Abstract
The surge in mobile colistin-resistant genes (mcr) has become an increasing public health concern, especially in carbapenem-resistant Enterobacterales (CRE). Prospective surveillance was conducted to explore the genomic characteristics of clinical CRE isolates harbouring mcr in 2015–2020. In this study, we aimed to examine the genomic characteristics and phonotypes of mcr-8 and mcr-9 harbouring carbapenem-resistant K. pneumoniae complex (CRKpnC). Polymerase chain reaction test and genome analysis identified CRKpnC strain AMR20201034 as K. pneumoniae (CRKP) ST147 and strain AMR20200784 as K. quasipneumoniae (CRKQ) ST476, harbouring mcr-8 and mcr-9, respectively. CRKQ exhibited substitutions in chromosomal-mediated colistin resistance genes (pmrB, pmrC, ramA, and lpxM), while CRKP showed two substitutions in crrB, pmrB, pmrC, lpxM and lapB. Both species showed resistance to colistin, with minimal inhibitory concentrations of 8 µg/ml for mcr-8-carrying CRKP isolate and 32 µg/ml for mcr-9-carrying CRKQ isolate. In addition, CRKP harbouring mcr-8 carried blaNDM, while CRKQ harbouring mcr-9 carried blaIMP, conferring carbapenem resistance. Analysis of plasmid replicon types carrying mcr-8 and mcr-9 showed FIA-FII (96,575 bp) and FIB-HI1B (287,118 bp), respectively. In contrast with the plasmid carrying the carbapenemase genes, the CRKQ carried blaIMP-14 on an IncC plasmid, while the CRKP harboured blaNDM-1 on an FIB plasmid. This finding provides a comprehensive insight into another mcr-carrying CRE from patients in Thailand. The other antimicrobial-resistant genes in the CRKP were blaCTX-M-15, blaSHV-11, blaOXA-1, aac(6′)-Ib-cr, aph(3′)-VI, ARR-3, qnrS1, oqxA, oqxB, sul1, catB3, fosA, and qacE, while those detected in CRKQ were blaOKP-B-15, qnrA1, oqxA, oqxB, sul1, fosA, and qacE. This observation highlights the importance of strengthening official active surveillance efforts to detect, control, and prevent mcr-harbouring CRE and the need for rational drug use in all sectors.
Similar content being viewed by others
Introduction
In 2017, the World Health Organization published a list of 12 globally important antimicrobial-resistant pathogens, with the highest priority being given to carbapenem-resistant Enterobacterales (CRE)1. The limited availability of treatments has resulted in the utilization of colistin, as a last-resort therapeutic drug for human infections caused by gram-negative bacteria. The widespread use of colistin in humans and animals has led to the emergence of colistin resistance, with continually increasing resistance rates observed in gram-negative bacteria2,3. Of particular concern is the spread of mobile colistin resistance genes (mcr) in CRE isolates, potentially giving rise to pan-drug resistant (PDR) strains. Currently, 10 variants of mcr (mcr-1 to mcr-10) have been documented4,5,6. The mcr gene encodes a phosphoethanolamine transferase that alters lipid A in the lipopolysaccharide of the bacterial outer membrane by adding a phosphoethanolamine4. This modification reduces the attachment of colistin to the bacterial outer membrane, thereby preventing cell lysis.
The presence of mcr has been reported in various gram-negative bacteria isolated worldwide from animal and human sources3,4. In Thailand, mcr-1-, mcr-2-, and mcr-3-carrying E. coli and K. pneumoniae strains have been isolated from patients7,8,9,10,11. However, other variants of mcr have not yet been documented. Since 2015, the Emerging Antimicrobial Resistant Bacterial Surveillance Program (EARB) have been established, focusing on carbapenem-resistant Enterobacterles (CRE) with or without colistin resistance isolated from patients in 11 hospitals across Thailand. Through this program, mcr-1 carrying CRE isolates have been identified12. The current study aimed to explore the genomic characterization and antimicrobial susceptibility of K. pneumoniae complex (KpnC) harbouring mcr-8 and mcr-9 isolated from patients during the EARB surveillance program to provide insights into the plasmidome and compare plasmids harbouring mcr-8 and mcr-9-in KpnC isolates.
Results
Identification and susceptibility of KpnC harbouring mcr-8 and mcr-9
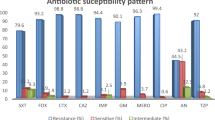
Of the 5,257 CRE isolates collected between 2015 and 2022, 17 carried the mcr-1 gene12. However, two isolates of CRKpnC harboured the mcr-8 and mcr-9 genes, respectively. During the EARB surveillance program, the prevalence of CRE carrying mcr was 0.36% (19/5257), with mcr-1 carriages accounting for 0.32% (17/5257), mcr-8 accounting for 0.02% (1/5257), and mcr-9 accounting for 0.02% (1/5257) carriage. Multiplex PCR analysis identified CRKpnC strain AMR20201034 as K. pneumoniae (CRKP) and strain AMR20200784 as K. quasipneumoniae (CRKQ), harbouring mcr-8 and mcr-9 genes, respectively (Table 1). Both species exhibited resistance to colistin, with MICs of 8 µg/ml for mcr-8-carrying CRKP isolate and 32 µg/ml for mcr-9-carrying CRKQ isolate (Table 1). Additionally, the mcr-8-carrying CRKP harboured blaNDM, while the mcr-9-carrying CRKQ harboured blaIMP, conferring carbapenem resistance. Further details on antimicrobial resistance profiles are presented in Table 1.
Genomic characteristics of mcr-8- and mcr-9-harbouring KpnC
Hybrid Nanopore-Illumina assemblies facilitated the generation of high-quality genomes of CRKP strain AMR20201034 and CRKQ strain AMR20200784. Comprehensive statistics on genome assemblies are provided in Table 1. The strain AMR20201034 was confirmed as K. pneumoniae using KmerFinder. It harboured mcr-8.1 on the InFIA-IncFII-type plasmid (96,575 bp; Fig. 1) and blaNDM-1 on the IncFIB-type plasmid (54,064 bp; Fig. 1). Other antimicrobial-resistant genes and virulence genes are shown in Table 1. The strain AMR20201034 was categorized as ST147 based on MLST profiling.
Plasmid map of mcr-8, mcr-9, blaNDM-1, and blaIMP-14 in this study.
KmerFinder identified CRKQ strain AMR20200784 as K. quasipneumoniae subsp. similispneumoniae that harboured mcr-9.1 on InFIB-IncHI1B-type plasmid (287,118 bp; Fig. 1), and blaIMP-14 on IncC-type plasmid (158,054 bp; Fig. 1). Other antimicrobial-resistant genes, plasmids, and virulence genes are presented in Table 1. This strain was designated as ST476.
The other antimicrobial-resistant genes identified were blaNDM-1, blaCTX-M-15, blaSHV-11, blaOXA-1, aac(6′)-Ib-cr, aph(3′)-VI, ARR-3, qnrS1, oqxA, oqxB, sul1, catB3, fosA, and qacE in the CRKP strain AMR20201034, and blaIMP-14, blaOKP-B-15, qnrA1, oqxA, oqxB, sul1, fosA, and qacE in the CRKQ strain AMR20200784. These antimicrobial-resistant genes were concordant with the observed resistance phenotypes in this strain (Table 1).
The CRKP ST147 (AMR20201034) possessed a set of virulence genes including fyuA, irp1, irp2, iutA, mrkA, mrkB, mrkC, mrkD, mrkF, mrkH, mrkI, mrkJ, ybtA, ybtE, ybtP, ybtQ, ybtS, ybtT, ybtU, and ybtX (Table 1). By contrast, the CRKQ ST476 (AMR20200784) carried a few virulence genes such as kfuA, kfuB, mrkA, mrkB, mrkC, mrkD, mrkF, mrkI, and mrkJ.
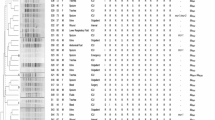
The SNP-phylogenetic tree showed the genetic relationship of the examined strains (Fig. 2). Strain AMR20201034 was closely related to strain SIKP147, which was isolated from an individual in Thailand. Interestingly, apart from mcr-8 in strain AMR20201034, the β-lactamase gene profiling was identical. On the contrary, strain AMR2020784 (ST476) exhibited a genetic relationship with strain ZKP111 (ST476), isolated from a patient in China.
Phylogenetic tree based on single nucleotide polymorphisms (SNPs) in K. pneumoniae complex using the neighbour-joining method. The tree was visualized and annotated using the Interactive Tree of Life. The tree is annotated based on sequence types (STs). Presentation of the genes is shown by filled symbols. The isolates used in this study are shown in red.
Plasmid harbouring carbapenemase genes
The circular plasmid carrying blaNDM-1 (pAMR1034-NDM) also harboured other antimicrobial-resistant genes such as aac(6′)-lb-cr, blaOXA-1, catB3, ARR-3, qacE, sula, blaCTX-M-15, qnrS, aph(3′)-VI, and bleO, all located on the same plasmid (IncFIB) (Fig. 1). The genetic arrangement of blaNDM-1 exhibited an upstream presence of the IS630 family, followed by blaNDM-1, bleO, anthranilate isomerase, protein disulfide reductase, cutA, and IS6-family downstream. As shown in Table 2 and Fig. 3, our pAMR1034-NDM was very similar to several plasmids carrying blaNDM-1 from humans in Thailand, Myanmar, China, Italy, Russia, USA, and Switzerland with 100% coverage and 99.95–99.96% identity. Alignment of these plasmids, almost revealed high similarity in their sequences and genetic organization among pAMR1034-NDM, p4-7008.20-NDM1, p5844-3, pC435-NDM1, pK5880C-NDM, pKP-26PI-FIB, pM321-NDM1, PSI0739-NDM, and plasmid unnamed-4, however, plasmid tig00000217 showed different in some genetic organization (Fig. 3).
Schematic representation of sequence alignment analysis of plasmids carrying blaNDM-1 genes. Homologous segments generated by a BLASTn comparison are shown as gray blocks that are connected across plasmids.
The circular plasmid carrying blaIMP-14 (pAMR784-IMP14) had IncC replicon type. It harbored sul1, qnrA1, and qacE (Fig. 1). The organization of blaIMP-14 consisted of the IS110 family upstream, followed by transposase, integrase, blaIMP-14, qacE, sul1, IS91-family transpoase, qnrA1, sul1, and IS110-family downstream. Notably, this plasmid exhibited another significant gene cluster comprising the tra gene cluster and mercury-resistant genes. The pAMR784-IMP14 showed similar to several plasmids harboured blaVIM-1 or blaNDM-1 from Vibrio spp., Aeromonas hydrophila, Salmonella enterica, Enterobacter chengduensis, Raoultella planticola, and E. coli from China, South Korea, Singapore, Viet Nam, and Switzerland (Table 2). Figure 4 showed alignment of these plasmids that revealed some parts of the genetic organization similar among these plasmids.
Schematic representation of sequence alignment analysis of of plasmid carrying blaIMP-14 gene to other related plasmids. Homologous segments generated by a BLASTn comparison are shown as gray blocks that are connected across plasmids.
Plasmid harbouring mcr-8 and mcr-9
As mentioned above, the CRKP strain AMR20201034 harboured mcr-8.1 on an IncFIA-IncFII-type plasmid with a size of 96,575 bp (circular plasmid; pAMR1034-mcr-8), while the CRKQ strain AMR20200784 harboured mcr-9.1 on an IncFIB-IncHI1B-type plasmid with a size of 287,118 bp (circular plasmid). Figure 1 illustrates the genetic map of these mcr-carrying plasmids. The mcr-8-harbouring plasmid exhibited mobile elements such as insertion sequences flanking both sites of the mcr-8. Another major gene cluster in this plasmid was the tra gene cluster involved in conjugation. In addition, the InFIA-InFII also contained the β-lactamase class C gene. As shown in Table 3 and Fig. 5, our pAMR1034-mcr8 was high similar to plasmids p2018N17-066-2-MCR8, p926-mcr8, pKA2-3-mcr8.1, pKP19-3138-3, pKP19288-mcr8, pSD21-MCR8 in the case of size and genetic organization. However, p2018C01-046-1-MCR8 showed different genetic organization and size when compare to others including our plasmid.
Schematic representation of sequence alignment analysis of of plasmids carrying mcr-8 genes. Homologous segments generated by a BLASTn comparison are shown as gray blocks that are connected across plasmids.
In contrast with the plasmid carrying mcr-9 (pAMR784-mcr-9), the mobile element was also flanked at the upstream and downstream of the mcr-9. This plasmid contained several metal-resistant genes such as copper, mercury, tellurium, and arsenic acid (Fig. 1). Enzymatic-coding genes were also present. Notably, the analysis of qseC and qseB genes involved in mcr-9 expression was not detected in CRKQ strain AMR20200784 (Fig. 1). Plasmid pAMR784-mcr9 showed coverage about 55–72% with high sequence identity to plasmids pKp2001, pAR8538-1, pVNCkp115, pKP308-2, pLH94-1, pKP20-558-2, and pINF156.1 from K. pneumoniae complex which isolated from humans, swine, and milk in China, Viet Nam, Laos, and Australia (Table 4). Among these related plasmids, we found pVNCKp115 contained mcr-8 whereas the rest plasmids have no any mcr genes (Table 4). As shown in Fig. 6, our plasmid revealed some parts of genetic organization similar to each other’s.
Schematic representation of sequence alignment analysis of plasmid carrying mcr-9 gene to related plasmids. Homologous segments generated by a BLASTn comparison are shown as gray blocks that are connected across plasmids.
Chromosomal mutations on colistin resistance
In addition to mcr-8 or mcr-9, we identified chromosomal gene mutations contributing to colistin resistance in both strains. CRKP strain AMR20201034 displayed one substitution in each of the following genes: crrB (C68S), lpxM (S253G), lapB (N212T), and pmrB (R256G), while multiple substitutions were only observed in pmrC (C27F, V50L, A135P, I138V) (Table 4). These mutations, along with the mcr-8.1, may contribute to moderate colistin resistance with an MIC of 8 ∞γ/µλ. When comparing our colistin-resistant K. quasipneumoniae AMR2020784 with colistin-susceptible K. quasipneumoniae ATCC700603, a few substitutions were detected in our strain (Table 5). Only one substitution was found in each gene following pmrB (S363I), pmrC (T224K), ramA (H38Y), and lpxM (G170D). High MIC value (32 ∞γ/µλ) of this strain may be contributed from these substitutions.
Discussion
Plasmid-mediated colistin resistance genes, known as mcr, represent a global concern. The majority of Enterobacterales harbouring mcr included E. coli, K. pneumoniae, Enterobacter spp., Salmonella spp., and C. freundii13. Of these, E. coli isolates carrying mcr exhibited the highest prevalence in humans, animals, and the environment13. By contrast, mcr-harbouring K. pneumoniae strains were predominantly observed in humans than in non-human subjects13. A previous study of the global prevalence of mcr genes revealed mcr-1 (4917/5191; 94.7%) as a common gene3. Non-mcr-1 harbouring Enterobacterales are also documented worldwide14.
Human infections with CRE isolates carrying mcr have been widely reported worldwide3,9,11,15,16,17,18,19. In this study, we reported the presence of mcr-8 harbouring CRKP and mcr-9-harbouring CRKQ detected from humans in Thailand. To the best of our knowledge, this study is the first to report such findings from our country. Human infections involving mcr-8-carrying K. pneumoniae have been documented in various countries worldwide including China20,21, Taiwan22, Myanmar23, Laos24, Saudi Arabia25, Bangladesh26, Lebanon27, Algeria28, Morocco29, Kenya30, Senegal31, and The Netherlands32. Our mcr-8-harbouring CRKP (strain AMR20201034) was identified as ST147, a high-risk clone with co-resistance to carbapenems that is prevalent in Europe and Asia13. This study revealed that our ST147 strain carried blaNDM-1 and was resistant to carbapenems. In Thailand, K. pneumoniae carrying mcr-8 strains have been detected in swine, including STs 35, 76, 999, 1746, 2791, 5229, 5231, and 523233. Notably, these isolates did not display carbapenem resistance. By contrast, the mcr-1 harbouring CRKP human isolates identified in Thailand were STs 15, 16, 45, 336, 340, 394, and 70910,11,12,34. This finding suggests a genetic diversity between human CRKP and swine K. pneumoniae carrying mcr isolates in Thailand.
The mcr-9-harbouring CRKQ in the current study was identified as ST476 and carried blaIMP-14, conferring carbapenems resistance. In Thailand, K. quasipneumoniae carrying mcr-7 and mcr-8 was detected in slaughtered pigs, while mcr-9 harbouring E. coli was also reported in pigs33,35. In Czech Republic, mcr-9 harbouring Enterobacterales isolated from humans were reported in Enterobacter hormaechei and Citrobacter freundii36. Globally, Klebsiella species harbouring mcr-9 were detected in K. pneumoniae, K. quasipneumoniae, K. oxytoca, K. grimontii, and K. michiganensis6,37,38,39,40. Human isolates of mcr-9-containing Klebsiella species have been reported in China, Argentina, Qatar, Ethiopia, Belgium, Denmark, Montenegro, Poland, Romania, Serbia, Slovenia, and Spain38,39,40,41,42,43. The CRKQ strain harbouring mcr-9 was initially reported in Argentina38; however, the Argentinian CRKQ belongs to the quasipneumoniae subspecies, while the Thai CRKQ (identified in our study) belongs to the similispneumoniae subspecies. In addition, our CRKQ exhibited fewer antimicrobial-resistant genes (qnrA1, oqxA, oqxB, sul1, fosA, and qacE) compared with Argentinian CRKQ (aac(6′)-IIc, aadA2, aph(3′)-Ia, aph(3′′)-Ib, blaDHA-1, blaSHV-12, catA2, dfrA12, ereA2, mphA, qnrB4, sul1, sul2, and tet(D)) 38.
The expression of mcr-9 is inducible and involves the presence of two-component regulators (qseB and qseC), which are typically located adjacent to mcr-9. However, our mcr-9-carrying CRKQ strain AMR20200784 lacked these two genes, similar to the K. quasipneumoniae strain M17277 and K. pneumoniae strain 162CM1 from Argentina and Thailand, respectively38,44. The genes downstream of mcr-9 displayed genetic diversity, especially with the absence of qseB/qseC two-component regulators in most plasmids45. This may indicate that the mcr-9 gene in our CRKQ strain may not be expressed; however, high-level colistin resistance was observed due to the multiple substitutions in chromosomal-mediated genes.
The combinations of mcr and chromosome-mediated colistin-resistant genes (pmrA, pmrB, pmrC, pmrD, phoP, phoQ, crrA, crrB, ramA, lpxM, lapB, rpoN, and mgrB) may contribute to the colistin resistance observed in our isolates. Notably, the substitutions in these chromosomal-mediated colistin-resistant genes in our CRKP and CRKQ strains differ from those reported in previous studies10,46,47. In our CRKP, only R256G of pmrB, C27F of pmrC, and N212T of lapB have been reported in the previously reports10,47. However, these substitutions were not yet confirmed by the experimental study10,47. Interestingly, when compared our CRKQ with K. quasipneumoniae ATCC700603 (colistin-susceptible strain) showed very few mutations in four genes (lpxM, pmrB, pmrC, ramA). These substitutions have not yet reported47. This suggests that substitutions contributing to colistin resistance in K. quasipneumoniae should be validated further.
Our study that the replicon types of plasmids carrying mcr-8 and mcr-9 were IncFIA-IncFII and IncFIB-IncHI1B, respectively. This suggests that hybrid plasmids (IncFIA-IncFII and IncFIB-IncHI1B) may represent another vehicle mediating the spread of mcr-8 and mcr-9 in Thailand. Notably, the plasmid replicon types carrying mcr-8 are single or multiple plasmid replicons including IncFII(K), IncFIA(HI1), IncFIA-IncFII, IncFIA(HI1)-IncFII(K), IncFIA(HI1-IncFII(pKP91), IncFII(K)-IncQ1, IncN-IncR, IncFII(K2)-IncFII(K7)-IncQ, and IncFIA-IncHI1B-IncR20,27,42,43,48,49,50,51,52. Our mcr-8.1 harbouring plasmid carries two plasmid replicons (IncFIA-IncFII), a configuration similar to what has been reported previously, particularly in China42.
According to previous studies, the plasmid replicon types containing mcr-9 were IncHI2, IncHI2A, IncFII, IncHI2-IncR, IncHI2-IncA/C2, and IncHI2-IncHI2A37,45,53,54. Of these, IncHI2 was the most predominant type (90.1%), accounting for 90.1% of plasmids harbouring mcr-9. A previous study conducted in Thailand demonstrated the presence of mcr-9 on an IncHI2 plasmid44. This suggests that IncHI2 and a hybrid plasmid IncFIB-IncHI1B may serve as disseminating vehicles of mcr-9 in Thailand.
The findings of the current study provide comprehensive insights into the characteristics of mcr-8- and mcr-9-carrying CRKpnC strains from humans in Thailand. The co-occurrence of mcr genes and carbapenemase genes was also demonstrated in CRE isolated from patients10. To mitigate the emergence of XDR or PDR strains, priority should be provided to strengthening official surveillance, active control, and prevention efforts to minimize the dissemination of mcr genes and chromosomal-mediated colistin-resistant genes among CRKpnC isolates in humans.
Methods
Identification of mcr-8 and mcr-9-harbouring K. pneumoniae complex
In total, 5257 CRE isolates were collected from individuals within a hospital network in 11 provinces throughout Thailand, as part of the EARB program from 2015 to 2022 (8 years). These isolates were identified by the hospital laboratories and then the isolates were sent to the Public Health Microbiological Laboratory of the Faculty of Public Health, Kasetsart University Chalermphrakiat Sakon Nakhon Province Campus for further confirmation and detection of antimicrobial-resistant genes.
The species and carbapenemase (blaIMP, blaKPC, blaNDM, and blaOXA-48-like) genes were determined using two multiplex polymerase chain reaction (PCR) tests described elsewhere56,57. The mcr genes (mcr-1 to mcr-10) were detected using a PCR test based on established protocols35,58. Of these CRE isolates, only carbapenem-resistant K. pneumoniae complex (CRKpnC) harbouring mcr-8 (strain AMR20201034) and mcr-9 (strain AMR20200784) were selected for further characterization (Table 1).
Antimicrobial susceptibility testing
Analysis of the minimal inhibitory concentration (MIC) of antimicrobials for mcr-8 and mcr-9-harbouring CPKpnC isolates was performed using the broth microdilution, and the results were interpreted according to the 2023 Clinical and Laboratory Standards Institute guidelines59, with E. coli ATCC 25922 as the control strain. Broth microdilution was conducted to determine the MIC of colistin, meropenem, ceftriaxone, ciprofloxacin, gentamicin, trimethoprim-sulfamethoxazole, and azithromycin. Meanwhile, disk diffusion was performed to determine the sensitivity of piperacillin-tazobactam, ceftazidime, amikacin, chloramphenicol, and nitrofurantoin to the examined pathogen.
Broth microdilution was used to determine the MIC of colistin using colistin sulphate (Merck, Germany) at concentrations of 1 µg/ml, 2 µg/ml, 4 µg/ml, 8 µg/ml, 16 µg/ml, and 32 µg/ml, respectively. According to the CLSI M100 33rd edition, an MIC of ≤ 2 µg/ml indicated intermediate susceptibility, while an MIC of ≥ 4 µg/ml indicated resistance.
Whole-genome sequencing
The DNA from mcr-8 and mcr-9-carrying CRKpnC isolates was extracted from tryptic soy agar plate cultures using ZymoBIOMICS DNA Kits (Tustin, CA, USA) following the manufacturer’s instructions. The concentration of the extracted DNA was determined using the Qubit dsDNA BR assay kit (Invitrogen, Oregon, USA). Subsequently, whole-genome sequencing was performed using short-read (Illumina) and long-read (Oxford Nanopore Technologies; ONT) platforms. Briefly, Illumina sequencing libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit for Illumina (New England Biolabs, Hitchin, UK) following the manufacturer’s recommendations. The genomic DNA was randomly fragmented to a size of 350 bp, and the fragments underwent A-tailing and ligation with adapters. The libraries were sequenced as paired-end reads (150 + 150 bp) using HiSeq 2500 instruments. The sequencing adapters were trimmed using Fastp v0.19.5 (https://github.com/OpenGene/fastp), and the quality of clean reads was determined using FastQC v0.11.8 (https://www.bioinformatics.babraham.ac.uk/projects/ fastqc/).
For ONT sequencing, the library preparation was conducted following the rapid barcoding DNA sequencing protocol with the SQK-RBK004 kit without DNA size selection, to preserve the plasmid DNA content. The libraries were then sequenced using a single R9.4.1/FLO-MIN106 flow cell on a MinION Mk1B sequencer. Base calling and demultiplexing of the raw data were achieved using Guppy v3.4.5 (ONT), specifying the high-accuracy model (-c dna_r9.4.1_450bps_hac.cfg). The ONT adapters were trimmed using Porechop v0.2.4 (https://github.com/rrwick/Porechop). Nanoplot v1.28.1 (https://github.com/wdecoster/NanoPlot) was employed to assess the quality of ONT reads. Hybrid assemblies with the ONT and Illumina data were performed using Unicycler v0.4.860. The genome sequences were submitted to the National Center for Biotechnology Information Prokaryotic Genome Annotation Pipeline (PGAP v4.12) for annotation. The default parameters were used for all software unless otherwise specified.
Bioinformatics analysis
The identification of isolates at the species level was carried out using KmerFinder 3.161. Antimicrobial resistance genes were identified using ResFinder 4.162 and the comprehensive antibiotic resistance database (CARD)63. Chromosome-mediated colistin resistance was explored by analysing the mgrB, crrA, crrB, phoP, phoQ, pmrA, pmrB, pmrC, pmrD, lapB, lpxM, ramA, and rpoN genes47. Gene sequences were scrutinized using local BLAST + and Clustal W, with K. pneumoniae MGH78578 (Genbank accession number: NC_009648.1) and K. quasipneumoniae subsp. similispneumoniae ATCC700603 (Genbank accession number: CP014696) genomes serving as colistin-susceptible references. Plasmid replicons were investigated using PlasmidFinder64. The virulence genes of KpnC were assessed using VirulenceFinder 2.065 and Institut Pasteur (https://bigsdb.pasteur.fr/klebsiella/klebsiella.html).
For multilocus sequence typing (MLST) analysis of the sequence types (STs) of KpnC, MLST 2.0 was used66. The genomic comparison of mcr-8 and mcr-9-harbouring KpnC isolates was accomplished through a modular single genome analysis to search for the genetically closest relatives in the database following the single nucleotide polymorphism (SNP) approach with BacWGSTdb67. A phylogenetic tree was constructed using REALPHY and MEGA X via the neighbour-joining method with 500 bootstrap replicates by applying the Tamura three-parameter model68,69. Visualization and annotation of the tree were carried out using Interactive Tree of Life (iTOL) v4 70. Klebsiella pneumoniae HS11286 (accession No. CP003200) was used as the reference sequences for SNP analysis. The sequence similarity search for four plasmids in the current study was performed using BLAST against the NCBI nucleotide database. In order to examine the distribution and relationships of other plasmids, plasmid sequences were downloaded from NCBI and compared. The Easyfig software (version 2.2.5) was used for the comparative and visualize the region of plasmids71.
Ethical approval
The stored bacterial isolates used in this study were collected as part of routine microbiological laboratory practice and were not considered to constitute human subjects research. The study was exempt from Kasetsart University's ethical guidelines because no human specimens or data were used. Informed consent is not a requirement. All experiments were performed in compliance with the relevant laws and guidelines.
Data availability
The datasets generated during the current study are available in the National Center for Biotechnology Information repository, with a Bioproject accession number PRJNA525849 with the Genbank accession numbers JAZGTD000000000 (K. quasipneumoniae subsp. similispneumoniae strain AMR20200784) and JAZGTE000000000 (K. pneumoniae strain AMR20201034).
References
World Health Organization. Global priority list of antibiotic-resistant bacteria to guide research, discovery, and development of new antibiotics (WHO, 2017).
Kempf, I., Jouy, E. & Chauvin, C. Colistin use and colistin resistance in bacteria from animals. Int. J. Antimicrob. Agents. 48, 598–606 (2016).
Elbediwi, M. et al. Global burden of colistin-resistant bacteria: Mobilized colistin resistance genes study (1980–2018). Microorganisms 7, E461 (2019).
Gharaibeh, M. H. & Shatnawi, S. Q. An overview of colistin resistance, mobilized colistin resistance genes dissemination, global responses, and the alternatives to colistin: A review. Vet. World 12, 1735–1746 (2019).
Wang, C. et al. Identification of novel mobile colistin resistance gene mcr-10. Emerg. Microbes Infect. 9, 508–516 (2020).
Wang, Y. et al. Detection of mobile colistin resistance gene mcr-9 in carbapenem-resistant Klebsiella pneumoniae strains of human origin in Europe. J. Infect. 80, 578–606 (2020).
Kamjumphol, W. et al. Draft genome sequence of a colistin-resistant Escherichia coli ST226: A clinical strain harbouring an mcr-1 variant. J. Glob. Antimicrob. Resist. 16, 168–169 (2019).
Malchione, M. D., Torres, L. M., Hartley, D. M., Koch, M. & Goodman, J. L. Carbapenem and colistin resistance in Enterobacteriaceae in Southeast Asia: Review and mapping of emerging and overlapping challenges. Int. J. Antimicrob. Agents 54, 381–399 (2019).
Paveenkittiporn, W. et al. Emergence of plasmid-mediated colistin resistance and New Delhi metallo-β-lactamase genes in extensively drug-resistant Escherichia coli isolated from a patient in Thailand. Diagn. Microbiol. Infect. Dis. 87, 157–159 (2017).
Paveenkittiporn, W., Kamjumphol, W., Ungcharoen, R. & Kerdsin, A. Whole-genome sequencing of clinically isolated carbapenem-resistant Enterobacterales harboring mcr genes in Thailand, 2016–2019. Front. Microbiol. 11, 586368 (2021).
Srijan, A. et al. Genomic characterization of nonclonal mcr-1-positive multidrug-resistant Klebsiella pneumoniae from clinical samples in Thailand. Microb. Drug Resist. 24, 403–410 (2018).
Boueroy, P. et al. Plasmidome in mcr-1 harboring carbapenem-resistant enterobacterales isolates from human in Thailand. Sci Rep. 12, 19051 (2022).
Biswas, U., Das, S., Barik, M. & Mallick, A. Situation report on mcr-carrying colistin-resistant clones of Enterobacterales: A global update through human-animal-environment interfaces. Curr. Microbiol. 81, 12 (2023).
Mmatli, M., Mbelle, N. M. & Osei Sekyere, J. Global epidemiology, genetic environment, risk factors and therapeutic prospects of mcr genes: A current and emerging update. Front. Cell Infect. Microbiol. 12, 941358 (2022).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Mediavilla, J. R. et al. Colistin- and carbapenem-resistant Escherichia coli harboring mcr-1 and blaNDM-5, causing a complicated urinary tract infection in a patient from the United States. mBio 7, e01191-16 (2016).
Mendes, A. C. et al. mcr-1 in carbapenemase-producing Klebsiella pneumoniae with hospitalized patients, Portugal, 2016–2017. Emerg. Infect. Dis. 24, 762–766 (2018).
Quan, J. et al. Prevalence of mcr-1 in Escherichia coli and Klebsiella pneumoniae recovered from bloodstream infections in China: A multicentre longitudinal study. Lancet Infect. Dis. 17, 400–410 (2017).
Zhong, L. L. et al. High rates of human fecal carriage of mcr-1-positive multidrug-resistant Enterobacteriaceae emerge in China in association with successful plasmid families. Clin. Infect. Dis. 66, 676–685 (2018).
Ge, H. et al. First report of Klebsiella pneumoniae co-producing OXA-181, CTX-M-55, and MCR-8 isolated from the patient with bacteremia. Front. Microbiol. 13, 1020500 (2022).
Li, T., Wang, L., Ko, W. & Lv, Y. Genetic and phenotypic characteristics of mcr-8 positive extended-spectrum β-lactamase-producing Klebsiella pneumoniae isolated from patient in China. Eur. J. Clin. Microbiol. Infect. Dis. 40, 901–904 (2021).
Chen, F. J. et al. Emergence of mcr-1, mcr-3 and mcr-8 in clinical Klebsiella pneumoniae isolates in Taiwan. Clin. Microbiol. Infect. 27, 305–307 (2021).
Tada, T. et al. Klebsiella pneumoniae co-harbouring blaNDM-1, armA and mcr-10 isolated from blood samples in Myanmar. J. Med. Microbiol. https://doi.org/10.1099/jmm.0.001750 (2023).
Hadjadj, L., Baron, S. A., Olaitan, A. O., Morand, S. & Rolain, J. M. Co-occurrence of variants of mcr-3 and mcr-8 Genes in a Klebsiella pneumoniae isolate from Laos. Front. Microbiol. 10, 2720 (2019).
Hala, S. et al. Co-occurrence of mcr-1 and mcr-8 genes in multi-drug-resistant Klebsiella pneumoniae from a 2015 clinical isolate. Int. J. Antimicrob. Agents. 57, 106303 (2021).
Farzana, R. et al. Emergence of mobile colistin resistance (mcr-8) in a highly successful Klebsiella pneumoniae sequence type 15 clone from clinical infections in Bangladesh. mSphere 5, e00023-20 (2020).
Salloum, T. et al. First report of plasmid-mediated colistin resistance mcr-8.1 gene from a clinical Klebsiella pneumoniae isolate from Lebanon. Antimicrob. Resist. Infect. Control. 9, 94 (2020).
Nabti, L. Z. et al. Development of real-time PCR assay allowed describing the first clinical Klebsiella pneumoniae isolate harboring plasmid-mediated colistin resistance mcr-8 gene in Algeria. J. Glob. Antimicrob. Resist. 20, 266–271 (2020).
Bonnin, R. A., Bernabeu, S., Jaureguy, F., Naas, T. & Dortet, L. MCR-8 mediated colistin resistance in a carbapenem-resistant Klebsiella pneumoniae isolated from a repatriated patient from Morocco. Int. J. Antimicrob. Agents. 55, 105920 (2020).
Kyany’a, C. & Musila, L. Colistin resistance gene mcr-8 in a high-risk sequence type 15 Klebsiella pneumoniae isolate from Kenya. Microbiol. Resour. Announc. 9, e00783-e820 (2020).
Sarr, H. et al. The emergence of carbapenem- and colistin-resistant enterobacteria in Senegal. Pathogens. 12, 974 (2023).
Dierikx, C. M. et al. Colistin-resistant Enterobacterales among veterinary healthcare workers and in the Dutch population. JAC Antimicrob. Resist. 4, dlac041 (2022).
Phetburom, N. et al. Klebsiella pneumoniae complex harboring mcr-1, mcr-7, and mcr-8 isolates from slaughtered pigs in Thailand. Microorganisms 9, 2436 (2021).
Shanmugakani, R. K. et al. PCR-dipstick-oriented surveillance and characterization of mcr-1- and carbapenemase-carrying Enterobacteriaceae in a Thai hospital. Front. Microbiol. 10, 149 (2019).
Khanawapee, A. et al. Distribution and molecular characterization of Escherichia coli harboring mcr genes isolated from slaughtered pigs in Thailand. Microb. Drug Resist. 27, 971–979 (2021).
Bitar, I. et al. Detection of five mcr-9-Carrying Enterobacterales isolates in four Czech Hospitals. mSphere 5, e01008-20 (2020).
Campos-Madueno, E. I., Moser, A. I., Risch, M., Bodmer, T. & Endimiani, A. Exploring the global spread of Klebsiella grimontii isolates possessing blaVIM-1 and mcr-9. Antimicrob. Agents Chemother. 65, e0072421 (2021).
Faccone, D. et al. Plasmid carrying mcr-9 from an extensively drug-resistant NDM-1-producing Klebsiella quasipneumoniae subsp. quasipneumoniae clinical isolate. Infect. Genet. Evol. 81, 104273 (2020).
Li, S. et al. A blaSIM-1 and mcr-9.2 harboring Klebsiella michiganensis strain reported and genomic characteristics of Klebsiella michiganensis. Front. Cell Infect. Microbiol. 12, 973901 (2022).
Tsui, C. K. M. et al. Draft genome sequence of an extended-spectrum β-lactamase-producing Klebsiella oxytoca strain bearing mcr-9 from Qatar. Microbiol. Resour. Announc. 9, e00429-20 (2020).
Legese, M. H. et al. Genomic epidemiology of carbapenemase-producing and colistin-resistant Enterobacteriaceae among sepsis patients in Ethiopia: A whole-genome analysis. Antimicrob. Agents Chemother. 66, e0053422 (2022).
Liu, M. C., Jian, Z., Liu, W., Li, J. & Pei, N. One health analysis of mcr-carrying plasmids and emergence of mcr-10.1 in three species of Klebsiella recovered from humans in China. Microbiol. Spectr. 10, e0230622 (2022).
Wang, X. et al. Emergence of colistin resistance gene mcr-8 and its variant in Raoultella ornithinolytica. Front. Microbiol. 10, 228 (2019).
Leangapichart, T. et al. Exploring the epidemiology of mcr genes, genetic context and plasmids in Enterobacteriaceae originating from pigs and humans on farms in Thailand. J. Antimicrob. Chemother. 78, 1395–1405 (2023).
Li, Y. et al. Characterization of the global distribution and diversified plasmid reservoirs of the colistin resistance gene mcr-9. Sci. Rep. 10, 8113 (2020).
Berglund, B. Acquired resistance to colistin via chromosomal and plasmid-mediated mechanisms in Klebsiella pneumoniae. Infect. Microbes Dis. 1, 10–19 (2019).
Binsker, U., Käsbohrer, A. & Hammerl, J. A. Global colistin use: A review of the emergence of resistant Enterobacterales and the impact on their genetic basis. FEMS Microbiol. Rev. 46, fuab049 (2022).
Kalová, A., Gelbíčová, T., Overballe-Petersen, S., Litrup, E. & Karpíšková, R. Characterisation of colistin -resistant Enterobacterales and Acinetobacter strains carrying mcr genes from Asian aquaculture products. Antibiotics (Basel) 10, 838 (2021).
Ma, K., Feng, Y., Liu, L., Yao, Z. & Zong, Z. A cluster of colistin- and carbapenem-resistant Klebsiella pneumoniae carrying blaNDM-1 and mcr-8.2. J. Infect. Dis. 221(Suppl 2), S237–S242 (2020).
Wang, Y. et al. Emergence of high-level colistin resistance mediated by multiple determinants, including mcr-1.1, mcr-8.2 and crrB mutations, combined with tigecycline resistance in an ST656 Klebsiella pneumoniae. Front. Cell Infect. Microbiol. 13, 1122532 (2023).
Yang, C. et al. Dissemination of blaNDM-5 and mcr-8.1 in carbapenem-resistant Klebsiella pneumoniae and Klebsiella quasipneumoniae in an animal breeding area in Eastern China. Front. Microbiol. 13, 1030490 (2022).
Zhao, J. et al. Convergence of MCR-8.2 and chromosome-mediated resistance to colistin and tigecycline in an NDM-5-producing ST656 Klebsiella pneumoniae isolate from a lung transplant patient in China. Front. Cell Infect. Microbiol. 12, 922031 (2022).
Fukuda, A., Nakano, H., Suzuki, Y., Nakajima, C. & Usui, M. Conjugative IncHI2/HI2A plasmids harboring mcr-9 in colistin-susceptible Escherichia coli isolated from diseased pigs in Japan. Access Microbiol. 4, acmi000454 (2022).
Hutton, W., Allman, E., McKeown, C., Singer, A. C. & Roberts, A. P. Complete genome sequence of mcr-9 containing Leclercia adecarboxylata. Microbiol. Resour. Announc. 12, e0048123 (2023).
Liu, Z. et al. Co-occurrence of blaNDM-1 and mcr-9 in a conjugative IncHI2/HI2A plasmid from a bloodstream infection-causing carbapenem-resistant Klebsiella pneumoniae. Front. Microbiol. 12, 756201 (2021).
Hatrongjit, R., Kerdsin, A., Akeda, Y. & Hamada, S. Detection of plasmid-mediated colistin-resistant and carbapenem-resistant genes by multiplex PCR. Methods X 5, 532–536 (2018).
Hatrongjit, R., Chopjitt, P., Boueroy, P. & Kerdsin, A. Multiplex PCR detection of common carbapenemase genes and identification of clinically relevant Escherichia coli and Klebsiella pneumoniae complex. Antibiotics (Basel) 12, 76 (2023).
Lei, C. W., Zhang, Y., Wang, Y. T. & Wang, H. N. Detection of mobile colistin resistance gene mcr-10.1 in a conjugative plasmid from Enterobacter roggenkampii of chicken origin in China. Antimicrob. Agents Chemother. 64, e01191-20 (2020).
Clinical and Laboratory Standards Institute CLSI. M100: Performance Standards for Antimicrobial Susceptibility Testing 33rd edn. (Clinical & Laboratory Standards Institute, 2023).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
Larsen, M. V. et al. Benchmarking of methods for genomic taxonomy. J. Clin. Microbiol. 52, 1529–1539 (2014).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Alcock, B. P. et al. CARD 2023: Expanded curation, support for machine learning, and resistome prediction at the comprehensive antibiotic resistance database. Nucleic Acids Res. 51(D1), D690–D699 (2023).
Carattoli, A. et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
Joensen, K. G. et al. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 52, 1501–1510 (2014).
Larsen, M. V. et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 50, 1355–1361 (2012).
Feng, Y., Zou, S., Chen, H., Yu, Y. & Ruan, Z. BacWGSTdb 2.0: A one-stop repository for bacterial whole-genome sequence typing and source tracking. Nucleic Acids Res. 3, gkaa821 (2020).
Bertels, F., Silander, O. K., Pachkov, M., Rainey, P. B. & van Nimwegen, E. Automated reconstruction of whole-genome phylogenies from short-sequence reads. Mol. Biol. Evol. 31, 1077–1088 (2014).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. Mega X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics. 27, 1009–1010 (2011).
Acknowledgements
We would like to thank the Editage for providing excellent English editing assistance.
Funding
This work was supported by the Health Systems Research Institute (HSRI Grant Number 64-105) and The Japan Agency for Medical Research and Development (AMED) with Grant Number 23wm0125010h004.
Author information
Authors and Affiliations
Contributions
R.H. and A.K. conceived and designed the study, performed data analysis, drafted the manuscript, and performed critical revisions of the manuscript for intellectual content. T.W., P.J., P.C., and P.B. performed laboratory experiments and data analysis. Y.A., T.I., and S.H. performed critical revision of the manuscript for intellectual content. All the authors read, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hatrongjit, R., Wongsurawat, T., Jenjaroenpun, P. et al. Genomic analysis of carbapenem- and colistin-resistant Klebsiella pneumoniae complex harbouring mcr-8 and mcr-9 from individuals in Thailand. Sci Rep 14, 16836 (2024). https://doi.org/10.1038/s41598-024-67838-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-67838-5
This article is cited by
-
Nationwide genomic surveillance of carbapenem- and colistin-resistant Klebsiella pneumoniae bloodstream isolates in Thailand (2020–2024)
Scientific Reports (2026)
-
mcr gene family evolution and structural mechanisms of colistin resistance: from mcr-1 to emerging variants
Archives of Microbiology (2026)
-
Co-occurrence of mcr-9 and blaNDM-5 in Multidrug-Resistant Enterobacter hormaechei Strain Isolated from a Patient with Bloodstream Infection
Current Microbiology (2025)