Abstract
Considering the high fatality of hepatocellular carcinoma (HCC), current prognostic systems are insufficient to accurately forecast HCC patients' outcomes. In our study, nine anoikis‑related genes (PTRH2, ITGAV, ANXA5, BIRC5, BDNF, BSG, DAP3, SKP2, and EGF) were determined to establish a risk scoring model using LASSO regression, which could be validated in ICGC dataset. Kaplan–Meier curves and time-dependent receiver operating characteristic (ROC) curve analysis confirmed the risk score possessed an accurate predictive value for the prognosis of HCC patients. The high-risk group showed a higher infiltration of aDCs, macrophages, T-follicular helper cells, and Th2 cells. Besides, PD-L1 was significantly higher in the high-risk group compared to the low-risk group. Several anoikis‑related genes, such as ANX5, ITGAV, BDNF and SKP2, were associated with drug sensitivity in HCC. Finally, we identified BIRC5 and SKP2 as hub genes among the nine model genes using WGCNA analysis. BIRC5 and SKP2 were over-expressed in HCC tissues, and their over-expression was associated with poor prognosis, no matter in our cohort by immunohistochemical staining or in the TCGA cohort by mRNA-Seq. In our cohort, BIRC5 expression was highly associated with the T stage, pathologic stage, histologic grade and AFP of HCC patients. In general, our anoikis-related risk model can enhance the ability to predict the survival outcomes of HCC patients and provide a feasible therapeutic strategy for immunotherapy and drug resistance in HCC. BIRC5 and SKP2 are hub genes of anoikis‑related genes in HCC.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) remains a great challenge in terms of global health, for it is the sixth most common type of cancer and the third leading cause of cancer-related mortality worldwide. Studies have shown that about 906,000 new cases (4.7%) and 830,000 deaths (8.3%) worldwide in 20201. At present, there are multiple therapeutic strategies for HCC, including surgical resection, liver transplantation, radiofrequency ablation, vascular catheterization and chemotherapy2,3. However, the prognosis of HCC remains poor, with a 5-year survival rate of 20%4. Therefore, early prediction for prognosis is of great significance to improve the survival outcomes of HCC patients. Considering that cirrhosis is the basis of HCC in most patients, the prognosis of HCC depends on the tumor burden, the degree of liver dysfunction, and the patient's performance status5. Several prognostic systems have been proposed for patients with HCC in the last three decades, including the BCLC system, CNLC classification, the MESH score, and the CLIP score6. Nevertheless, it is rather difficult to confirm the best prognostic system that could be widely used for all HCC patients in clinical practices7. As a result, current prognostic systems are not sufficient to accurately forecast the outcomes of HCC patients. In addition, the identification of molecular biomarkers associated with HCC prognosis attracted considerable attention, but the number of prognostic model is limited. Hence, it is necessary to develop a new tool based on prognostic biomarkers to achieve an optimal evaluation of survival outcomes in HCC patients.
Anoikis is a type of programmed apoptosis initiated by detachment of cells from the extracellular matrix (ECM). It is a mechanism that contributes to tissue homeostasis by eliminating detached or misplaced cells under physiological or pathological conditions. Besides, anoikis can limit cancer progression by preventing the dissemination of cancer cells to distant organs8. Studies have shown that the induction of the anoikis occurs through the interaction of the intrinsic pathway (mitochondria approach) and extrinsic pathway (death receptor approach). It has been reported cancer cells develop anoikis resistance by escaping the death signals and activating the pro-survival signals9,10. Meanwhile, acquisition of anoikis resistance is a hallmark of cancer cell invasiveness, metastasis, treatment resistance, and relapse11. In addition, researchers have identified that different players, including cell adhesion molecules, growth factors, hypoxia, stemness, autophagy, long non-coding RNAs, and signaling pathways, are associated with anoikis resistance, allowing cell survival in the cellular environment12,13,14,15. However, the precise mechanisms involved in anoikis resistance are not fully understood. Therefore, the early investigation into the mechanisms of anoikis resistance may greatly help to improve the prognostic assessment of cancer and further efficacious treatments for cancer.
Currently, enormous efforts have shown that many anoikis-related genes are associated with the proliferation of several tumors, including gastric cancer, colorectal cancer, lung cancer, and pancreatic cancer16,17,18. For example, GDH1 provides signals of anoikis resistance and tumor proliferation in LKB1-deficient lung cancer. Also, GDH1 knockdown can not only improve cell sensitivity to anoikis induction but also reduce tumor metastasis19. In high-grade serous ovarian carcinoma, CBX2 has been verified to drive anoikis resistance and tumor metastasis20. TMPRSS4, a cell surface-anchored serine protease, is reported to promote anoikis resistance, enhancing survival of circulating tumor cells and leading to early metastasis in prostate cancer21. Therefore, anoikis‑related genes are considered as promising diagnostic markers and potential therapeutic targets. However, the construction of a prognostic risk model based on anoikis‑related genes remains unclear in HCC. Hence, we exploit the relationship between anoikis‑related genes and HCC clinical information to forecast the outcomes of HCC patients.
Given the limited effects of standard treatments, increasing attention is focused on immunotherapy in recent years22,23. Studies highlighted immune cells producing cytokines and components of the ECM as response to liver injury, which promotes the survival of cancer stem cells24. Currently, immunotherapy is considered as an additional treatment and witness substantial progress25,26. Thus, exploring the underlying mechanism between the immune system and cancer to enhance the immunotherapy of HCC is urgently required.
Herein, multiple mRNA expression databases were utilized to develop and validate a risk scoring model related to anoikis for HCC. Moreover, we explored the potential relevance between the prognostic risk score and tumor stemness, immunity as well as drug sensitivity. Finally, we identified BIRC5 and SKP2 as hub genes in HCC. They are over-expressed in HCC tissues, and were associated with poor prognosis, no matter in the TCGA cohort or our TMA cohort. All of these might not only promote the capability to forecasting the survival outcomes of HCC patients, but also provide prominent reference for molecular mechanisms of immunotherapy and drug resistance in HCC.
Methods
Data collection process
The mRNA expression matrix of 374 liver hepatocellular carcinoma samples and 50 adjacent normal samples, and the corresponding clinicopathological information were downloaded from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/). Besides, we also downloaded the mRNA expression data and the clinical information of another 231 HCC patients from the International Cancer Genome Consortium (ICGC) database (https://dcc.icgc.org/). From the GeneCards (https://www.genecards.org/), 82 anoikis‑related genes were extracted by screening with a relevance score > 1.8.
Identification of prognostic anoikis‑related DEGs and functional analysis
To obtain differentially expressed genes (DEGs), we used the “limma” package of R language to conduct differential analysis on the expression of the above 82 anoikis‑related genes in normal and tumor samples from the TCGA database. Thirty-eight anoikis‑related DEGs were obtained, and the screening criteria were as follows: |log (fold change)| > 1 and adjusted p < 0.0527. Through univariate Cox regression analysis, 26 anoikis‑related DEGs were associated with the prognosis of HCC (prognostic anoikis‑related DEGs, p < 0.05).
STRING Database (Version 11.0, https://STRINGDb.org/) was used to map the protein–protein interaction (PPI) network of these 26 prognostic anoikis‑related DEGs. The Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses were performed based on the aforementioned prognostic anoikis‑related DEGs28.
Establishment and verification of anoikis risk scoring model
The least absolute contraction and selection operator (LASSO) method was used to punish the factor data. The risk score was calculated according to the standardized expression level of each gene and its corresponding regression coefficient. Then an anoikis risk scoring model was established with nine genes. According to the median risk score, all cases were divided into the high-risk scoring group and the low-risk scoring group. The “stats” and “Rtsne” packets of R language were used for principal component analysis (PCA) and t-distribution random neighbor embedding (t-SNE) to detect the internal correlation between the two groups. Kaplan–Meier (K-M) curve was used to evaluate the survival difference between the two groups. The reliability of the risk model was analyzed by using the “time-dependent receiver operating characteristic (ROC)” R package. The statistical significance of all analyses was set as p < 0.05. The prognostic accuracy of the anoikis risk score was validated in the ICGC cohort. Univariate and multivariate Cox analyses were used to analyze the independent prognostic role of age, gender, grade, stage, and the anoikis risk score in the TCGA cohort and the ICGC cohort.
Associations of the anoikis risk scoring model and immune cells, immune-related functions, stemness score and PD-L1 expression
We used single sample gene set enrichment analysis (ssGSEA) via the R packages "GSVA" and "GSEABase" to evaluate the filtering scores of 16 immune cell types and the activity of 13 immune-related functions. The meaningful value is p < 0.05. The R packages "limma" and "ggpubr" were used to further visualize the relationship between each immune subtype and risk score. Spearman’s correlation test was used to test the correlation between the risk scores and tumor stemness, immune, stromal, and ESTIMATE scores. Pearson analysis was used to test the association of risk scores with the expression of the key immunomodulator PD-L1.
Drug sensitivity analysis
The data of RNA-seq expression profiles and DTP NCI-60 compound activity data were downloaded from CellMiner (https://discover.nci.nih.gov/cellminer) to analyze the drug sensitivity of nine model genes in HCC. Then we chose drugs approved by FDA to further analyze the correlation between nine model genes expression and drug sensitivity through R package “impute”, “limma”, “ggplot2”, and “ggpubr”. Pearson correlation analysis was carried out to screen the first 16 gene drugs according to the Pearson correlation coefficient, to study the relationship between prognosis anoikis-related genes and drug sensitivity.
Patients and tissue samples
This study was approved by the Human Ethics Committee of the Second Hospital of Dalian Medical University to collect tissue samples. We collected paraffin-embedded liver cancer tissues and adjacent non-tumor tissues from 88 HCC patients who underwent surgical treatment at the Second Hospital of Dalian Medical University from 2017 to 2019. These HCC patients ranged from 35 to 77 years old, including 19 females and 69 males. All patients signed an informed consent form as required by the ethics committee. Complete clinical data and follow-up information were available for all patients.
Immunohistochemistry
We first produced tissue microarrays (TMAs) of these 88 HCC tissues and matched adjacent non-cancerous tissues. Three μm sections were obtained from paraffin-embedded tissues for BIRC5 and SKP2 immunohistochemical studies according to standard procedures. Briefly, a brief protein hydrolysis digestion was performed using IHC enzyme antigen recovery agent and peroxidase blocking was performed using 3% H2O2. Then sections were incubated overnight at 4 °C with BIRC5 (Survivin) polyclonal antibody (1:200, Bioss, China) or SKP2 polyclonal antibody (1:100, Bioss, China). After PBS wash to remove the unbound primary antibody, the secondary antibody (enzyme-labeled goat anti-rabbit IgG, dilution, 1:1000) was coupled with peroxidase at incubate for 30 min at 37 °C. Sections were stained with 3,3′diaminobenzidine (DAB) substrate and washed with water. Finally, sections were counterstained with hematoxylin for 20 s, then washed with hydrochloric acid and ammonia, dehydrated, cleared, and cover-stained.
Evaluation of staining
Stained sections were scanned using an Aperio GT450 section scanner (Leica Biosystems Imaging, USA), and markers were pathologically evaluated in the Aperio ImageScope image viewer according to the percentage of positive cells and staining intensity. Protein expression was scored by two independent and experienced pathologists. The degree of staining was defined as the percentage of hepatocytes or paracancerous tissue in the positively stained area as a percentage of the entire tissue area, scored on a scale of 1–4, 1 (0–25%); 2 (26–50); 3 (51–75%); 4 (76–100%). Staining intensity was defined as a score of 0–3, 0 (negative); 1 (weakly positive); 2 (positive); 3 (strongly positive). The percentages and intensity scores were summed to give each patient a final immunoreactivity score (IRS). Low expression of BIRC5 was judged when the IRS was 1 and high expression was judged when the score was 2–3. Low expression of SKP2 was judged when the IRS was 1–3 and high expression was judged when the IRS was 4–6.
Statistical analysis
SPSS 26.0 software was used for statistical analysis, and the chi-square test was used to analyze the relationship between protein/mRNA expression and clinicopathological characteristics of HCC patients. Univariate and multivariate Cox regression analyses were used to analyze the factors influencing survival in HCC patients. The Kaplan–Meier method was used for the detection of survival analysis and the log-rank test was used to assess the differences. The relative risk of death was expressed as an adjusted hazard ratio (HR) with a 95% confidence interval (CI). If p < 0.05 all tests were considered statistically significant.
Institutional review board statement
The study was conducted in accordance with the Declaration of Helsinki, and The Ethics Committee of the Second Affiliated Hospital of Dalian Medical University approved this research.
Results
Screening and identification of prognostic anoikis‑related DEGs in HCC
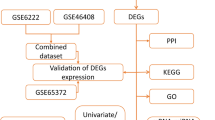
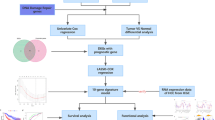
The general strategy for the present study is shown in Supplementary Fig. S1. We screened anoikis-related genes whose relevance score > 1.8 from the GeneCards and obtained 82 genes (Supplementary Table S1). Subsequently, a total of 40 genes related to HCC prognosis were obtained by univariate Cox regression analysis in the TCGA cohort (Fig. 1A). Among the above 82 genes, 38 genes were DEGs between tumor tissues and normal tissues. We intersected them and finally got 26 prognostic anoikis-related genes (Fig. 1B). Then, we performed the heatmap analysis on the expression of 26 overlapping genes in tumor tissues and normal tissues. In general, the 26 anoikis‑related genes were overexpressed in tumor tissues compared with normal tissues (Fig. 1C). Next, we used the PPI network to find the intrinsic relationship between the 26 genes. In Fig. 1D, the correlation network containing all the anoikis-related genes was presented; PTRH2, ITGAV, ANXA5, BIRC5, BDNF, BSG, DAP3, SKP2 and EGF were identified as hub genes.
Identify prognostic anoikis-related DEGs and construct the prognostic model in HCC in the TCGA cohort. (A) P-value and hazard ratio (HR) of 40 anoikis genes related to HCC prognosis by univariate Cox regression analysis. (B) Venn diagram showed anoikis-related DEGs and prognostic anoikis-related genes. (C) Heatmap showed the expression of 26 overlapping genes between tumor tissues (red) and normal tissues (blue). (D) Correlation network of 26 prognostic anoikis-related DEGs. The red lines indicated positive correlations. (E) LASSO coefficient profiles of the expression of anoikis-related DEGs. (F) Penalty value obtained through LASSO cross-validation. An optimal log λ value was indicated by the vertical black line in the figure.
Gene biological function and pathway analysis
We used GO annotation and KEGG pathway enrichment analysis to analyze the biological functions and pathways of the 26 prognostic anoikis-related DEGs. GO enrichment analysis indicated that the 26 DEGs are enriched in biological processes such as anoikis, regulation of anoikis, and regulation of apoptotic signaling pathway. The DEGs mainly have the molecular functions of integrin binding, virus receptor activity, exogenous protein binding, and so on. Besides, they were related to cell components such as focal adhesion and cell-substrate (Supplementary Fig. S2). KEGG pathway analysis revealed that these DEGs were associated with the regulation of human papillomavirus (HPV) infection, focal adhesion, regulation of actin cytoskeleton, and hepatitis (Supplementary Fig. S2). In general, the prognostic anoikis-related DEGs have momentous biological effects in HCC.
Construction of the prognostic model based on anoikis‑related genes in the TCGA cohort
Subsequently, we developed the risk score model based on 26 prognostic anoikis-related genes (Supplementary Fig. S3). The Lasso coefficient map of 26 candidate gene expression levels was shown in Fig. 1E. The 26 candidate genes were subjected to Lasso regression analysis for dimensionality reduction, and cross-validation was used to obtain the optimal penalty value (λ) in Lasso regression (Fig. 1F). An anoikis prognostic risk score model was established according to the regression coefficient, and the risk score was calculated using the following formula: risk score = 0.093040 × PTRH2 + 0.014600 × ITGAV + 0.071630 × ANXA5 + 0.099028 × BIRC5 + 0.145664 × BDNF + 0.176856 × BSG + 0.156909 × DAP3 + 0.130661 × SKP2 + 0.041248 × EGF. Based on the median risk score calculated by the risk score formula, HCC patients in the TCGA cohort were divided into high-risk or low-risk groups (Fig. 2A). The results of PCA and t-SNE analysis showed that HCC patients in different risk groups were well-separated (Fig. 2B,C). In addition, we further investigated the relationship between survival status, survival time and the risk score. As shown in Fig. 2D, HCC patients presented higher mortality and shorter survival time in the high-risk group than that in the low-risk group. The prognostic significance of this risk model in HCC was further identified utilizing K-M analysis. In Fig. 2E, the OS of the high-risk group was greatly lower than that of the low-risk group (p = 2.338e−05). We also used the time-ROC analysis curve to evaluate the sensitivity and specificity of the prognostic risk score model. The area under the receiver operating characteristic curve (AUC) was 0.778 at 1 year, 0.713 at 2 years, and 0.697 at 3 years (Fig. 2F).
Validation of anoikis prognostic risk model in the TCGA cohort. (A–D) Risk score (A) distribution, (B) PCA plot, (C) t-SNE, and (D) OS status analysis of the TCGA cohort. (E) Kaplan–Meier survival curves were used to compare OS between the low-risk and the high-risk groups. (F) ROC curves verified the prognostic performance of the risk model.
External validation of anoikis prognostic risk model in the ICGC cohort
To test the reliability of the prognostic risk model established in the TCGA cohort, we selected HCC patients from the ICGC cohort and also divided them into the low-risk group or high-risk group by the median risk score calculated in the TCGA cohort (Fig. 3A). Similarly, PCA and t-SNE analysis exhibited excellent separation between the two groups (Fig. 3B,C). HCC patients in the high-risk group also have a shorter survival time compared with the low-risk group (Fig. 3D). Besides, the K-M survival curve showed the OS of the high-risk group was vastly shorter compared with that in the low-risk group (p = 3.712e−08, Fig. 3E). At the same time, the time-ROC curve revealed the strong predictive power of our model in the ICGC cohort, and the AUC predictive values for 1-year, 2-years, and 3-years survival rates were 0.733, 0.700, and 0.722 respectively (Fig. 3F). These results confirmed that our prognostic risk model was reliable and exportable.
External Validation of anoikis prognostic risk model in the ICGC cohort. (A–D) Risk score (A) distribution, (B) PCA plot, (C) t-SNE, and (D) OS status analysis of the ICGC cohort. (E) Kaplan–Meier survival curves were used to compare OS between the low-risk and the high-risk groups. (F) ROC curves verified the prognostic performance of the risk model.
Independent prognostic value of the risk model
To determine whether the risk score could be an independent prognostic indicator, we used the univariate and multivariate Cox regression analysis to evaluate it and other clinical characteristics (age, gender, pathological TNM stage and grade). According to the results of univariate regression analysis, the risk score and pathological TNM stage were significantly correlated with OS of HCC patients in the TCGA cohort (all p values < 0.001, Fig. 4A). The risk score, pathological TNM stage, and gender were significantly associated with OS of HCC patients in the ICGC cohort (p = 0.031 in gender, and other p values < 0.001, Fig. 4B). Furthermore, we conducted the multivariate Cox regression analysis to correct the influencing factors, and the risk score was an independent predictor of OS (TCGA cohort: HR = 3.300, 95% CI = 2.154–5.056, p < 0.001; ICGC cohort: HR = 3.461, 95% CI = 1.572–7.622, p = 0.002, Fig. 4C,D). In particular, the HR of our risk score was higher than that of the pathological TNM stage.
Univariate and multivariate Cox regression analyses for the anoikis risk score and their levels between different clinicopathologic characteristics. (A,B) Univariate Cox regression analysis of OS in (A) the TCGA cohort and (B) the ICGC cohort. (C,D) Multivariate Cox regression analysis of OS in (C) the TCGA cohort and the (D) ICGC cohort. (E–I) Anoikis risk score levels of age (≤ 65 and > 65), gender (female and male), groups (group 1–2 and group 3–4), and pathological TNM stages (stage I-II and stage III-IV) in the TCGA cohort (E, G, and H) and the ICGC cohort (F and I).
In addition, we compared the risk score of HCC patients in different clinical characteristic groups. The score was equivalently insignificant between the different age and gender groups (TCGA cohort: p = 0.150 and 0.071, Fig. 4E; ICGC cohort: p = 0.87 and 0.63, Fig. 4F). In the TCGA cohort, the risk score of grade 3–4 patients was higher than that of grade 1–2 (p = 4.2e−09, Fig. 4G). The risk score of pathological TNM III-IV was also found to be higher than that of stage I-II (p = 0.0027, Fig. 4H). In the ICGC cohort, the risk score of stage III-IV was also found to be higher than that of stage I-II (p = 0.00031, Fig. 4I). These results suggested the high risk score of our model was connected with high-level (grade 3–4) and advanced stage HCC patients.
Relationship between immune infiltration and anoikis risk score of HCC
To explore the potential clinical relevance between immune status and anoikis, it is worth conducting ssGSEA to calculate the enrichment scores of various immune cells, related functions, or pathways in the TCGA and ICGC cohorts. As shown in Fig. 5A and B, aDCs, macrophages, T-follicular helper cells, and Th2 cells all showed high infiltration in the high-risk groups in the TCGA and ICGC cohorts (p < 0.05). Regarding immune-related pathways, MHC class I was significantly upregulated in the high-risk group, while type I IFN response and type II IFN response were downregulated in the TCGA cohort and the ICGC cohort (all adjusted p < 0.05, Fig. 5C,D) and the. In addition, we explored the risk scores of the four groups of tumor-infiltrating immune cell subtypes (C1, C2, C3, C4), as shown in Fig. 5E. We found that the highest risk score was closely related to group C1, and the risk score of group C3 was the lowest.
Relationship between immune infiltration and anoikis risk score of HCC. (A–D) Box plots presented 16 remarkable immune cells and 13 immune-related functions between two groups in the TCGA cohort and the ICGC cohort. The adjusted p-value was shown as ns, which is not significant; *p < 0.05; **p < 0.01; and ***P < 0.001. (E) Comparison of risk scores in four immune cell subtypes.
Correlation between tumor stemness, tumor microenvironment, PD-L1 expression and anoikis risk score
Tumor stemness, including RNAss and DNAss, is a key regulator of tumor progression. DNA methylation-based stem cell index (Manasi) can reflect epigenetic stem cell characteristics, and mRNA expression-based mRNAs represent stem cell expression of the transcriptome. The risk score had no relationship with DNAss, but positively correlated with RNAss (Supplementary Fig. S4A,B). Besides, immune and stromal scores were used to evaluate tumor microenvironment (TME). There was a negative correlation between the risk score and stromal score, but no relationship between the risk score and immune score (Supplementary Fig. S4C,D). The expression level of PD-L1 was significantly higher in the high-risk group compared with the low-risk group (p = 0.00048, Supplementary Fig. S4E), and it was positively correlated with the risk score (R = 0.18, p = 0.00047, Supplementary Fig. S4F).
Relationship between the expression of anoikis-related model genes and drug sensitivity
Anoikis-related genes play crucial roles in regulating drug resistance. We used the CellMiner database to explore the relationship between nine model genes and drug sensitivity. The top 16 gene-drug pairs ranked by Pearson correlation coefficient were shown in Fig. 6, which includes Dexrazoxane, LDK-378, Ibrutinib, Erlotinib, Oxaliplatin, Dasatinib, Lomusti, Tamoxifen, Tyrothricin, Nelarabine, Bleomycin, Ifosfamide, Palbociclib and Lenvatinib, respectively. We found that a total of 4 genes were associated with drug sensitivity, namely ANX5, ITGAV, BDNF, and SKP2. SKP2 was positively correlated with the sensitivity of chemotherapy, while ANXA5 was negatively correlated with the sensitivity of targeted drugs. Interestingly, ANXA5 was negatively associated with seven drug sensitivity tests, including chemotherapy drugs and targeted drugs.
Scatter plots for the correlation between nine model genes expression and drug sensitivity (top 16 pairs ranked by p value).
WGCNA identified hub genes in our anoikis model
To find the most critical genes among the 9 model genes for further study, we performed WGCNA analysis on the TCGA database. Based on the scale-free topological model and average connectivity of the WGCNA analysis, the soft threshold was set to 2 (Supplementary Fig. S5A). The clustered gene dendrogram showed HCC patient co-expression modules, with each leaf representing a gene and branches on the tree representing co-expression modules. (Supplementary Fig. S5B). Eleven correlations were determined by WGCNA analysis between modules and normal samples and HCC, with the pink module having the highest positive correlation in HCC (cor = 0.4, p = 3e−25, Fig. 7A). We identified two hub genes (BIRC5 and SKP2) by intersecting the pink module genes with our model genes through Venn diagrams (Fig. 7B).
WGCNA identified hub genes in our anoikis model. (A) Heatmap of the correlation of gene modules with normal tissues and HCC tissues. (B) Venn diagram analysis showed that the overlap of WGCNA analysis and LASSO model led to two hub genes: BIRC5 and SKP2. (C,D) Differential expression of (C) BIRC5 and (D) SKP2 in normal tissues and HCC tissues based on TCGA database. (E,F) K-M curves showing the difference of OS between the (E) BIRC5/(F) SKP2 high and low expression groups. (G,H) K-M curves showing the difference of PFI between the (G) BIRC5/(H) SKP2 high and low expression groups.
Based on the TCGA database, we explored the expression differences of these 2 hub genes between normal tissues and HCC tissues. The results revealed that both BIRC5 and SKP2 were significantly highly expressed in HCC tissues compared to normal tissues (Fig. 7C,D). K-M curves showed that both BIRC5 and SKP2 high expression groups were associated with poor prognosis in patients with HCC compared to low expression groups, both in terms of OS (Fig. 7E,F) and progression-free interval (PFI, Fig. 7G,H).
Correlation between the expression levels of hub genes and clinicopathological characteristics in the TCGA cohort
The correlation between BIRC5 and SKP2 mRNA expression and clinicopathological characteristics of HCC was assessed based on the TCGA database. Supplementary Table S2 showed that BIRC5 mRNA expression levels were significantly correlated with T-stage, pathologic stage, histologic grade, and Alpha-fetoprotein (AFP). In addition, Supplementary Table S3 displayed that SKP2 mRNA expression levels were significantly correlated with age, histologic grade, and AFP.
Our study further investigated the effects of BIRC5 and SKP2 mRNA expression on OS and PFI in HCC by univariate Cox and multivariate Cox regression analysis. Univariate Cox regression showed that the T stage, M stage, pathologic stage, BIRC5 mRNA expression and SKP2 mRNA expression were correlated with OS in HCC (Supplementary Table S4A). Multivariate Cox regression showed that BIRC5 mRNA expression was an independent factor (Supplementary Table S4B). As for PFI, univariate Cox regression suggested that the T stage, M stage, pathologic stage, vascular invasion, BIRC5 mRNA expression and SKP2 mRNA expression were correlated with it in HCC (Supplementary Table S5A). BIRC5 mRNA expression was an independent factor influencing PFI in HCC (Supplementary Table S5B).
We also analyzed the expression of these two factors in several other cancers, including lung adenocarcinoma (LUAD), breast invasive carcinoma (BRCA), as well as most other digestive tumors, including cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), esophageal carcinoma (ESCA), pancreatic adenocarcinoma (PAAD) and rectum adenocarcinoma (READ), to verify the specificity of their prognostic value in HCC. The median expression levels in these cancers themselves were used as cutoff. K-M curves showed that only BIRC5 high expression group were associated with poor OS in PAAD, and BIRC5 has no prognostic value in other cancers. SKP2 has no prognostic value in all of the above cancers (Supplementary Fig. S6). So overall, the value of these two factors for poor prognosis in HCC is somewhat specific.
Immunohistochemical staining of BIRC5 and SKP2 and correlation with clinicopathological characteristics in our TMA cohort
To further experimentally verify the expression levels of BIRC5 and SKP2, 88 HCC and adjacent non-cancerous tissue samples were collected in this study, and the expression levels of BIRC5 and SKP2 were detected by immunohistochemical staining (Fig. 8A,F). BIRC5 was mainly expressed in the nucleus of HCC cells. The IRS of BIRC5 protein was significantly higher in HCC than in adjacent non-cancerous tissues (Fig. 8B–E). In addition, SKP2 was mainly expressed in the cytoplasm of HCC cells. SKP2 protein levels were also significantly higher in HCC tissues than in adjacent non-cancerous tissues (Fig. 8G–K).
Experimental validation of immunohistochemical staining of BIRC5 and SKP2 in oure TMA cohort and their prognostic value. (A) Immunohistochemical staining of BIRC5 in the HCC TMA cohort. (B) The IRS of BIRC5 in HCC and adjacent non-cancerous tissues. ****p < 0.0001; Data are presented as the mean ± standard error of mean. (C–E) The intensities of BIRC5 immunostaining were (C) positive, (D) weakly positive and (E) negative, no strongly staining was observed in HCC tissues. Left image, magnification ×100; right image, magnification ×400; the red squares indicate the area shown in higher magnification. (F) Immunohistochemical staining of SKP2 in the HCC TMA cohort. (G) The IRS of SKP2 in HCC and adjacent non-cancerous tissues. ****p < 0.0001; Data are presented as the mean ± standard error of mean. (H–K) The intensities of SKP2 immunostaining were (H) strongly positive, (I) positive, (J) weakly positive and (K) negative. Left image, magnification ×100; right image, magnification ×400; the red squares indicate the area shown in higher magnification. (L,M) Kaplan–Meier survival curves to compare (L) OS and (M) PFI between the high-BIRC5 expression and low-BIRC5 expression patients. (N,O) Kaplan–Meier survival curves to compare (N) OS and (O) PFI between the high-SKP2 expression and low-SKP2 expression patients.
We collected detailed information on the clinical characteristics of the TMA cohort to further validate the correlation between BIRC5 or SKP2 expression and clinicopathological characteristics of HCC. Table 1 showed that BIRC5 expression levels were significantly correlated with T-stage, pathologic stage, histologic grade, and AFP. For SKP2, its expression levels were only significantly correlated with age and smoking history (Supplementary Table S6).
In order to further investigate the potential differential expressions of BIRC5 or SKP2 in liver tissues affected by chronic liver diseases such as hepatitis and cirrhosis, as compared to normal liver tissues, we re-classified our original liver control tissues based on the presence or absence of hepatitis and cirrhosis. We first conducted an analysis using Pearson's Chi-squared test and Fisher's exact test. The results revealed that the expression level of BIRC5 was found to be independent of the history of cirrhosis and hepatitis among HCC patients (Table 2A). On the other hand, the expression level of SKP2 showed a significant correlation with the cirrhosis history of HCC patients (p = 0.042), but not with their hepatitis history (Table 2B). However, when examining the IRS scores, no significant difference was observed in the expression of BIRC5 or SKP2 between tissues with or without cirrhosis, as well as between tissues with or without hepatitis (Supplementary Fig. S7). These findings suggest that although there is a higher prevalence of SKP2 positivity among patients with cirrhosis, there are still individuals without cirrhosis who exhibit high SKP2 expression. Consequently, there is no significant difference in the mean values between these two groups. Therefore, further investigation is required to elucidate the role of SKP2 in cirrhosis.
Prognostic value of BIRC5 and SKP2 in our TMA cohort
The K-M survival curves of the TMA cohort revealed that high expression of BIRC5 was associated with poor prognosis in HCC (Fig. 8L, PFI, p < 0.05; Fig. 8M, OS, p < 0.05). Similarly, PFI (p < 0.05, Fig. 8N) and OS (p < 0.01, Fig. 8O) were significantly lower in SKP2-high expressing HCC compared to SKP2-low expressing HCC. These findings are consistent with the results of our bioinformatics analysis.
Univariate Cox regression showed that T-stage, pathologic stage, histologic grade, BIRC5 expression, and SKP2 expression affected OS in the HCC TMA cohort (Table 3A). Multivariate Cox regression showed that pathologic stage and histologic grade were independent factors affecting OS (the above clinicopathological characteristics and BIRC5 or SKP2 expression, Table 3B). As for PFI, T-stage, pathologic stage, histologic grade, BIRC5 expression, and SKP2 expression were correlated to PFI by univariate Cox regression (Supplementary Table S7A). Multivariate Cox regression showed that the pathologic stage was an independent factor influencing PFI (Supplementary Table S7B).
Discussion
Given the high heterogeneity observed in HCC patients, it is significant to develop a new prognostic system to predict the outcomes of the patients more accurately. It has been proved that anoikis can block cancer development and resistance in anoikis leads to cancer progression and metastasis. So we obstructed and verified a prognostic risk model based on anoikis-related genes in our study. In addition, we identified BIRC5 and SKP2 as hub genes and preliminarily validated them in the TCGA cohort and our own TMA cohort.
The TNM system, widely applied in the majority of substantial tumors, fails to account for the degree of liver dysfunction and performance status of patients29. Although the BCLC system may provide more prognostic information, it still lacks universal applicability in clinical practices6. Under the situation that prognostic systems for HCC patients have certain shortages, several signatures based on hub genes have been established in recent research30,31. Here, our model has higher predictive power in contrast to previous studies, for our model was successfully tested in both internal and external databases. Particularly, the time-ROC curve shows that AUC predictive values for the 1-year survival rate are 0.778 in internal validation and 0.733 in external validation, respectively. We noted the AUCs of the prognostic model based on DNA methylation-driven genes for 1 year, 2 years, and 3 years were 0.650, 0.696, and 0.66432. Hence, our model is probably more robust and effective. We found that the high-risk score was correlated with adverse clinical characteristics, such as low tumor differentiation and advanced stage, which contribute to poor survival outcomes of high-risk patients. Besides, univariate and multivariate Cox regression analyses revealed that our prognostic signature is capable of serving as an independent prognostic factor for patients with HCC. Furthermore, We not only investigated the role of anoikis in the immune microenvironment but also explored the relationship between anoikis-related DEGs expression and drug sensitivity.
In the present study, we identified a risk model consisting of the nine anoikis-related genes (PTRH2, ITGAV, ANXA5, BIRC5, BDNF, BSG, DAP3, SKP2, and EGF) for predicting the prognosis of HCC. Recently, Chen et al. constructed a prognostic anoikis model in HCC using part of the TCGA database as a training dataset33. Since we used higher criteria for screening the genes of anoikis (relevance score > 1.8) and the whole TCGA dataset as a training set, only BSG is the same gene among our 9 model genes and the above model. Therefore, our model and their model are completely different. More importantly, we have identified hub genes, BIRC5 and SKP2 in HCC anoikis model using the WGCNA method, which is an innovative finding. Furthermore, we validated the overexpression and prognostic value of these two hub genes in liver cancer through immunohistochemical staining in our cohort, providing further confirmation.
Multiple studies have reported that most of the nine anoikis-related genes are related to cancer. IGTAV, involved in cell–cell and ECM adhesions, serves as an independent prognostic factor in esophageal adenocarcinoma (EAC) without neoadjuvant therapy34. In colon cancer cells, activation of TP53 suppresses ITGAV expression, resulting in cell survival35. Weiler et al. confirmed that overexpression of IGTAV is associated with poor clinical outcomes in patients with HCC36. Cilengitide is an ITGAV antagonist, and Mas-Moruno et al. identified that cilengitide reduces angiogenesis and inhibits breast cancer bone metastasis in vivo preliminary results37,38. Annexin A5 (ANXA5) was reported to participate in HCC pathogenesis via integrin and MEK-ERK pathways39. Kang et al. suggested that the administration of ANXA5 alleviates the immunosuppressive properties of TME generated by chemotherapy and improves the efficacy of anti-tumor therapy. So the ANXA5 administration following chemotherapy could be considered as a promising immune checkpoint inhibitor for cancer treatment40. Basigin (BSG), also known as cluster of differentiation 147 (CD147), is overexpressed and considered to be a prognostic marker in various human tumors41,42,43,44,45. It has been observed that CD147 increases cathepsin B expression by activating β-catenin signaling, which mediates CD147-induced invasive phenotype in HCC46. Studies have shown that BSG-directed monoclonal antibodies excite clinical success in HCC treatment47. Additionally, the overexpression of brain-derived neurotrophic factor (BDNF) and tropomyosin-related kinase B (TrkB) receptors is associated with lower OS in HCC48. It was reported that the BDNF/TrkB axis can promote several important biological processes in several types of cancer, such as cell proliferation, tumor immunosuppression, drug resistance, resistance to anoikis, activation of PI3K/AKT and JAK/STAT3 pathways, EMT, and angiogenesis49. Several studies have suggested that the downregulation of BDNF suppresses cell viability and metastasis in multiple solid tumors50,51. Therefore, the BDNF/TrkB axis can be considered as a potential therapeutic target for several types of cancer. DAP3 was reported to function as a positive mediator of anoikis. Han et al. found that the expression of DAP3 is significantly associated with the malignant properties of HCC cells. DAP3 suppresses adenosine-to-inosine (A-to-I) RNA epitome to further promote cancer progression52. It was observed that EGF is highly expressed in HCC and positively relates to higher tumor grade. In addition, EGF promotes the migration and aggressiveness of HCC cells mainly via the induction of fibronectin (FN) in vitro53.
To identify pivotal genes, we utilized the WGCNA algorithm to screen key genes and acquired the overlapping genes among these key genes and our model genes. We obtained the two hub genes: BIRC5 and SKP2. It is necessary to further investigate the potential mechanisms of the two hub genes.
Survivin (BIRC5), a member of the inhibitor of the apoptosis family, can promote cell division and tumor proliferation54. Hori et al. found that BIRC can predominantly suppress anoikis55. Xu et al. suggested that BIRC5 was upregulated in HCC and associated with a poor prognosis56. Co-suppression of OCT4 and BIRC5 can inhibit cancer proliferation by inducing cancer cell apoptosis and cell cycle arrest in HCC57. Also, the downregulation of BIRC5 leads to the inhibition of migration and invasion in penile cancer58. Thereby, BIRC5 should be regarded as a potential target for anti-cancer drugs and prognosis prediction59.
S-phase kinase-associated protein 2 (SKP2), also known as p45, is involved in cancer progression, migration, and invasion60. High expression of SKP2 was widely observed in various human tumors, indicating the close correlation between tumor progression and SKP261,62,63. It is reported that the knockdown of the endogenous SKP2 expression by RNA interference reduces cell proliferation, blocks the cell cycle at the G1 phase, and increases apoptosis in HCC64. Delogu et al. reported that SKP2 cooperates with N-Ras and AKT proto-oncogenes to promote carcinogenesis in HCC65. Currently, SKP2 is not only an independent prognostic factor for HCC but also a new target for anti-tumor drugs and gene therapy52,66.
We further preliminarily verified the two hub genes based on the TCGA database and HCC tissues we collected. We not only analyzed the mRNA expression of the two hub genes in the TCGA cohort, but also performed immunohistochemical staining to explore the protein expression in our TMA cohort. In the TCGA cohort, we observed the upregulation of the two hub genes in HCC patients, accompanied by poor survival outcomes. The expression level of BIRC5 mRNA was associated with the T stage, pathologic stage, histologic grade and AFP. SKP2 mRNA expression level was closely related to age, histologic grade and AFP. In our TMA cohort, all of the HCC patients have indications for surgery, so the percentage of early-stage patients is more than the TCGA cohort (pathologic stage III & IV is 25.7% in the TCGA cohort, and it is 15.9% in our cohort). Therefore, the OS of HCC patients in our TMA cohort seems relatively longer than that in the TCGA cohort. Besides, HCC tissues and adjacent non-cancerous tissues in our cohort are paired, from the same patients. Even though there are some differences between our patients and the TCGA cohort patients, BIRC5 and SKP2 were still both overexpressed in HCC tissues compared with tumor-adjacent tissues in our TMA cohort. Moreover, high expression of BIRC5 or SKP2 protein also indicated poor prognosis in our TMA cohort. As previously mentioned, Fang et al. suggested that silencing of AFP may reduce intracellular BIRC5 mRNA expression to promote apoptosis in HepG2 cells67. In lung adenocarcinoma, the high level of BIRC5 was related to advanced AJCC stage, T and N stages68. Finally, Taken together, the results above indicated that the expression of the hub genes has a strong correlation with the prognosis and clinical features in HCC, no matter mRNA or protein, which further validated the stability and accuracy of our risk model.
We further employed functional analysis and found that the prognostic model was enriched in anoikis regulation and tumor-related pathways, including PI3K-Akt, ErbB, and P53 pathways. Additionally, our model was significantly associated with virus infection, such as HPV, Human immunodeficiency virus (HIV), Hepatitis B virus (HBV), and Hepatitis C virus (HCV). As revealed by Kakavandi and his colleagues, viral infection promotes resistance to anoikis and provides the opportunity for cancer cells to spread to distant organs, which is in line with our research results69,70,71.
Currently, immunotherapy opens a new chapter for cancer treatment. Investigation of the potential mechanism between immune microenvironment and tumor progression is needed. Studies revealed that macrophages have two extreme functional phenotypes, including M1 and M2 macrophages. Tumor-associated macrophages (TAMs) usually acquire an M2-like phenotype and play vital roles in the TME72. TAMs produce several cytokines to suppress other immune cells. Besides, TAMs could actively enhance the remodeling of the tumor stroma and facilitate the metastatic process of cancer cells by producing several proteolytic enzymes73,74,75. In addition, Li et al. found that the high level of Th2 cells is significantly correlated with poor prognosis in HCC76. In our study, the high-risk group has higher infiltration of macrophages and Th2 cells, which means macrophages and Th2 cells are risky immune cells for HCC. We also discovered that PD-L1 was dramatically upregulated in the high-risk group, pointing out that PD-L1 inhibitors could be more effective in HCC patients in the high-risk group. Besides, it is important to consider the contributions of BIRC5 and SKP2 to the tumor microenvironment. BIRC5 is a potential biomarker and inducer of myeloid-derived suppressor cell (MDSC) infiltration in HCC, leading to T cell rejection or dysfunction of the tumor immune microenvironment, ultimately reducing the response to immune checkpoint inhibitors (ICIs)77. Cytotoxic T lymphocytes (CTLs) are the primary lymphocytes responsible for killing cancer cells in liver cancer78. They primarily restrict tumor growth by inducing G1 phase cell cycle arrest, a mechanism that may involve downregulation of SKP279. Additionally, SKP2 knockdown has been found to significantly increase the expression of immunoinfiltration-related genes in osteosarcoma (OS) mouse models, suggesting that SKP2 may mediate immune rejection in the tumor microenvironment80.
Resistance of cancer cells to chemotherapy remains a big challenge in cancer treatment. Resistance to cisplatin treatment is a major clinical issue in HCC81. Studies have demonstrated that suppressing BIRC5 can enhance the cytotoxicity of chemotherapy agents like cisplatin82. Asechi et al. discovered that BIRC5 expression, mediated by PI3K, enables rat hepatoma cell lines to resist cisplatin-induced apoptosis83. Sorafenib, the first-line chemotherapy agent for advanced HCC, sensitizes resistant HCC cells to radiation-induced apoptosis by inhibiting STAT3 phosphorylation and reducing BIRC5 expression84. Furthermore, the combination of Sorafenib (SF) and BIRC5 shRNA has shown a synergistic effect in reversing multidrug resistance (MDR). Regarding SKP2, studies have reported that both lonafarnib and troglitazone induce G1-S phase block by down-regulating SKP2. Conversely, ectopic overexpression of SKP2 leads to resistance to troglitazone, resulting in a decrease in the number of G1-phase blocked cells85,86. Hepatitis B virus X has been found to downregulate SHIP, promoting HCC metastasis and chemical resistance by inducing SKP2 expression87. Oxaliplatin (OXA) is one of the most common chemotherapy drugs for patients with HCC88. In HCC, cyanidin significantly increases OXA sensitivity and inhibits the EMT induced by OXA via PI3K/Akt signaling89. In addition, PKI-587, the dual PI3K/mTOR inhibitor, increases the chemosensitivity of OXA in HCC90. In our study, the high level of ANXA5 and ITGAV is negatively related to the low sensitivity to OXA.The underlying mechanism between low sensitivity to OXA and ANXA5 and ITGAV needs further exploration. Currently, Lenvatinib is considered a new first-line targeted therapy for patients with advanced HCC. In our study, high expression of ITGAV is positively related to the high sensitivity of lenvatinib, which indicates that HCC patients with high expression of ITGAV could benefit from lenvatinib. In summary, the combination of specific target inhibitors and chemotherapy drugs appears to be a promising option for the treatment of HCC.
However, a few limitations of this study should be taken into consideration. First, the construction and validation of our prognostic model are based on retrospective databases, so our results should be further confirmed by prospective clinical research. Second, further experimental studies of the nine anoikis-related genes in HCC progression are needed.
Conclusions
Our study established a novel risk signature based on nine anoikis-related genes, which perform a good capability in predicting the prognosis of HCC patients. In addition, our study reflected that risk score was closely related to the immune microenvironment and analyzed the relationship between anoikis-related genes and drug resistance. Finally, we identified BIRC5 and SKP2 as hub genes among the nine model genes, and verified them at the mRNA and protein levels. The results confirmed that BIRC5 and SKP2 should be considered as potential prognostic predictors in HCC. Altogether, our study can not only provide potential therapeutic targets for HCC but also bring new insights into drug resistance and ameliorate the effect of HCC treatment.
Data availability
The datasets generated and/or analysed during the current study are available in the TCGA repository (https://portal.gdc.cancer.gov/) and ICGC repository (https://dcc.icgc.org/).
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249. https://doi.org/10.3322/caac.21660 (2021).
Anwanwan, D., Singh, S. K., Singh, S., Saikam, V. & Singh, R. Challenges in liver cancer and possible treatment approaches. Biochim. Biophys. Acta Rev. Cancer 1873(1), 188314. https://doi.org/10.1016/j.bbcan.2019.188314 (2020).
Forner, A., Reig, M. & Bruix, J. Hepatocellular carcinoma. Lancet 391(10127), 1301–1314. https://doi.org/10.1016/s0140-6736(18)30010-2 (2018).
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 71(1), 7–33. https://doi.org/10.3322/caac.21654 (2021).
Marrero, J. A. et al. Diagnosis, staging, and management of hepatocellular carcinoma: 2018 practice guidance by the American Association for the Study of Liver Diseases. Hepatology 68(2), 723–750. https://doi.org/10.1002/hep.29913 (2018).
Maida, M., Orlando, E., Cammà, C. & Cabibbo, G. Staging systems of hepatocellular carcinoma: A review of literature. World J. Gastroenterol. 20(15), 4141–4150. https://doi.org/10.3748/wjg.v20.i15.4141 (2014).
Vitale, A. et al. Italian Liver Cancer Ita Li Ca study: Overview of prognostic systems for hepatocellular carcinoma and ITA.LI.CA external validation of MESH and CNLC classifications. Cancers https://doi.org/10.3390/cancers13071673 (2021).
Taddei, M. L., Giannoni, E., Fiaschi, T. & Chiarugi, P. Anoikis: An emerging hallmark in health and diseases. J. Pathol. 226(2), 380–393. https://doi.org/10.1002/path.3000 (2012).
Adeshakin, F. O. et al. Mechanisms for modulating anoikis resistance in cancer and the relevance of metabolic reprogramming. Front. Oncol. 11, 626577. https://doi.org/10.3389/fonc.2021.626577 (2021).
Paoli, P., Giannoni, E. & Chiarugi, P. Anoikis molecular pathways and its role in cancer progression. Biochim. Biophys. Acta 1833(12), 3481–3498. https://doi.org/10.1016/j.bbamcr.2013.06.026 (2013).
Kim, Y. N., Koo, K. H., Sung, J. Y., Yun, U. J. & Kim, H. Anoikis resistance: An essential prerequisite for tumor metastasis. Int. J. Cell Biol. 2012, 306879. https://doi.org/10.1155/2012/306879 (2012).
Hu, P. et al. Acidosis enhances the self-renewal and mitochondrial respiration of stem cell-like glioma cells through CYP24A1-mediated reduction of vitamin D. Cell Death Dis. 10(1), 25. https://doi.org/10.1038/s41419-018-1242-1 (2019).
Vander Linden, C. & Corbet, C. Therapeutic targeting of cancer stem cells: Integrating and exploiting the acidic niche. Front. Oncol. 9, 159. https://doi.org/10.3389/fonc.2019.00159 (2019).
Wang, C. et al. Elevated level of mitochondrial reactive oxygen species via fatty acid β-oxidation in cancer stem cells promotes cancer metastasis by inducing epithelial-mesenchymal transition. Stem Cell Res. Ther. 10(1), 175. https://doi.org/10.1186/s13287-019-1265-2 (2019).
Simpson, C. D., Anyiwe, K. & Schimmer, A. D. Anoikis resistance and tumor metastasis. Cancer Lett. 272(2), 177–185. https://doi.org/10.1016/j.canlet.2008.05.029 (2008).
Wang, Y. N. et al. CPT1A-mediated fatty acid oxidation promotes colorectal cancer cell metastasis by inhibiting anoikis. Oncogene 37(46), 6025–6040. https://doi.org/10.1038/s41388-018-0384-z (2018).
Ye, G. et al. Nuclear MYH9-induced CTNNB1 transcription, targeted by staurosporin, promotes gastric cancer cell anoikis resistance and metastasis. Theranostics 10(17), 7545–7560. https://doi.org/10.7150/thno.46001 (2020).
Seitz, A. K. et al. Profiling of long non-coding RNAs identifies LINC00958 and LINC01296 as candidate oncogenes in bladder cancer. Sci. Rep. 7(1), 395. https://doi.org/10.1038/s41598-017-00327-0 (2017).
Jin, L. et al. The PLAG1-GDH1 axis promotes anoikis resistance and tumor metastasis through CamKK2-AMPK signaling in LKB1-deficient lung cancer. Mol. Cell 69(1), 87-99.e7. https://doi.org/10.1016/j.molcel.2017.11.025 (2018).
Wheeler, L. J. et al. CBX2 identified as driver of anoikis escape and dissemination in high grade serous ovarian cancer. Oncogenesis 7(11), 92. https://doi.org/10.1038/s41389-018-0103-1 (2018).
Lee, Y. et al. TMPRSS4 promotes cancer stem-like properties in prostate cancer cells through upregulation of SOX2 by SLUG and TWIST1. J. Exp. Clin. Cancer Res. 40(1), 372. https://doi.org/10.1186/s13046-021-02147-7 (2021).
Greten, T. F., Lai, C. W., Li, G. & Staveley-O’Carroll, K. F. Targeted and immune-based therapies for hepatocellular carcinoma. Gastroenterology 156(2), 510–524. https://doi.org/10.1053/j.gastro.2018.09.051 (2019).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359(6382), 1350–1355. https://doi.org/10.1126/science.aar4060 (2018).
Hernandez-Gea, V., Toffanin, S., Friedman, S. L. & Llovet, J. M. Role of the microenvironment in the pathogenesis and treatment of hepatocellular carcinoma. Gastroenterology 144(3), 512–527. https://doi.org/10.1053/j.gastro.2013.01.002 (2013).
Zhang, Y. & Zhang, Z. The history and advances in cancer immunotherapy: Understanding the characteristics of tumor-infiltrating immune cells and their therapeutic implications. Cell Mol. Immunol. 17(8), 807–821. https://doi.org/10.1038/s41423-020-0488-6 (2020).
Finn, R. S. et al. Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N. Engl. J. Med. 382(20), 1894–1905. https://doi.org/10.1056/NEJMoa1915745 (2020).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47. https://doi.org/10.1093/nar/gkv007 (2015).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Amin, M. B. et al. The eighth edition AJCC cancer staging manual: Continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J. Clin. 67(2), 93–99. https://doi.org/10.3322/caac.21388 (2017).
Huo, J., Wu, L. & Zang, Y. Development and validation of a CTNNB1-associated metabolic prognostic model for hepatocellular carcinoma. J. Cell Mol. Med. 25(2), 1151–1165. https://doi.org/10.1111/jcmm.16181 (2021).
Long, J. et al. Development and validation of a TP53-associated immune prognostic model for hepatocellular carcinoma. EBioMedicine 42, 363–374. https://doi.org/10.1016/j.ebiom.2019.03.022 (2019).
Long, J. et al. DNA methylation-driven genes for constructing diagnostic, prognostic, and recurrence models for hepatocellular carcinoma. Theranostics 9(24), 7251–7267. https://doi.org/10.7150/thno.31155 (2019).
Chen, Y., Huang, W., Ouyang, J., Wang, J. & Xie, Z. Identification of anoikis-related subgroups and prognosis model in liver hepatocellular carcinoma. Int. J. Mol. Sci. https://doi.org/10.3390/ijms24032862 (2023).
Loeser, H. et al. Integrin alpha V (ITGAV) expression in esophageal adenocarcinoma is associated with shortened overall-survival. Sci. Rep. 10(1), 18411. https://doi.org/10.1038/s41598-020-75085-7 (2020).
Janouskova, H. et al. Activation of p53 pathway by Nutlin-3a inhibits the expression of the therapeutic target α5 integrin in colon cancer cells. Cancer Lett. 336(2), 307–318. https://doi.org/10.1016/j.canlet.2013.03.018 (2013).
Weiler, S. M. E. et al. TAZ target gene ITGAV regulates invasion and feeds back positively on YAP and TAZ in liver cancer cells. Cancer Lett. 473, 164–175. https://doi.org/10.1016/j.canlet.2019.12.044 (2020).
Bäuerle, T. et al. Cilengitide inhibits progression of experimental breast cancer bone metastases as imaged noninvasively using VCT, MRI and DCE-MRI in a longitudinal in vivo study. Int. J. Cancer 128(10), 2453–2462. https://doi.org/10.1002/ijc.25563 (2011).
Mas-Moruno, C., Rechenmacher, F. & Kessler, H. Cilengitide: The first anti-angiogenic small molecule drug candidate design, synthesis and clinical evaluation. Anticancer Agents Med. Chem 10(10), 753–768. https://doi.org/10.2174/187152010794728639 (2010).
Sun, X. et al. Annexin A5 regulates hepatocarcinoma malignancy via CRKI/II-DOCK180-RAC1 integrin and MEK-ERK pathways. Cell Death Dis. 9(6), 637. https://doi.org/10.1038/s41419-018-0685-8 (2018).
Kang, T. H. et al. Annexin A5 as an immune checkpoint inhibitor and tumor-homing molecule for cancer treatment. Nat. Commun. 11(1), 1137. https://doi.org/10.1038/s41467-020-14821-z (2020).
Kumar, D., Vetrivel, U., Parameswaran, S. & Subramanian, K. K. Structural insights on druggable hotspots in CD147: A bull’s eye view. Life Sci. 224, 76–87. https://doi.org/10.1016/j.lfs.2019.03.044 (2019).
Lucchetti, D. et al. CD147 promotes cell small extracellular vesicles release during colon cancer stem cells differentiation and triggers cellular changes in recipient cells. Cancers https://doi.org/10.3390/cancers12020260 (2020).
Huang, H. K. et al. Isoorientin decreases cell migration via decreasing functional activity and molecular expression of proton-linked monocarboxylate transporters in human lung cancer cells. Am. J. Chin. Med. 48(1), 201–222. https://doi.org/10.1142/s0192415x20500111 (2020).
Guo, W., Abudumijiti, H., Xu, L. & Hasim, A. CD147 promotes cervical cancer migration and invasion by up-regulating fatty acid synthase expression. Int. J. Clin. Exp. Pathol. 12(12), 4280–4288 (2019).
Fan, X., Wu, W., Shi, H. & Han, J. RNA interference targeting CD147 inhibits the invasion of human cervical squamous carcinoma cells by downregulating MMP-9. Cell Biol. Int. 37(7), 737–741. https://doi.org/10.1002/cbin.10104 (2013).
Wang, S. J. et al. CD147 promotes collective invasion through cathepsin B in hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 39(1), 145. https://doi.org/10.1186/s13046-020-01647-2 (2020).
Xiong, L., Edwards, C. K. 3rd. & Zhou, L. The biological function and clinical utilization of CD147 in human diseases: A review of the current scientific literature. Int. J. Mol. Sci. 15(10), 17411–17441. https://doi.org/10.3390/ijms151017411 (2014).
Lam, C. T. et al. Brain-derived neurotrophic factor promotes tumorigenesis via induction of neovascularization: Implication in hepatocellular carcinoma. Clin. Cancer Res. 17(10), 3123–3133. https://doi.org/10.1158/1078-0432.Ccr-10-2802 (2011).
Serafim Junior, V., Fernandes, G. M. M., Oliveira-Cucolo, J. G., Pavarino, E. C. & Goloni-Bertollo, E. M. Role of tropomyosin-related kinase B receptor and brain-derived neurotrophic factor in cancer. Cytokine 136, 155270. https://doi.org/10.1016/j.cyto.2020.155270 (2020).
Li, F., Wang, X. & Yang, L. MicroRNA-147 targets BDNF to inhibit cell proliferation, migration and invasion in non-small cell lung cancer. Oncol. Lett. 20(2), 1931–1937. https://doi.org/10.3892/ol.2020.11715 (2020).
Wang, Y. H., Huo, B. L., Li, C., Ma, G. & Cao, W. Knockdown of long noncoding RNA SNHG7 inhibits the proliferation and promotes apoptosis of thyroid cancer cells by downregulating BDNF. Eur. Rev. Med. Pharmacol. Sci. 23(11), 4815–4821. https://doi.org/10.26355/eurrev_201906_18067 (2019).
Han, J. et al. Suppression of adenosine-to-inosine (A-to-I) RNA editome by death associated protein 3 (DAP3) promotes cancer progression. Sci. Adv. 6(25), eaba5136. https://doi.org/10.1126/sciadv.aba5136 (2020).
Liu, Z., Chen, D., Ning, F., Du, J. & Wang, H. EGF is highly expressed in hepatocellular carcinoma (HCC) and promotes motility of HCC cells via fibronectin. J. Cell Biochem. 119(5), 4170–4183. https://doi.org/10.1002/jcb.26625 (2018).
Duffy, M. J., O’Donovan, N., Brennan, D. J., Gallagher, W. M. & Ryan, B. M. Survivin: A promising tumor biomarker. Cancer Lett. 249(1), 49–60. https://doi.org/10.1016/j.canlet.2006.12.020 (2007).
Hori, M. et al. The detergent-soluble cytoplasmic pool of survivin suppresses anoikis and its expression is associated with metastatic disease of human colon cancer. PLoS ONE 8(2), e55710. https://doi.org/10.1371/journal.pone.0055710 (2013).
Xu, R. et al. Identification of prognostic markers for hepatocellular carcinoma based on the epithelial-mesenchymal transition-related gene BIRC5. BMC Cancer 21(1), 687. https://doi.org/10.1186/s12885-021-08390-7 (2021).
Cao, L. et al. OCT4 increases BIRC5 and CCND1 expression and promotes cancer progression in hepatocellular carcinoma. BMC Cancer 13, 82. https://doi.org/10.1186/1471-2407-13-82 (2013).
Zhao, Y. et al. BIRC5 regulates inflammatory tumor microenvironment-induced aggravation of penile cancer development in vitro and in vivo. BMC Cancer 22(1), 448. https://doi.org/10.1186/s12885-022-09500-9 (2022).
Zhang, L. et al. Identification of potential prognostic biomarkers for hepatocellular carcinoma. J. Gastrointest. Oncol. 13(2), 812–821. https://doi.org/10.21037/jgo-22-303 (2022).
Wang, G., Chan, C. H., Gao, Y. & Lin, H. K. Novel roles of Skp2 E3 ligase in cellular senescence, cancer progression, and metastasis. Chin. J. Cancer 31(4), 169–177. https://doi.org/10.5732/cjc.011.10319 (2012).
Shin, E., Kim, S. H., Jeong, H. Y., Jang, J. J. & Lee, K. Nuclear expression of S-phase kinase-associated protein 2 predicts poor prognosis of hepatocellular carcinoma. APMIS 120(5), 349–357. https://doi.org/10.1111/j.1600-0463.2011.02838.x (2012).
Yokoi, S. et al. Down-regulation of SKP2 induces apoptosis in lung-cancer cells. Cancer Sci. 94(4), 344–349. https://doi.org/10.1111/j.1349-7006.2003.tb01444.x (2003).
Liu, J. et al. Cytoplasmic Skp2 expression is associated with p-Akt1 and predicts poor prognosis in human breast carcinomas. PLoS ONE 7(12), e52675. https://doi.org/10.1371/journal.pone.0052675 (2012).
Zhang, S. H. et al. Clinicopathologic significance and function of S-phase kinase-associated protein 2 overexpression in hepatocellular carcinoma. Hum. Pathol. 43(7), 1084–1093. https://doi.org/10.1016/j.humpath.2011.08.019 (2012).
Delogu, S. et al. SKP2 cooperates with N-Ras or AKT to induce liver tumor development in mice. Oncotarget 6(4), 2222–2234. https://doi.org/10.18632/oncotarget.2945 (2015).
Hershko, D. D. Oncogenic properties and prognostic implications of the ubiquitin ligase Skp2 in cancer. Cancer 112(7), 1415–1424. https://doi.org/10.1002/cncr.23317 (2008).
Fang, Z. L. et al. Effects of AFP gene silencing on Survivin mRNA expression inhibition in HepG2 cells. Genet. Mol. Res. 14(2), 3184–3190. https://doi.org/10.4238/2015.April.10.30 (2015).
He, D., Huang, K. & Liang, Z. Prognostic value of baculoviral IAP repeat containing 5 expression as a new biomarker in lung adenocarcinoma: A meta-analysis. Expert Rev. Mol. Diagn. 21(9), 973–981. https://doi.org/10.1080/14737159.2021.1947798 (2021).
Kakavandi, E., Shahbahrami, R., Goudarzi, H., Eslami, G. & Faghihloo, E. Anoikis resistance and oncoviruses. J. Cell Biochem. 119(3), 2484–2491. https://doi.org/10.1002/jcb.26363 (2018).
Tu, W., Gong, J., Tian, D. & Wang, Z. Hepatitis B virus X protein induces SATB1 expression through activation of ERK and p38MAPK pathways to suppress anoikis. Dig. Dis. Sci. 64(11), 3203–3214. https://doi.org/10.1007/s10620-019-05681-9 (2019).
Nahand, J. S. et al. Possible role of HPV/EBV coinfection in anoikis resistance and development in prostate cancer. BMC Cancer 21(1), 926. https://doi.org/10.1186/s12885-021-08658-y (2021).
Murray, P. J. et al. Macrophage activation and polarization: Nomenclature and experimental guidelines. Immunity 41(1), 14–20. https://doi.org/10.1016/j.immuni.2014.06.008 (2014).
Zhang, S. et al. Tumor-associated macrophages promote tumor metastasis via the TGF-β/SOX9 axis in non-small cell lung cancer. Oncotarget 8(59), 99801–99815. https://doi.org/10.18632/oncotarget.21068 (2017).
Liguori, M., Solinas, G., Germano, G., Mantovani, A. & Allavena, P. Tumor-associated macrophages as incessant builders and destroyers of the cancer stroma. Cancers 3(4), 3740–3761. https://doi.org/10.3390/cancers3043740 (2011).
Biswas, S. K., Allavena, P. & Mantovani, A. Tumor-associated macrophages: Functional diversity, clinical significance, and open questions. Semin. Immunopathol. 35(5), 585–600. https://doi.org/10.1007/s00281-013-0367-7 (2013).
Li, X. et al. Establishment of a lncRNA-based prognostic gene signature associated with altered immune responses in HCC. Front. Immunol. 13, 880288. https://doi.org/10.3389/fimmu.2022.880288 (2022).
Liu, Y. et al. Identification and validation of Birc5 as a novel activated cell cycle program biomarker associated with infiltration of immunosuppressive myeloid-derived suppressor cells in hepatocellular carcinoma. Cancer Med. https://doi.org/10.1002/cam4.6271 (2023).
Hao, X. et al. Targeting immune cells in the tumor microenvironment of HCC: New opportunities and challenges. Front. Cell Dev. Biol. 9, 775462. https://doi.org/10.3389/fcell.2021.775462 (2021).
Matsushita, H. et al. Cytotoxic T lymphocytes block tumor growth both by lytic activity and IFNgamma-dependent cell-cycle arrest. Cancer Immunol. Res. 3(1), 26–36. https://doi.org/10.1158/2326-6066.CIR-14-0098 (2015).
Ferrena, A. et al. SKP2 knockout in Rb1/p53 deficient mouse models of osteosarcoma induces immune infiltration and drives a transcriptional program with a favorable prognosis. bioRxiv https://doi.org/10.1101/2023.05.09.540053 (2023).
Terazawa, T. et al. Transarterial infusion chemotherapy with cisplatin plus S-1 for hepatocellular carcinoma treatment: A phase I trial. BMC Cancer 14, 301. https://doi.org/10.1186/1471-2407-14-301 (2014).
Hayashi, N. et al. Adenoviral infection of survivin antisense sensitizes prostate cancer cells to etoposide in vivo. Prostate 65(1), 10–19. https://doi.org/10.1002/pros.20232 (2005).
Asechi, H. et al. Resistance to cisplatin-induced apoptosis via PI3K-dependent survivin expression in a rat hepatoma cell line. Int. J. Oncol. 37(1), 89–96 (2010).
Huang, C. Y. et al. Sorafenib enhances radiation-induced apoptosis in hepatocellular carcinoma by inhibiting STAT3. Int. J. Radiat. Oncol. Biol. Phys. 86(3), 456–462. https://doi.org/10.1016/j.ijrobp.2013.01.025 (2013).
Koga, H. et al. Troglitazone induces p27Kip1-associated cell-cycle arrest through down-regulating Skp2 in human hepatoma cells. Hepatology 37(5), 1086–1096. https://doi.org/10.1053/jhep.2003.50186 (2003).
Shen, J. et al. Simultaneous inhibition of tumor growth and angiogenesis for resistant hepatocellular carcinoma by co-delivery of sorafenib and survivin small hairpin RNA. Mol. Pharm. 11(10), 3342–3351. https://doi.org/10.1021/mp4006408 (2014).
Su, K. J. & Yu, Y. L. Downregulation of SHIP2 by hepatitis B virus X promotes the metastasis and chemoresistance of hepatocellular carcinoma through SKP2. Cancers https://doi.org/10.3390/cancers11081065 (2019).
Liu, L., Zheng, Y. H., Han, L. & Qin, S. K. Efficacy and safety of the oxaliplatin-based chemotherapy in the treatment of advanced primary hepatocellular carcinoma: A meta-analysis of prospective studies. Medicine 95(40), e4993. https://doi.org/10.1097/md.0000000000004993 (2016).
Li, X. et al. Cyanidin inhibits EMT induced by oxaliplatin via targeting the PDK1-PI3K/Akt signaling pathway. Food Funct. 10(2), 592–601. https://doi.org/10.1039/c8fo01611a (2019).
Zhang, Y. et al. PKI-587 enhances chemosensitivity of oxaliplatin in hepatocellular carcinoma through suppressing DNA damage repair pathway (NHEJ and HR) and PI3K/AKT/mTOR pathway. Am. J. Transl. Res. 11(8), 5134–5149 (2019).
Funding
The study was supported by National Natural Science Foundation of China (81800203), a project admitted in the Dalian Deng Feng Program: key medical specialties in construction funded by the People’s Government of Dalian Municipality (No. 243, 2021) and Doctoral Start-up Foundation of Liaoning Province (20180540088).
Author information
Authors and Affiliations
Contributions
Z.Z., F.X. and J.Y. performed the data analysis and interpreted the data. K.G. and Q.W. collected samples and contributed to the experiments of this study. B.T., H.Z. and Y.Z. prepared the draft. B.T. and Z.Z. performed the visualization and revised the draft. B.T., Q.W. and R.L. designed the research and supervised all the work. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhong, Z., Xie, F., Yin, J. et al. Development of a prognostic model for anoikis and identifies hub genes in hepatocellular carcinoma. Sci Rep 13, 14723 (2023). https://doi.org/10.1038/s41598-023-41139-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-023-41139-9
This article is cited by
-
Identification of a novel anoikis-related gene signature to predict prognosis and tumor microenvironment in intrahepatic cholangiocarcinoma carcinoma
Discover Oncology (2025)
-
Integrative analysis of multi-omics data identified PLG as key gene related to Anoikis resistance and immune phenotypes in hepatocellular carcinoma
Journal of Translational Medicine (2024)