCMs and 137 TCM-derived ingredients spanning 26 human disease contexts, re-processed via a unified bioinformatics pipeline. This atlas captures TCM-induced genome-wide alterations in both protein-coding genes and long non-coding RNAs. We confirmed the dataset’s biological fidelity by validating the high reproducibility of the dataset, the enrichment of known pharmacological targets, and recapitulated the well-established therapeutic associations between TCM and disease treatment. This standardized dataset serves as a foundational resource for researchers to systematically investigate therapeutic mechanisms and predict clinical indications of TCM.
Similar content being viewed by others
Background & Summary
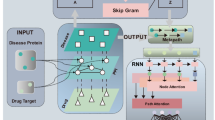
Traditional Chinese Medicine (TCM) plays a pivotal role in global health management, particularly in oncology, recognized for its cost-effectiveness, widespread accessibility, and well-documented efficacy in improving patient prognosis and quality of life1,2. Its therapeutic effects are mediated through a broad spectrum of biological mechanisms, including the regulation of the immune microenvironment3, induction of tumor cell apoptosis4, and inhibition of angiogenesis5. For instance, specific agents such as Artesunate have been shown to trigger mitochondrial dysfunction and ROS-mediated cell cycle arrest in colorectal cancer, while Ginsenoside Rh2 modulates the phenotype of tumor-associated macrophages to impede metastasis6,7. Furthermore, bioactive macromolecules like TCM polysaccharides contribute significantly to immunomodulation8,9. To comprehensively decipher these intricate molecular activities, transcriptomic profiling has emerged as an essential methodology. This high-throughput technology enables the simultaneous quantification of genome-wide expression changes, facilitating the systematic exploration of how herbal medicines and their active ingredients modulate signaling networks and biological pathways to exert their therapeutic effects10.
Although numerous TCM-related transcriptomic datasets have accumulated in public repositories such as Gene Expression Omnibus (GEO), these valuable resources remain fragmented and heterogeneous11,12. Independent studies typically utilize varying experimental platforms, distinct control conditions, and inconsistent metadata standards, introducing significant technical variations and batch effects that hinder cross-study comparison and large-scale data reuse.
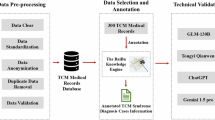
To fill this gap, we constructed a harmonized transcriptomic atlas of TCM that unifies these scattered resources into a cohesive landscape. We present a robust transcriptomic resource consisting of 362 harmonized datasets, encompassing 27 distinct TCMs (e.g., Astragali Radix, Ginkgo biloba) and 137 TCM-derived ingredients (e.g., Curcumin, Quercetin) across 26 distinct disease contexts. Processed through a unified workflow, this dataset serves as a valuable resource for both the multi-scale mechanistic exploration of TCM and the development of personalized therapies for human diseases (Fig. 1).
Workflow for the harmonized TCM transcriptomic atlas. (A) Data acquisition integrating 362 datasets from 27 TCMs and 137 ingredients across 26 disease contexts. (B) Unified harmonization pipeline generating expression matrices, differential statistics, and functional enrichment results. (C) Validation of data quality, including sample correlation analysis, assessment of phenotype reversal, and analysis of pathway enrichment.
Methods
Data acquisition and curation
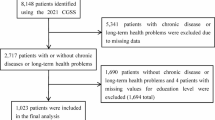
Transcriptomic datasets pertaining to TCM were sourced and systematically curated from public repositories, with the GEO as the primary source. We employed a comprehensive search strategy using keywords including “Traditional Chinese Medicine,” “TCM,” “herb,” and specific names of TCMs and active ingredients (e.g., Astragali Radix, Britanin). Inclusion criteria were as follows: (1) samples derived from human tissues or cell lines; (2) studies containing both TCM-treated groups and appropriate solvent/vehicle control groups; (3) datasets containing accessible raw count data or pre-processed series matrix files. Detailed metadata, including organism, cell line, treatment duration, and dosage, were manually curated and standardized. In total, 362 independent datasets encompassing 1,471 samples were retained for downstream analysis.
Data preprocessing and standardization
Expression data were standardized using log2 transformation. Subsequently, gene annotations were unified by converting all identifiers to official Gene Symbols according to the human reference genome GRCh38 to ensure genomic consistency.
Differential expression analysis
To identify TCM-induced gene signatures, including protein-coding genes (PCGs) and long non-coding RNAs (lncRNAs), differential expression analysis was performed on log2-transformed expression data. Statistical testing was conducted using the limma package with an empirical Bayes approach when more than two samples were available across treatment and control groups. Genes with a log2 fold change (logFC) > 1 or <−1 and an adjusted P-value < 0.05 were considered differentially expressed (Up or Down). For experiments lacking biological replicates, differential expression was assessed using log2 fold change only, with a stricter threshold of |logFC| > 1.5. Genes not meeting these criteria were labeled as stable13.
Functional and pathway enrichment analysis
To systematically elucidate the biological functions and signaling mechanisms modulated by TCM agents, Gene Ontology (GO) biological process and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed using the clusterProfiler R package. The Benjamini-Hochberg method was utilized to control the False Discovery Rate (FDR), and terms with an adjusted P-value < 0.05 were considered significantly enriched.
Data Records
The harmonized transcriptomic atlas in this study has been deposited in the Figshare14 repository and is accessible at https://doi.org/10.6084/m9.figshare.31094347. The dataset is organized into four primary file types to facilitate data reuse and downstream analyses.
First, the sample metadata file (TCM_Atlas_Metadata.xlsx) provides detailed experimental descriptions and sample grouping information for all 362 harmonized transcriptomic datasets. These comparisons cover perturbations induced by 27 TCMs and 137 TCM-derived ingredients across 26 disease contexts (Fig. 2). To enable precise sample selection, this file includes comprehensive set of column variables, categorized as follows: identifiers (customized ID, GSE_id, Plat_info), TCM information (TCM/ingredient_name, TCM/ingredient classification), biological model (Organism, Cell_Type, Cell_line), experimental design (Experiment_type, Sequence_type, Treatment_condition, Control_condition), sample type (Treatment_samples, Control_samples), and disease context (Disease Classification).
Statistical distribution of therapeutic agents (A) and disease contexts (B) in the TCM transcriptomic atlas.
Second, the repository includes systematically processed gene expression matrices derived from the original GEO datasets, which are organized within the Expression_Matrices directory. The individual dataset files follow a standardized naming convention: [ID]_expression.csv. In these matrices, row identifiers represent gene symbols annotated according to the human reference genome GRCh38.p13, and columns correspond to the unique sample identifiers (GSM IDs) listed in the metadata.
Third, the results of differential expression analysis are provided in the file [ID]_DEGs.csv for each experimental comparison. This file reports the statistics of differentially expressed PCGs and lncRNAs, with key columns including Symbol, logFC, P.Value, adj.P.Val, and State.
Finally, to support biological interpretation, functional enrichment analysis results are organized within the Functional_and_Pathway_Enrichment_Analysis directory. This folder contains the results of GO and KEGG pathway analysis using differential expression gene lists. Individual files follow a standardized naming convention based on unique experiment identifiers: [ID]_GO.csv and [ID]_KEGG.csv.
Technical Validation
The TCM-related transcriptional datasets were curated and validated by multiple independent researchers through a rigorous manual selection process. This process guaranteed that the selected datasets were both relevant and of high quality. To verify the reliability and reproducibility of the dataset, we calculated the pairwise Pearson’s correlation coefficient among independent biological replicates for each TCM treatment condition and disease context, to quantify the similarity between these samples. This revealed a high degree of biological reproducibility, with an average correlation coefficient of 0.98 across all conditions (Fig. 3A). To validate the translational potential of the generated atlas, we performed Gene Set Enrichment Analysis (GSEA) to systematically evaluate the association between TCM-induced transcriptomic alterations and cancer-associated expression patterns using The Cancer Genome Atlas (TCGA) data15. The results demonstrated that TCM induction could significantly reverse the disease signatures of cancer types for which the corresponding TCMs have been clinically validated in peer-reviewed literature. For example, Huang Qi (Astragali Radix) significantly reversed the colon adenocarcinoma (COAD) gene signature (NES = −1.37, P = 3.56e-3; Fig. 3B). It aligns with recent experimental evidence for the ability of Huang Qi-containing formulations to promote tumor blood vessel normalization in colon cancer16. Similarly, consistent with its known anti-cancer properties, Ren Shen (Ginseng) significantly downregulated the bladder urothelial carcinoma (BLCA) gene signature (NES = −1.41, P = 4.61e-03; Fig. 3B). This finding is corroborated by studies on ginsenosides, the key active component of Ren Shen, which induces apoptosis in human bladder cancer cells17. Cang Zhu (Atractylodis Rhizoma) significantly reversed the breast cancer (BRCA) gene signature (NES = −1.56, P = 1.24e-03; Fig. 3B). This finding aligns with recent experimental evidence demonstrating that atractylenolides, the major bioactive compounds in Cang Zhu, can inhibit the tumor growth of breast cancer cells18,19. To verify that TCM treatments induced transcriptomic changes in their known pharmacological targets, an overlap analysis was performed. We observed statistically significant overlaps (P < 0.05) between the observed DEGs and known TCM targets across multiple herbs (Fig. 3C). For example, Cang Zhu treatment altered the expression of its known target genes. KEGG pathway analysis revealed that the IL-17 and TNF signaling pathways mediate its therapeutic effects against human diseases (Fig. 3D), a finding corroborated by published literature20. Together, these findings provide strong functional support for the reliability of the identified DEGs, reinforcing the quality of our TCM-induced transcriptomic dataset and highlighting their translational potential in guiding TCM-mediated disease treatment. In the future, the emergence of large-scale in vivo TCM transcriptomic datasets will further expand the data resource and deepen our understanding of the molecular mechanisms underlying TCM therapies.
Biological validation of the TCM-induced transcriptomic atlas. (A) Distribution of Pearson’s correlation coefficients for biological replicates. (B) GSEA revealed the potential reversal effects of specific TCMs on disease signatures, including Huang Qi in COAD, Ren Shen in BLCA, and Cang Zhu in BRCA. (C) Statistically significant overlap between the DEGs of TCMs and the known target sets. (D) KEGG pathway enrichment analysis of genes perturbed by Cang Zhu.
Data availability
The harmonized gene-level expression matrices and associated metadata generated in this study are publicly available at Figshare: https://doi.org/10.6084/m9.figshare.31094347.
Code availability
The bioinformatic analyses were conducted using R statistical software (version 4.4.3). No custom algorithms or software were developed for this study; all analyses utilized standard functions from publicly available R packages. Data cleaning and manipulation were performed using dplyr (v1.1.4) and stringr (v1.6.0). Differential expression analysis was conducted using limma (v3.62.2). GO and KEGG pathway enrichment analyses, as well as Gene Set Enrichment Analysis (GSEA), were performed using clusterProfiler (v4.14.0). Visualizations were generated using ggplot2 (v4.0.0) and enrichplot (v1.26.1).
References
Wang, J., Wong, Y.-K. & Liao, F. What has traditional Chinese medicine delivered for modern medicine? Expert Reviews in Molecular Medicine 20, e4 (2018).
Zhang, X., Qiu, H., Li, C., Cai, P. & Qi, F. The positive role of traditional Chinese medicine as an adjunctive therapy for cancer. Bioscience trends 15, 283–298 (2021).
Gao, S. et al. Novel Natural Carrier‐Free Self‐Assembled Nanoparticles for Treatment of Ulcerative Colitis by Balancing Immune Microenvironment and Intestinal Barrier. Advanced healthcare materials 12, 2301826 (2023).
Guo, W. et al. Aloperine Suppresses Cancer Progression by Interacting with VPS4A to Inhibit Autophagosome‐lysosome Fusion in NSCLC. Advanced Science 11, 2308307 (2024).
Liu, X. et al. Natural medicines of targeted rheumatoid arthritis and its action mechanism. Frontiers in Immunology 13, 945129 (2022).
Huang, Z. et al. Artesunate inhibits the cell growth in colorectal cancer by promoting ROS-dependent cell senescence and autophagy. Cells 11, 2472 (2022).
Li, H. et al. Modulation the crosstalk between tumor-associated macrophages and non-small cell lung cancer to inhibit tumor migration and invasion by ginsenoside Rh2. BMC cancer 18, 579 (2018).
Guo, C. et al. Novel Chinese angelica polysaccharide biomimetic nanomedicine to curcumin delivery for hepatocellular carcinoma treatment and immunomodulatory effect. Phytomedicine 80, 153356 (2021).
Li, J. et al. Purification, structural characterization, and immunomodulatory activity of the polysaccharides from Ganoderma lucidum. International journal of biological macromolecules 143, 806–813 (2020).
Wang, K. et al. Inhibition of inflammation by berberine: Molecular mechanism and network pharmacology analysis. Phytomedicine 128, 155258 (2024).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic acids research 41, D991–D995 (2012).
Zhao, H. et al. So3D: a comprehensive three-dimensional spatial omics resource for decoding tissue architecture in physiology and disease. Nucleic Acids Research 54, D1281–D1290 (2026).
Zhao, H. et al. LncTarD 2.0: an updated comprehensive database for experimentally-supported functional lncRNA–target regulations in human diseases. Nucleic acids research 51, D199–D207 (2023).
Zhao, H. et al. An in-depth transcriptomic atlas deciphering traditional Chinese medicine mechanisms and disease associations. figshare https://doi.org/10.6084/m9.figshare.31094347.v6 (2026).
Cancer Genome Atlas Research Network, J. The cancer genome atlas pan-cancer analysis project. Nat. Genet 45, 1113–1120 (2013).
Liang, Y. et al. Astragali Radix-Curcumae Rhizoma normalizes tumor blood vessels by HIF-1α to anti-tumor metastasis in colon cancer. Phytomedicine 140, 156562 (2025).
Li, X. et al. Gypenoside‐induced apoptosis via the PI3K/AKT/mTOR signaling pathway in bladder cancer. BioMed Research International 2022, 9304552 (2022).
Long, F., Wang, P., Ma, Y., Zhang, X. & Wang, T. Chemopreventive effects of atractylenolide-III on mammary tumorigenesis via activation of the Nrf2/ARE pathway through autophagic degradation of Keap1. Biomedicine & Pharmacotherapy 176, 116852 (2024).
Xu, H. et al. Atractylenolide‐1 affects glycolysis/gluconeogenesis by downregulating the expression of TPI1 and GPI to inhibit the proliferation and invasion of human triple‐negative breast cancer cells. Phytotherapy Research 37, 820–833 (2023).
Nguyen, L. T. H., Nguyen, N. P. K., Tran, K. N., Shin, H.-M. & Yang, I.-J. Network Pharmacology and Experimental Validation to Investigate the Antidepressant Potential of Atractylodes lancea (Thunb.) DC. Life 12, 1925 (2022).
Acknowledgements
We are grateful to all contributors to this study and acknowledge the funding sources that provided financial support. This work was supported by the National Natural Science Foundation of China (62372144, 62573169, 62572155) and Outstanding Youth Foundation of Heilongjiang Province of China (YQ2023F004).
Author information
Authors and Affiliations
Contributions
H.Y.Z.: Study conception and design, methodological design, data processing and analysis, manuscript drafting and revision. P.Q.B: Data processing and analysis, figure and table visualization, manuscript drafting and revision. Z.M.L.: Data collection and collation. M.R.G.: Data processing. L.L.: Data collection and collation. D.C.H.: Data processing. J.C.G.: Study conception and design. L.W.: Study conception and design, methodological design, resource provision and support, supervision and guidance.All authors read, reviewed, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, H., Ben, P., Liu, Z. et al. An in-depth transcriptomic atlas deciphering traditional Chinese medicine mechanisms and disease associations. Sci Data 13, 608 (2026). https://doi.org/10.1038/s41597-026-06988-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06988-9