Abstract
The effects of random dislocations on the vibrational properties of finite-length RNA and DNA macro-structures have been investigated by means of a harmonic Hamiltonian and the Green’s function method. The RNA molecule has been modeled using a half ladder model, and three models (a fishbone model and two different strand models) have been employed to model the structure of DNA. The lengths of the finite and cyclic systems are gradually increased to more accurately approximate the structures of RNA and DNA. Springs whose behaviors are governed by Hooke’s constitutive law have been used to represent the bonds between the masses, with the stiffness of the vertical springs randomly changing along the length of each model. This results in a more realistic representation of the inherent randomness of the studied structures. To investigate the effect of random dislocations on the mechanical response of the studied systems, it has been assumed that a random mass-spring ensemble has been knocked out of place by an external force. It has been found that increasing the number of building blocks along the length of the models suppresses the influence of dislocations on the vibration spectra of RNA and DNA. It was also observed that at low frequencies, the influence of dislocations becomes more pronounced. Besides, taking into account the collective mass of the sugar-phosphate backbone results in the appearance of gaps in the vibration spectra. By introducing dislocations into the models, additional dislocation-induced vibrational states appear in the DOS curves. What stands out from the results is that the responses of the studied systems to these changes are strictly nonlinear. The employed methodology can be applied to investigate the mechanical response of damaged RNA and DNA.
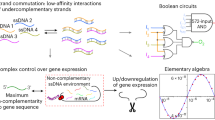
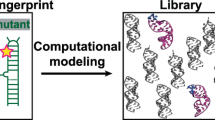
Similar content being viewed by others
Introduction
Ribonucleic acid (RNA)1,2,3,4and deoxyribonucleic acid (DNA)5are the genetic material found in all living organisms and comprise the building blocks of life. They are how genetic information is stored and read. Within the last few decades, these two macro-molecules have come to be treated as natural nanowires, and due to their interesting properties- e.g., self-assembly, easy modification, etc.- have attracted a great deal of attention from the scientific community6,7. DNA consists of two strands wrapped together in a double-helical structure, while RNA is a single-stranded molecule (Fig. 1). These strands are made up of subunits called nucleotides. Each nucleotide contains a nitrogenous base, a carbon-based sugar (S) group (deoxyribose in DNA and ribose in RNA), and a phosphate (P) group, which is attached to the S molecule. Nitrogenous bases in DNA contain one of four possible bases: adenine (A), thymine (T), cytosine (C), and guanine (G). These bases usually occur in pair units, commonly known as base-pairs, with A always binding with T (AT), and C always pairing up with G (CG). RNA does not contain any T bases. The molecule instead makes use of another base known as Uracil (U), which attach to A. Both DNA and RNA feature a sugar-phosphate backbone, but compared to RNA’s ribose, the deoxyribose in DNA lacks one hydroxyl group.
Both of these macro-structures run the risk of being damaged by the environment and if not immediately repaired, the resulting mutations can lead to undesirable effects and ultimately disease. These damages can be mechanical and take the form of dislocations in the structure of RNA/DNA. Dislocations are environment-caused changes in the crystalline structure of a system. An example of the impact of the environment on DNA is skin cancer, which can be brough on by overexposure to the sun’s ultraviolet rays8,9. Similarly, any disorder in the structure of RNA can also lead to such problems as pathogenesis of cancer, cardiac diseases, various types of infections, and autoimmune/inflammatory illnesses10,11,12. In addition to environmental factors, DNA and RNA can also be exposed to oxidative damage from metabolic byproducts such as free radicals13. Quantitative and qualitative investigations into the electronic and mechanical properties of DNA and RNA sequences can be very effective in the diagnosis and treatment of diseases, case-specific medicine, environmental analysis, drug discovery, and so many other fields14,15,16. In recent years, the applications of DNA and RNA in electronic biosensors capable of diagnosing diseases have been studied17,18,19,20,21. For example, the conductance and charge transport properties of single microRNA chains associated with autism have been investigated by Oliveira et al20.. Santhanam et al. introduced DNA/RNA electrochemical biosensing devices that can replace the existing polymerase chain reaction methods for the fast containment of widespread outbreaks like the Covid-19 pandemic21.
Although the electronic properties of isolated RNA and DNA molecules have been the subject of exhaustive research and study22,23,24,25,26,27,28,29,30, the analogous vibrational properties remain somewhat unexplored. Vibrating on timescales from femtoseconds to nanoseconds, RNA and DNA are by no means static structures31. Having an in-depth knowledge of the dynamical properties of DNA is central to the development of theoretical models of biological processes such as base-pair opening, replication, and transcription32. The sequencing pattern of the base-pairs and how it influences the mechanical properties of the DNA molecule has previously been investigated33. It was demonstrated that the sequencing pattern influences the DNA’s flexibility and its ability to form links. This happens when certain patterns slide past others to produce non-native contacts. This behavior is inherently nonlinear. In the investigation presented herein, this nonlinearity manifests itself in both the density of states (DOS) and vibration curves. The same is also true for the influence of a dislocation, which is essentially a nonlinear phenomenon. These alterations to the structures of the systems are, surprisingly, not met by a linear response from the system.
Understanding DNA transcription begins with studying DNA thermal denaturation, a process explored by Dauxois and Peyrard34. They introduced a nonlinear dynamical model for DNA denaturation, incorporating cooperativity effects via anharmonic nearest-neighbor stacking interactions. Their transfer-integral calculations reveal that this one-dimensional model, even without long-range interactions, undergoes a sharp denaturation akin to a first-order transition at a temperature lower than that of a similar model with harmonic coupling. Self-consistent phonon calculations emphasize the critical importance of nonlinear effects in this mechanism. Zoli35presents an intermediate (mesoscopic) approach to constructing a three-dimensional Hamiltonian model that captures the primary interactions responsible for the stability of helical molecules. This method demonstrates that the Hamiltonian’s complexity can be incrementally increased by incorporating additional elements, such as new degrees of freedom, thereby offering a more precise depiction of the double helix in three dimensions. The 3D mesoscopic model provides a detailed description of the helix at the base pair level and enables the prediction of the thermodynamic and structural properties of molecules in solution. Demonstrating a phase transition, DNA melting has been extensively researched within the field of statistical physics36, resulting in a significant body of literature that explores the helix-coil transition and the thermally induced creation of denaturation bubbles. The flexibility characteristics of DNA helices have also been the subject of thorough examination, including aspects such as force-extension behavior, persistence length, and cyclization probability-the likelihood that a portion of linear chain ensembles will form loops. To investigate these properties, numerous computational techniques have been employed, including transfer integrals37,38and transfer matrix methods39, Monte Carlo simulations40, molecular dynamics41, and path integrals42,43,44. These methods have been applied to both mesoscopic Hamiltonian and polymer physics models.
Future advancements in technology may see conventional solid-state devices replaced by molecular components, leveraging the mechanical properties of a few particles. DNA and RNA molecules are prime candidates for this role, thanks to their flexibility, self-assembling capabilities, and well-defined, reproducible architectures. While the electronic properties of these molecules are important, their elastic properties also play a crucial role in numerous biological processes, as highlighted by Barbi et al. in their 1999 study45. Over the years, the vibrational properties of DNA and RNA have been extensively studied using Raman and IR spectroscopy. Detailed atomic-level descriptions are necessary to fully understand the fine structure observed in Raman spectroscopy, but simple mechanical models can effectively describe macromolecular motion. This is essential for investigating the mechanical properties of DNA and RNA macrostructures, as our work demonstrates.
When analyzing the effects of hydrogen bonds, single-chain models fall short, necessitating the use of double-chain models. These models, widely discussed in the literature, represent RNA and DNA as single- and double-chain structures with nucleotides depicted as effective sites connected by effective elastic constants46,47,48. The double-chain model enables the examination of interchain coupling’s impact on vibrational properties. Since DNA lacks periodicity, disorder effects such as dislocations-which significantly influence DNA behavior-are incorporated into the models. Simple elastic rod models provide valuable insights into elastic properties but overlook the local polymorphism due to base-pair sequencing. In contrast, models using Hamiltonian and equations of motion, as employed in our study, treat atoms as lattice points within the crystalline structure, forming the entire DNA and RNA sequences. Intermediate heuristic models, which account for local inhomogeneity due to base pairing, offer a desirable balance by maintaining numerical stability and microscopic detail while simplifying fully microscopic structures.
The sequence-dependent nature of DNA and RNA, along with inherent randomness, affects the mechanical response of these structures. Variations in the masses of bases and the elastic force between pairs influence the systems’ potential energy and vibrational behavior. Our models, which represent DNA and RNA molecules without their natural twisting, consider these molecules as quasi-1D systems with randomly placed base pairs or building blocks. By applying these mechanical models, we aim to understand macromolecular motions in the presence of random dislocations. Our primary goal is to present a methodology for investigating the mechanical response of such macromolecules from a physical perspective. This methodology is adaptable to other biopolymers with similar structures. Our analyses are based on two main propositions: formulating an analytical model to predict the vibrational response of DNA- and RNA-like structures, and incorporating an increasing number of blocks to achieve a more accurate representation of these macro-structures.
In this theoretical study, a framework has been presented to investigate the influence of random dislocations on the mechanical properties of different quasi-one-dimensional (1D) macro-structures and the nonlinear consequences of this interaction. It needs to be mentioned that the best known way to account for the hydrogen bonds between the pairs and the elements of the sugar-phosphate backbone is to use the Morse potential35. In the present work, however, these bonds have been mimicked using the linear-elastic relationship described by Hooke’s law of elasticity. Although this approach imposes an approximation on the results, it is nonetheless a tried and tested procedure employed by many experts49. We have adopted a quantum mechanical framework for our analysis. The derivation of the DOS curves for the lattice structures of DNA and RNA relies on quantum mechanical principles. In our computations, the Hamiltonian functions as an operator derived from displacement and momentum operators, which adhere to a commutation relationship. The equation of motion is derived using the Heisenberg equation. As the Green’s function is obtained directly from this equation of motion-a fundamentally quantum mechanical relationship-both the Green’s function and the equation of motion are inherently quantum mechanical. To that end, a half ladder model (HLM) of RNA and three different models of DNA - i.e. a fishbone model (FBM) and two double-strand models (DSM) labeled 1DSM and 2DSM- have been constructed. It is worth noting that all of the models have been studied in two different configurations (finite and cyclic). The calculations have been carried out utilizing a harmonic approximation50,51and the Green’s function method52,53,54,56,57,58. The nitrogenous bases in the RNA model and the base-pairs in the two DNA models have been arranged in three different randomly-arranged configurations (32, 64, and 128 bases/base-pair). Each model features a single nonlinear imperfection (dislocation) at a random location along its length. This paper is organized as follows: In section II, the general characteristics of the different models are described. In section III, the equation of motion for the dynamical Green’s function is obtained, the DOS formula is introduced in terms of the Green’s function, and the section is capped off with the assembly of the dynamic matrix. Finally, in sections IV and V, the results are discussed and the conclusions are presented.
Schematic structures of the (a) RNA and (b) DNA strands. DNA comprised of repeated stacks of either adenine (A)-thymine (T) or guanine (G)-cytosine (C) base-pairs, coupled by hydrogen bonds, twisted in a double-helix structure with a sugar-phosphate backbone. RNA has common A, G, and C bases with DNA, but in place of T, it features Uracil (U).
Model
To study the effects of random dislocations on the mechanical properties of finite-length RNA and DNA, the following four models have been constructed: The HLM model for RNA and the FBM, 1DSM, and 2DSM models for DNA. These models, along with schematic representations of the effects of dislocations, are illustrated in Fig. 2(a-d). Fig. 2a shows the structure of the HLM model of the RNA strand. In this model, the nitrogenous bases are modeled as a quasi-1D chain of masses linked together in a repetitive structure. Each base is connected vertically to the SP backbone. A schematic view of the structure of the FBM model of DNA is given in Fig. 2b. In this model, AT and CG base-pairs are considered as a chain of masses. Consecutive base-pairs are linked together and have been randomly distributed along the DNA molecule. The SP backbones of the DNA have been modeled by assemblies of masses on the upper and lower boundaries. Backbone sites are linked to the base-pairs but share no connection. The 1DSM model has been built based on the ladder-like structure of DNA and contains only two rows of mass (Fig. 2c). Each site represents a nitrogenous base (A, T, C, and G) plus the adjacent SP backbone. In the 2DSM model, the nitrogenous bases and the SP backbone have been modeled by individual sites. Each base is connected to its nearest neighbors via horizontal and vertical links and interacts with the SP backbone, similar to the FBM model (Fig. 2d). The 2DSM model is essentially a combination of the FBM and 1DSM models. In all of the models, the interaction potential between the masses is modeled using a series of horizontal and vertical springs whose behaviors have been defined using Hooke’s constitutive equation. To investigate how a randomly-placed dislocation affects the vibrational spectra of finite-length RNA and DNA, a random mass-spring ensemble was eliminated from each model (Fig. 2(a-d)). In both configurations (finite and cyclic), a limited number of bases/base-pairs has been considered for each model. The number of bases is increased in three steps, \(N_{bp}(N_{b})=32, 64, 128\), to calculate the DOS and dispersion diagrams of the DNA and RNA macro-molecules. Since DNA and RNA are not periodic systems, disorder effects - which determine key features of DNA and RNA behavior - are introduced in the models. Evidently, comparing the responses of perfect and imperfect models (regardless of their finite/cyclic nature) would yield important insights into the responses and the consequences thereof, even at a theoretical level. Finally, a comparison is drawn between the results of the perfect and imperfect structures.
The imperfect models- i.e., models with dislocations- of the HLM model (for modeling the RNA strand) and the FBM, 1DSM, and 2DSM (for modeling the DNA strand) are presented in panels (a), (b), (c), and (d), respectively. \(j_{0}\), \(j_{n}\) (\(n=1,2\)), and \(j_{n^{\prime }}\) (\(n^{\prime }=3,4\)) represent the stiffnesses of the linear springs mimicking the molecular bonds between the subsites of the building blocks. The masses of the sub-sites inside the BLUCs of the models are labeled from \(M_{1}\) to \(M_{N_{s}}\).
Formulation
As depicted in Fig. 2(a-d), the interaction forces between the building blocks of the RNA and DNA structures are represented by equivalent springs whose behaviors are governed by Hooke’s constitutive law. The following equation would give Hamiltonian of the vibrating system:
in which \(M_{\alpha }\) represents the direction-independent mass of the \(\alpha\) sub-site. Also, \(\hat{p}^{\alpha }_{\mu }\) and \(\hat{x}^{\alpha }_{\mu }\) are, respectively, the Cartesian momentum and displacement operators of the \(\alpha\) sub-site along the \(\mu\) direction. These operators are used to satisfy Eq. (2):
a commutation relationship in which \(\delta\) represents the Kronecker symbol. In Eq. (1), \(N_{d}\) and \(N_{s}\) stand for the dimension and the number of sub-sites in each system, respectively. Furthermore, \(K^{\alpha \beta }_{\mu \nu }\) is the force-constant tensor for sub-site \(\alpha\) along the direction \(\mu\) due to a small displacement in sub-site \(\beta\) along the direction \(\nu\). The tensor can be calculated by:
wherein U represents the interaction potential of the system, a function of the instantaneous positions of all the atoms in the system. The subscript “zero” on the right-hand side of the equation corresponds to a potential minimum. The mass-normalized displacements can be defined as \(u^{\alpha }_{\mu }=\sqrt{M_{\alpha }}x^{\alpha }_{\mu }\). Hence, \({\varvec{u}}\) is the normalized column matrix of the displacements containing all of the displacements whose transpose can be assembled into the form of the following \(1\times (N_{d}N_{s})\) row matrix:
where \(N_{s}\) for the HLM, FBM, 1DSM, and 2DSM models is \(N_{s}=2N_{b}\), \(N_{s}=3N_{bp}\), \(N_{s}=2N_{bp}\), and \(N_{s}=4N_{bp}\), respectively. In this notation, the harmonic Hamiltonian assumes the following matrix form:
wherein \(\dot{{\varvec{u}}}=\textrm{d}{\varvec{u}}/\textrm{d}t\) is the corresponding conjugate momentum, with t standing for time and \({\varvec{\varPhi }}\) representing the mass-normalized force-constant tensor. The latter can be obtained using Eq. (6):
in which \(m_{\alpha \beta }\equiv \sqrt{M_{\alpha }M_{\beta }}\). The matrix form of \({\varvec{\varPhi }}\) would be as follows:
which is a \((N_{d}N_{s})\times (N_{d}N_{s})\) matrix. It should be mentioned that \({\varvec{u}}\) and \(\dot{{\varvec{u}}}\) satisfy the following commutation relationship:
with \({\varvec{I}}\) standing for a unit matrix. Note that all of the calculations have been carried out using dimensionless units, i.e. all of the physical constants are set equal to unity.
Now, we proceed to describe the Green’s function approach, a tool suitable for obtaining the vibrational spectrum of the system in terms of its DOS. The retarded Green’s function is defined as59:
in which \(\Theta (t)\) represents the Heaviside step function for time t and \(\langle ...\rangle\) represents the average over the canonical density matrix. By definition, \({\varvec{D}}\) is a square matrix that is equal to zero when \(t\le 0\). Taking the derivative of the retarded Green’s function in Eq. (9) with respect to time is the first step in obtaining the equation of motion:
which results in the derivation of the following expression:
in which \(\delta (t)=\textrm{d}\Theta \left( t\right) /\textrm{d} t\), with \(\delta (t)\) representing the Dirac \(\delta\)-function. At \(t=0\), the square brackets in the first term on the right-hand side of Eq. (11) takes on the value of zero, rendering the accompanying \(\delta (t)\), and therefore the entire term preceding the plus sign, equal to zero. Eq. (11) can therefore be expressed as:
To obtain the equation of motion, the first derivative of Eq. (11) would not suffice. Therefore, taking the second derivative of the equation with respect to time yields:
Eq. (14) gives the first term on the right-hand side of Eq. (13):
where multiplying \(\delta (t)\) by the term inside the angle brackets guarantees the calculation of the commutation relation at time t. The Heisenberg equation for the conjugate momentum can be employed to calculate the second term:
wherein the Hamiltonian (Eq. (5)) should be expressed at time t. Hence, \(\textrm{d}\dot{{\varvec{u}}}(t)/\textrm{d} t\) can be written as:
the first term on the right-hand side of Eq. (16) is equal to zero and the second term can be obtained by the following equation:
Using Eqs. (13), (14), (16), and (17), the equation of motion would be derived as:
The Green’s function depends solely on changes in time t, or on the time-translation invariant. We can therefore present the frequency-domain Fourier transform of the Green’s function in the following form:
Multiplying Eq. (18) by \(\exp (-\textrm{i}\omega t)\) and evaluating the integral over time t, the frequency domain of the Green’s function would be obtained as:
in which the following identity has been employed:
In Eq. (20), \(\omega\) has been replaced by \(\omega +\textrm{i}0^{+}\). This operation results in the retarded Green’s function to be analytic in the upper half of the complex frequency plane. The vibrational DOS60 can therefore be written as:
a relationship wherein the trace is taken over all quantum numbers which label the Hamiltonian. Thus, Eq. (22) can be rewritten as:
in which the value of the parameter \(D^{\alpha \alpha }\) is given by Eq. (20). The vibrational spectrum of each finite/cyclic system with no imperfections can be numerically computed via the following characteristic equation:
Since all of the models studied herein are systems with finite lengths, assuming that each model contains \(N_{s}\) sub-sites, the dynamic matrix would be a \(N_{s}\times N_{s}\) square matrix. Therefore, the first square block of Eq. (7) would be sufficient to represent the entire dynamic matrix:
where the Cartesian coordinate subscripts (\(\mu\) and \(\nu\)) have been excluded. Using Eqs. (3), (6) and (25), the general form of the dynamic matrix for the cyclic perfect HLM model of the RNA structure would be obtained as:
in which \(j_{0}\) is the force constant representing the stiffness of the springs between the bases and \(j_{n^{\prime }}\) (\(n^{\prime }=3,4\)) stands for the force constant of the vertical springs connecting the AU/GC base-pair to the SP backbones (Fig. 2a). The masses of all different sub-sites are given by a unique label ranging from \(M_{1}\) to \(M_{N_{s}}\) (Fig. 2(a-d)). The dynamic matrix of the finite-length HLM model is similar to that of the cyclic model, with the exception that \(\varPhi _{2N_{s}}=\varPhi _{N_{s}2}=0\) and
To investigate the effects of random dislocations, one mass (along with its corresponding equivalent spring) has been removed from the SP backbone. As demonstrated by Fig. 2a, it is assumed that the ith mass has been removed, in which case the dynamic matrix elements undergo the following change:
The dynamic matrices of the other cyclic and finite systems are introduced in the Supplementary Material.
Results and discussion
The formulism developed in the previous sections is employed to investigate the effects of random dislocations on the vibrational spectrum of the HLM model of RNA and the FBM, 1DSM, and 2DSM models of DNA. The results will then be compared to those of the perfect and imperfect finite and cyclic configurations. For the finite and cyclic cases, three different lengths have been considered for each system (\(N_{bp}(N_{b})=32, 64, 128\)). Also, a mass (and its corresponding spring(s)) has been randomly removed from each structure to investigate the effects of dislocations. We first introduce the necessary parameters to obtain the numerical results. Based on the shapes of their structures, the four models have been allocated the following mass distributions: the HLM model of the RNA molecule (Fig. 2a) contains five different masses, i.e., the individual masses of the A, U, C, and G bases, and the collective mass of the SP backbone. This model contains three spring constants, \(j_{0}\), \(j_{3}\), and \(j_{4}\). The horizontal springs are labeled with \(j_{0}\), and \(\{j_{3},j_{4}\}\) dub the vertical springs. The finite and cyclic configurations include three different masses in the FBM model (Fig. 2b): (1) the mass of the AT base-pair, (2) the mass of the CG base-pair, and (3) the collective mass of the SP backbone. These three masses are linked to each other by an ensemble of three springs with different stiffnesses. The force constants of the horizontal springs connecting the AT and GC blocks are represented in our calculations by \(j_{0}\). Also, \(j_{1}(j_{2})\) is the force constant of the perpendicular springs connecting the AT (GC) blocks to the SP backbones. The 1DSM model contains four different masses (Fig. 2c): (1) the mass of base A plus the mass of the SP backbone, (2) the mass of base T plus the mass of the SP backbone, (3) the mass of base C plus the mass of the SP backbone, and (4) the mass of base G plus the mass of the SP backbone. The model also includes three types of springs, with \(j_{0}\) representing the force constant of the horizontal springs and \(j_{1}(j_{2})\) standing for the force constant of the vertical springs connecting the bases of the AT (GC) base-pairs. The 2DSM model includes five different masses (Fig. 2d): (1) the mass of base A, (2) the mass of base T, (3) the mass of base C, (4) the mass of base G, and (5) the mass of the SP backbone. The five force constants in this model include \(j_{0}\), \(j_{1}\), \(j_{2}\), \(j_{3}\), and \(j_{4}\). Similar to the previous models, \(j_{0}\) is the force constant of the horizontal springs and the remaining four are the force constants of the vertical springs. The atomic masses of the building blocks of the models are given in Table 161. These masses are randomly distributed throughout the HLM model of RNA in the form of double A-SP, U-SP, C-SP, and G-SP sets. Furthermore, the masses appear in the form of triple SP-AT-SP, SP-CG-SP, SP-TA-SP, SP-GC-SP, double A-T, C-G, T-A, G-C, and quadruple SP-A-T-SP, SP-C-G-SP, SP-T-A-SP, SP-G-C-SP sets in the FBM, 1DSM, and 2DSM models of DNA, respectively. Each of these mass groups contains a set of force constants also appearing randomly inside the models.
In the calculations presented herein, two different force constants have been employed: \(j_{0}=0.090\;\textrm{mdyne}/{\text{\AA }}\) and \(j_{1}=0.075\;\textrm{mdyne}/{\text{\AA }}\)62. Besides, we define \(j_{2}\), \(j_{3}\), and \(j_{4}\) in terms of \(j_{0}\), whose values are equal to \(0.75j_{0}\), \(0.70j_{0}\), and \(0.60j_{0}\), respectively. Also, in the numerical calculations, we have taken into account the mass of the SP backbone in the DNA (\(M_{0}\)) and the force constants of the horizontal springs (\(j_{0}\)). Furthermore, the intra-pair distances have been set equal to unity. These are scales in terms of which the remaining parameters will be expressed. Also, the relationships \(q_{max}=\pi /a\) and \(\omega _0=\sqrt{j_{0}/M_{0}}\) have been defined to make the wave vector and the frequency axes dimensionless.
The effect of random dislocations on the dispersion curves and the DOS of the RNA and DNA structures are illustrated in Figs. 3-6. To capture the effect of the randomness inherent to RNA and DNA, the size of each finite/cyclic system is gradually increased and then the DOS of each system is calculated. The number of block groups along the length of the finite and cyclic models have been assumed equal to \(N_{b}=32,64,128\) in the RNA model and \(N_{bp}=32,64,128\) in the DNA models. Although, for a fixed number of \(N_{bp}(N_{b})\), all of the finite and cyclic models have the same length, with the number of sub-sites inside each model differing from the rest (\(N_{s}=2N_{b}\) for the HLM, \(N_{s}=3N_{bp}\) for the FBM, \(N_{s}=2N_{bp}\) for the 1DSM, and \(N_{s}=4N_{bp}\) for the 2DSM). Using the equations presented in the previous sections, the dispersion curves and the DOS of the two different configurations (i.e., finite and cyclic) of the RNA and DNA models have been calculated. The energy eigenvalues of the dynamic matrices of the finite and cyclic RNA and DNA models with \(N_{b}(N_{bp})=32\) have been plotted as a function of their respective site indices and the results are shown in Fig. 3(a-h) and Fig. 4(a-h), respectively. In these figures, the left- and right-hand sides of the panels show, respectively, the dispersion curves of the perfect (without random dislocation (a,c,e,g panels)) and imperfect (with random dislocation (b,d,f,h panels)) states of all four models. Fig. 5(a-l) and Fig. 6(a-l) present the DOS of the four models in, respectively, the finite and cyclic configurations for \(N_{bp}(N_{b})=\{32,64,128\}\). In these two figures, the HLM model is represented by panels (a,b,c), the FBM model by panels (d,e,f), the 1DSM model by panels (g, h, i), and the 2DSM model by panels (j, k, l). Also, the solid and dashed lines show the DOS of the perfect and imperfect systems, respectively.
The vibrational dispersion curves for the finite configuration of (a, b) HLM model (c, d) FBM model (e, f) 1DSM model (g, h) 2DSM model with \(N_{bp}(N_{b})=32\). The left panels (a, c, e, g) show the dispersion curve for the perfect systems, while the right panels (b, d, f, h) relate to the imperfect systems.
The vibrational dispersion curves for cyclic configuration of the (a, b) HLM, (c, d) FBM, (e, f) 1DSM, and (g, h) 2DSM models with \(N_{bp}(N_{b})=32\). The left panels (a, c, e, g) show the dispersion curve for the perfect systems, while the right panels (b, d, f, h) relate to the imperfect systems.
According to Fig. 3(a-h) and Fig. 4(a-h), there is \(N_{s}\)branches. This wide range of atomic motions in DNA, from low (collective motion, confirmed by Raman spectroscopy63) to high frequency (local vibration), has been theoretically explained64. The low frequency vibration in DNA molecules might play an important role in DNA ’breathing’ and drug intercalation65. Also, the degenerate bands in the dispersion curves of the HLM, FBM, and 2DSM models are fewer than those in the 1DSM model. The reason is the low symmetry of these models which results in the appearance of fewer branches in the dispersion diagrams. Several gaps can be seen between the branches of the finite and cyclic configurations of all four models. A large gap has also taken form between the branches in both configurations (finite and cyclic) of the HLM, FBM, and 2DSM models. The gap is due to the characteristics inherent to the structure of these models that keeps this forbidden frequency range almost constant for every configuration. This large gap can also be attributed to the taking into account of the individual masses of the SP backbones in the HLM, FBM, and 2DSM models, causing varying vibrational behaviors. In both the finite and cyclic configurations, there are differences in the dispersion curves of the perfect and imperfect systems. In the imperfect models (models with dislocations), some extra bands are observed within the forbidden frequency range. These differences are less pronounced in the 2DSM model, primarily due to the larger number of masses the model contains.
As seen in Fig. 5(a-l) and Fig. 6(a-l), increasing the number of building blocks in each model alters the fluctuations in the DOS diagrams. Additionally, by increasing the number of the random building blocks (\(N_{bp}/N_{b}\)) along the length of each finite and cyclic model, the fluctuations in the DOS curves also increases, which is accompanied by a decrease in the fluctuations in these curves. An increase in the number of building blocks increases the likelihood of the occurrence of repetitive blocks, thereby augmenting the degeneracy of the systems. Comparing the dispersion and DOS curves of the perfect and imperfect systems shows that the presence of random dislocations results in the appearance of extra bands in the dispersion curves, and these manifest themselves as peaks in the DOS curves. These differences are greater at low frequencies and by increasing the number of building blocks, the differences in the perfect and imperfect DOS curves decrease. As seen in Figs. 3-6, there is a degree of dissimilarity among the dispersion and DOS diagrams of all four models in all of their states (different lengths and random distribution of building blocks). However, despite the overall differences in the appearance of the bands for various random configurations, the boundaries of the band gaps remain almost unaffected. By gradually increasing the lengths of the finite and cyclic cases, the discrepancy between the DOS diagrams of the perfect and imperfect systems starts to decline. It can be said that the effect of dislocation has decreased, but it has not disappeared completely. These incremental size enhancements increase the accuracy of the models, making them better tools for the approximation of the mechanical properties of actual RNA and DNA molecules. So, considering a medium-sized model containing a fixed number of randomly-shuffled building blocks would be a logical first step to not only obtain a better estimation of the vibrational spectrum of very long systems, but also to save computation time. Following this approach, \(N_{bp}(N_{b})=128\) blocks were considered for both configurations (finite cyclic) of each model, making the number of subsites of the HLM, FBM, 1DSM, and 2DSM models equal to \(N_{s}=256\), \(N_{s}=384\), \(N_{s}=256\), \(N_{s}=512\), respectively. What all the systems have in common in their responses to the above-mentioned stimuli is that they behave in a nonlinear fashion. To demonstrate this, one needs only consider how a system responds to a simple base-pair increase. For instance, the DOS of the HLM model with 64 base-pairs does not appear to have undergone a linear change as compare to the model with a 32 base-pair ensemble.
The DOS diagrams of the HLM, FBM, 1DSM, and 2DSM models with finite configuration. In each panel, the solid line is the DOS curve of the perfect system and the dashed line relates to the imperfect systems.
The DOS diagrams of the HLM, FBM, 1DSM, and 2DSM models with cyclic configuration. In each panel, the solid line is the DOS curve of the perfect system and the dashed line relates to the imperfect systems.
The finite-length 1DSM (32 base pairs) model was reanalyzed assuming an invariant linear spring stiffness throughout the perfect and dislocated systems. The results indicate that in the perfect configuration, the lower frequencies are seldom affected by the invariable spring stiffnesses, whereas in the imperfect (dislocated) system, the peaks of the low-frequency part of the DOS diagram show a noticeable level of incongruence. In the high-frequency region, however, both the probability of the occurrence of the frequencies (heights) and the frequencies themselves (locations) have experienced a shift. These discrepancies means that the constant base pair stiffnesses - which is an unrealistic modeling approach - introduces a significant level of inaccuracy into the results. These observations and results can be seen in Fig. 7. To ensure that the results remain relatively stable over multiple reruns (realizations), we analyzed the finite-length 1DSM model six more times - three analyses for each of the perfect and imperfect systems. The DOS curves for the perfect and imperfect cases, along with their average curves, can be seen in Fig. 8. As it can be observed, the overall shapes of the average curves align suitably with the other DOS curves in both systems, confirming that the results still remain consistent over multiple iterations.
The DOS diagrams of the perfect (a) and imperfect (b) finite 1DSM model for two different cases of vertical springs and \(N_{bp}=32\).
Comparison between the average and single-run DOS curves of the perfect (a) and imperfect (b) finite 1DSM model for \(N_{bp}=32\).
Conclusion
By using a harmonic Hamiltonian and the Green’s functions approach, the effects of random dislocations on the vibrational spectra of RNA and DNA strands have been investigated for two different configurations (finite and cyclic). The building blocks have been randomly distributed along the length of these systems. The sequence-dependent nature of DNA and RNA, along with their inherent randomness, can significantly influence and alter the mechanical response of these structures. This is because the masses of the bases and the elastic forces between each pair vary along the length of these systems. Such characteristics can affect the potential energy, thereby changing their vibrational behavior. The models we studied are simplified elastic representations of DNA and RNA molecules, designed without considering their twisted structures. Due to the intrinsic variability in the base pairs/building blocks along their lengths, native DNA and RNA are translationally variant. Consequently, our models are treated as quasi-1D systems with a specific number of randomly placed base pairs/building blocks. We used these mechanical models to understand macromolecular motions in the presence of random dislocations. Our primary goal is to present a methodology for investigating the mechanical response of such macromolecules from a physical perspective. As far as we know, such an analytical quantum mechanical approach is largely missing in DNA and RNA research, which is where our study finds its significance. The methodology we introduce could also be applied to other biopolymers with similar structures. Our analyses are based on two main propositions: 1) Developing an analytical model to predict the vibrational response of DNA- and RNA-like structures, and 2) increasing the number of blocks in the systems to achieve a more accurate representation of these macromolecules. Based on obtained the results, a series of dispersion curves and DOS diagrams have been derived. Each configuration was investigated using four models (the HLM model of the RNA and the FBM, 1DSM, and 2DSM models of the DNA). Using an appropriate set of parameters for the models, it was observed that the influence of random dislocations decreases as the number of random building blocks along the length of the finite/cyclic system increases, but the effect does not disappear completely. It was also observed that the influence of the dislocations becomes more amplified at low frequencies. Besides, it was found that due to a large number of masses in the structure of the 2DSM model, the effect of the dislocation becomes less pronounced in this model. Also, taking the mass of the SP backbone into account has created a large gap and a wide forbidden frequency range in the DOS and dispersion curves of both configurations of each model. These gaps change by the introduction of the dislocation into the structure, but they remain unaltered by changes in the size of the building blocks in both configurations. The same large gap does not appear in the dispersion curves of the 1DSM model, whose dispersion curves show a high degree of degeneracy. An interesting observation is that the systems do not exhibit a linear behavior in response to different changes applied to their structures, as evidenced by the dispersion and DOS curves. A reanalysis of a 32-base pair model showed that assuming a constant spring stiffness throughout the system leads to inaccuracies. In the perfect system, low frequencies are not affected, but in the dislocated system, low frequencies are affected. In the high-frequency region, both the probability and frequency values are shifted, indicating that the constant stiffness assumption is unrealistic and introduces significant errors. The finite-length 1DSM model was rerun six times to check the stability of results, and the analysis of DOS curves for perfect and imperfect systems showed that the average curves align well with other DOS curves, confirming consistent results over multiple iterations. Overall, the results obtained herein can be useful in understanding the mechanical properties of similar macro-structures. The employed methodology can also be used to study damaged RNA and DNA.
Data availability statement
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Langridge, R. & Gomatos, P. J. The Structure of RNA: Reovirus RNA and transfer RNA have similar three-dimensional structures. Science 141, 694 (1963).
Naito, S. & Ishihama, A. Function and structure of RNA polymerase from vesicular stomatitis virus. Journal of Biological Chemistry 251, 4307 (1976).
Lee, D. N. & Landick, R. Structure of RNA and DNA chains in paused transcription complexes containing Escherichia coli RNA polymerase. Journal of molecular biology 228, 759 (1992).
Haurwitz, R. E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J. A. Sequence-and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355 (2010).
Watson, J. D. & Crick, F. H. Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171, 737 (1953).
Chhabra, R., Sharma, J., Liu, Y., Rinker, S. & Yan, H. DNA self-assembly for nanomedicine. Advanced Drug Delivery Reviews 62, 617 (2010).
Sharma, J. et al. Control of self-assembly of DNA tubules through integration of gold nanoparticles. Science 323, 112 (2009).
Van Steeg, H. & Kraemer, K. H. Xeroderma pigmentosum and the role of UV-induced DNA damage in skin cancer. Molecular Medicine Today 5, 86–94 (1999).
Cadet, J. & Douki, T. Formation of UV-induced DNA damage contributing to skin cancer development. Photochemical and Photobiological Sciences 17, 1816 (2018).
He, L. G. & Hannon, J. MicroRNAs: small RNAs with a big role in gene regulation. Nature Reviews Genetics 5, 522 (2004).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nature Reviews Cancer 6, 857 (2006).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nature Reviews Genetics 11, 597 (2010).
Lodish, H. et al. Molecular cell biology, 5nd edi (Freeman, New York, 2004).
Debouck, C. & Goodfellow, P. N. DNA microarrays in drug discovery and development. Nature Genetics 21, 48 (1999).
Yoo, S. M., Choi, J. H., Lee, S. Y. & Yoo, N. C. Applications of DNA microarray in disease diagnostics. Journal of Microbiology and Biotechnology 19, 635–638 (2009).
Petrosyan, K. G. & Hu, C. K. Fluctuation effects in gene regulation by microRNAs and correlations between gene and pseudogene mRNAs in the control of cancer. Journal of Statistical Mechanics: Theory and Experiment 2015, 07019 (2015).
Baeumner, A. J., Cohen, R. N. & Miksic, V. RNA biosensor for the rapid detection of viable Escherichia coli in drinking water. Biosensors and Bioelectronics 18, 405 (2003).
Vercoutere, W. and Akeson, Biosensors for DNA sequence detection. Current Opinion in Chemical Biology, 6, 816-822 (2002)
McCauley, T. G., Hamaguchi, N. & Stanton, M. Aptamer-based biosensor arrays for detection and quantification of biological macromolecules. Analytical biochemistry 319, 244 (2003).
Oliveira, J. I. N., Albuquerque, E. L., Fulco, U. L., Mauriz, P. W., Sarmento, R. G., Caetano, E. W. S., and Freire, V. N. Conductance of single microRNAs chains related to the autism spectrum disorder. EPL (Europhysics Letters), 107, 68006 (2014)
Santhanam, M., Algov, I. & Alfonta, L. DNA/RNA electrochemical biosensing devices a future replacement of PCR methods for a fast epidemic containment. Sensors 20, 4648 (2020).
Kelley, S. O. & Barton, J. K. Electron transfer between bases in double helical DNA. Science 283, 375 (1999).
Cuniberti, G., Craco, L., Porath, D. & Dekker, C. Backbone-induced semiconducting behavior in short DNA wires. Physical Review B 65, 241314 (2002).
Kubar, T., Woiczikowski, P. B., Cuniberti, G. & Elstner, M. Efficient calculation of charge-transfer matrix elements for hole transfer in DNA. The Journal of Physical Chemistry B 112, 7937 (2008).
Paez, C. J., Schulz, P. A., Wilson, N. R. & Römer, R. A. Robust signatures in the current-voltage characteristics of DNA molecules oriented between two graphene nanoribbon electrodes. New Journal of Physics 14, 093049 (2012).
Klotsa, D., Römer, R. R. & Turner, M. S. Electronic Transport in DNA. Biophysical Journal 89, 2187 (2005).
Mousavi, H., Khodadadi, J. & Grabowski, M. Electronic properties of long DNA nanowires in dry and wet conditions. Solid State Communications 222, 42 (2015).
Mousavi, H. & Grabowski, M. Nonlinear electron transport across short DNA segment between graphene leads. Solid State Communications 279, 30–33 (2018).
Mousavi, H., Jalilvand, S., Sani, S. S., Hartman, J. A. L. & Grabowski, M. Electronic properties of different configurations of double-strand DNA-like nanowires. Solid State Communications 319, 113974 (2020).
Lambropoulos, K. & Simserides, C. Tight-binding modeling of nucleic acid sequences: Interplay between various types of order or disorder and charge transport. Symmetry 11, 968 (2019).
Jiménez, M. G. et al. Observation of coherent delocalized phonon-like modes in DNA under physiological conditions. Nature Communications 7, 11799 (2016).
Reiss, C. Nonlinear Excitations in Biomolecules, edited by M (Peyrard, Les Editions de Physique, Springer, Berlin, 1994).
Ortiz, V. & de Pablo, J. J. Molecular origins of DNA flexibility: sequence effects on conformational and mechanical properties. Physical Review Letters 106, 238107 (2011).
Dauxois, T. & Peyrard, M. Entropy-driven DNA denaturation. Physical Review E 47, R44 (1993).
Zoli, M. Non-linear Hamiltonian models for DNA. European biophysics journal 51, 431 (2022).
Poland, D., Scheraga, and H. Phase Transitions in One Dimension and the Helix?Coil Transition in Polyamino Acids. Journal of Chemical Physics, 45, 1456 (1966)
Srivastava, S. & Singh, N. The probability analysis of opening of DNA. Journal of Chemical Physics 134, 115102 (2011).
Sulaiman, A., Zen, F. P., Alatas, H. & Handoko, L. T. The thermal denaturation of the Peyrard?Bishop model with an external potential. Physica Scripta 86, 015802 (2012).
Padinhateeri, R., and Menon, G. I. Stretching and Bending Fluctuations of Short DNA Molecules. Physical Biology, 104, 463 (2013) (2013)
Apostolaki, A. & Kalosakas, G. Targets of DNA-binding proteins in bacterial promoter regions present enhanced probabilities for spontaneous thermal openings. Biophysical Journal 104, 463 (2013).
Joshi, H., Dwaraknath, A. & Maiti, P. K. Structure, stability and elasticity of DNA nanotubes. Physical Chemistry Chemical Physics 17, 1424 (2015).
Zoli, M. Helix untwisting and bubble formation in circular DNA. Journal of Chemical Physics 138, 205103 (2013).
Zoli, M. Twisting and bending stress in DNA minicircles. Soft Matter 10, 4304 (2014).
Zoli, M. Flexibility of short DNA helices under mechanical stretching. Physical Chemistry Chemical Physics 18, 17666 (2016).
Barbi, M., Cocco, S., Peyrard, M. & Ruffo, S. A twist opening model for DNA. Journal of Biological Physics 24, 97 (1999).
Powell, J. W., Edwards, G. S., Genzel, L., Kremer, F., and Wittlin. A. Investigation of far-infrared vibrational modes in polynucleotides. Physical Review A, 35, 3929 (1987)
Van Zandt, L. L. & Saxena, V. K. Millimeter-microwave spectrum of DNA: Six predictions for spectroscopy. Physical Review A 39, 2672 (1989).
Wittlin, A., Genzel, L., Kremer, F., Häseler, S. & Poglitsch, A. Far-infrared spectroscopy on oriented films of dry and hydrated DNA. Physical Review A 34(493), 34 (1986).
Páez, C. J., Rey-González, R. & Schulz, P. A. Macroscopic localization lengths of vibrational normal modes in a heuristic DNA model. Physical Review B 81, 024203 (2010).
Li, J. J. & Zhu, K. D. Coherent optical spectroscopy in a biological semiconductor quantum dot-DNA hybrid system. Nanoscale Research Letters 7, 133 (2012).
Sales, M. O., Lyra, M. L., de Moura, F. A. B. F., Fulco, U. L. & Albuquerque, E. L. Sub-diffusive electronic transport in a DNA single-strand chain with electron-phonon coupling. Journal of Physics: Condensed Matter 27, 035104 (2015).
Jalilvand, S., Mousavi, H., and Irani M. Metal-semiconductor transition in \(\Psi\)-graphene. “Materials Today Communications, 33, 104833 (2022)
Jalilvand, S. & Mousavi, H. Multi-band tight-binding model of \(MoS_{2}\) monolayer. Journal of Electronic Materials 49, 3599 (2020).
Mousavi, H. & Bagheri, M. Controlling the bandgap of boron nitride nanotubes with carbon doping. Journal of Electronic Materials 44, 2693 (2015).
Mousavi, H. Sublattice superconductivity in boron nitride nanotube. Journal of superconductivity and novel magnetism 26, 2905 (2013).
Jalilvand, S. & Mousavi, H. Vibration spectra of DNA and RNA segments. European Biophysics Journal 53, 95 (2024).
Mousavi, H., Mirzaei, M., Jalilvand, S. & Sani, S. S. Vibrational properties of DNA in different models. Mechanics of Advanced Materials and Structures 29, 3950 (2022).
Mousavi, H., Mirzaei, M. & Jalilvand, S. Mechanical response of double-stranded DNA to dynamic excitation. Journal of Vibration and Control 29, 214 (2023).
Bruus, H. & Flensberg, K. Many-Body Quantum Theory in Condensed Matter Physics: An Introduction 2nd edn. (University Press, UK, 2004).
Sandonas, L. M., Gutierrez, R., Pecchia, A., Croy, A. & Cuniberti, G. Quantum phonon transport in nanomaterials: Combining atomistic with non-equilibrium green’s function techniques. Entropy 21, 735 (2019).
Peyrard, M. Melting the double helix. Nature Physics 2, 13–14 (2006).
Powell, J. W., Edwards, G. S., Genzel, L., Kremer, F. & Wittlin, A. Investigation of far-infrared vibrational modes in polynucleotides. Physical Review A 35, 3929 (1987).
Gorelik, V. S., Krylov, A. S. & Sverbil, V. P. Local Raman spectroscopy of DNA. Bulletin of the Lebedev Physics Institute 41, 310 (2014).
Jo, S. et al. Vibrational characteristics of DNA nanostructures obtained through a mass-weighted chemical elastic network model. RSC Advances 7, 47190 (2017).
Chou, K. C. Low-frequency vibrations of DNA molecules. Biochemical Journal 221, 27 (1984).
Funding
This study is in no way funded by any organization and is an independent investigation carried out exclusively by the authors of the study.
Author information
Authors and Affiliations
Contributions
All three authors have contributed, in equal measure, to the study’s conception and findings. Material preparation, data collection and analysis were performed by Samira Jalilvand, Moein Mirzaei and Hamze Mousavi. The first draft of the manuscript was written by Samira Jalilvand and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. Formulation and formal analysis; coding; results interpretation and writing the original draft were performed by Samira Jalilvand. Formulation and formal analysis; coding; results interpretation; and editing were performed by Moein Mirzaei. Formulation and formal analysis; results interpretation were performed by Hamze Mousavi.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jalilvand, S., Mirzaei, M. & Mousavi, H. Nonlinear mechanical response of finite-length soft composites with random dislocations. Sci Rep 14, 27121 (2024). https://doi.org/10.1038/s41598-024-78623-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-78623-9