Abstract
Chronic liver diseases (CLD) account for more than 2% of deaths worldwide. Extensive research has been conducted to better understand CLD, generating vast amounts of data. However, only a small fraction of raw preclinical data are publicly available, posing a significant challenge for transparency, reproducibility, and data reuse. Therefore, we built a preclinical liver imaging dataset, the first of its kind to our knowledge. The database contains longitudinal liver MRI scans from mice with hepatocellular carcinoma, metabolic dysfunction-associated steatohepatitis (MASH, formerly NASH), and fibrosis, as well as CT scans of mice with MASH and mice carrying a dysfunctional ICAM-1 gene. Superimposable MRI and CT scans bridge the gap between the modalities. Some of the 222 murine scans have annotated segmentations. Metadata containing both scan and mouse parameters are organized using a tailored metadata profile in ISA-Tabs. This dataset enables advanced image analysis, such as building tools for automated segmentation, train radiomics analysis tools, or can be used as a reference control dataset.
Similar content being viewed by others
Background & Summary
Chronic liver diseases (CLD) account for more than 2% of deaths worldwide1. The major causes of CLD are viral hepatitis (Hepatitis B virus, Hepatitis C virus) and steatotic liver diseases such as metabolic dysfunction-associated steatotic liver disease and alcohol-associated liver disease, with risk factors including low childhood vaccination rates, exposure to infected blood, obesity, and increased alcohol consumption. In CLD, chronic inflammation and fibrosis can over time lead to liver cirrhosis which has its own complications and consequences such as portal hypertension, hepatocellular insufficiency, and hepatocellular carcinoma2,3.
Medical imaging can be used to diagnose and monitor liver cirrhosis as an alternative to invasive biopsies. In the clinic, ultrasound is the most commonly used imaging modality due to its wide availability and low cost4,5, but also elastography, CT, and MRI are applied to substantiate and refine the diagnosis6. In preclinical investigations, MRI is mainly used due to its high versatility7,8. MRI can grade fibrosis and steatosis, besides accurately assessing liver size and detect and differentiate between different types of lesions. In addition, MRI can reduce the need for contrast agents, which are not always well tolerated.
Existing preclinical imaging databases include CT data from healthy mice9 or include different types of disease models and imaging modalities, such as the Preclinical Image DAtaset Repository (PIDAR)10. In addition, initiatives to create a mouse atlas are helpful to the research community11, such as published by Dogdas et al.12, which is based on a single animal, or Wang et al.13, who propose a deformable atlas. In contrast to these examples and the liver CT data of Fiebig et al.14, this dataset focuses on multimodal preclinical liver and whole-body imaging data from CLD mice and healthy controls.
To improve the understanding of CLD, scientists have been performing extensive research, resulting in the generation of large data amounts. Out of all the publicly available data, however, only a small fraction of (raw) data is available which remains a major concern. As data availability is necessary for open science and data reuse, we intend to build a dataset of preclinical liver imaging data, which is the first of its type to our knowledge. Via this way, the data can be reused by the community. Possible data reuse strategies are for example, i) training computational models, like generative adversarial networks15, ii) creating an attenuation correction atlas for PET by use of overlapping CT and MRI data16, iii) training segmentation tools17, iv) detecting malignancies or pathological changes by computational models18, v) predicting disease progression19, and vi) reducing animal numbers by using control animals of one of the studies in case of a similar setup20. To give a more concrete example on computational research, livers of healthy and diseased mice might be segmented and analyzed to apply or develop neural networks for segmentation or contour extraction (see Fig. 1 liver segmentation as an example)21,22.
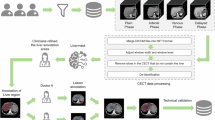
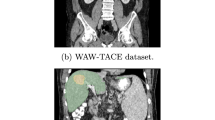
Representative CT and MRI liver scans. In (a–d) scanning direction and intervals and imaging modalities are shown in addition to study groups. In (e) longitudinal native T2w MRI of the liver of an hepatocellular carcinoma model from 22 to 28 weeks is shown, (f) displays T1w MRI of healthy, MASH/HFD, and toxic liver injury model (left to right) of the multiparametric MRI study7, (g) shows CT (left) to image liver size and body fat analysis segmentation (right) of HFD wild-type mice, (h) presents whole-body CT and (water and fat) MRI in axial (left) and coronal (right) views of healthy mice29. DEN: diethylnitrosamine, HFD: high-fat diet.
Methods
There are five subsets of studies that have been pooled into one dataset to increase disease spectra and variety of modalities, including (functional) murine MRI data of hepatocellular carcinoma, liver fibrosis and steatosis, as well as whole-body CT data from mice fed with high-fat diet, and combined CT and MRI scans of healthy mice. The latter scans are also provided with segmentations of the liver and other organs. See Table 1 and Fig. 1 for more information on these studies, a study overview, and sample images. In addition, imaging parameters and information on segmentation labels can be found in Tables 1–6, 8–11.
MRI assessment of hepatocellular carcinoma progression
To study the dynamics of the cellular immune compartment within the tumor microenvironment of a chronically inflamed and damaged liver during both progression and regression of HCC, the toxin-based DEN/CCl₄ model was employed23,24. The fibrosis-associated HCC mouse model relies on the induction of spontaneous tumor formation through a single administration of the carcinogen diethylnitrosamine (DEN) followed by repeated injections of carbon tetrachloride (CCl₄). The single dose of the tumor initiator DEN was administered to C57BL/6 J mice (WT, RWTH Aachen University) 14 days after birth at a concentration of 25 mg/kg body weight (BW) (diluted in PBS). At 4 weeks of age, the animals also received CCl4 injections (0.5 ml/kg BW, diluted in germ oil) once a week over a period of 22 weeks. Both agents were applied by i.p. injection. Following the manifestation of HCC at 26 weeks of age, a combined therapy consisting of immune checkpoint blockade and anti-angiogenic therapy was implemented to induce tumor regression. The efficacy of the therapeutic approach in promoting HCC regression was assessed by comparing treated groups with control cohorts receiving isotype antibodies. Isotype antibodies utilized in the study were IgG1 (BioXcell, BE0088; 40 mg/kg BW) and IgG2b (BioXcell, BE0090; 200 µg/mouse). Control groups received either IgG1 isotype antibody alone (WT, n = 7) or a combination of both isotypes (WT, n = 7). The first control group received 6 i.p. injections of IgG1 over two weeks, while the second control group received 5 i.p. injections of the combined antibodies over the same time frame. Please note that as this study has not yet been published, only data from control animals are included in the dataset.
To monitor tumor growth and size during HCC progression and regression, MRI was performed using a 7 T Biospec 70/20 scanner (Bruker BioSpin, Ettlingen, Germany) with an RF RES 300 1H 075/040 QSN TR volume coil (T13161V3), at (22,) 26, 27, and 28 weeks of age under isoflurane anesthesia. To obtain structural information about the liver, a T2-weighted sequence was used. Additionally, to assess Brownian motion and thus extracellular matrix composition and cellularity within the liver tissue, a diffusion-weighted sequence was included (scan parameters are shown in Table 2). Both sequences were respiration-triggered to minimize breathing artifacts.
Multiparametric MRI to assess hepatic fibrosis and steatosis
The objective of Baskaya et al.7 was to explore the feasibility of CLD classification using longitudinal multiparametric MRI. For this purpose, 8–10-week-old male C57BL/6 J wild-type mice (Janvier Labs, Le Genest‐Saint‐Isle, France) were randomly assigned to two distinct CLD models, a high-fat diet (HFD) group (n = 7) and a CCl4 group (n = 10), as well as an untreated control group (n = 7). To induce metabolic dysfunction associated steatohepatitis (MASH), mice were fed with HFD chow containing 40% fat, 20% fructose, and 2% cholesterol (Research Diets Inc, New Brunswick, NJ) for 24 weeks. In the CCl4 group, inflammation-induced fibrotic remodeling of the liver was induced by i.p. injections of CCl4 (diluted in corn oil, Sigma Aldrich) at a dose of 0.6 ml/kg BW twice a week for 8 weeks. In this group, four out of ten mice died. The untreated control group was fed with standard chow (ssniff, Soest, Germany) for 24 weeks.
MRI was performed using a 7 T Biospec 70/20 scanner (Bruker BioSpin, Ettlingen, Germany) with an RF RES 300 1H 075/040 QSN TR volume coil (T13161V3). Anatomical and functional MRI measurements under isoflurane anesthesia were conducted at 0, 4, 8, 12, 16, 20, and 24 weeks in the HFD and control groups, and at 0, 4, and 8 weeks in the CCl4 group (at least two days post CCl4 injection). For organizational reasons, the time points in the database must be read as time points after disease induction and not as injection time points, therefore, 0, 4, 8, 12, 16, 20, and 24 weeks correspond to 0, 40320, 80640, 120960, 161280, 201600, and 241920 min, respectively.
To avoid interference between the two contrast agents gadoxetic acid (Primovist, Bayer, Germany) and ferucarbotran (VivoTrax™, Magnetic Insight Inc., Alameda, USA), two separate MRI protocols, labeled MR1 and MR2, were performed following a 48-hour washout period (see Tables 3–5). The first contrast agent was used to evaluate hepatocyte function, while the latter was used to assess macrophage function. As respiration can cause artifacts in MRI, all liver scans were acquired with respiration gating. In both the MR1 and MR2 protocols, liver anatomy was assessed by T1-weighted gradient echo and T2-weighted spin echo sequences (with fat suppression). Detailed structural information at the cellular level and extracellular matrix composition was obtained by apparent diffusion coefficient mapping using a diffusion-weighted echo planar imaging sequence. Lower apparent diffusion coefficients correlated with fibrosis. Structural imaging was followed by functional sequences to assess hepatocyte function (MR1), liver damage (MR2), and macrophage activity (MR2).
In the MR1 protocol, a fat-selective T1-weighted gradient echo sequence with water suppression was acquired to determine the liver fat content. Paraffin oil-filled tubes were placed in the field of view for reference. T1 relaxometry was performed before and after gadoxetic acid injection (0.025 mmol/kg BW in 100 μL 0.9% saline). Between the relaxometry mapping, the hepatocyte function was assessed, and the contrast agent was injected manually at approximately 10 µL/s, a dynamic contrast enhanced MRI scan of a central slice in the liver (1600 acquisitions) was performed after 30 s of baseline scans. Lower gadoxetic acid uptake correlated with reduced hepatocyte function. Finally, the anatomical T1-weighted gradient echo sequence was acquired again, but this time contrast-enhanced. The MR1 protocol took 50–55 min to complete.
The MR2 protocol continued with T2 relaxometry mapping before and after ferucarbotran injection (8 μmol Fe/kg BW ferucarbotran in 100 μL 0.9% saline) using a multi-slice multi-echo T2 mapping sequence. Low R2 values correlated with higher liver damage demonstrated by serum glutamic-oxaloacetic transaminase levels. To assess macrophage phagocytic activity, a dynamic susceptibility contrast MRI was performed between these scans, with contrast agent injected at 10 µL/s after 30 s of dummy scans. Lower ferucarbotran uptake indicated decreased macrophage function. The MR2 protocol ended with a contrast-enhanced T2-weighted spin echo sequence (as before). The MR2 acquisition time was completed after 40–45 min.
Liver size measurements by native CT to test the effect of beta-glucans on steatosis
To study the benefit of oat beta-glucan in CLD, more specific, metabolic dysfunction-associated steatotic liver disease, amongst others, liver volume and fat distribution were measured by µCT measurements25. Hence, male 8 weeks old C57BL/6 J mice (Janvier labs, Le Genest-Saint-Isle, France) were fed with a chow diet (control, n = 3) or with a western style high fat diet containing 40% fat, 20% fructose, and 2% cholesterol (Research Diets Inc, New Brunswick, NJ, HFD, n = 6) for 24 weeks, as described in Jaeger et al.25. Solubilized oat beta-glucan (Garuda, Exeter, USA) was administered with the drinking water (control + beta-glucan, n = 3; HFD + beta-glucan, n = 6). The body fat distribution was assessed under isoflurane anesthesia with a total body normal scan in a hybrid micro-CT optical imaging system (MILabs B.V., Houten, the Netherlands) with an X-ray tube voltage of 55 kV, tube current of 0.17 mA, an isotropic voxel size of 140 μm, and a scan time of 4 minutes and 11 seconds (Table 6). To analyze fat composition, an interactive segmentation protocol was loaded and livers were segmented manually for volume measurements (Imalytics Preclinical, Gremse-IT GmbH, Aachen, Germany)26.
Liver size measurements by native CT to test the role of ICAM-1
To assess the role of intercellular adhesion molecule 1 (ICAM-1, or CD54), a cell surface protein with a role in inflammation and immunity27, in metabolic dysfunction, WT mice and mice with an ICAM-1 mutation were studied in their fat distribution and liver size28. Therefore, 8–12-week-old male WT (C57BL/6 J, RWTH Aachen University) and ICAM-1 mutant mice (B6.129S7-Icam1tm1Bay/J, RWTH Aachen University)27 were fed a HFD (40% fat, 20% fructose, 2% cholesterol, Brogaarden, Lynge, Denmark) for 12 weeks. µCT scans were performed under isoflurane anesthesia in ultra-focus fast mode in a hybrid micro-CT optical imaging system (MILabs B.V., Houten, the Netherlands) with a tube voltage of 65 kV, a tube current of 0.13 mA, an isotropic voxel size of 140 μm, and a scan time of 37 s (Table 6). Segmentations (Imalytics Preclinical, Gremse-IT GmbH, Aachen, Germany)26 of body fat (subcutaneous and visceral), liver, bones, and the lungs are included.
Whole-body CT-MRI of healthy mice
This study tracked the biodistribution of immunoglobulins29, therefore healthy female Crl:SKH1-Hrhr nude mice (Charles River Laboratories, Wilmington, MA) of 6–8 weeks were imaged directly after injection as well as 4.5 h after injection of fluorescently labeled immunoglobulins. However, the optical fluorescence data are not included in the dataset as it is outside the scope of the latter. Imaging was performed under isoflurane anesthesia at the two time points inside the hybrid imaging animal holder (MILabs B.V., Houten, the Netherlands) first in the hybrid micro-CT optical imaging system (MILabs B.V.), then the MRI scan was performed. First the optical fluorescence imaging was performed, then the mouse holder was moved to the CT unit, and mice were scanned with the total body normal protocol with tube voltage of 55 kV, tube current of 0.17 mA, an isotropic voxel size of 140 μm (Table 6). Then, mice kept inside the animal holder were transferred to perform MRI in a 7 T BioSpec 70/20 USR MRI scanner (Bruker BioSpin, Ettlingen, Germany) equipped with an RF Res 300 1H 112/086 QSN TO AD volume coil (T12053V3). For whole-body imaging a Dixon-based T2-weighted fat-water-separated turbo RARE (rapid acquisition with relaxation enhancement) with the in Table 1 specified parameters was performed. During fluorescence tomography reconstructions, the vaseline-filled inner marker holes in the animal holder enabled overlap of CT and MRI data. For CT and MRI scans, segmentations of bladder, bone (only CT), brain (only MRI), caecum, colon, heart, intestine, kidney, liver, lungs, spinal cord (only MRI), spleen, and stomach are included. Fluorescence tomography data are not included in the dataset.
Animal housing
All experiments were approved by the North Rhine-Westphalia ‘Stage agency for nature, environment, and consumer protection (Landesamt für Natur-, Umwelt- und Verbraucherschutz Nordrhein-Westfalen, LANUV). Mice were socially housed with a conventional light cycle of 12 hours (7 am to 7 pm), at 20–24 °C, humidity of 45–65% and bedding was changed weekly. All animals were kept in sterile, individually ventilated cages. Sterilized drinking water and (standard) chow were available to the mice ad libitum.
Data Records
The dataset is available at Zenodo30, where the data are stored together with the metadata in ISA-Tabs as specified in the metadata profile, which is shown in Tables 7, 8, 9, 10, 11. In the ISA-Tabs, file names are linked with all the relevant metadata related to the mice and imaging modalities used. The same metadata as displayed in the assay files (a_*.text) can also be found on the research data management platform Coscine, which is accessible upon request.
The file structure is flat. Studies were named as follows and can be searched in the search bar: i) hepatocellular carcinoma MRI, ii) multiparametric liver MRI, iii) beta-glucans liver CT, iv) ICAM-1 CT fat segmentations, and v) CT-MRI whole body. 3D imaging data are saved as.dcm or.nii files, files from iii)-v) are accompanied by 3D segmentations in.seg or.segff files (see study descriptions).
Data Overview
This dataset contains data from 5 studies, namely, i) a study to monitor HCC development with MRI, ii) multiparametric MRI to evaluate hepatic fibrosis and steatosis with various structural and functional scans, iii/iv) liver volume measurements with CT to investigate the interplay of beta-glucans or ICAM-1 with a HFD including body fat and liver segmentations, and v) whole-body MRI and CT including segmentations of healthy mice.
Technical Validation
DEN and CCl₄ are commonly used agents to induce HCC, as demonstrated in previous studies through histological analysis23,24. In this study, the livers exhibited multiple HCC lesions on MRI, which were further confirmed by the presence of tumors macroscopically observable upon excision. In the secondly described study, by Baskaya et al.7, MASH and liver fibrosis progression were confirmed by histology. Similarly, Jaeger et al.25 display by liver histology the effects of beta-glucan and high fat diet, which are in line with the imaging results.
The mice with the dysfunctional ICAM-1 gene used in the study by Eswaran et al.28 were originally described in Sligh et al.27 and it was reported that these mice only showed residual membrane-bound ICAM-1 in thymus, lung, and spleen tissues on staining, but none in gut and liver. Before the animals were used in experiments, they were tested for the presence of the mutation by polymerase chain reaction with primers for ICAM-17 (5′CTGAGCCAGCTGGAGGTCTCG3′, ICAM-18 (5′GAGCGGCAGAGCAAAAGAAGC3′), and ICAM-19 (5′AGGACAGCAAGGGGGAGGATT3′), as described in Eswaran et al.28. All WT and B6.129S7-Icam1tm1Bay/J mice used were healthy and came from the same barrier.
In the combined CT-MRI study29, healthy animals were used and CT- and MRI-based segmentations were validated with dice score analyses.
Data availability
The dataset is available at https://doi.org/10.5281/ZENODO.17130082.
Code availability
No custom code has been used to curate the dataset. The data can be downloaded directly without the need for any specific software.
References
Cheemerla, S. & Balakrishnan, M. Global Epidemiology of Chronic Liver Disease. Clin Liver Dis (Hoboken) 17, 365–370 (2021).
Sharma, A. & Nagalli, S. Chronic Liver Disease. (StatPearls Publishing, Treasure Island (FL), 2023).
Iwakiri, Y. & Trebicka, J. Portal hypertension in cirrhosis: Pathophysiological mechanisms and therapy. JHEP Reports 3, 100316 (2021).
Singh, S., Hoque, S., Zekry, A. & Sowmya, A. Radiological Diagnosis of Chronic Liver Disease and Hepatocellular Carcinoma: A Review. J Med Syst 47, 73 (2023).
Vernuccio, F. et al. Advances in liver US, CT, and MRI: moving toward the future. Eur Radiol Exp 5, 52 (2021).
Taouli, B., Ehman, R. L. & Reeder, S. B. Advanced MRI Methods for Assessment of Chronic Liver Disease. AJR Am J Roentgenol 193, 14–27 (2009).
Baskaya, F. et al. Pathophysiologic Mapping of Chronic Liver Diseases With Longitudinal Multiparametric MRI in Animal Models. Invest Radiol 59, 699–710 (2024).
Chae, Y. J. et al. Preclinical Long-term Magnetic Resonance Imaging Study of Silymarin Liver-protective Effects. J Clin Transl Hepatol 000, 000–000 (2022).
Rosenhain, S. et al. A preclinical micro-computed tomography database including 3D whole body organ segmentations. Sci Data 5, 180294 (2018).
FAIRsharing Team. FAIRsharing record for: Preclinical Image Dataset Repository. FAIRsharing https://doi.org/10.25504/FAIRSHARING.F64055.
Khmelinskii, A. et al. Articulated Whole-Body Atlases for Small Animal Image Analysis: Construction and Applications. Mol Imaging Biol 13, 898–910 (2011).
Dogdas, B., Stout, D., Chatziioannou, A. F. & Leahy, R. M. Digimouse: a 3D whole body mouse atlas from CT and cryosection data. Phys Med Biol 52, 577–587 (2007).
Wang, H., Stout, D. B. & Chatziioannou, A. F. A Deformable Atlas of the Laboratory Mouse. Mol Imaging Biol 17, 18–28 (2015).
Fiebig, T. et al. Three-Dimensional In Vivo Imaging of the Murine Liver: A Micro-Computed Tomography-Based Anatomical Study. PLoS ONE 7, e31179 (2012).
Han, T. et al. Breaking medical data sharing boundaries by using synthesized radiographs. Sci Adv 6, eabb7973 (2020).
Krokos, G., MacKewn, J., Dunn, J. & Marsden, P. A review of PET attenuation correction methods for PET-MR. EJNMMI Phys 10, 52 (2023).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18, 203–211 (2021).
Magnuska, Z. A. et al. Combining Radiomics and Autoencoders to Distinguish Benign and Malignant Breast Tumors on US Images. Radiology 312, e232554 (2024).
Han, T. et al. Image prediction of disease progression by style-based manifold extrapolation. Nat Mach Intell 4, 1029–1039 (2022).
Bonapersona, V. et al. Increasing the statistical power of animal experiments with historical control data. Nat Neurosci 24, 470–477 (2021).
Ansari, M. Y. et al. A lightweight neural network with multiscale feature enhancement for liver CT segmentation. Sci Rep 12, 14153 (2022).
Dakua, S. P. & Sahambi, J. S. Automatic Left Ventricular Contour Extraction from Cardiac Magnetic Resonance Images Using Cantilever Beam and Random Walk Approach. Cardiovasc Eng 10, 30–43 (2010).
Zhang, Q. et al. Proteomic analysis of DEN and CCl4-induced hepatocellular carcinoma mouse model. Sci Rep 14, 8013 (2024).
Huang, Y. et al. The hepatic senescence-associated secretory phenotype promotes hepatocarcinogenesis through Bcl3-dependent activation of macrophages. Cell Biosci 11, 173 (2021).
Jaeger, J. W. et al. Microbiota modulation by dietary oat beta-glucan prevents steatotic liver disease progression. JHEP Reports 6, 100987 (2024).
Gremse, F. et al. Imalytics Preclinical: Interactive Analysis of Biomedical Volume Data. Theranostics 6, 328–341 (2016).
Sligh, J. E. et al. Inflammatory and immune responses are impaired in mice deficient in intercellular adhesion molecule 1. Proc Natl Acad Sci USA 90, 8529–8533 (1993).
Eswaran, S. et al. Intercellular adhesion molecule-1 protects against adipose tissue inflammation and insulin resistance but promotes liver disease activity in western-diet fed mice. Sci Rep 15, 25884 (2025).
Schraven, S. et al. CT- and MRI-aided fluorescence tomography reconstructions for biodistribution analysis. Invest Radiol 59, 504–512 (2024).
Cat. catgonzftw1/preclinical_imaging. Preclinical Imaging Static Site v1.0.0. Zenodo https://doi.org/10.5281/ZENODO.17130082 (2025).
Acknowledgements
This work has been funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – Project-ID 403224013 – SFB 1382 The Gut-Liver Axis and Project ID 321137804 - FOR2591 Severity assessment. The data used in this publication was managed using the research data management platform Coscine with storage space granted by the Research Data Storage (RDS) of the DFG and Ministry of Culture and Science of the State of North Rhine-Westphalia (DFG: INST222/1261-1 and MKW: 214-4.06.05.08 - 139057). In addition, the authors thank Jasmin Groß and Jan Schumacher for their technical support.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
S.S. – data acquisition, data processing, validation, project conception, writing. C.G. – data processing, data management, project conception. F.B. – data acquisition. L.K. – data processing. A.B. – data acquisition. R.M.G. – writing: original draft (revision), review & editing. R.B. – data acquisition. D.M. – data acquisition. M.L.B. – experimental design. K.M.S. – experimental design. A.S. – experimental design. F.K. – experimental design, project conception, supervision. All authors revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
SS and DM are employed at Gremse-IT GmbH.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schraven, S., Gonzalez, C., Baskaya, F. et al. A preclinical CT and MRI Liver Imaging Dataset with Anatomical, Functional and Segmentation Data. Sci Data 13, 446 (2026). https://doi.org/10.1038/s41597-026-07003-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-07003-x