Abstract
Left ventricular (LV) global longitudinal strain (LVGLS) is versatile; however, it is difficult to obtain. We evaluated the potential of an artificial intelligence (AI)-generated electrocardiography score for LVGLS estimation (ECG-GLS score) to diagnose LV systolic dysfunction and predict prognosis of patients with heart failure (HF). A convolutional neural network-based deep-learning algorithm was trained to estimate the echocardiography-derived GLS (LVGLS). ECG-GLS score performance was evaluated using data from an acute HF registry at another tertiary hospital (n = 1186). In the validation cohort, the ECG-GLS score could identify patients with impaired LVGLS (≤ 12%) (area under the receiver-operating characteristic curve [AUROC], 0.82; sensitivity, 85%; specificity, 59%). The performance of ECG-GLS in identifying patients with an LV ejection fraction (LVEF) < 40% (AUROC, 0.85) was comparable to that of LVGLS (AUROC, 0.83) (p = 0.08). Five-year outcomes (all-cause death; composite of all-cause death and hospitalization for HF) occurred significantly more frequently in patients with low ECG-GLS scores. Low ECG-GLS score was a significant risk factor for these outcomes after adjustment for other clinical risk factors and LVEF. The ECG-GLS score demonstrated a meaningful correlation with the LVGLS and is effective in risk stratification for long-term prognosis after acute HF, possibly acting as a practical alternative to the LVGLS.
Similar content being viewed by others
Introduction
Recent developments in echocardiography have emphasized the significance of the left ventricular (LV) global longitudinal strain (GLS; LVGLS) as a prognosticator of various heart diseases1, including heart failure (HF)2, valvular heart diseases3,4,5, and cardiomyopathies6. Despite its vast clinical utility, there are barriers to the widespread use of LVGLS. For instance, the measurement and interpretation of the LVGLS are time-consuming and complex compared to those of the ejection fraction and require specific machines, software, and skilled personnel.
Electrocardiography (ECG) is a simple, cost-effective, and well-established method for heart-disease screening. Advances in artificial intelligence (AI) have expanded the capabilities of ECG, enhancing its diagnostic scope in the cardiovascular field7. AI can also assist in the diagnosis of subclinical atrial fibrillation, HF, hypertrophic cardiomyopathy (HCM), and valvular heart diseases8. Previous studies have indicated the potential of AI-assisted analysis of ECG patterns and features not only for the detection of certain diseases but also for the estimation of LV function. For example, AI-enabled ECG analysis can distinguish ST-segment elevation, myocardial infarction, or LV systolic dysfunction among patients who visit the emergency department9,10.
Given the expanding clinical utility of LVGLS, with its superior sensitivity over that of the LV ejection fraction (LVEF) and robust prognostic value across various heart diseases, leveraging AI technology to estimate LVGLS from ECG features has merits, as this could enhance its clinical efficacy and cost-effectiveness. In the present study, we generated an ECG-derived GLS (ECG-GLS) score using an AI deep-learning algorithm and evaluated its potential in diagnosing LV systolic dysfunction and predicting the prognosis of patients with HF.
Methods
Study population and data management
This multicenter retrospective cohort study involved two tertiary hospitals (Hospitals A and B). ECG and echocardiographic data were obtained from the following four registries: (1) the STRATS-AHF cohort, which included patients who were admitted for acute HF with a range of ejection fractions from June 2009 to June 2015 (n = 1,608);2 (2) angiotensin receptor/neprilysin inhibitor (ARNI) baseline cohort, which included patients with HF with reduced ejection fraction (HFrEF) who were prescribed ARNI from September 2015 to April 2020 (n = 409); (3) ARNI follow-up cohort from December 2020 to January 2021 (n = 409);11 and (4) three-chamber strain cohort, which included patients who underwent echocardiography with GLS data from February 2020 to November 2022, irrespective of diagnosis (n = 984). The validation cohort comprised patients in the STRATS-AHF registry from Hospital B2. Patients with inadequate images for strain analysis were excluded. A total of 2,882 and 1,186 echocardiography and ECG pairs were included in the training and validation cohorts, respectively (Fig. 1). All ECG recordings used consistent settings (25 mm/s speed and 10 mm/mV voltage gain), and the input method for our analysis involved ECG image files (jpg format). Echocardiography and 12-lead ECG pairs performed within a 7-day interval were included in the analysis. Clinical data were collected by reviewing electronic medical records. The study outcomes were the 5-year all-cause death (ACD) and a composite of the 5-year ACD and hospitalization for HF (HHF)2.
Composition of the training and validation cohorts. Echocardiography and 12-lead electrocardiography (ECG) pairs performed within a 7-day interval were included in the analysis.
The study protocol was approved and the written informed consent was waived by the Seoul National University Bundang Hospital Institutional Review Board (IRB No. B-2212-801-102) and the Seoul National University Hospital Institutional Review Board (IRB No. J-2302-117-1407), given the retrospective nature of this study. All clinical investigations were conducted according to the principles of the Declaration of Helsinki.
AI algorithm
The AI algorithm was developed using a transfer-learning approach that leveraged the encoder component from an existing deep-learning system. This system, designed to classify cardiac rhythms and provide various risk scores for emergency conditions, utilizes a common ECG encoder based on a modified ResNet-152 architecture. The architecture was further enhanced with squeeze excitation blocks and a non-local network to capture both local and global features from the ECG signals.
We retained the head of the original convolutional neural network (CNN) and, replaced the tail portion with two multi-layer perceptron (MLP) layers. Each layer consists of 128 neurons with ReLU activation functions, and dropout layers were added to mitigate overfitting. These layers were designed to map features from the encoder to a single numerical output, the ECG-GLS score.
The model was fine-tuned using the Adam optimizer with a learning rate of 0.0001, over a maximum of 20 epochs, incorporating early stopping to prevent overfitting. During training, data augmentation techniques, including random shift, rotation, random color jitter, and Gaussian noise, were applied to improve the model robustness. The model was developed using the PyTorch framework and trained on a single RTX A6000 GPU, completing the process in under three hours. Each inference takes less than 1 s per case in routine use. The loss function used for training was the mean squared error (MSE), appropriate for this regression task (Supplementary Figure S1).
Echocardiography and strain analysis
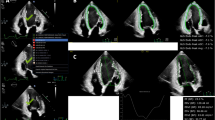
Echocardiographic images were obtained using comprehensive echocardiography in accordance with the guidelines of the American Society of Echocardiography12. LVEF was measured from apical four- and two-chamber views using the Modified Simpson’s method. LVGLS was measured using a built-in software in each echocardiography machine. For the training cohorts 2 and 3, the Image-Arena system (TomTec Imaging Systems, Munich, Germany) was utilized to measure LVGLS. For strain analysis, the endocardial borders were traced on the end-systolic frame defined by the QRS complex. The software tracked speckles along the endocardial border and myocardium throughout the cardiac cycle. The peak longitudinal strain was automatically computed by averaging regional strain values. The LVGLS was obtained from apical three-, four-, and two-chamber views. Because the LVGLS values are expressed as absolute values to avoid confusion, higher values represent better function. For patients with sinus rhythm, analyses were performed on a single cardiac cycle. For patients with atrial fibrillation, strain values were calculated as the average of three cardiac cycles. Representative cases are described in Fig. 2.
Representative case of LVGLS and ECG-GLS score pair. (A) A 43-year-old female who underwent echocardiography for routine checkup. (B) A 78-year-old female with ischemic cardiomyopathy.
Statistical analysis
Continuous variables are presented as means and standard deviations or medians with interquartile ranges and were compared using the independent two-sample t-test or Mann–Whitney U test. Categorical variables are presented as frequencies with percentages and were compared using the χ2 test. The primary measure of model performance was the AUROC for the prediction of LV systolic dysfunction estimated by the LVGLS. AUROC curves were compared using DeLong’s test.
Kaplan–Meier curves for study outcomes were plotted to indicate the discrimination capacities of the ECG-GLS score and LVGLS, which were compared using the log-rank test. We used Cox proportional hazards modeling to compare the predictability for study outcomes between the ECG-GLS score and the echocardiography-derived GLS (LVGLS). The HR was adjusted for parameters that showed an association with study outcomes on univariate analysis (p < 0.1) or had relevant clinical significance, excluding those with > 10% missing data or multicollinearity with other variables. The Harrell’s C-index (C-index) was calculated to compare the predictive performance of Cox proportional hazards models using the bootstrapping method. A two-sample t-test was conducted on 1,000 sets of bootstrapped C-indices to compare the two models.
Statistical significance was defined as a two-sided p value < 0.05. All statistical analyses were performed using R software, version 4.1.2 (https://www.R-project.org).
Results
Baseline characteristics and echocardiographic parameters
A total of 2,882 and 1,186 patients were included in the training and validation cohorts, respectively. The baseline characteristics are presented in Table 1. The training cohort comprised four registries, including patients with acute HF, chronic HF, various heart diseases, and normal heart function from Hospital A. The validation cohort was also drawn from one of the four registries, but the patients were from Hospital B. Age and sex were comparable between the training and validation cohorts; however, other features, including comorbidities, serum creatinine levels, and echocardiographic parameters, were significantly different. The mean interval between ECG and echocardiography was 0.58 ± 1.26 and 1.00 ± 2.49 days in the training and validation cohorts, respectively.
ECG-GLS performance
The LVGLS and ECG-GLS score were significantly correlated, with a correlation coefficient of 0.64 (p < 0.001, Fig. 3A) in the validation cohort. In the training cohort, the areas under the receiver operating characteristic curve (AUROCs) of the ECG-GLS scores for detecting LVGLS ≤ 16% (indicating LV systolic dysfunction) and LVGLS ≤ 12% (indicating more severe LV systolic dysfunction) were 0.93 and 0.90, respectively. In the validation cohort, the AUROCs of the ECG-GLS score for detecting LVGLS ≤ 16% and LVGLS < 12% were 0.85 and 0.82, respectively (Fig. 3B). The precision, recall, and F1 score of the predictive model using the ECG-GLS score to identify patients with LVGLS ≤ 12% were 0.80, 0.84, and 0.82, respectively. The area under the precision-recall curve was 0.90 (Supplementary Figure S2). The diagnostic performance of the ECG-GLS score for identifying patients with HFrEF (LVEF < 40%) was comparable to that of the LVGLS (AUROC, 0.85 vs. 0.83, respectively; p = 0.08; Fig. 4).
ECG-GLS evaluations. (A) In the validation cohort, the LVGLS and ECG-GLS score were significantly correlated (correlation coefficient = 0.64, p < 0.001). (B) Receiver-operating characteristic curves of the ECG-GLS score for the prediction of LVGLS ≤ 12% in the training cohort (blue) and validation cohort (red) are shown. Abbreviations: AUROC, area under the receiver operating characteristic curve; ECG-GLS,electrocardiography-derived global longitudinal strain; LVGLS, left ventricular global longitudinal strain on echocardiography.
Receiver-operating characteristic curves. The performance of the ECG-GLS score (solid line) and LVGLS (dashed line) for the prediction of heart failure with reduced ejection fraction are shown. Abbreviations: AUROC, area under the receiver operating characteristic curve; ECG-GLS,electrocardiography-derived global longitudinal strain; LVGLS, left ventricular global longitudinal strain on echocardiography.
Prediction of study outcomes
Five-year ACD and the composite of ACD and HHF were significantly higher in patients with ECG-GLS scores ≤ 12 than in patients with ECG-GLS scores > 12 (log-rank p = 0.002 and p < 0.001, respectively; Fig. 5A). An analogous pattern was observed for the Kaplan–Meier curves using an LVGLS cutoff of 12% (log-rank p < 0.001 and p = 0.001, respectively; Fig. 5B).
Survival curve comparisons. (A) Event-free survival curves for ACD and the composite outcome of ACD and HHF according to an ECG-GLS score > 12 (blue) and ≤ 12 (red). (B) Event-free survival curves for ACD and the composite outcome of ACD and HHF according to an LVGLS > 12% (green) and ≤ 12% (purple). Abbreviations: ACD, all-cause death; ECG-GLS, electrocardiography-derived global longitudinal strain; HHF, hospitalization for heart failure; LVGLS, left ventricular global longitudinal strain onechocardiography.
LVGLS ≤ 12% (hazard ratio, HR 1.41, 95% confidence interval [CI] 1.16–1.71) and ECG-GLS score ≤ 12 (HR 1.36, 95% CI 1.12–1.66) were associated with a significantly higher 5-year ACD on univariate analysis (Supplementary Table S1). After adjustment for relevant clinical risk factors and echocardiographic parameters, a low ECG-GLS score (≤ 12) was a significant predictor of ACD (HR 1.38, 95% CI 1.11–1.73), with results comparable to those for LVGLS ≤ 12% (HR 1.43, 95% CI 1.14–1.79; p value for bootstrapped mean of Harrel’s C-indices = 0.109; Table 2). Similar results were observed for the composite outcome of the 5-year ACD and HHF. LVGLS ≤ 12% (HR 1.29, 95% CI 1.09–1.54) and ECG-GLS score ≤ 12 (HR 1.34, 95% CI 1.12–1.60) were both significant indicators of the composite outcome (p value for bootstrapped mean of Harrell’s C-indices = 0.423; Table 3; results of the univariate analysis are provided in Supplementary Table S2).
Discussion
In the present study, we demonstrated a strong correlation between the ECG-GLS (estimated from ECG features using an AI-based algorithm) and the LVGLS (measured using echocardiography). We also found that the performance of ECG-GLS in predicting the long-term prognosis of patients with HF and LVGLS was similar to that of the LVGLS, suggesting the relevance of our AI-based algorithm for estimating GLS from ECG features and its potential as a practical alternative to LVGLS. To our knowledge, the present study is the first to attempt to estimate the LVGLS from ECG data using an AI-based algorithm.
Research on AI-based algorithms that use ECG images to detect a range of heart diseases has been reported, expanding the role of AI in cardiology8. The utilization of AI-based ECG algorithms has focused on the early detection of diseases that can be easily diagnosed using ECG but are often missed, such as atrial fibrillation13 or ST-segment elevation myocardial infarction14. However, the rapid growth of AI has expanded the potential of ECG to an extent that it rivals that of echocardiography by revealing previous unseen pieces of information. Acceptable AI-enabled ECG performance has been demonstrated in the diagnosis of various types of HF10,15,16 and myocardial diseases, such as HCM17 and cardiac amyloidosis18. Moreover, AI-enabled ECG HCM scores were correlated with decreases in LV outflow tract gradients and N-terminal pro-B-natriuretic peptide levels over time in patients with obstructive HCM who were prescribed mavacamten19. Most recently, Lee et al. showed improved performance in prognostication related to diastolic dysfunction with AI-enabled ECG20. In the present study, we attempted to extend the capability of AI-enabled ECG to predict LVGLS, a state-of-the-art technology in the echocardiography field. Using ECG features of 2,881 patients from four retrospective registries, our AI-based algorithm for the estimation of LVGLS produced ECG-GLS scores that showed a good correlation with echocardiographic measurements of the LVGLS and were able to predict the long-term prognosis in patients who were admitted for HF. As the ECG-GLS score is a unitless value and is not numerically identical to the LVGLS (%) measured by echocardiography, the correlation coefficient of 0.64 seems lower than expected. However, a small number of patients were distributed out of the limit of agreements in the Bland-Altman analysis (Supplementary Figure S3). Additionally, the prognostic predictability of GLS in HF patients was preserved with a similar cutoff value, which means the distribution of ECG-GLS score is similar to that of LVGLS.
In the present study, the AUROCs of the ECG-GLS score for detecting LVGLS ≤ 16% and ≤ 12% were 0.85 and 0.82, respectively, which were not as high as we expected. This is because the validation cohort comprised patients admitted for HF, whereas patients with all ranges of LVEF and LVGLS were included in the training cohort. In addition, the number of patients included in our study was smaller than that in other studies on AI-enabled ECG because of the limited collection of GLS data. Nonetheless, this study is the first to predict LVGLS using AI-enabled ECG. Our ECG-GLS score is comparable to the LVGLS in identifying patients with HFrEF. Furthermore, the ECG-GLS score is consistent with the familiar unit of the LVGLS. Thus, the ECG-GLS score is user-friendly and can be used interchangeably with the LVGLS in real-world settings.
Furthermore, the ECG-GLS score was comparable to the LVGLS score in predicting the long-term prognosis of patients. LVGLS has been shown to be a better indicator than LVEF for predicting cardiovascular outcomes in hospitalized patients with acute HF2. Previous studies on AI-enabled ECG have predominantly emphasized its diagnostic capabilities, specifically for detecting various heart diseases or conditions. Although the diagnostic potential of AI-enabled ECG is considerable and holds significant promise for clinical use, augmenting the output of AI-enabled ECG algorithms with prognostic information can further enhance their utility. This integration of prognostic data reinforces and validates the diagnostic accuracy of the algorithm. Consistent with this, our findings indicate that AI-enabled ECG serves not only as an accurate indicator of the LVGLS, but also as a valuable tool for prognostication, thereby extending its role beyond mere diagnostics.
ECG has many advantages: it is ubiquitous, inexpensive, rapid, and does not require special training. Recently, AI researchers have expanded the role of ECG in the diagnosis of various heart diseases8. Although the LVGLS has recently been recognized as a valuable tool, echocardiographic LVGLS measurements require additional time and effort from experts, as well as dedicated software or machines, which makes GLS underutilized compared to its recommendations and demonstrated utility21. The LVGLS also has the troublesome issue of between-vendor variability22. Moreover, measuring the LVGLS using echocardiography may be limited in patients with poor echocardiographic windows, atrial fibrillation, and videos with low frame rates23. In contrast, the ECG-GLS score is free from almost all of these limitations. Using the ECG-GLS score, non-experts can also predict the value of the LVGLS, diagnose LV systolic dysfunction, and forecast the outcome of patients easily, making it usable outside the echocardiography laboratory and significantly reducing time and costs. Integrating AI in interpreting ECG results could potentially improve the cardiac diagnostic process, making it more accessible and less reliant on specialized training and equipment.
Limitations
This study has several limitations. First, the ECG and echocardiography pairs used in this study were insufficient to provide better prediction. Although we included diverse population with a wide range of LVGLS values in the training cohort, the potential for overfitting remains. Future studies should incorporate a larger dataset, with more diverse populations to further improve the model’s performance. Additionally, evaluating the utility of the ECG-GLS score in a more diverse population across various clinical settings will be essential to fully assess its generalizability and robustness. Second, although various vendors were used in the measurement of the LVGLS, the number of ECG-echocardiography pairs used in the training cohort significantly differed among vendors. Hence, the generalization of our ECG-GLS score prediction to all strain vendors requires further investigation. This may limit the generalizability of the findings to broader populations. In addition, there was no formal sample size calculation for this study, as we used all available data to enhance model performance. Finally, the validation cohort included patients with HF, which made the predictive ability of our ECG-GLS score lower than expected. Further studies are needed to demonstrate the utility of the ECG-GLS score in various populations.
Conclusions
The ECG-GLS score, estimated from ECG features using an AI-based algorithm, shows a meaningful correlation with the LVGLS measured on echocardiography and is effective in risk stratification for the long-term prognosis after acute HF. While ECG-GLS score is not a direct substitute for echocardiographic LVGLS, it offers a practical and accessible alternative, particularly in settings where echocardiography is not readily available. With further refinement and inclusion of additional datasets, the ECG-GLS score has the potential to become a valuable tool in clinical practice, complementing traditional imaging modalities.
Data availability
The data used in the present study cannot be made publicly available owing to ethical restrictions set by the Seoul National University Bundang Hospital Institutional Review Board (https://e-irb.snubh.org), as patient and participant privacy would be compromised. A minimally anonymized dataset can be requested by contacting the corresponding authors.
References
Potter, E. & Marwick, T. H. Assessment of left ventricular function by echocardiography: The case for routinely adding global longitudinal strain to ejection fraction. JACC Cardiovasc. Imaging. 11, 260–274. https://doi.org/10.1016/j.jcmg.2017.11.017 (2018).
Park, J. J., Park, J. B., Park, J. H. & Cho, G. Y. Global longitudinal strain to predict mortality in patients with acute heart failure. J. Am. Coll. Cardiol. 71, 1947–1957. https://doi.org/10.1016/j.jacc.2018.02.064 (2018).
Magne, J. et al. Distribution and prognostic significance of left ventricular global longitudinal strain in asymptomatic significant aortic stenosis: An individual participant data meta-analysis. JACC Cardiovasc. Imaging. 12, 84–92. https://doi.org/10.1016/j.jcmg.2018.11.005 (2019).
Zhu, D. et al. Left ventricular global longitudinal strain is associated with long-term outcomes in moderate aortic stenosis. Circ. Cardiovasc. Imaging. 13, e009958. https://doi.org/10.1161/CIRCIMAGING.119.009958 (2020).
Alashi, A. et al. Incremental prognostic utility of left ventricular global longitudinal strain in asymptomatic patients with significant chronic aortic regurgitation and preserved left ventricular ejection Fraction. JACC Cardiovasc. Imaging. 11, 673–682. https://doi.org/10.1016/j.jcmg.2017.02.016 (2018).
Tower-Rader, A. et al. Prognostic value of global longitudinal strain in hypertrophic cardiomyopathy: A systematic review of existing literature. JACC Cardiovasc. Imaging. 12, 1930–1942. https://doi.org/10.1016/j.jcmg.2018.07.016 (2019).
Siontis, K. C., Noseworthy, P. A., Attia, Z. I. & Friedman, P. A. Artificial intelligence-enhanced electrocardiography in cardiovascular disease management. Nat. Rev. Cardiol. 18, 465–478. https://doi.org/10.1038/s41569-020-00503-2 (2021).
Attia, Z. I., Harmon, D. M., Behr, E. R. & Friedman, P. A. Application of artificial intelligence to the electrocardiogram. Eur. Heart J. 42, 4717–4730. https://doi.org/10.1093/eurheartj/ehab649 (2021).
Choi, Y. J. et al. Artificial intelligence versus physicians on interpretation of printed ECG images: Diagnostic performance of ST-elevation myocardial infarction on electrocardiography. Int. J. Cardiol. 363, 6–10. https://doi.org/10.1016/j.ijcard.2022.06.012 (2022).
Adedinsewo, D. et al. Artificial intelligence-enabled ECG algorithm to identify patients with left ventricular systolic dysfunction presenting to the Emergency Department with Dyspnea. Circ. Arrhythm. Electrophysiol. 13, e008437. https://doi.org/10.1161/CIRCEP.120.008437 (2020).
Moon, M. G. et al. Reverse remodeling assessed by left atrial and ventricular strain reflects treatment response to Sacubitril/Valsartan. JACC Cardiovasc. Imaging. 15, 1525–1541. https://doi.org/10.1016/j.jcmg.2022.03.019 (2022).
Lang, R. M. et al. Recommendations for cardiac chamber quantification by echocardiography in adults: An update from the American Society of Echocardiography and the European Association of Cardiovascular Imaging. J. Am. Soc. Echocardiogr. 28, 1–39 e14. https://doi.org/10.1016/j.echo.2014.10.003 (2015).
Attia, Z. I. et al. An artificial intelligence-enabled ECG algorithm for the identification of patients with atrial fibrillation during sinus rhythm: A retrospective analysis of outcome prediction. Lancet. 394, 861–867. https://doi.org/10.1016/s0140-6736(19)31721-0 (2019).
Kim, D. et al. A retrospective clinical evaluation of an artificial intelligence screening method for early detection of STEMI in the emergency department. J. Korean Med. Sci. 37, e81. https://doi.org/10.3346/jkms.2022.37.e81 (2022).
Attia, Z. I. et al. Screening for cardiac contractile dysfunction using an artificial intelligence-enabled electrocardiogram. Nat. Med. 25, 70–74. https://doi.org/10.1038/s41591-018-0240-2 (2019).
Cho, Y. et al. Artificial intelligence-based electrocardiographic biomarker for outcome prediction in patients with acute heart failure: Prospective cohort study. J. Med. Internet Res. 26, e52139. https://doi.org/10.2196/52139 (2024).
Ko, W. Y. et al. Detection of hypertrophic cardiomyopathy using a convolutional neural network-enabled electrocardiogram. J. Am. Coll. Cardiol. 75, 722–733. https://doi.org/10.1016/j.jacc.2019.12.030 (2020).
Tison, G. H., Zhang, J., Delling, F. N. & Deo, R. C. Automated and interpretable patient ECG profiles for disease detection, tracking, and discovery. Circulation Cardiovasc. Qual. Outcomes. 12 https://doi.org/10.1161/circoutcomes.118.005289 (2019).
Tison, G. H. et al. Assessment of disease status and treatment response with artificial intelligence-enhanced electrocardiography in obstructive hypertrophic cardiomyopathy. J. Am. Coll. Cardiol. 79, 1032–1034. https://doi.org/10.1016/j.jacc.2022.01.005 (2022).
Lee, E. et al. Artificial intelligence-enabled ECG for left ventricular diastolic function and filling pressure. NPJ Digit. Med. 7. https://doi.org/10.1038/s41746-023-00993-7 (2024).
Barbieri, A. et al. Current use of echocardiography in cardio-oncology: Nationwide real-world data from an ANMCO/SIECVI joint survey. Eur. Heart J. Imaging Methods Pract. 2, qyae081. https://doi.org/10.1093/ehjimp/qyae081 (2024).
Farsalinos, K. E. et al. Head-to-Head comparison of global longitudinal strain measurements among nine different vendors: The EACVI/ASE inter-vendor comparison study. J. Am. Soc. Echocardiogr. 28, 1171–1181. https://doi.org/10.1016/j.echo.2015.06.011 (2015).
Voigt, J. U. et al. Definitions for a common standard for 2D speckle tracking echocardiography: Consensus document of the EACVI/ASE/Industry Task Force to standardize deformation imaging. J. Am. Soc. Echocardiogr. 28, 183–193. https://doi.org/10.1016/j.echo.2014.11.003 (2015).
Funding
This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), which is funded by the Ministry of Health & Welfare, Republic of Korea (grant number: RS-2023-00265933).
Author information
Authors and Affiliations
Contributions
All authors were involved in the conceptualization and design of this research. JP, H-KK, HJC, and YEY were involved in data collection, and H-MC, JK, J-BP, G-YC, YC, and I-CH analyzed the data. JK was responsible for the generation of the ECG-GLS score. H-MC wrote the manuscript draft and prepared figures, and JK, YC, and I-CH reviewed and corrected the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Joonghee Kim developed the algorithm and is the founder and CEO of the startup company, ARPI Inc. Youngjin Cho has worked at ARPI Inc. as a research director. The remaining authors declare no conflicts of interest.
Independent data access and analysis
YC had full access to all the data in the study and takes responsibility for its integrity and the data analysis.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Choi, HM., Kim, J., Park, J. et al. AI derived ECG global longitudinal strain compared to echocardiographic measurements. Sci Rep 14, 26458 (2024). https://doi.org/10.1038/s41598-024-78268-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-78268-8
Keywords
This article is cited by
-
Evolving role of echocardiography across the heart failure spectrum: from prevention to advanced management
Heart Failure Reviews (2025)
-
Current State of Artificial Intelligence in Assessing Cardiac Function
Current Cardiology Reports (2025)