Abstract
Moringa is the sole genus in the family Moringaceae used for medicinal and nutrient purposes. Morphological features, phytochemical attributes, and molecular characterization were used for the genetic association and classification among Moringa oleifera, M. peregrina, and M. stenopetala. Moringa peregrina recorded a similarity of 84% lonely and placed M. stenopetala with M. oleifera into a cluster score with a similarity of 95.3%. M. peregrina is characterized by phenolic content (243 mg/100 g), flavonoids (7 mg/100 g), and antioxidant activity (1226.85 mg/100 g). GC-MS analysis revealed that M. oleifera contained twenty compounds with 2-decenal (E) (39.14%), 2-undecenal (15.51%), nonanal (3.60%), and 2-octenal, (E) (2.48%), while M. peregrina identified eighteen compounds with 2-decenal (Z) (25.42%), 2-docecen-1-al (9.35%), and 13-Docosenoic acid, methyl ester, (Z) (4.16%). M. stenopetala identified fifteen compounds containing 2-decenal (E) (26.67%), 2-undecenal (24.10%), and nonanal (4.40%). A broad sense of similarity has been scored between M. oleifera and M. stenopetala by the phytochemical compositions, especially the similarity in the main compounds such as 2-decenal (E), 2-undecenal, and nonanal. It can be concluded that efforts need to be expanded to pay attention to study Moringa taxa, due to the rarity of Moringa peregrina, and the focus should be on sustainable utilization and conservation. The potential of these taxa would greatly benefit indigenous species in terms of their maintenance, and there is a need for more comprehensive bio-prospecting studies. Therefore, this study evaluates the variability among Moringa and highlights the significance of leaf and seed ultrastructure to provide more information and evaluate potential approaches.
Similar content being viewed by others
Introduction
Moringa is the sole genus in the family Moringaceae. It contains 13 species, which are distributed in Africa, Arabia, Southeast Asia, and South America1,2 and cultivated in all tropical and subtropical areas3,4,5. The M. oleifera Lam. and M. peregrina (Forssk.) Fiori are the most economical species in agricultural and medicinal uses6,7. Moringa is one of the groups producing mustard oil (glucosinolates), i.e., Brassicaceae and Caricaceae8. Generally, plants in the Moringa genus are used for various applications and utilizations, including animal feed, human consumption, medicinal use, fertilization, antimicrobial agents, biofuel and biogas production, purification of water, foliar nutrients, green manures, and ornamental purposes9,10,11,12,13,14,15. M. oleifera and M. peregrina are widely cultivated and studied species11, in contrast, other species have not been studied in detail, and their potential applications and uses have not been given little research and development16. Most research in Moringa is concentrated mostly on medical applications17,18, anatomical identification of plants19, and anti-viral activity20. Many authors have used the morphological and anatomical characteristics of plants to identify them21,22,23. Plant breeding efforts have been based on taxonomic identification to attribute diagnostic features to particular or varietal parentage24. M. oleifera, “the drum stick,” is the most well-known and investigated, particularly of the 13 species of genus Moringa used in traditional medicine due to the phytochemicals found in its different parts25. M. stenopetala, “cabbage tree,” is important as it is a major drought-resistant vegetable plant with medicinal and nutritional properties similar to or surpassing those found in M. oleifera. However, little research and development has been done on this species15,26,27. M. peregrina “the Arabian tree of Moringa” is one of the most economically important and valuable medicinal plants in the Egyptian desert. It is threatened by vulnerable habitat, over-grazing as fuel and seed collection, emphasizing the need for habitat protection and conservation15,28.
The Moringa is known for its quick growth rate, reaching up to 10–12 m in height, and adapts well to humid and dry climates. When ripe, it is identified by its pinnate leaves and long, woody pods because it opens into three valves containing globular seeds. The nutritional and medical benefits of Moringa, as well as its usage as a decorative plant and for animal feed, have drawn the attention of farmers, scientists, and researchers7,29. Moringa has bisexual flowers that are highly pollinated, and pollination is mainly facilitated by animals such as bees and sunbirds. The seeds are also strongly winged, which may allow them to be spread short distances from the parent tree by wind, helping pollination30. The Moringa oleifera plant grows quickly; it can reach a height of 12 m at the crown and, depending on the cultivar, begins to produce its first fruit 6 to 12 months after planting. In many regions worldwide, trees can flower and bear fruit in one year, and multiple seed harvests are possible. Fruits (pods) ripen about three months after flowering and have to be collected promptly. Each pod contains approximately 26 seeds with a diameter of 1 cm7,31.
Natural products extracted from medicinal plants offer the potential for the discovery of new drugs because of their chemical diversity and wide availability. The medical usefulness of these plants results from the bioactive phytochemicals that have specific physiological functions in humans. Alkaloids, flavonoids, tannins, terpenoids, saponins, and phenolic compounds are well-documented as major secondary metabolites32,33 which provide a broad spectrum of biological activities like antioxidant, anticancer, antiviral, anti-inflammatory, antiviral, and antimicrobial properties. Contextually, the chemical diversity of these secondary metabolites and their unique chemical structures also had a major impact on biological activities. It is worth noting that the biological activities of plant extracts may be due to a single compound or to the participation of several compounds together in what is known as synergistic activity. In addition, they play an important role in plant’s defense mechanism to mitigate harmful environmental effects. Moreover, by oxidation of fatty acids these phenolic compounds prevent detrimental changes in living organisms34,35. M. oleifera is one of the traditional medicinal plants, popularly called drumstick tree or horseradish tree, has been widely researched for medicinal uses of bioactive compounds isolated from the plant. All plant parts are used for either human and livestock uses. Moringa leaves are consumed as food as well as nutritional supplements. In spite of their beneficial uses, studies on these bioactive compounds of Moringa are limited in literature.
Antioxidants are of crucial importance in preserving food by avoiding oxidative damage and deterioration in food quality36. Prior studies conducted on M. peregrina seeds concentrated on the composition of fatty acid and fixed oil content37. However, published studies on the essential oil content of M. peregrina seeds are limited. Hence, the current study aims at investigating the chemical constituents and antioxidant activity of oil extracted from M. peregrina seeds. This study reports antioxidant properties of Moringa seeds using hexane and ethanolic extracts.
Morphological and genetic diversity assessments are important for plant improvement programs. The majority of studies evaluated the genetic diversity of M. oleifera using various techniques to assess inter-species variations38,39. However, studies on local genotypes of M. peregrina are limited. Variations in M. peregrina were reported for plants grown in the South Sinai of Egypt40 and Western Saudi Arabia41.
Multi-locus DNA markers, such as inter-simple sequence repeat (ISSR) and start codon target (SCoT), are simple and common PCR-based techniques and are largely used for determining genetic diversity42,43,44. Start codon targeted polymorphism (SCoT) is a PCR-based technique depending on the short-conserved region of plant genes surrounding the ATG translation start codon45,46 using single primers. Hence, this SCoT technique is similar to RAPD or ISSR because a single primer is used in all these techniques47,48.
Understanding the genetic diversity of a species in a district is essential when developing management strategies for conservation and improvement activities. Estimation of Moringa diversity estimation is significant in clarifying the relationships between individuals within and among the different populations, which impacts conservation management greatly. The general objective of this study is to evaluate the variability among three Moringa species: M. oleifera, M. peregrina, and M. stenopetala. The leaves and seeds of Moringa sp. were investigated using morphological characterization, SEM (scanning electron microscopy) examination, and numerical analysis of the studied characteristics. This study highlighted the significance of the leaf and seed ultrastructure in exploring species variability. This study highlights the different approaches, viz., micro- and macro-morphological, chemical, and molecular attributes in discrimination and valuation to determine the genetic relationship and classification among species. The attributes generated and provided more information on the association of Moringa species. The current study aims to classify the investigated three species of Moringa. This study documents morphological and genetic differences among the studied Moringa species.
Materials and methods
Plant materials
Samples of the studied Moringa species were collected from a Moringa collection by Dr. Aboelfetoh M. Abdalla, a botanical garden in Belbes, Al-Sharkia Governorate of Egypt that is a genetic repository for Moringa species in Egypt. Collections were submitted to the procedures and under the authority of the National Gene Bank (NGB), Agricultural Research Center (ARC), Giza, Egypt. The NGB Herbarium conducted the formal identification and referral of plant materials as presented in Table 1. All taxa are deposited in the National Gene Bank (NGB), Agricultural Research Center (ARC), 9 Gamaa St., 12619, Giza, Egypt. All samples were taxonomically checked to validate the identification methods suggested by Bailey49. An additional identification check was conducted using the plant materials from the Cairo University Herbarium and The Agriculture Museum in Giza, Egypt. All samples were submitted to the Herbarium at Al-Azhar University on November 17, 2022 and the Herbarium of NGB (Table 1). This study compiles information on species, their locations, the number of individuals collected, their distribution, conservation status, and evaluation.
Thirty-one morphological traits of the leaf and seed were scored and coded to build a numerical data matrix. Statistical analysis with PRIMER (Software, Version 6.0; https://www.primer-e.com) was used to compare the investigated taxa. Five individuals were used to represent each species in estimating the mean and standard error (SE) for quantitative traits and used in the numerical analysis of traits. All the experiments were performed following the relevant guidelines and legislations of the National Gene Bank (NGB).
Phytochemical analysis
Phytochemical constituents
The protocol devised by Nagata and Yamashita50 was followed to determine chlorophyll a, b, and carotenoid contents. The leaf samples (0.2 g) of Moringa were ground in acetone (80%) and then filtered using Whatman glass microfiber filters (GF/F filter discs: 0.7 μm pore size).
The absorbance of the extract was recorded at wave lengths of 663, 645, and 446 nm by using a UV-spectrophotometer.
Chlorophyll a and b, as well as carotenoid contents, were calculated using the following equations (I-III):
-
I.
Chlorophyll a = 0.999 × A663–0.0989 × A645.
-
II.
Chlorophyll b = -0.328 × A663 + 1.77 × A645.
-
III.
Carotenoid = (0.216 × A663–1.22 × A645) – (0.304 × A505 + 0.452 × A453).
Where, A663, A645, A505, and A453 are the readings at wavelengths of 663, 645, 505, and 446 nm, respectively50.
Two grams of green leaves were finely ground, after that the dry powder was defatted using petroleum ether (60–80 °C), subsequently the residue was extracted with 20 ml methanol (80%). Then, the extract was filtered. All measurements were conducted using an ultraviolet-visible spectrophotometer-MA9523-SPEKOL 211 (ISKRA, Slovenia). The total phenolic content (TPC) of the extract was determined following Singleton et al.51. Woisky and Salation52 method was used to estimate the total flavonoid content (TFC). The total antioxidant capacity (TAC) of the extracts was determined according to Prieto et al.53, whereas ascorbic acid was estimated using the procedure of Klein and Perry54. The absorbances were measured at 765 nm, 415 nm, and 695 nm for TPC, TFC, and TAC, respectively.
GC-MS analysis
Extraction
Ten grams (10 g) from each seed dry powder were extracted separately with 150 ml n-hexane at room temperature with shaking daily, followed by filtration and extraction again four times. Then, each extract was filtered using Whatman filter paper No.1 and concentrated by using a rotatory evaporator (Buchi, Switzerland) at (40ºC). The obtained extracts were collected and stored at room temperature in the dark for further processing.
Analysis
Gas chromatography-mass spectrometry (GC-MS) analysis was conducted using a Thermo Scientific, Trace GC Ultra/ISQ Single Quadrupole MS instrument coupled with TG-5MS fused silica capillary column (30 m, 0.251 mm, 0.1 mm film thickness). A 70 eV electron ionization system was used, with Helium gas serving as the carrier gas at a flow rate of 1 ml/min for the GC-MS analysis. The temperature for both MS transfer line and injector was adjusted to 280 °C. The oven temperature was set to 50 °C initially, with a hold time of 2 min. It then increased to 150 °C at a rate of 7 °C per minute, then to 270 °C at a rate of 5 °C per minute, with a hold time of 2 min. Finally, it reached 310 °C as the final temperature, with a hold time of 10 min, and increased at a rate of 3.5 °C per minute. The investigation focused on quantifying all the detected components by calculating the percentage relative peak area. The compounds were tentatively identified by comparing their retention time and mass spectra with the NIST and WIL-LY library data acquired from the GC-MS instrument55,56,57,58.
Molecular analysis
Start codon target (SCoT) markers
Ten fresh leaflets for each taxon were collected in silica gel bags (5:1 silica gel: tissue) for the step of DNA extraction. The samples were powdered using immersion in liquid nitrogen and then crushed via a sterile mortar and pestle. The isolation was done using the DNeasy plant mini kit (Biobasic) and stored at − 80 °C for subsequent steps. The quality was checked using a Nanodrop 8000 Spectrophotometer (Thermo Scientific, Wilmington, USA). The DNA extraction was done according to Mahdy et al.59.
Fifteen SCoT primers (Table 2) were used according to the design of Collard and Mackill60 and procured from Biobasic Com. PCR amplification was performed using Techni TC-512 Thermal Cycler. The temperature and time were programmed as one cycle at 94 °C for 4 min, then 40 cycles for one minute at 94 °C, 1 min for annealing at 57 °C and two minutes at 72 °C, and finally, 72 °C for 10 min. The PCR products were stored at 4 °C for further analysis. The products were screened on a 1.2% agarose gel staining with ethidium bromide. A ladder marker of 100 bp was used. The run was conducted for 30 min at 100 V in mini-submarine gel BioRad. Gels were photographed by a transilluminator and analyzed by Bio-Rad video Gel documentation 2000. Profiles were recorded as 1 if present or 0 if absent on standard marker via Alpha Ease FCTM (version 4.9.1) software.
Banding profiles were recorded as one if present or zero if absent based on standard marker using Alpha Ease FCTM (version 4.0.1) software. Genetic similarity was determined using the Jaccard coefficient61. Algorithms of the un-weighted pair group method with arithmetic (UPGMA) averages are used to build trees62 to study the relationship among populations.
Statistical analysis
A randomized complete block design (RCBD) was used for the experimental design. Data were analyzed using the factorial method of Sendecor and Cochran63 where the means were compared using the Least Significant Difference (LSD). The DNA bands produced by each primer were counted, and the molecular sizes of these bands were compared with those of the ladder DNA markers. The bands scored from DNA banding patterns generated by the primers were pooled together. Then, the presence or absence of each DNA band was recorded as a binary character in a data matrix (coded 1 (present) and 0 (absent) to calculate genetic similarity. Calculations were done using Dice similarity coefficients64 by using the SPSS software (v. 10). The genetic similarity matrix was then used to construct a dendrogram tree among the studied Moringa species using PRIMER software (www.primer-e.com).
Results and discussion
Morphological traits of leaves, seeds, chemical features, and molecular markers were examined to assess the taxonomic relationship between the three species of the genus Moringa under study: M. peregrina, M. oleifera, and M. stenopetala. The numerical analysis method utilizing these attributes was also applied to assess how similar or unlike these species are to one another.
Leaf and seed morphology
The growth forms perennial shrubs; the stem is erect, much branched, and cylindrical in the studied species. The leaflet in the studied species was examined concerning shape, type, texture, apex, and the number of leaflets (Table 3; Fig. 1). The leaflet is sub sessile in the studied species; the petiole surface is sparsely hairy in M. peregrina and pubescent in M. oleifera and M. stenopetala. The leaves of the studied species are compound and recorded in two types: pinnate in M. peregrina and imparipinnate in M. oleifera and M. stenopetala. The leaflets of all species are entire, and for blade outline (leaflet shape), two main types were recorded: oblong, found in M. peregrina, and elliptic-obovate, found in M. oleifera and M. stenopetala. Leaflet apices ranged from acute in Moringa peregrina, to emarginate in M. oleifera. We also observed an obtuse-emarginated apex in M. stenopetala. Leaflets are glabrous in the upper leaf in the studied species. In contrast, they are in the lower leaf sparsely hairy in M. peregrina and pubescent in M. oleifera and M. stenopetala. The leaflet length is (1.7–2.1 cm) in M. peregrina, (1.5–2.3 cm) in M. oleifera and M. stenopetala. The leaflet width is (0.4–0.5 cm) in M. peregrina, and (1–1.5 cm) in M. oleifera and M. stenopetala. Seed shape (Fig. 2) is ovoid-trigonous in M. peregrina, globose or sub-globose with three papery wings in M. oleifera, and elliptical in M. stenopetala. Seed color is brown in M. peregrina and M. oleifera and creamy white in M. stenopetala. Seed surface sculpture is reticulate and anticlinal wall shape straight in studied species. Seed length varied among the studied species; 1.7–2.1 cm in M. peregrine, 2.5–2.7 cm in M. oleifera, and 2.3–2.7 cm in M. stenopetala. Seed width varied among the studied species; 1.2–1.5 cm in M. peregrina, 2–2.6 cm in M. oleifera, and 1.2–1.5 cm in M. stenopetala. The anticlinal wall shape of the seed is straight or undulated in M. peregrina and straight in M. oleifera and Moringa stenopetala, anticlinal walls raised in M. peregrine, M. oleifera, and M. stenopetala, outer periclinal walls convex in M. peregrina concave in M. oleifera and M. stenopetala, inner periclinal walls concave in M. peregrine, M. oleifera, and M. stenopetala. Epidermal cell wall straight to undulate, stomata leveling depressed and stomata type anomocytic in M. peregrine, M. oleifera, and M. stenopetala (Table 3; Fig. 3).
Our results indicated that the growth forms perennial shrubs; the stem is erect, much-branched, and cylindrical in the studied species. The leaflet in the studied species was examined in terms of shape, type, texture, apex, and number of leaflets. The leaflet is subsessile in the studied species; the petiole surface is sparsely hairy in M. peregrina and pubescent in M. oleifera and M. stenopetala. The leaves of the studied species are compound and recorded in two types: pinnate in M. peregrina and imparipinnate in M. oleifera and M. stenopetala. The leaflets of all species are entire and for blade outline (leaflet shape), oblong, in M. peregrina and elliptic-obovate in M. oleifera and M. stenopetala. Leaflet apices ranged from acute in M. peregrina, to emarginate in M. oleifera, and obtuse-emarginated apex in M. stenopetala. Leaflets are glabrous in the upper leaf whereas in the lower leaf sparsely hairy in M. peregrina and pubescent in M. oleifera and M. stenopetala. These were found to be important diagnostic characteristics as they conformed with the results obtained by Azza65, who reported that M. oleifera leaflet shape is obocordate, with emarginate apex, symmetric base, leaflet upper surface hairy, colliculate sculpture of leaflet upper surface, tuberculate-reticulate on lower one, also Rangnath et al.66 reported that M. oleifera leaves generally tripinnate, pinnate and pinnules opposite, evanescent; circulars − 2 cm long and 0.6–1 cm wide. The side elliptic, the terminal obviates. M. stenopetala leaves imparipinnate. The shape is the elliptic, obtuse apex, symmetric base. The leaflet’s upper surface is a hairy, lower smooth, rugose sculpture of the leaflet surface upper, reticulate-verrucate on the lower one. M. peregrina leaves are pinnate, leaflet shape is linear, acute apex, symmetric base, hairy leaflet upper surface (non-glandular, glandular), smooth lower surface, anomocytic stomata on the upper epidermis at superficial level, unclear on lower one, the rugose-tuberculate sculpture of upper leaflet surface, and rugose-tuberculate on lower one. Also, Zhigila67 observed that the leaves in M. oleifera varieties are alternate, composite, bipinnate, or tripinnate, with 2 to 6 pairs of opposite pinnae bearing opposite leaflets in 3 to 7 pairs. It also has a broader terminal leaflet, green leaf lamina color, and an entire leaflet margin.
The seed shape is ovoid-trigonous in M. peregrina, globose or sub-globose with three papery wings in M. oleifera, and elliptical in M. stenopetala. The seed color is brown in M. peregrina and M. oleifera and creamy white in M. stenopetala. Seed surface sculpture is reticulate and anticlinal wall shape straight in the studied species. The anticlinal wall shape of the seed is straight or undulated in M. peregrina and straight in M. oleifera and M. stenopetala, anticlinal walls raised in M. peregrine, M. oleifera, and M. stenopetala, outer periclinal walls convex in M. peregrina concave in M. oleifera and the M. stenopetala, inner periclinal walls concave in M. peregrine, M. oleifera and M. stenopetala. Epidermal cell wall straight to undulate, stomata leveling depressed and stomata type anomocytic in M. peregrine, M. oleifera, and M. stenopetala. These results are in agreement with Azza65 who characterized M. oleifera as seeds are brown, round with tan edges, rough texture, reticulate epidermal cell walls, anticlinal walls raised (4–6 gonal)-straight, and concave outer periclinal walls. M. stenopetala seeds are brown with rough texture, reticulate-foveate epidermal cell walls, raised anticlinal walls (5–6 gonal)-straight, and concave outer periclinal walls.
Leaf and leaflet morphology of Moringa species: (A) Moringa peregrina, (B) Moringa oleifera and (C) Moringa stenopetala. Photographs were taken by the co-author, Prof. Dr. Ahmed M. El-Taher (eltaher69@azhar.edu.eg).
Microphotographs of seed morphology of the Moringa studied species: D1-D2. Moringa peregrina, E1-E2. Moringa oleifera and F1-F2. Moringa stenopetala. Photographs were taken by the co-author, Prof. Dr. Ahmed M. El-Taher (eltaher69@azhar.edu.eg).
20
Microphotographs of leaf epidermal of the Moringa studied species: G1-G2. Moringa peregrina, H1-H2. Moringa oleifera and I1-I2. Moringa stenopetala.
Numerical analysis
Taxonomic relatedness of the studied species was analyzed using morphological characterization. The numerical analysis was conducted using the attributes of Moringa species, including their 31 morphological characteristics. A dendrogram was constructed and revealed two main clusters. M. peregrina was separated in the first cluster at an 83.8% similarity level. The second cluster comprised M. oleifera and M. stenopetala at a 95.35% similarity level (Fig. 4).
The M. oleifera and M. stenopetala are more related and separated in cluster I, and M. peregrina is separated in cluster II. This study agrees with the results of Azza65 who observed that the studied species can be split into two main clusters based on similarity or dissimilarity distance. The first cluster included both M. oleifera and M. stenopetala, linked together at a 0.5 similarity level. Whereas, the second cluster included a single species, M. peregrina, (with a similarity level of 2.0). All three species are linked in the main cluster at similarity level of 2.0. This is attributed to their belonging to the Moringa genus.
Dendrogram showing the interrelationships between three species of Moringa based on 31 morphological characters by using the Primer program (www.primer-e.com).
Phytochemical study
Chemical constituents
Methanolic extracts of leaves were derived to determine and discriminate the presence of significant secondary metabolites in three Moringa species under study. Table 4 presents the phytochemical contents (mg/100 g) for three Moringa species. The highest content of chlorophyll a and b were found in M. oleifera (40.98 and 12.2 mg/100 g), followed by stenopetala (40.19 and 11.01 mg/100 g) and M. peregrina (29.2 and 3.57 mg/100 g and), respectively. While, the M. peregrina contains the highest content of phenols (243 mg/100 g) and antioxidant capacity (1226.85 mg/100 g), M. peregrina is characterized by the highest flavonoid content (7 mg/100 g) only.
GC-MS analysis of Moringa species seed extracts
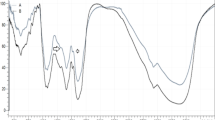
GC-MS analysis of M. oleifera seed n-hexane extract identified twenty compounds (Table 5; Fig. 5A) representing 75.27% of the total extract composition. The main identified compounds are 2-decenal, (E) (39.14%), 2-undecenal (15.51%), nonanal (3.60%), and 2-octenal, (E) (2.48%). While, GC-MS analysis of M. peregrina seed n-hexane extract led to the identification of eighteen compounds (Table 6; Fig. 5B), representing 55.89% of the total extract composition. The main identified compounds are 2-decenal, (Z) (25.42%), 2-docecen-1-al (9.35%), and 13-Docosenoic acid, methyl ester, (Z) (4.16%). Moreover, GC-MS analysis of M. stenopetala seed n-hexane extract led to identifying fifteen compounds (Table 7; Fig. 5C) representing 74.95% of the total extract composition. The main identified compounds are 2-decenal, (E) (26.67%), 2-undecenal (24.10%), and nonanal (4.40%). Taken together, the current finding showed that there is a slight variation in the chemical composition between the three investigated Moringa species, with a very large similarity between M. oleifera and M. stenopetala in the chemical composition, especially the similarity in the major compounds [e.g., 2-decenal, (E), 2-undecenal, and nonanal (4.40%)].
GC-MS chromatograms of the investigated Moringa species seed extracts; 1 A) M. oleifera, 1 B) M. peregrina, and 1 C) M. stenopetala.
The significant difference between leaf and seed extracts is mainly due to the extraction method. The type of solvent used in extraction plays a major role in determining the type of chemical compounds to be extracted. In the present study, the leaves were extracted using polar solvents like acetone and methanol, which resulted in polar extracts comprising some specific chemical classes such as phenols (flavonoids and phenolic acids), anthraquinones, coumarins and other related compounds. On the other hand, extracting seeds using n-hexane as a non-polar solvent resulted in targeting a specific class of non-polar compounds like fatty acids and their derivatives. Therefore, we can conclude that the choice of solvent used in extraction has a major impact on the chemical composition and biological activities of the investigated species.
The current study showed that the variation in these phytoconstituents (Tables 5, 6 and 7) in Moringa species contributes to the various medicinal uses and biological activities68. This is also useful in taxonomical studies for classification and systematics. Moringa possesses high antioxidant activity primarily because of its high flavonoid contents in flavanol and glycoside forms69. The flavonoids are one of the secondary metabolites that have various bioactivities at harmless concentrations70. Dietary flavonoids are valuable in preventing against several cancer diseases71,72 and have curative properties against various pathogens. Furthermore, they are traditionally utilized for their analgesic, antimicrobial, and soothing properties73,74,75.
The high antioxidant activity in Moringa plants is attributed to their high content of phenolics. For example, a high content occurs in M. stenopetala with phenolic content level of 243 mg/100 and antioxidant level of 1226.75 mg/100 g (Table 4). The free radicals produced in cells are stabilized by phenolic compounds by donating or accepting electrons, acting as antioxidants. High concentrations of phenols in the extracts might induce caspase and apoptosis. It corresponds to those of Abdel Baky and El-Baroty76. El-Alfy et al.77, Teixera et al.78, and Saini et al.79,80 reported that the major carotenoid content detected in the M. oleifera and M. peregrina. Moringa leaves in general are reported to have relatively high levels of protein, vitamin A, potassium, vitamin C, and calcium2,81. Zaghloul et al.82 reported that M. peregrina was also used as fodder to increase animal weight. M. oleifera contains a high amount of zeatin that has been used as a natural plant growth enhancer and helps to increase crop yields7. GC-MS technique has been widely used to identify the chemical constituents of Moringa species seed extracts. Previous reports revealed the presence of a broad array of volatile and non-volatile compounds among the tested Moringa extracts83,84,85.
Molecular characterization and genetic diversity
Genetic analysis was conducted using 16 primers of start codon target (SCoT) markers to examine the molecular polymorphism among three Moringa taxa (see Table 1 for more details about taxa) (Fig. 6). The PCR reactions produced 112 amplicons with a length of 110–1341 bp, 18 of which were polymorphic, while 77 were monomorphic amplicons (Table 8). The polymorphism oscillated from no polymorphism by SCoT-14 to 80% by SCoT-05, with an average of 31%.
Profiles of fifteen SCoT markers for three Moringa Taxa. From left to right: Lane 1: DNA ladder (100 bp), Lane 2: Moringa oleifera, Lane 3: Moringa peregrina, and Lane 4: Moringa stenopetala. for more details about taxa, see Table 1.
Unique amplicons summed up 17 bands plus 15 of which were negative unique bands. The marker SCoT2 registered the highest number of unique bands, while 3 were unique negative bands (SCoT11). This variance can be utilized to distinguish the studied markers revealed as unique bands.
Genetic-based identification of Moringa using SCoT markers enabled polymorphism detection to reveal the relationship between the studied taxa. Genetic characterization is crucial for managing the entire verification, classification, and genetic improvement43. The SCoT markers presented a high capacity for the determination of intra- and inter-genomic diversity because of their resultant polymorphism59,86,87,88. In this trend, some conclusions were obtained by Mahdy et al.59 on cowpea and El-Taher et al.89 on Cichorium taxa. The number of genetic parameters was estimated to evaluate the discriminatory power and diversity as detailed in Table 8. Heterozygosity (H) averaged 0.124 ± 0.02 and Shannon’s information index (I) averaged 0.18 ± 0.03. Notably, the SCoT-16, SCoT7, and SCoT9 recorded zero, and the SCoT-12 scored the highest value of 0.30 in both genetic estimations. The heterozygosity (H) is significant in polymorphism information content, which ranges from zero to one90. The derived data appeared to have a moderate informative value of diversity (Table 8). The Shannon index (I) is one of the most common measurements to estimate genetic diversity91. Previous studies are in general agreement with our results59. These genetic estimations could be used for determining diversity occurring between and within species and mirror heterozygosity in them. The heterozygosity is estimated based on the allele frequency.
Genetic correlation (Table 9) was summed up with the highest value of 0.798 between taxon 2 and taxon 3, followed by between taxon 1 and taxon 2 and between taxon 1 and taxon 3, 0.776 and 0.769, respectively. The genetic distance and variance were estimated for more classification (Table 9). The variance was recorded at 11.8, while the genetic distance oscillated from 2.69 to 2.92 with an average of 2.79. Genetic correlation revealed a very close genetic relationship among the studied taxa, and all dropped into the same class. That may be due to the variance decomposition targeting different genome loci by the SCoT technique68,92. Additionally, it may be due to the makeup of the taxon’s genome or the effect of evolution naturally.
Noteworthy, Moringa species have economic uses in Egypt for their antioxidant, antimicrobial, and anti-diabetic activities. Moringa is traditionally used for treating wounds, fever, constipation, muscle pains, burns, labor pain, hypertension, malaria, stomach disorder, asthma, skin problems, and to expel a retained placenta37. Typically, taxonomic diversity refers to the genetic relationship between and within species. Three Moringa species were recorded in Egypt. Diversity measurements of Moringa were estimated as presented in Table 1. All Moringa taxa in the present study have at least one aspect of health benefit or environmental service. Health benefits include medicinal and pharmaceutical values, while environmental services are through shading, phytoremediation, and reducing soil loss. Despite the importance of Moringa, they are threatened by over-harvesting and over-cutting, climate change, and habitat loss.
For more diversity measurements, Dominance, Simpson diversity, Simpson reciprocal indices were calculated as presented in Table 10. Diversity is the variety and involves counting or listing species at its simplest level. Simpson index was estimated at 0.37, Dominance index was 0.63, and the Reciprocal index of Simpson was at 2.7. Simpson index for within species diversity was recorded at 0.145 (M. stenopetala), 0.28 (M. peregrina), and 0.855 (M. oleifera). Moringa stenopetala (6.875) scored the highest value of Reciprocal index, while the Moringa oleifera (0.852) recorded the highest value of Simpson. The results represented a high diversity for M. oleifera compared to others. The diversity index is a quantitative measure of the diversity of different organisms/species and how evenly the individuals are distributed among those species. The value of diversity index increases when the number of types increases and evenness increases93. Simpson index is weighted arthimetic average of proportional abundance and measures the probability that two individuals selected randomly from a sample will belong to the same species. This average will increase when the number of species decreases, and abundance of the most abundant species increases. Therefore, the Simpson index has small values in datasets of high diversity and vice versa. For this, Simpson index is usually expressed as a Dominance (its inverse) or reciprocal (its compliment) which is also called the Gini-Simpson index93,94. These results are in agreement with Shaltout and Bedair95. Moringa species are classified for conservation as: vulnerable (VU), very common (VC), and common (C). Propagation of M. peregrina had been successfully established in the Orman Botanic Garden and Shehab Mazhar Botanic Garden (private sector) in Giza95. A successful plantation of M. peregrina was done in South Sinai96. However, for a reason, M. peregrina was completely over-grazed in Orman Botanic Garden. Mortality of populations of M. peregrina is due to over-grazing82. Climate change impacts are becoming more evident, and mitigating its effects on localized species is a major challenge95.
Conclusion
Despite the important properties and potential uses of Moringa species among tropical plants, little research and development attention has been given to it. Besides its pharmaceutical use, Moringa is also a beneficial bio-material in farming. Several phytochemicals are yet to be identified in order to consider for all the potential therapeutic benefits. More attention is needed to Moringa as a priority plant to lessen malnutrition and as a source of farm income. This emphasizes the need for detailed and rigorous scientific study to assess the attributes of Moringa and utilize its full potential.
The obtained results reported that phenological features were assessed in the taxonomical association, which separate M. peregrina lonely (84%) and placed M. stenopetala with M. oleifera into a cluster score with a 95.3% similarity. A wide similarity has been observed between M. oleifera and M. stenopetala by the phytochemical compositions, especially the similarity in the major compounds such as 2-decenal, (E), 2-undecenal, and nonanal. Additionally, genetic estimations, i.e., heterozygosity (0.124), Shannon index (0.18), and Simpson index, confirmed the integration of these approaches.
Plant identification is the first step towards the determination of the basis for sustainable utilization of a taxon. Plant verification is based on the integration of micro- and macro-morphological assessment, molecular characterization, and phytochemical evaluation, which play a main role in the classification of within and between taxa. The morphology-based characterization is still the first step toward classification, genetic association, and changing climates. These distinct taxa and their phylogenetic associations were confirmed and evaluated in the current study using morphological, phytochemical, and molecular approaches. These methodologies proved helpful for classification, systematics, and phylogenetic studies at the taxonomical ranks of Moringa. We report that the leaf characteristics were the most important attributes in constructing the indented key for Moringa spp. We report that the genetic-based identification confirms the studies coming from morphological attributes. The integration of genetic and morphological along with phytochemical studies is helpful in the identification and verification of Moringa taxa. Finally, the genome assembly of Moringa among different countries diverges visibly with a firm amount of gene flow.
These taxa have the potential to be used as rich genetic resources for future studies in genetic diversity and germplasm preservation. The potential of these species can be benefited by; it would greatly benefit many countries that rely thoroughly on indigenous plants as a strength source for their maintenance. The rarity Moringa species, including M. peregrina, are also genetically vulnerable in their respective habitats due to several factors, ranging from unsustainable utilization, little scientific research and studies, and various threats. Further research is critically needed for Moringa species and should focus mainly on sustainable utilization and conservation. Efforts need to be expanded to pay the attention Moringa species rightfully deserve. If the genus Moringa, which consists of more potentially valuable species, can be further evaluated for its various characteristics, it can contribute to conservation and sustainable use. To sum up, the investigated Moringa species are a prolific source of bioactive secondary metabolites. There is an urgent need for more intensive bioprospecting studies and to establish a link between chemical composition and biological activities via molecular modeling experiments. Therefore, UPLC-QTOF-MS/MS metabolite profiling of the leaf extract is strongly recommended in future perspectives accompanied by docking studies.
Data availability
The taxa used are available for distribution to those interested in it. Requests for material should be mailed to National Gene Bank (NGB), Agricultural Research Center (ARC), 9 Gamaa St., PO Box: 12619; Giza, Egypt, or emailed to the corresponding authors: ehab.mahdy@arc.sci.eg, fatma.hamada@aswu.edu.eg.
References
Amaglo, N. K., Bennett, R. N., Curto, L. & Rosa, R. B. Lo Turco V. Profiling selected phytochemicals and nutrients in different tissues of the multipurpose tree Moringa oleifera L., grown in Ghana. Food Chem. 122, 1047–1054 (2010).
Rani, N. Z., Husain, K. & Kumolosasi, E. Moringa ganus: A review of phytochemistry and pharmacology. Front. Pharmacol. 9, 108 (2018).
Olson, M. Intergeneric relationships within the Caricaceae-Moringaceae Clade (Brassicales) and potential morphological synapomorphies of the clade and its families. Int. J. Plant. Sci. 163, 51–65 (2002).
Mallenakuppe, R. H., Homabalegowda, M. D. & Gouri, P. History, taxonomy and propagation of Moringa oleifera- a review. SSR Inst. Int. J. Life Sci. 5, 2322–2327 (2019).
Ruiz, A. I., Mercado, M. I., Guantay, M. E. & Ponessa, G. I. Anatomía E histoquímica foliar y caulinar de Moringa oleifera (Moringaceae). Bol. Soc. Argent. Bot. 54, 325–343 (2019).
Migahid, A. M. Flora of Saudi Arabiavol. 1101 (Riyadh University Publication, 1978).
Leone, A. et al. Cultivation, genetic, ethnopharmacology, phytochemistry ad pharmacology of Moringa oleifera leaves: An overview. Int. J. Mol. Sci. 16, 12791–12835 (2015).
Chase, M. et al. An ordinal classification for the families of flowering plants. Ann. Missouri Bot. Gard. 85, 531–553 (1998).
Anwar, F., Latif, S., Ashraf, M. & Gilani, A. Moringa oleiaera: A food plant with multiple medicinal uses. Phytother Res. 21, 17–25 (2007).
Boukandoul, S., Casal, S. & Zaidi, F. The potential of some Moringa species for seed oil production. Agriculture. 8, 150 (2018).
El-Awady, M., Hassan, M., Abdel-Hameed, E. & Gaber, A. Comparison of the antimicrobial activities of the leaves-crude extracts of Moringa peregrina and Moringa oleifera in Saudi Arabia. Int. J. Curr. Microbiol. App Sci. 4, 1–9 (2015).
Gopalakrishnan, L., Doriya, K. & Kumar, D. Moringa oleifera: A review on nutritive importance and its medicinal application. Food Sci. Hum. Wellness. 5, 49–56 (2016).
Rashid, U., Anwar, F., Moser, B. & Knothe, G. Moringa oleiferaaoil: A possible source of biodiesel. Bioreso Technol. 99, 8175–8179 (2008).
Robiansyah, I., Hajar, A., Al-kordy, M. & Ramadan, A. Current status of economically important plant Moringa Peregrina (Forrsk.) Fiori in Saudi Arabia: A review. Int. J. Theore Appl. Sci. 6, 79–86 (2014).
Padayachee, B. & Baijnath, H. An overview of the medicinal importance of Moringaceae. J. Med. Plant. Res. 6, 5831–5839 (2012).
Seifu, E. Actual and potential applications of Moringa stenopetala, underutilized indigenous vegetable of southern Ethiopia: A review. Int. J. Agric. Food Res. 3, 8–19 (2014).
Ezeamuzie, T., Amberkedeme, A., Shode, F. & Ekwebelem, S. C. Antiinflammatory effects of Moringa oleifera root. Int. J. Pharmacogn. 34, 207–212 (1996).
Mohammed, S. et al. An overview of natural plant antioxidants: Analysis and evaluation. Adv. Biochem. J. 1, 44–72 (2013).
Jayeola, A. Anatomical identification of plant fragments in the powdered samples of Moringa oleifera Lam. (Moringaceae), in The National Summit on Moringa Development, Abuja (2010).
Okoye, E., Ezeifeka, G. & Nworu, C. Evaluation of the antiviral activity of Moringa oleifera on three RNA viruses, in The National Summit on Moringa Development, Abuja (2010).
Noraini, T. & Cutler, D. Leaf anatomical micromorphological characters of some Malaysian Parashorea (Dipterocarpaceae). J. Trop. Sci. 21, 156–167 (2009).
Sharma, A., Sehrawai, S., Singhrot, R. & Tele, A. Morphological chemical characterization of Psidium species. Notulae Boanicae Horti Agrobotanici Cluj-Napoca (2010).
Soladoye, M., Sonobare, M. & Chukwuma, E. Morphometric study of the Genus Indigofera Linn. (Leguminosae-Papilionoideae) in south-western Nigeria. Int. J. Bot. 6, 227–234 (2010).
Abubakar, B., Mua’zu, S., Khan, A. & Adamu, A. Morpho-anatomical variation in some accessions of Moringa oleifera Lam. From northern Nigeria. Afr. J. Plant. Sci. 5, 742–748 (2011).
Kumar, P., Singh, K. & Kumar, A. Hepatoprotective studies on aerial parts of Moringa oleifera Lam. On carbon tetrachloride induced liver cell damage in albino rats. Ann. Biol. Res. 1, 27–35 (2010).
Nibret, E. & Wink, M. Trypanocidal and antileukaemic effects of the essential oils of Hagenia Abyssinica, Leonotis Ocymifolia, Moringa Stenopetala, and their main individual constituents. Phytomedicine. 17, 911–920 (2010).
Walter, A., Samuel, W., Peter, A. & Joseph, O. Antibacterial activity of Moringa oleifera and Moringa stenopetala methanol and n-hexane seed extracts on bacteria implicated in water borne diseases. Afr. J. Microbiol. Res. 5, 153–157 (2011).
Tahany, M. et al. Study on combined antimicrobial activity of some biologically active constituents from wild Moringa peregrina Forssk. J. Yeast Fungal Res. 1, 015–024 (2010).
Lalas, S. & Tsaknis, J. Characterization of Moringa oleifera seed oil variety periyakulam 1. J. Food Compos. Anal. 15, 65–78 (2002).
Gandji, K. et al. Status and utilisation of Moringa oleifera Lam: A review. Afr. Crop Sci. J. 26, 137–156 (2018).
Shindano, J. & Kasase, C. Moringa (Moringa oleifera): A source of food and nutrition, medicine and industrial products, in In African Natural Plant Products: New Discoveries and Challenges in Chemistry and Quality, ACS Symposium Series, Washington, DC, American Chemical Society, 421–467 (2009).
Uilah, N., Zahoor, M., Khan, F. & Khan, S. Review on general introduction to medicinal plants, its phytochemicals and roles of heavy metal and inorganic constituents. Life Sci. J. 11, 520–527 (2014).
El-Dahiyat, F., Rashrash, M., Abuhamdah, S., Farha, R. & Babar, Z. Herbal medicines: A cross-sectional study to evaluate the prevalence and predictors of use among Jordanian adults. J. Pharm. Policy Pract. 3, 2 (2020).
Mahdy, E., El-Sharabasy, S. & El-Dawayati, M. In vitro production of quinones, In Nutraceuticals Production from Plant cell Factory, Singapor, Springer, Singapore, 345–374 (2022).
Gong, H. et al. Effects of several quinones on insulin aggregation. Sci. Rep. 4, 5648 (2014).
Lorenzo, J. et al. Main characteristics of peanut skin and its role for the preservation of meat products. Trends Food Sci. Technol. 77, 1–10 (2018).
Senthilkumar, A., Karuvantevida, N., Rastrelli, L., Kurup, S. & Cheruth, A. Traditional uses, pharmacological efficacy, and phytochemistry of Moringa peregrina (Forssk.) Fiori.- a review. Front. Pharmacol. 9, 465 (2018).
Ganesan, S. et al. Genetic diversity and population structure study of drumstick (Moringa oleifera Lam.) Using morphological and SSR markers. Ind. Crop Prod. 60, 316–325 (2014).
Saini, R. K., Saad, K. R., Ravishankar, G. A., Giridhar, P. & Shetty, N. P. Genetic diversity of commercially grown Moringa oleifera Lam. Cultivars from India by RAPD, ISSR and cytochrome P450-based markers. Plant. Syst. Evol. 299, 1205–1213 (2013).
Zaghloul, M., El-Wahab, A., Moustafa, A. & R., & Ecological assessment and phenotypic and fitness variation of Sinai’s populations of Moringa Peregrina. Appl. Ecol. Environ. Res. 8, 351–336 (2010).
Osman, H. E. & Abohassan, A. A. Morphological and analytical characterization of Moringa Peregrina populations in western Saudi Arabia. Int. J. Theor. Appl. Sci. 4, 174–184 (2012).
Sharla, N., Bobba, S. & Siddiq, E. ISSR and SSR markers based on AG and GA repeats delineate geographically diverse Oryza Nivara accessions and reveal rare alleles. Curr. Sci. 84, 683–690 (2003).
Mahdy, E., Ibrahim, S., EL-Shaer, H. & Mansour, M. Genetic diversity and relationship between Egyptian Vigna (Vigna spp. (L.) Walp.) Taxa populations via phenotypic and molecular profiling. Vegetos. https://doi.org/10.1007/s42535-024-00827-1 (2024).
Sanchez, H., Loarce, J. & Ferrer, E. Simple sequence repeat primers used in polymerase chain reaction amplifcation to study genetic diversity in barley. Genome. 39, 112–117 (1996).
Joshi, C., Zhou, H., Huang, X. & Chiang, V. Context sequences of translation initiation codon in plants. Plant. Mol. Biol. 35, 993–1001 (1997).
Sawant, S., Singh, P., Gupta, S., Madnala, R. & Tuli, R. Conserved nucleotide sequences in highly expressed genes in plants. J. Genet. 78, 123–131 (1999).
Gupta, M., Chyi, Y., Romero-Severson, J. & Owen, J. Amplification of DNA markers from evolutionary diverse genomes using single primers of simple-sequence repeats. Theor. Appl. Genet. 89, 998–1006 (1994).
Williams, J., Kubelik, A., Livak, K., Rafalski, J. & Tingey, S. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res. 18, 6531–6539 (1990).
Bailey, L. in Manual of Cultivated Plants. 601–609 (eds Bailey, L.) (New York, Macmillan Co., 1924).
Nagata, M. & Yamashita, I. Simple method for simultaneous determination of chlorophyll and carotenoids in tomato fruit. J. Japanese Soc. Food Sci. Technol. 39, 925–928 (1992).
Singleton, V., Orthofer, R. & Lamuela-Raventos, R. Analysis of total phenols and other oxidation substances and antioxidants by means of Folin Ciocalteu reagent. Methods Enzymol. 299, 152–178 (1999).
Woisky, R. & Salation, A. Analysis of propolis: Some parameters and procedures for chemical quality control. J. Agric. Res. 37, 99–105 (1998).
Prieto, P., Pineda, M. & Aguilar, M. Spectrophotometric quantitation of antioxidant capacity through the formation of a phosphomolybdenum complex: Specific application to the determination of vitamin E. Anal. Biochem. 269, 337–341 (1999).
Klein, B. & Perry, A. Ascorbic acid and vitamin A activity in selected vegetables from different geographical areas of the United States. J. Food Sci. 47, 941–945 (1982).
Abdel-Wareth, M., El-Hagrassi, A., Abdel-Aziz, M., Nasr, S. & Ghareeb, M. Biological activities of endozoic fungi isolated from Biomphalaria alexandrina snails maintained in different environmental conditions. Int. J. Environ. Sci. 76, 780–799 (2019).
Madkour, H. et al. Gas chromatography-mass spectrometry analysis, antimicrobial, anticancer and antioxidant activities of n-hexane and methylene chloride extracts from Senna italica. J. App Pharm. Sci. 7, 023–032 (2017).
Shawky, B. et al. Evaluation of antioxidants, total phenolics and antimicrobial activities of ethyl acetate extracts from fungi grown on rice straw. J. Renew. Mater. 7, 667–682 (2019).
Khalaf, O., Abdel-Aziz, M., El-Hagrassi, A., Osman, A. & Ghareeb, M. Biochemical aspect, antimicrobial and antioxidant activities of Melaleuca and Syzygium species (Myrtaceae) grown in Egypt. J. Phys. Conf. Ser. 1879, 022062 (2021).
Mahdy, E., El-Shaer, H., Sayed, A. & El-Halwagi, A. Genetic diversity of local cowpea (Vigna spp. (L.) Walp.) Accessions cultivated in some regions of Egypt. Jordan J. Biol. Sci. 14, 775–789 (2021).
Collard, B. & Mackill, D. Start codon targeted (SCoT) polymorphism: A simple novel DNA marker technique for generating gene-targeted markers in plants. Plant. Mol. Biol. Rep. 27, 86–93 (2009).
Jaccard, P. Nouvelles recherches sur la distribution florale. Lausanne, Rouge. Bull. Soc. Vaud Sci. Nat. 44, 223–270 (1908).
Nei, M. & Li, M. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Nat. Acad. Sci. 76, 5269–5273 (1979).
Sendecor, G. & Cochran, W. Statistical methods, 7th Edition. ed., Oxford and J.B.H. Publishing com (1990).
Dice, L. Measures of the amount of ecologic association between species. Ecology. 26, 297–302 (1945).
Azza, S. Morpho-anatomical variations of leaves and seeds among three Moringa species. Life Sci. J. 11, 827–832 (2014).
Rangnath, K., Radhakrushna, K. & Narayanlal, S. Moringa oleifera: Morphology and medicinal use. Ijariie. 9, 4395–4396 (2023).
Zhigila, D., Mohammed, S., Oladele, F. & Sawa, F. Numerical analyses of leaf and fruit external morphology in Moringa oleifera Lam. J. Teknol (Sci Eng). 77, 123–131 (2015).
El-Ghadban, E., Abou El-leel, O. & Mahdy, E. Morphological, phytochemical and molecular characterization on some Jatropha species cultivated in Egypt. Int. J. Pharm. Sci. Scient Res. 3, 1–13 (2017).
Wang, Y. et al. Subcritical ethanol extraction of flavanoids from Moringa oleifera leaf and evaluation of antioxidant activity. Food Chem. 218, 152–158 (2017).
Irshad, M., Ahmed, I., Goel, H. & Rizvi, M. Phytochemical screening and high performance TLC analysis ofsome cucurbits. Res. J. Phytochem. 4, 242–247 (2010).
Ren, W., Oiao, Z., Wang, H., Zhu, L. & Zhang, L. Promising anticancer agents. Med. Res. Rev. 23, 519–534 (2003).
Aggarwal, B. & Shishodia, S. Molecular targets of dietry agents for prevention and therapy of cancer. Biochem. Pharmacol. 71, 1397–1421 (2006).
Thirunavukkarasu, P., Ramanathan, T., Ramkumar, I. & Shanmugapriya, R. Anti-ulcer effect of Avicennia officinalis leaves in albino rats. World Appl. Sci. J. 9, 55–58 (2010).
Singh, A., Duggal, S. & Suttee, A. AcaIlicifoliusfolius Linn-lesser known medicinal plants with significant pharmacological activities. Int. J. Phytomed. 1, 1–3 (2009).
Ganesh, S. & Vennila, J. Phytochemical analysis of Acanthus Ilififolius and Avicennia officinalis by GC-MS. Res. J. Phytochem. 5, 60–65 (2011).
Abdel Baky, H. & El-Baroty, G. Characterization of Egyptian Moringa peregrine seed oil and its bioactivities. Int. J. Manage. Sci. Bus. Res. 2, 98–108 (2013).
El-Alfy, T., Ezzat, S., Hegazy, A., Amer, A. M. & Kamel, G. Isolation of biologically active constituents from Moringa peregrina (Forssk.) Fiori. (family: Moringaceae) growing in Egypt. Pharmacogn. Mag. 7, 109–115 (2011).
Teixera, E. M., Carvalho, M. R., Neves, V. A., Silva, M. A. & Arantes-Pereira, L. Chemical characteristics and fractionation of proteins from Moringa oleifera Lam. Leaves. Food Chem. 147, 51–54 (2014).
Saini, R., Shetty, N. & Giridhar, P. Carotenoid content in vegetative and reproductive parts of commercially grown Moringa oleifera Lam. Cultivars from India by LC-APCI-MS. Eur. Food Res. Technol. 238, 971–978 (2014).
Saini, R., Sivanesan, I. & Keum, Y. Phytochemicals of Moringa oleifera: A review of their nutritional, therapeutic and industrial significance. 3 Biotech. 6, 1–14 (2016).
Mathur, B. Moringa Book. St. Louis, MI Trees for Life International. (2005).
Zaghloul, M., Hamrick, J. & Moustafa, A. Conservation genetics of Sinai’s remnant populations of Moringa Peregrina, an economically valuable medicinal plant. Conserv. Genet. 13, 9–19 (2012).
Melaku, Y., Arnold, N., Schmidt, J. & Dagne, E. Analysis of the husk and kernel of the seeds of Moringa Stenopetala. Bull. Chem. Soc. Ethiop. 31, 107–113 (2017).
Maqbul, M. et al. Comparative study of Moringa oleifera with Moringa peregrine seed oil using GC-MS and its antimicrobial activity against Helicobacter pylori. Orient. J. Chem. 36, 481–492 (2020).
El Sayed, A., Omar, F., Emam, M., Farag, M. & UPLC-MS/MS GC-MS based metabolites profiling of Moringa oleifera seed with its anti-Helicobacter pylori and anti-inflammatory activities. Nat. Prod. Res. 36, 6433–6438 (2022).
Jain, A., Bhitia, S., Banga, S., Prakash, S. & Laxmikumaran, M. Potential use of RAPD technique to study the genetic diversity in Indian mustard (Brassica juncea) and its relationship to heterosis. Theor. Appl. Genet. 88, 116–122 (1994).
Gupta, S. et al. Analogy of ISSR and RAPD markers for comparative analysis of genetic diversity among different Jatropha curcas genotypes. Afr. J. Biotechnol. 7, 4230–4243 (2008).
Tatikonda, L. et al. AFLP-Based molecular characterization of an elite germplasm collection of Jatropha curcas L., a biofuel plant. Plant. Sci. 176, 505–513 (2009).
El-Taher, A. et al. Characterization of some Cichorium taxa grown under mediterranean climate using morphological traits and molecular markers. Plants. 12, 388 (2023).
Botstein, D., White, R., Skolnick, M. & Davis, R. Construction of a genetic linkage map in man using restriction fragment length polymorphism. Am. J. Hum. Genet. 32, 314–331 (1980).
Shannon, C. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Mahdy, E. & Rizk, R. Genetic variation of arta populations (Calligonum Polygonoides subsp. comosum) in Egypt: Genepools for biodiversity and afforestation genetic variation of arta populations. J. Water Land. Dev. 56, 81–90 (2023).
Simpson, E. Measurement of diversity. Nature. 163, 688 (1949).
Pielou, E. The measurement of diversity in different types of biological collections. J. Theoret Biol. 13, 131–144 (1966).
Shaltout, K. & Bedair, H. Diversity, distribution and regional conservation status of the Egyptian tree flora. Afr. J. Ecol. 60, 1155–1183 (2022).
Abd El-Wahab, M. Reproductive ecology of wild trees and shrubs in southern Sinai, M.Sc. thesis, Suez Canal University (1995).
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
Conceptualization, O.F.A.E., F.A.H., and T.R.; methodology, O.F.A.E., H.S.A., F.A.H., M.A.G., A.T.A., and S.S.S.; software, E.M.B.M., A.T.A., F.A.H., and A.M.E.; validation, A.M.E., F.A.H., A.M.E., and A.T.A.; formal analysis, H.S.A., A.T.A., A.M.E., R.R., F.A.H., and M.A.G.; investigation, M.A.G., F.A.H., R.R., S.S.S., E.M.B.M., and H.S.A.; resources, O.F.A.E., and A.M.E.; data curation, A.M.E., H.S.A., and S.S.S.; writing—original draft preparation, A.M.E., R.R., F.A.H., O.F.A.E., T.R., and E.M.B.M.; writing—review and editing, O.F.A.E., S.S.S., R.R., E.M.B.M., F.A.H., A.M.E., M.A.G., and T.R.; visualization, E.M.B.M., M.A.G. and H.S.A.; supervision, T.R., F.A.H., and E.M.B.M.; project administration, T.R, and A.M.E.; funding acquisition, F.A.H. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hamada, F.A., Sabah, S.S., Mahdy, E.M. et al. Genetic, phytochemical and morphological identification and genetic diversity of selected Moringa species. Sci Rep 14, 30476 (2024). https://doi.org/10.1038/s41598-024-79148-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79148-x