Abstract
This study aimed to construct and assess a machine-learning algorithm designed to forecast survival rates and risk stratification for patients with gastric neuroendocrine neoplasms (gNENs) after diagnosis. Data on patients with gNENs were extracted and randomly divided into training and validation sets using the Surveillance, Epidemiology, and End Results database. We developed a prediction model using 10 machine learning algorithms across 101 combinations to forecast cancer-related mortality in patients with gNENs, selecting the best model using the highest mean over a sequence of time-dependent area under the receiver operating characteristic (ROC) curve (AUC). The performance of the final model was assessed through time-dependent ROC curves for discrimination and calibration curves for calibration. The maximum selection rank method was used to determine the best prognostic risk score threshold for classifying patients into high- and low-risk groups. Afterward, Kaplan–Meier analysis and log-rank test were used to compare survival rates among these groups. Our study examined 775 patients with gNENs, dividing them into training and validation sets. A training set comprised 543 patients, with a median follow-up of 42 months and cumulative mortality rates of 40.0% at 1 year, 48.6% at 3 years, and 54.0% at 5 years. A validation set comprised 232 patients, with cumulative mortality rates of 29.1% at 1 year, 43.5% at 3 years, and 53.2% at 5 years. The optimal random survival forest (RSF) model (mtry = 4, node size = 5) achieved an AUC of 0.839 for survival prediction in the training set. Comprising 11 variables such as demographics, treatment details, tumor characteristics, T staging, N staging, and M staging, the RSF model revealed high predictive accuracy with AUCs of 0.92, 0.96, and 0.96 for 1-, 3-, and 5-year survival, respectively, which was consistently reflected in the validation set with AUCs of 0.88, 0.92, and 0.89, respectively. Moreover, patients were risk-stratified. Although our RSF model effectively stratified patients into different prognostic groups, it needs external validation to confirm its utility for noninvasive prognostic prediction and risk stratification in gNENs. Further research is required to verify its broader clinical applicability.
Similar content being viewed by others
Introduction
Gastric neuroendocrine neoplasms (gNENs) are heterogeneous tumors originating from neurons and neuroendocrine cells. These cells belong to the widespread neuroendocrine system tasked with producing various hormones that manage the operations of the digestive system. The incidence of gNENs in gastric tumors is relatively low; however, it has been increasing, attributed partly to the improved gNEN detection rates and awareness1. According to a recent study, the incidence of gNENs has increased over the past 40 years from 0.309 to 6.149 per 1,000,000 people2. Despite their rarity, gNENs possess significant clinical importance due to their potential for aggressive behavior, especially in higher-grade forms. Patients with gNENs have low median survival rates compared with those with other digestive NENs3,4,5. However, there is currently no ideal predictive model or biomarker to predict their prognosis.
The tumor node metastasis(TNM) staging system, as recommended by the American Joint Committee on Cancer (AJCC), has become a crucial prognostic element for gNENs6,7. Nonetheless, other factors, such as age, therapeutic interventions, or tumor grade, which were not accounted for in the AJCC staging framework, could influence the gNEN prognosis8. The mitotic count and Ki-67 proliferation index, key components of the World Health Organization (WHO) classification, facilitate the evaluation of prognoses for patients with gNENs9. While classifications by the AJCC or WHO can typically be employed to forecast overall survival, their applicability for precise prognostication on an individual level is limited. Consequently, there is a pressing need to develop an updated model or framework that consolidates all pertinent clinicopathological factors, which can greatly improve the accuracy of prognostic forecasts for patients diagnosed with gNENs.
The survival rates for patients with gNENs vary and are influenced by factors such as tumor stage at diagnosis, histological subtype, and the treatments applied. Accurate prediction of survival rates is paramount, as it guides clinical decision-making, including selecting treatment modalities and managing patient care. Predictive models that accurately estimate survival outcomes can facilitate personalized medicine, enabling healthcare providers to tailor treatment plans to individual patient profiles, thereby improving prognosis for patients with gNENs. However, most predictive models for gNENs prognosis are currently constructed using traditional Cox survival analysis10,11,12, which has average predictive performance and is difficult to apply widely in clinical practice. With the development of artificial intelligence, the application of machine learning technology in the medical field is rapidly becoming a key driving force for improving the effectiveness of disease diagnosis, treatment, and prevention. Therefore, it is necessary to explore and compare multiple machine-learning methods in constructing models for predicting the survival rate of patients with gNENs to select the optimal machine-learning model. The purpose of this study was to identify important prognostic factors and develop and validate an individual machine learning model for gNENs based on the Surveillance, Epidemiology, and End Results (SEER) database, predicting survival rates for 1-, 3-, and 5-years.
Methods and materials
We followed the transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD) statement13 to report the development and validation of this prediction model.
Participants
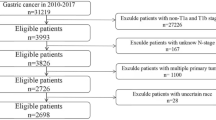
The SEER database, known for its accessibility and being the most extensive public cancer dataset, covers approximately 28% of the US population. It compiles comprehensive data on cancer incidence and mortality across all 18 SEER cancer registries. The SEER*Stat software (version 8.4.3), using the SEER Research Plus Data, 17 registries from November 2021 Sub (2000–2019), was utilized for patient identification. Patients diagnosed with gNENs were pinpointed using the International Classification of Diseases for Oncology site codes. Exclusion criteria were set to omit cases with ambiguous follow-up details or tumor locations and instances where gNENs were not identified as the initial primary tumor. Afterward, the study participants were randomly allocated to either the training set, which was utilized for both model fitting and parameter tuning, or the validation set, which was employed for the final evaluation on unseen data, adhering to a 7:3 ratio (Fig. 1). Data.
Flowchart of patient selection and grouping process.
Based on information extracted from previously released studies and their clinical significance14,15,16,17, the following patient demographics and clinical details were collected directly from the SEER database: age, race, sex, the 8th AJCC staging, T staging, N staging, M staging, differentiation (the proliferative marker Ki-67 was applied to determine the tumors’ differentiation status, regarding the degree of differentiation as unknown, poorly differentiated, moderately differentiated, and well-differentiated), marital status (including single, married, widowed, or divorced), information on radiotherapy and chemotherapy during follow-up, survival status (dead or alive), and follow-up time. Follow-up was considered to start at the time of cancer diagnosis.
Missing data handling
In the SEER database, missing data is a common issue. However, excluding patients with incomplete data can introduce significant bias into the study. In our analysis, the age variable was the sole variable exhibiting missing data, with a missing proportion of 8.4%. All other variables used in the study were complete, with no missing information. Since age was found to have a skewed distribution, we addressed the missing data by imputing it with the median18.
Definition of outcome
Patient death due to a tumor and its complications was defined as the event of interest.
-
1.
Death due to tumor progression: When the cancer itself progresses to the point that it causes the patient’s death due to organ failure, metastasis, or other direct effects of the tumor.
-
2.
Death due to cancer-related complications: This can include complications directly related to the cancer, such as hemorrhage from a tumor, obstruction (for example, gastrointestinal), or paraneoplastic syndromes.
Statistical analysis
Data analysis was conducted using R software (version 4.3.3), and major R codes during analyses were provided in supplemental files. The approach to statistical testing was two-tailed, with a P-value threshold < 0.05 set for determining statistical significance unless stated otherwise. Normally distributed variables are presented using mean (standard deviation), while non-normally distributed variables are presented using median (interquartile ranges). Variables with categorical values are expressed as percentages. Wilcoxon rank sum and chi-square tests were conducted to compare training and validation sets. The reverse Kaplan–Meier method was used to calculate the median follow-up time.
Screening of machine learning algorithms: To construct a survival prediction model for patients with gNENs with high precision and consistent performance, we amalgamated ten machine learning algorithms along with their 101 unique combinations19,20. The comprehensive list of algorithms encompassed the random survival forest (RSF), least absolute shrinkage and selection operator(Lasso), elastic net (Enet), Ridge, Cox boost, stepwise Cox, survival support vector machine (survival-SVM), generalized boosted regression modeling (GBM), supervised principal component analysis (SuperPC), and Cox partial least squares regression (plsRcox). The survival prediction models were constructed using the leave-one-out cross-validation (LOOCV) approach. The integrated area under the receiver operating characteristic (ROC) curve (AUC) was calculated for the training and validation sets. The optimal model was determined by identifying the one with the highest mean time-dependent AUC value across the training and validation sets (Fig. 1S). After the final calculation, the RSF model obtained the highest AUC value.
The RSF model was constructed using 11 features, including race, sex, age, chemotherapy, radiation, differentiation, the extent of the tumor, marital status, and T, N, and M stages in 8th editor AJCC staging. In the development of the RSF model, we optimized three key parameters to enhance the model’s performance: the number of trees, the number of variables randomly selected at each split (mtry), and the minimum node size. These parameters were selected based on their impact on prediction accuracy and model stability.While constructing 200 survival trees, the prediction error rate tended to be low and stable (Fig. 2A). Additionally, after completing the construction of the RSF model in 1000 trees (mtry = 4, node size = 5), the variable importance (VIMP) of all features for tree growth was generated, as depicted in Fig. 2B, with higher VIMP indicating that the variable exerted a greater predictive effect on survival probability (cause-specific survival).
Random survival forest model performance and variable importance analysis. (A) The error rate plot as a function of the number of trees in the model shows the out-of-bag error rate declining and stabilizing as more trees are added. (B) Variable importance plot indicating the relative contribution of each variable to the model’s predictive accuracy. The extent of the tumor has the highest importance, followed by the differentiation and M staging. Variables such as marital status have a minimal impact on the model’s predictive performance.
Performance evaluation of random forest model: We constructed an RSF model to predict the 1-, 3-, and 5-year survival probabilities and prognostic risk score of patients with gNENs. The performance of the RSF model was evaluated using discrimination and calibration21. For the evaluation of model discrimination, a time-dependent ROC curve analysis was conducted21,22,23. A calibration curve was performed using 1000 bootstrap resamples to evaluate model calibration21. Calibration was evaluated by focusing on the calibration intercept and slope, where an intercept of 0 and a slope of 1 signify ‘ideal’ calibration24.
A prognostic risk score for patients with gNENs was calculated to indicate patient prognosis using the RSF model, which revealed that a higher prognostic risk score was associated with a worse outcome. In the training data, a cutoff prognostic risk score was chosen using the maximum choice ranking statistic in the “maxstat” R package, which was an outcome-oriented approach providing the cutoff point value corresponding to the most significant relationship with survival25. By dichotomizing patients into distinct risk groups, this method facilitated more actionable clinical decision-making, allowing for clearer stratification between high-risk and low-risk patients. The Kaplan–Meier method was used to estimate survival distributions in different risk groups, and the log-rank test was performed to compare survival probability.
Results
Patient characteristics
In this study, 775 patients from the SEER database were analyzed, with a median survival time of 48 months. Among these patients, 543 were allocated to the training set and 232 to the validation set. The demographic and clinical characteristics of the patients in both sets are outlined in Table 1. No statistically significant variations were observed between the two sets in terms of general characteristics, thereby supporting their suitability for use as training and validation sets.
In the training set, the cumulative mortality rates were 40.0% at one year, 48.6% at three years, and 54.0% at five years, with a median follow-up time of 42 months (interquartile ranges: 9, 95). In the validation set, the cumulative mortality rates were 29.1% at one year, 43.5% at three years, and 53.2% at five years, with a median follow-up time of 48 months (interquartile ranges: 9, 93). The cumulative mortality rates did not exhibit a statistically significant difference between the training and validation sets.
Construction of the RSF model in the training set
Based on the training set data, we fitted 101 prediction models through the LOOCV framework and further calculated the AUC value of each model in the validation data set (Fig. 1S). Combining the AUC values of each model in the training and the validation sets, the most accurate model for predicting the survival of patients with gNENs was the classification RSF model (Fig. 1S), with a mean AUC of 0.839, higher than other models.
Assessing the model performance
In the training set, the RSF model demonstrated higher AUC values for predicting 1-, 3-, and 5-year survival in gNEN patients compared to the 8th AJCC staging system(Fig. 3A). Specifically, the RSF model had AUCs of 0.92 (95% CI: 0.89–0.93), 0.96 (95% CI: 0.94–0.97), and 0.96 (95% CI: 0.94–0.97), while the AJCC had AUCs of 0.81 (95% CI: 0.77–0.85), 0.83 (95% CI: 0.80–0.86), and 0.82 (95% CI: 0.79–0.86) for the same time points. The RSF model demonstrated statistically significant superiority at 1 year (p < 0.001), 3 years (p < 0.001), and 5 years (p < 0.001). The calibration curves presented an acceptable agreement between the training set (Fig. 4A–C). For 1-, 3-, and 5-year survival probabilities, the calibration intercepts were 0.007, -0.017, and − 0.023, and the calibration slopes were 0.93, 1.14, and 1.29, respectively.
Time-dependent ROC curves for RFS model. (A) Demonstrates the ROC curves for the training set at 1-year, 3-year, and 5-year time points with AUCs of 0.92, 0.96, and 0.96, respectively, indicating strong predictive performance. (B) Time-dependent ROC Curves for the validation set at 1-year, 3-year, and 5-year time points with AUCs of 0.88, 0.92, and 0.89, respectively, suggesting good generalizability of the prognostic model.
Calibration plots for the RSF model. (A-C) Calibration plots for 1-year, 3-year, and 5-year survival probabilities in the training set. (D-F) Calibration plots for 1-year, 3-year, and 5-year survival probabilities in the validation set. Each plot compares the actual observed survival rates (y-axis) against the survival probabilities predicted by the RSF model (x-axis). The 45-degree dotted line represents perfect calibration where predicted probabilities match the actual rates. The closer the solid line is to the dotted line, the better the model’s calibration.
In the validation set, the RSF model demonstrated higher AUC values for predicting 1-, 3-, and 5-year survival in gNEN patients compared to the 8th AJCC staging system(Fig. 3B). Specifically, the RSF model had AUCs of 0.88 (95% CI: 0.84–0.93), 0.92 (95% CI: 0.88–0.96), and 0.89 (95% CI: 0.85–0.94), while the AJCC had AUCs of 0.85 (95% CI: 0.81–0.90), 0.88 (95% CI: 0.84–0.93), and 0.84 (95% CI: 0.79–0.89) for the same time points. The RSF model’s superiority was statistically significant at 3 years (p = 0.043) and 5 years (p = 0.007), but not at 1 years (p = 0.174). The calibration curves presented an acceptable agreement in the validation set (Fig. 4D–F). For 1-year, 3-year, and 5-year survival probabilities, the calibration intercepts were 0.014, − 0.021, and − 0.026, and the calibration slopes were 0.94, 1.16, and 1.25, respectively.
Risk stratification
As presented in Fig. 5, the threshold for dividing patients with high and low risk of death according to the training set was 53.84. The patient’s death risk exceeding 53.84 indicates that the patient has a poor prognosis.
Prognostic risk score distribution and maximal rank statistic analysis. The top panel displays the distribution of the prognostic risk score with two distinct groups indicated by blue (low risk) and red (high risk) bars. The bottom panel shows the maximally selected rank statistics, plotting the standardized log-rank statistic against the prognostic risk score. The optimal cut-point, determined by the maximal statistic, is marked by a dashed vertical line at a score of 53.84, effectively stratifying patients into low- and high-risk categories for survival outcomes.
The validation set was subsequently divided into high-risk and low-risk groups based on the cutoff point. As demonstrated in Fig. 6A, the Kaplan–Meier curves for both groups revealed significantly different survival times based on the log-rank test (P < 0.001). In the high-risk group, the 1-, 3-, and 5-year cumulative mortality rates were 54.1%, 76.2%, and 84.4%. In the low-risk group, the 1-, 3-, and 5-year cumulative mortality rates were 0.9%, 6.5%, and 17.8%.
Kaplan-Meier survival curves comparing risk stratification by RSF model and AJCC staging system in the validation set. (A) Kaplan-Meier survival curves stratified by the RSF (Random Survival Forest) model risk groups. Patients were categorized into high-risk (yellow) and low-risk (blue) groups based on their RSF scores. The survival probability over time is shown, with a significant difference between the two groups (p < 0.0001). The RSF model effectively distinguishes between patients with different survival outcomes, as reflected by the clear separation between the survival curves and minimal overlap of the 95% confidence intervals (shaded areas). (B) Kaplan-Meier survival curves stratified by the AJCC (American Joint Committee on Cancer) staging system. Patients were categorized into stages I (yellow), II (blue), III (green), and IV (red). Although the overall comparison between the stages is statistically significant (p < 0.0001), there is substantial overlap of the 95% confidence intervals between the different stages. This overlap indicates that the AJCC staging system is less effective in differentiating the survival outcomes of patients across different stages compared to the RSF model.
Figure 6B reveals the survival curves of patients with different AJCC stages. The survival curves for different AJCC stages demonstrated a statistically significant difference in survival probabilities across the follow-up period (P < 0.001). However, at certain time points, especially where the CIs overlapped, the ability of AJCC staging to clearly differentiate between patient risk levels was less effective, indicating limitations in its overall performance.
we performed a subgroup survival analysis based on the AJCC staging system, stratifying patients into high-risk and low-risk groups according to their risk scores. The results showed a significant survival difference between high-risk and low-risk groups in AJCC Stage I patients, with the low-risk group having better outcomes (Fig. 7A, p < 0.0001). In AJCC Stage II, the high-risk group had significantly lower survival than the low-risk group (Fig. 7B, p = 0.001). In AJCC Stage III, although the high-risk group had lower survival rates, the difference was not statistically significant (Fig. 7C, p = 0.51). It is important to note that survival curves for AJCC Stage IV patients are not presented, as all Stage IV patients were classified into the high-risk group.
Kaplan-Meier survival curves Stratified by AJCC stages and risk groups. (A) Survival probability for AJCC Stage I patients. (B) Survival probability for AJCC Stage II patients. (C) Survival probability for AJCC Stage III patients. Survival curves for AJCC Stage IV patients are not displayed, as all Stage IV patients were classified in the high-risk group.
Discussion
Our study focused on developing and evaluating predictive models for patient prognosis in gNENs using various machine-learning algorithms. Among the 100 model combinations tested, including Enet, Ridge, Cox Boost, stepwise Cox, survival-SVM, GBM, SuperPC, and plsRcox, the RSF model emerged as the most effective, incorporating 11 predictors. The RSF model demonstrated superior discriminative ability and calibration in predicting 1-, 3-, and 5-year survival rates of patients with gNENs, outperforming the 8th edition AJCC staging system. These findings suggested that the RSF model could significantly improve prognosis prediction for gNENs, offering a more accurate tool for guiding treatment and follow-up care.
To the best of our knowledge, few prognostic models for patients with gNENs have been developed. B. Zhang et al. constructed a prognostic prediction model for patients with gNENs using six characteristic proteins: ENTPD1, TNXB, EML1, DMD, SORBS2, and S100B26. Nevertheless, the implementation of this model in clinical practice is hindered by the challenges associated with detecting these specific proteins. Nomograms were constructed incorporating clinicopathological features to predict the prognosis of patients with gNENs, and higher predictive accuracy was demonstrated compared to AJCC staging and WHO staging systems10,27,28,29. However, the few existing studies are either based on genomics or involve small, single-center cohorts with predictors unavailable in the SEER database. Owing to these differences, a direct comparison between our RSF model and existing Cox regression models was unfeasible. Yang Zhihao et al. proposed that deep learning radiomics analysis could be used as a potential noninvasive tool for prognosis prediction and risk stratification in patients with gNENs12, which was a model built based on small sample data, and the discrimination was low. It did not risk-stratify patients, which was not conducive to individualized clinical management of patients. The Ki-67 marker index is recognized as a prognostic indicator for gNENs and plays a role in determining tumor grade. This index is traditionally calculated by analyzing tumor tissue that has been single-immunostained for Ki-67, involving the counting of both Ki-67-positive and Ki-67-negative tumor cells within a hot spot chosen subjectively. However, variability in observer assessments and challenges in differentiating between tumor and non-tumor cells can result in inaccurate Ki-67 index measurements, potentially leading to misclassification of tumor grades. With the development of artificial intelligence, several calculation methods for extracting the Ki-67 index have emerged in recent years30,31,32. However, there are some limitations to all of the above-mentioned methods, such as failure to distinguish between neoplastic and non-neoplastic cells, manual selection of hot spots (which is subject to error), or lack of scalability33. In the end, the effectiveness of these methods must be evaluated based on their ability to predict outcomes and forecast prognoses in patient groups that have been monitored over long periods through comprehensive clinical follow-up data. Consequently, it would be beneficial to develop a better system for gNEN prognosis prediction and risk stratification that uses stronger calculation methods and large multi-center queues.
Our RSF model demonstrated exceptional apparent discriminatory power, consistently achieving a differentiation level exceeding 0.9 for distinguishing 1-, 3-, and 5-year survival rates in patients with gNENs. While this high level of discrimination indicates the model’s effectiveness in stratifying patient risk, we did not conduct a comparative analysis of calibration metrics, including slope and intercept. Instead, we followed the approach adopted by the majority of literature, focusing on discrimination to select models. Importantly, the calibration of the RSF model is within an acceptable range, which supports its clinical relevance. Thus, while we can assert that the RSF model exhibits superior discrimination in our dataset, it is essential to recognize that this evaluation does not encompass a comprehensive comparison of model calibration. Therefore, we cannot definitively conclude that the RSF model outperforms the Cox model or other models. This limitation highlights the need for caution in interpreting our findings. Future studies should explore additional measures such as log loss or Brier score to provide a more nuanced evaluation of both discrimination and calibration, ensuring a more robust understanding of model effectiveness in clinical practice.
Compared with the survival curves of the AJCC staging system, the survival curves of the RSF model exhibited clearer separation throughout the follow-up period, with no overlap in the 95% CIs between risk groups. This indicated that the RSF model possessed higher accuracy and discriminative ability in distinguishing survival probabilities among different risk groups, making it a more effective tool for predicting patient prognosis. Moreover, our subgroup survival analysis based on the AJCC staging system further underscored the limitations of the AJCC model, which suggests that the AJCC system may not fully capture the prognostic heterogeneity within certain stages. In contrast, the RSF model consistently differentiated between high-risk and low-risk groups across all stages, highlighting its superior ability to stratify patients accurately according to their survival probabilities. Furthermore, the superior performance of the RSF model was reflected in its higher AUC, further demonstrating its potential application in prognostic assessment. Generally, the data suggested that the RSF model might provide added benefit when evaluating survival for patients with gNENs. Besides, we stratified patients according to prognostic risk and developed a straightforward, facilitating reliable stratification of patients’ prognoses. However, it is important to note that the RSF model is based on data from a specific cohort and may not fully account for variations in patient populations or clinical practices outside the study context. Clinicians are advised to exercise caution when utilizing the model across diverse patient populations, given the variability in individual treatment responses. Moreover, the model currently lacks consideration of all potential confounding factors, underscoring the importance of employing clinical judgment alongside its application. Future developments of this model could be improved by incorporating more extensive datasets and additional variables, thereby enhancing its generalizability and predictive accuracy. In conclusion, although the RSF model constitutes a substantial advancement in the application of machine learning for personalized medicine, it should be regarded as a supplement to, rather than a substitute for, clinical expertise and thorough patient assessment.
It is worth mentioning that the lack of detailed follow-up treatment information in the SEER database limited the accuracy and generalizability of the study’s conclusions on patient outcomes and may contribute to a bias related to immortal time, as treatment exposures are often assessed after diagnosis. Immortal time bias is a significant concern in survival analyses, particularly when evaluating the impact of treatments initiated after diagnosis. In our study, the timing of treatment relative to diagnosis may inadvertently influence the interpretation of prognostic outcomes. Specifically, patients who survive long enough to receive chemotherapy or radiation may demonstrate better survival outcomes not solely due to the efficacy of these treatments but also because their survival to treatment initiation reflects more favorable underlying health status or disease characteristics. This limitation highlights the risk of misrepresenting treatment effects, as prognostic assessments based on post-diagnosis treatment may not fully capture the complexities of disease progression and treatment timing. Therefore, careful consideration of immortal time bias is essential in interpreting our findings and understanding their implications for clinical practice.
This register-based study has several limitations that should be acknowledged. First, using SEER data, which primarily reflects the US population, might introduce selection bias and limit the generalizability of our findings to other populations, including those in different countries or regions. Additionally, while our study employed RSF to predict patient outcomes, we acknowledge the potential risk of overfitting due to the relatively small sample size and the complexity of the RSF model, which considers interactions between variables. Although RSF incorporates ensemble method to reduce overfitting, it is possible that we exceeded the 10 events per variable rule, as RSF models do not rely solely on the number of candidate predictors but also model potential interactions. The absence of external validation further underscores this limitation, and future studies are needed to confirm the model’s generalizability.These limitations underscore the need for further external validation and calibration studies to confirm the model’s accuracy and reliability before it can be widely implemented in clinical practice. Furthermore, while our model exhibited promising performance, its clinical utility remains to be fully established, and additional studies are required to ensure that it can be effectively applied in real-world settings.
Conclusions
Our RSF model demonstrated efficacy in stratifying individual patients into distinct prognostic groups, suggesting its potential utility as a noninvasive tool for prognostic prediction and risk stratification in patients with gNENs.
Data availability
The data analyzed during the current study are available from the corresponding author on reasonable request.
References
Delle Fave, G. et al. ENETS consensus guidelines update for gastroduodenal neuroendocrine neoplasms. Neuroendocrinology 103 (2), 119–124 (2016).
Hu, P. et al. Trends of incidence and prognosis of gastric neuroendocrine neoplasms: A study based on SEER and our multicenter research. Gastric Cancer 23 (4), 591–599 (2020).
Dasari, A. et al. Trends in the incidence, prevalence, and survival outcomes in patients with neuroendocrine tumors in the United States. JAMA Oncol. 3 (10), 1335–1342 (2017).
Man, D., Wu, J., Shen, Z. & Zhu, X. Prognosis of patients with neuroendocrine tumor: A SEER database analysis. Cancer Manag. Res. 10, 5629–5638 (2018).
Dasari, A., Mehta, K., Byers, L. A., Sorbye, H. & Yao, J. C. Comparative study of lung and extrapulmonary poorly differentiated neuroendocrine carcinomas: A SEER database analysis of 162,983 cases. Cancer 124 (4), 807–815 (2018).
Xie, J. W. et al. Evaluation of clinicopathological factors related to the prognosis of gastric neuroendocrine carcinoma. Eur. J. Surg. Oncol.: J. Eur. Soc. Surg. Oncol. Br. Assoc. Surg. Oncol. 42 (10), 1464–1470 (2016).
Xie, J. W. et al. Modified AJCC staging of gastric neuroendocrine carcinoma based on T staging can improve the capacity of prognosis assessment. J. Cancer Res. Clin. Oncol. 144 (12), 2391–2397 (2018).
Zhong, Q. et al. Incidence trend and conditional survival estimates of gastroenteropancreatic neuroendocrine tumors: A large population-based study. Cancer Med. 7 (7), 3521–3533 (2018).
Nagtegaal, I. D. et al. The 2019 WHO classification of tumours of the digestive system. 76 (2), 182–188 (2020).
Zhang, S. et al. A novel and validated nomogram to predict overall survival for gastric neuroendocrine neoplasms. J. Cancer 10 (24), 5944–5954 (2019).
Yang, Z. H. et al. Prognostic value of computed tomography radiomics features in patients with gastric neuroendocrine neoplasm. Front. Oncol. 13, 1143291 (2023).
Yang, Z. et al. Deep learning radiomics analysis based on computed tomography for survival prediction in gastric neuroendocrine neoplasm: A multicenter study. Quant. Imaging Med. Surg. 13 (12), 8190–8203 (2023).
Moons, K. G. et al. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): Explanation and elaboration. Ann. Intern. Med. 162 (1), W1–73 (2015).
Perri, G., Prakash, L. R. & Katz, M. H. G. Pancreatic neuroendocrine tumors. Curr. Opin. Gastroenterol. 35 (5), 468–477 (2019).
Lee, L., Ito, T. & Jensen, R. T. Prognostic and predictive factors on overall survival and surgical outcomes in pancreatic neuroendocrine tumors: Recent advances and controversies. Expert Rev. Anticancer Ther. 19 (12), 1029–1050 (2019).
Zhang, X. F. et al. Margin status and long-term prognosis of primary pancreatic neuroendocrine tumor after curative resection: Results from the US neuroendocrine tumor study group. Surgery 165 (3), 548–556 (2019).
Zhou, Y. J. et al. Marital status, an independent predictor for survival of gastric neuroendocrine neoplasm patients: A SEER database analysis. BMC Endocr. Disord. 20 (1), 111 (2020).
Zhang, Z. Missing data imputation: Focusing on single imputation. Ann. Transl. Med. 4 (1), 9 (2016).
Liu, Z. et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat. Commun. 13 (1), 816 (2022).
Liu, H. et al. Mime: A flexible machine-learning framework to construct and visualize models for clinical characteristics prediction and feature selection. Comput. Struct. Biotechnol. J. 23, 2798–2810 (2024).
Alba, A. C. et al. Discrimination and calibration of clinical prediction models: Users’ guides to the medical literature. Jama 318 (14), 1377–1384 (2017).
Blanche, P., Dartigues, J. F., Jacqmin-Gadda, H. Estimating and comparing time‐dependent areas under receiver operating characteristic curves for censored event times with competing risks. 32(30):5381–5397. (2013).
Kamarudin, A. N., Cox, T., Kolamunnage-Dona, R. Time-dependent ROC curve analysis in medical research: Current methods and applications. 17, 1–19 (2017).
Van Calster, B., McLernon, D. J., van Smeden, M., Wynants, L. & Steyerberg, E. W. Calibration: The Achilles heel of predictive analytics. BMC Med. 17 (1), 230 (2019).
Hothorn TaL, B. On the exact distribution of maximally selected rank statistics. Comput. Stat. Data Anal. 43, 121–137 (2003).
Zhang, B., Zhang, K. & Chen, L. J. A. O. PD-3 prognosis evaluation and molecular typing of gastric neuroendocrine tumors based on proteomics and non-negative matrix factorization model. 34, S1–S2 (2023).
Cao, L. L. et al. Nomogram based on tumor-associated neutrophil-to-lymphocyte ratio to predict survival of patients with gastric neuroendocrine neoplasms. World J. Gastroenterol. 23 (47), 8376–8386 (2017).
Wang, Y. L. et al. Establishment and validation of a nomogram to predict overall survival of patients with gastric neuroendocrine neoplasms. Zhonghua Wei Chang. Wai Ke Za Zhi = Chin. J. Gastrointest. Surg. 24 (10), 883–888 (2021).
Cao, L. L. et al. A novel predictive model based on preoperative blood neutrophil-to-lymphocyte ratio for survival prognosis in patients with gastric neuroendocrine neoplasms. Oncotarget 7 (27), 42045–42058 (2016).
Shi, P., Zhong, J., Hong, J., Huang, R. & Wang, K. Chen, Y. Automated Ki-67 quantification of immunohistochemical staining image of human nasopharyngeal carcinoma xenografts. 6 (1), 32127 (2016).
Zhong, F., Bi, R., Yu, B., Yang, F. & Yang, W. Shui, R. A comparison of visual assessment and automated digital image analysis of Ki67 labeling index in breast cancer. 11 (2), e0150505 (2016).
Tuominen, V. J., Ruotoistenmäki, S., Viitanen, A. & Jumppanen, M. Isola, J. ImmunoRatio: A publicly available web application for quantitative image analysis of estrogen receptor (ER), progesterone receptor (PR), and Ki-67. 12, 1–12 (2010).
Govind, D. et al. Improving the accuracy of gastrointestinal neuroendocrine tumor grading with deep learning. 10 (1), 11064 (2020).
Acknowledgements
Not applicable.
Funding
This work was supported by grants from the Natural Science Foundation of Guangxi (NO. 2021GXNSFAA220036).
Author information
Authors and Affiliations
Contributions
Study design: Lu-Huai Feng and Tianbao Liao; data collection: Yang Lu, Wei‑Yuan Wei, Lina Huang, Tingting Su and Tianbao Liao; manuscript preparation: Tingting Su and Tianbao Liao; data analysis and interpretation: Lu-Huai Feng, Tingting Su and Tianbao Liao; all authors confirm that they contributed to manuscript reviews—revising it critically for important intellectual content—and read and approved the final draft for submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
We accessed SEER database data following protocol and obtained ethics exemption from Youjiang Medical University for Nationalities, as the data used were publicly available and contained no personal identifying information.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liao, T., Su, T., Lu, Y. et al. Random survival forest algorithm for risk stratification and survival prediction in gastric neuroendocrine neoplasms. Sci Rep 14, 26969 (2024). https://doi.org/10.1038/s41598-024-77988-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-77988-1