Abstract
Automatic and accurate tooth segmentation on 3D dental point clouds plays a pivotal role in computer-aided dentistry. Existing Transformer-based methods focus on aggregating local features, but fail to directly model global contexts due to memory limitations and high computational cost. In this paper, we propose a novel Transformer-based 3D tooth segmentation network, called PointRegion, which can process the entire point cloud at a low cost. Following a novel modeling of semantic segmentation that interprets the point cloud as a tessellation of learnable regions, we first design a RegionPartition module to partition the 3D point cloud into a certain number of regions and project these regions as embeddings in an effective way. Then, an offset-attention based RegionEncoder module is applied on all region embeddings to model global context among regions and predict the class logits for each region. Considering the irregularity and disorder of 3D point cloud data, a novel mechanism is proposed to build the point-to-region association to replace traditional convolutional operations. The mechanism, as a medium between points and regions, automatically learns the probabilities that each point belongs to its neighboring regions from the similarity between point and region features, achieving the goal of point-level segmentation. Since the number of regions is far less than the number of points, our proposed PointRegion model can leverage the capability of the global-based Transformer on large-scale point clouds with low computational cost and memory consumption. Finally, extensive experiments demonstrate the effectiveness and superiority of our method on our dental dataset.
Similar content being viewed by others
Introduction
Artificial Intelligence (AI) technology is improving by leaps and bounds. For stomatology tasks, like digital dentistry, orthodontics and dental implants, learning-based 3D tooth segmentation is a crucial step for a computer-aided-design (CAD) system to automatically and accurately identify individual tooth and gingiva based on 3D real oral scanning data or dental model. This will facilitate the subsequent diagnosis and treatment and free the dentists from tedious manual segmentation work.
However, due to noise, irregularity, high redundancy, nonuniform distribution and disorder of 3D point cloud data, 3D point cloud segmentation is full of challenges. Traditional 3D point cloud segmentation methods1,2,3 have poor accuracy and meanwhile cannot be extended to large-scale data. Driven by the tremendous success of 2D computer vision, deep learning-based technology has become the preferred choice for 3D segmentation tasks.
Recently, Transformer architecture has been successfully applied in natural language processing4,5,6 and 2D computer vision tasks7,8 due to its strong advantage in learning long-range dependencies of input sequences, and exhibits better performance than convolutional neural networks (CNNs). As the core of Transformer, self-attention mechanism5 has been proven suitable for processing 3D point clouds, and some Transformer-based works have emerged for point cloud learning. Most of these works9,10,11,12,13,14,15 use Transformer as an auxiliary module with the aim of achieving feature aggregation in the local patch, and only a small proportion16,17,18 propose Transformer-based architectures that apply global attention on the whole point cloud while avoiding the overhead of constructing local neighborhoods. Nevertheless, the global attention mechanism makes these approaches unsuitable for processing large-scale point clouds, e.g., dental point clouds with a scale of \(10^5\), due to the heavy computation burden and unacceptable memory consumption.
In this paper, we propose a novel Transformer-based 3D tooth segmentation network, called PointRegion, which successfully lowers the computation complexity and memory consumption from the perspective of reducing the input sequence length. Our work is inspired by 2D image segmentation. To reduce inference time and memory consumption while retaining the accuracy, Zhang et al.19 proposed a decoder-free Vision Transformer (ViT) based segmentation method, which treats an image as a set of learnable regions and obtains pixel-level result by a convolution-based correlation mechanism. Compared to per-point prediction on dense regular grids, interpreting an image as a set of interrelated regions is closer to the essence of semantic segmentation. We believe that the same applies to 3D point cloud semantic segmentation. However, since point cloud and image are quite different data, the mechanisms of patch embedding in ViT and correlation computation using standard convolutions are not directly applicable to 3D point clouds. Therefore, we make some analysis and adjustments listed as follows:
-
RegionPartition module Unlike image pixels, point clouds are disordered and unstructured. Therefore, we first need to explore a partitioning method suitable for point clouds and then project the point cloud into a sequence of region embeddings. Although the related works14,17,18 introduced in Section Grid-based Methods for Point Cloud Learning have the practice of converting a point cloud into local patches, the points in the overlapping area of these patches will cause ambiguity when obtaining per-point label. Based on the fact that the closer points generally have closer semantic information, we propose an effective non-overlapping partitioning method in the RegionPartition module. Moreover, we expand the receptive field of each point during feature aggregation to compensate for the information loss caused by region partition, which will be introduced in detail in Section RegionPartition Module.
-
RegionEncoder module In our PointRegion framework, after the RegionPartition module, the region-level embedding instead of the point-level embedding is input to the attention layer, i.e., the RegionEncoder module, which means the computation complexity is quadratic with respect to the number of regions instead of points. It is worth noting that the number of regions in our method is far less than the number of points, which makes our method suitable for processing large-scale dental point clouds. Taking a point cloud with N=10240 points as an example, existing Transformer-based methods typically treat it as an input sequence for feature learning, with the input sequence being N. Although mechanisms like ViT exist to reduce the length of the input sequence, they are not suitable for processing irregular 3D point clouds. Our PointRegion employs the RegionPartition module to divide the point cloud into 1024 regions, leveraging the powerful local and non-local feature extraction capabilities of EdgeConv to preserve as much structural information of the point cloud as possible. Subsequently, the RegionEncoder module accepts the 1024 region embeddings as the input sequence, predicting category probabilities for each region. At this point, the input sequence is reduced by a factor of ten compared to the original input point cloud. Since the computational complexity and spatial occupancy of the self-attention mechanism grow quadratically with the input sequence, the computational complexity and spatial occupancy of PointRegion when using Transformers for point cloud feature extraction are also significantly reduced. The experimental results in Table 2 confirm this point.
-
Point-to-region association After modeling inter-region relations using the global Transformer, the main difficulty lies in generating per-point segmentation result with per-region prediction. The affinity head is a mechanism based on standard convolution to describe region geometries by local pixel-region association as a corresponding solution in this work19. However, it is not suitable for 3D point clouds. To this end, a novel mechanism is designed to establish point-to-region association by utilizing information similarity between points and regions. On the basis of region logits generated by the RegionEncoder module, our point-to-region association mechanism can help our model learn the probabilities of each point belonging to the current reference region and its neighbourhoods, which enables us to segment teeth in a per-region instead of per-point prediction fashion as opposed to conventional UNet-style models9,20,21,22.
Based on the above analyses, we propose a PointRegion network, which successfully extends the 2D segmentation method19 to 3D large-scale point cloud segmentation task. By extensive experimental evaluation, the experimental results demonstrate the excellent performance of the proposed deeping learning network on our dental dataset. Furthermore, to the best of our knowledge, none of the previous works explored the use of relatively complex Transformer structures in 3D tooth segmentation tasks.
In summary, the main contributions of our work include the followings:
-
We propose a novel, accurate, and efficient Transformer-based 3D tooth segmentation network, namely, PointRegion, which is capable of learning on large-scale dental data without expensive computational cost and huge memory consumption.
-
We design a RegionPartition module to partition the 3D point cloud into a certain number of non-overlapping regions and then learn region embeddings in an effective local-nonlocal way.
-
Additionally, we propose a novel mechanism for establishing association between points and regions. This point-to-region mechanism enables our model to achieve point-level tooth segmentation in a more effective per-region prediction fashion.
-
Last but not least, extensive experiments and ablative analysis are conducted on our collected and annotated dental dataset. Compared to other state-of-the-art methods, our proposed method performs well and achieves superior results.
Related work
Considering whether to pre-process 3D point cloud data, most existing learning-based 3D point cloud segmentation methods can be roughly classified into grid-based methods23,24,25,26,27,28,29,30,31 and point-based methods9,10,11,12,13,14,15,16,17,18,20,21,22,32,33,34. Due to the disorder and irregularity of point cloud data, the grid-based methods typically convert point clouds into regular 2D/3D grid representations through projection or voxelization, and then utilize CNNs for feature learning. In contrast, the point-based methods learn directly on the original structure of point clouds, effectively avoiding the time penalty and inevitable information loss caused by grid transformation.
Grid-based methods for point cloud learning
As the foundation of deep learning, CNN has demonstrated powerful learning ability in various 2D visual tasks35,36,37,38,39. In order to explore its potential on 3D point clouds, some researchers attempt to transform irregular points into intermediate grid structure. The 2D grid-based methods23,26,27,28,29,31,40,41,42 typically project a point cloud into 2D images. Among them, Lawin et al.26 proposed to first leverage virtual cameras to obtain multi-view images of a point cloud, and then performed pixel-wise prediction on the images of each view using a multi-stream CNN. These predicted scores are further fused to obtain the final semantic labels for each point. Instead of choosing the positions of multiple cameras, Squeezeseg29 and its improved version SqueezesegV231 put forward projecting the point cloud onto a sphere and processing its spherical representation with an end-to-end pipeline based on CNNs. Although the good scalability of these methods enables them to handle large-scale point clouds, their performance is sensitive to the selection of projection plane, and meanwhile occlusion issues inevitably result in information loss. It is difficult to select an appropriate projection plane in practice.
In the case of 3D grid-based methods24,25,30,43, point clouds are usually voxelized to dense grid representations. However, using standard 3D convolutions directly on voxels makes the memory consumption and computation grow cubically with the voxel resolution, thereby limiting the performance of this type of methods. In response to this challenge, approaches based on sparse convolution44,45 and tree structures46,47,48,49 have emerged. These solutions have indeed reduced learning costs and enhanced performance, but the loss of geometric details during voxelization cannot be ignored.
Point-based methods for point cloud learning
Compared with the grid-based methods, the point-based methods construct a deep network architecture learned directly on the point cloud, which can preserve geometric structure of the point cloud to the greatest extent. As a milestone in deep learning-based point cloud processing, PointNet32 uses shared multi-layer perceptrons (MLPs) to aggregate features independently on each point followed by a permutation-invariant max-pooling operation. However, PointNet is unable to capture local features and cannot perform well in complex tasks like segmentation, therefore, a hierarchical PointNet++20 has been designed. Inspired by PointNet32 and PointNet++20, many subsequent works50,51,52,53,54 borrow from hierarchical structures and aggregate local neighborhood information in a pooling manner.
A different approach redefines effective convolution operators for point clouds. Li et al.21 proposed \({\mathcal {X}}\)-Conv consisting of \({\mathcal {X}}\)-transformation and typical convolution operators in PointCNN. It successfully extends CNN to feature learning of irregular point clouds. PointConv22 treats convolution kernels as nonlinear functions of the local coordinates of 3D points comprised of weight and density functions. Thomas et al.33 presented KPConv, in which a set of kernel points are used to carry weights, and correlation coefficients are determined by the positional relationship between the input points and the kernel points. In addition, the deformable version of KPConv is more flexible, robust, and suitable for complex tasks.
According to the operating scale, Transformer-based methods can be classified into local-based Transformers and global-based Transformers. Local-based Transformers9,10,11,12,13,14,15,55,56,57,58 make full use of the attention mechanism that is a natural fit for point clouds, and design an attention-based module for local geometric feature learning. Zhao et al.9 proposed Point Transformer (PT) to capture the local information by the self-attention vector, which is proven to be more effective compared to the scalar self-attention. Inspired by Swin Transformer8, Zhang et al.14 and Lai et al.15 proposed to convert point cloud into independent 3D grids through voxelization and then build the local self-attention within each window. On the contrary, global-based Transformers16,17,18,34,59,60 are used to learn the global context features. Inspired by the nonlocal operation, Yan et al.34 presented a local-nonlocal module to capture the neighbor and long-range dependencies of sampled points. Point Cloud Transformer (PCT) proposed by Guo et al.16 stacks four global attention layers to learn semantic information, and the offset-attention in PCT instead of the original self-attention is able to sharpen the attention weights and reduce the influence of noise. Following ViT7, Point-BERT17 and Point-MAE18 first grouped point cloud into several local patches and adopted a mini-PointNet32 to project them into point embeddings, which are then fed to pre-train the global-based Transformers. Overall, local-based Transformers are applicable to large-scale point cloud processing, but multiple construction of local patches for irregular data has become a bottleneck in efficiency. In contrast, the huge memory and computational costs make it difficult for global-based Transformers to deal with large-scale data.
Deep learning segmentation methods for dental point clouds
In addition to general-purpose point cloud segmentation methods, various deep learning models have been proposed for tooth segmentation on 3D intraoral scanned (IOS) data recent years. To preserve the topological structure and geometric details of 3D ISO data as much as possible, most of these methods are implemented based on points, reflecting the current research trend in the field.
A majority of these methods are fully supervised61,62,63,64,65,66,67 with the exception of a few which are weakly supervised or semi-supervised68,69,70. Zanjani et al.63 introduced a Mask-MCNet framework which localizes each tooth by predicting its 3D bounding box and simultaneously segments the points that belong to each individual tooth instance. Lian et al.61 evaluated MeshSegNet which learns global information from the tooth point cloud using a PointNet similar network and the local information is learnt using different scale of adjacency matrices. This network borrows heavily from PointNet architecture but stands out by its use of adjacency matrix for understanding the local geometrics. Zhang et. al62 proposed a method which learns separately from the coordinates and the normals information by employing a parallel network. These methods discussed herein have been trained and evaluated on relatively small dataset of annotated 3D intraoral scans (starting 30 from to 120), with a dearth of sample diversity, predominantly focusing on fully aligned normal dentition. This limitation hampers their applicability in real-world clinical scenarios, where a broader range of dental configurations and anomalies are encountered. In addition, recent weakly supervised and self-supervised methods have garnered widespread attention due to their ability to learn from limited data. DArch68 proposes a weakly annotated training approach to train the segmentation network using only a few annotated teeth from each dental model. However, the first stage of DArch still requires complete centroid annotations, which is challenging to obtain in the absence of dense point-wise annotations. Despite the reduced dependence on manual annotation provided by weakly supervised or semi-supervised methods, supervised approaches generally demonstrate stronger generalization capabilities. Furthermore, when evaluating the outcomes, the segmentation results from supervised are notably more precise, both in terms of accuracy and visual performance, which is essential for clinical dental diagnostics and treatment processes. Due to the fact that most fully supervised methods are based on MLP and convolution, in order to fully leverage the advantages of Transformer, this paper attempts a point cloud segmentation method based on Transformer.
Materials and methods
Theoretical foundation
Deep Learning, a subfield of machine learning, has made significant advancements in recent years across various domains such as image recognition, speech recognition, and natural language processing. The core of this technology lies in constructing multi-layer neural network models that can automatically learn complex features from data through extensive training. With the evolution of deep learning techniques, their potential in processing 3D data has also become increasingly evident, particularly in the field of point cloud segmentation.
Point cloud and mesh are two common methods for representing 3D data. Point cloud, as a collection of points in 3D space, describes the geometric shape of an object by recording the coordinates of each point (typically x, y, z coordinates). Mesh, on the other hand, describe the surface of an object by constructing a network of polygons through vertices, edges, and faces.
In the application of point cloud segmentation, deep learning models typically require a large amount of annotated data for training. This data includes not only the point cloud itself but also the corresponding segmentation labels, indicating which part of the object each point belongs to. Through supervised learning, deep learning models can learn the mapping relationship from point cloud data to segmentation labels. As training progresses, the model’s segmentation capabilities are gradually strengthened, ultimately enabling accurate segmentation of new point cloud data.
Overview
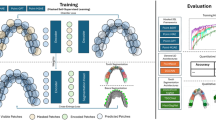
In this paper, we design a PointRegion model for 3D tooth segmentation. By interpreting the point cloud as a tessellation of learnable regions, our model not only exploits Transformer to learn global context information, but also segments tooth in an effective way. The overview of our PointRegion model is shown in Fig. 1.
Overall architecture of PointRegion. The dental mesh is first transformed into point cloud through Mesh2Point, then the RegionPartition module divides the point cloud into several regions and projects these regions into region embeddings. The region embeddings are input to the RegionEncoder module to predict region logits. To establish the association between points and regions, the probabilities that each point belongs to neighboring regions are learned from the intermediate output features (red dashed arrow) of the two modules. Based on the probability matrix, the point-wise class logits are output in a weighted summation manner. Finally, the graph-cut algorithm is further applied to improve segmentation performance.
Since our original dental model is mesh data, we need to convert it into point cloud. In Section Mesh2Point, we first detail the process of converting mesh into points. Then, Section RegionPartition module and Section RegionEncoder module respectively detail our RegionPartition module and RegionEncoder module for region feature learning. In Section Point level segmentation based on point and region association, we describe the method of mapping region-wise prediction to point-wise prediction and the solution to further refine segmentation results.
Mesh2Point
Given an input mesh dental model, we sample it to obtain the point cloud consisting of N points, each of which has d-Dimensional attributes. In the simplest case of \(d=3\), each point is described by 3-Dimensional coordinates of the corresponding mesh cell center. Moreover, it is also possible to include additional information such as the 3-Dimensional normal vector of the cell surface and the 9-Dimensional coordinates of the cell’s three vertices. The values of different attributes vary greatly, therefore, in order to reduce data differences and achieve more stable network optimization, we perform min-max normalization along each dimension of the point cloud data. We denote the normalized point cloud as \({\mathcal {P}}=\{p_i:p_i\in \mathbb {R}^{d},i=1,\dots , N\}\), which serves as the input to the network.
RegionPartition module
Details of RegionPartition module. Black dots represent the sampled points, green solid circles represent the regions which the point cloud is divided into.
Inspired by Dynamic Graph CNN (DGCNN)50, we design a RegionPartition module to divide a point cloud into multiple non-overlapping regions and learn region embeddings from the local-nonlocal point features. As shown in Fig. 2, this module is composed of two branches, the upper branch is used for dividing the point cloud into regions and the lower branch is responsible for extracting regional feature embedding.
Division of regions
Firstly, according to the 3D coordinates of points, we select G points from the original point cloud \({\mathcal {P}}\in \mathbb {R}^{N\times d}\) respectively as the initial center points of G disjoint regions by using the farthest point sampling (FPS) algorithm. We denote the set of these G initial center points as \({\mathcal {C}}^r=\{p_i:p_i\in \mathbb {R}^3,i=1,\dots ,G\}\subset {\mathcal {P}}\), where \(p_i\) is the 3D coordinates of the i-th point. Then, G disjoint regions are formed by different number of points nearest to each selected center point. For now, point cloud \({\mathcal {P}}\) can be represented as \({\mathcal {R}}=\{R_i:i=1,\dots ,G\}\), where \(R_i\cap R_j=\emptyset , \forall i,j\in [1,G], i\ne j\). This region division method is referred to as nearest neighbor clustering (NNC) in our work. This method can help our model avoid conflicts during the point-to-region mapping phase mentioned in Section Point level segmentation based on point and region association.
Extraction of regional feature embedding
Since the point cloud is divided into several regions, it is desirable to significantly increase the receptive field for each point before aggregating point features into embeddings, such that more geometric details of the input point cloud can be more likely preserved. To this end, we adopt cascaded EdgeConv block proposed in DGCNN to learn point features \(F^p=\{f_i^p:f_i^p\in \mathbb {R}^{d_e},i=1,\dots , N\}\), where \(f_i^p\) is a \(d_e\)-Dimensional feature vector for each point \(p_i \in {\mathcal {P}}\). Thanks to dynamic graph update of DGCNN, each point can have a greater and nonlocal receptive field by concatenating multiple EdgeConv blocks. More specifically, suppose that the range of receptive field of each point after the first EdgeConv block is k, then each point can extract information from \(k^2\) points after the second concatenated block. Then, based on the partition of the points from the division of regions branch, we average all the point features in one region as the final region embeddings \(F^e=\{f_i^e:f_i^e\in \mathbb {R}^{d_e},i=1,\dots ,G\}\), where \(f_i^e\) is the region embedding of the region \(R_i\). This can be formulated as Eq. (1) :
where \(m_i\) is the number of points in the region \(R_i\). If the feature \(f_j^p\) in Eq. (1) is 3D coordinates of point \(p_j\), we can update the set of regional center points \({\mathcal {C}}^r=\{p_i^r:p_i^r\in \mathbb {R}^3,i=1,\dots ,G\}\).
RegionEncoder module
In order to predict the class logits for each region, we propose a RegionEncoder module that takes region embeddings as its input. The module is a Transformer-based structure, and we use offset-attention mechanism16 instead of self-attention5 to learn the global context of the point cloud by directly modeling inter-region relations.
Let \(F_{in}\) and \(F_{out}\) be the input and output of offset-attention (OA) block, the OA block can be represented as follows:
where \(W_q\), \(W_k\), \(W_v\) are the shared learnable linear transformation, and Q, K, V are respectively the query, key and value matrices. LBR is a feed-forward neural network composed of Linear, BatchNorm and ReLU layers in sequence. For the normalization of weights in Eq. (3), different from the standard self-attention that adopts a scaling strategy in the batch dimension and softmax in the column dimension, OA uses softmax operations on the row dimension and \(l_1\)-norm on the column dimension. Moreover, the attention features in OA is replaced by the offset between the attention features and the input features in Eq. (4) inspired by graph convolution networks71.
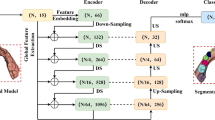
Figure 3 shows the implementation details of RegionEncoder module. The region embeddings \(F^e\) are first projected to a high-dimensional latent space using a shared MLP for the input of a cascaded L layers of OA blocks with residual connection. Then the outputs of all OA blocks are concatenated to be the input of a shared MLP. Hereafter, we can get the region-wise feature representation \(F^{mr}\) formulated as Eq. (5):
where OA\(_l\) represents the l-th OA block as described in Eqs. (2)-(4), and its output \(F_l^{r}=\{f_{l,i}^r,i=1,\dots ,G\}\) serves as the input for the next OA block.
In order to extract global feature vectors g, we introduce global prior information of maxillary and mandible encoded as a one-hot categorical vector according to our dental dataset. This vector is first processed by MLP and then concatenated with the max-pooling and mean-pooling vectors of \(F^{mr}\). In this way, we can effectively avoid assigning maxillary (or mandible) labels to mandible (or maxillary). Following most other segmentation works16,20,32,50, the global features g are repeated G times first, and then concatenated with region-wise representation \(F^{mr}\). The concatenated feature vectors are processed by a region classifier to predict region logits \(Y^r=\{y_i^r:y_i^r\in \mathbb {R}^C,i=1,\dots ,G\}\), where C is the number of semantic labels.
Details of RegionEncoder Module. MaxP and MeanP represent max pooling and mean pooling operations, respectively.
Point level segmentation based on point and region association
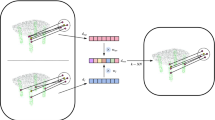
To achieve the goal of point level segmentation, we propose a learnable mechanism to establish the association between each point and all G non-overlapping regions obtained in Section RegionPartition module. This method is suitable for unstructured point clouds and also crucial for achieving fine-grained point cloud segmentation based on coarse-grained region prediction. For each point \(p_i \in {\mathcal {P}}\), we can build its association with \(R_j \in {\mathcal {R}}, j=1,\dots ,G\) and calculate its class logits through the following three steps. The implementation details of the first two steps are shown in Fig. 4.
Searching for K-nearest neighbor regions (SKNR)
For each point \(p_i\), we define its neighboring regions according to Euclidean distances between the point and the centers \({\mathcal {C}}^r\) of all G regions obtained from Eq. (1). Via K-nearest neighbor (KNN) algorithm, we can find K neighboring regions of point \(p_i\) in \({\mathcal {R}}\). We use \({\mathcal {N}}_{p_i} = \{N_k:N_k \in \{1,\dots , G\},k=1,\dots ,K\}\) formulated as Eq. (8) to record the indices of the K neighboring regions of point \(p_i\):
Learning point-to-region probability (LP)
After that, to quantify point-region association, the probabilities \(\{s_k(p_{i},R_{N_{k}}): k=1,\dots ,K\}\) that \(p_{i}\) belongs to its neighbouring regions \(\{R_{N_{k}}:R_{N_{k}}\in {\mathcal {R}},k=1,\dots ,K\}\) need to be learned.19 took the convolution module followed by a softmax to produce normalized probabilities, but it is not feasible for irregular point clouds. In our work, we design a shared function \({\mathcal {M}}(\cdot )\) to learn the probability for each point-region pair based on the similarity between the point feature and its neighboring regional features, and the similarity operation is denoted as \(sim(\cdot )\). This step is formally defined as follows:
where the function \({\mathcal {M}}(\cdot )\) is a shared MLP followed by softmax, W is the learnable weights of the shared MLP and \(f_i^{pm}\) is the point feature of the point \(p_{i}\) obtained from \(f_i^p\) (see Section RegionPartition module) through shared MLPs for efficiency. The feature of the region with the k-th neighbouring region index \(N_k \in {\mathcal {N}}_{p_{i}}\) is selected from one of the outputs of L OA blocks in the RegionEncoder module, for one, \(f_{l,N_k}^{r}\), i.e., the k-th neighboring region feature output by the l-th OA block. All \(s_k(p_{i},R_{N_k})\) form a probability matrix \(S\in \mathbb {R}^{N\times K}\).
Calculating point-wise class logits through weighted summation
Finally, with the help of the region logits \(Y^r\) introduced in Section RegionEncoder module, we calculate the class logits for each point \(p_{i}\), denoted as \(Y^p=\{y_i^p:y_i^p\in \mathbb {R}^C,i=1,\dots ,N\}\), through weighted summation as Eq. (10):
The point \(p_i\) is labeled as the class with the maximum class logits value. To further improve segmentation performance and refine the coarse segmentation boundaries, we use the graph-cut algorithm72 to post-process the segmentation results.
Details of the mechanism of point-to-region association. (a) Illustration of searching for K-nearest neighbor regions, where the red dot stands for a 3D point \(p_{i}\) within the region \(R_g\). (b) Illustration of learning point-to-region probability.
Experimental evaluation
Dataset and metrics
To evaluate our model, we collected a set of tooth mesh models from the real-world clinics, which contain a variety of dental cases, such as missing teeth, tooth deformities and post orthodontic teeth. The dental model dataset is first manually labeled by ten people using Mesh Labeler software (version 3.4)73, and then two people validate the annotated data. The entire dataset consists of 916 dental models (403 maxillaries and 513 mandibles), with each mesh model containing an average of 100,000 faces. We randomly and evenly split it into 815 models for training and 101 for evaluation.
To evaluate the performance of our method, we use the mean intersection over union (mIoU), the overall accuracy (OAcc) as effectiveness evaluation criteria. Besides, we also use the Giga floating point operations (GFLOPs) to measure the efficiency of different methods.
Implementation details
Our segmentation networks are trained with PyTorch on a NVIDIA TITAN RTX 24GB GPU for 200 epochs, we use Adam optimizer with an initial learning rate of \(10^{-3}\) and a weight decay of \(10^{-4}\) to minimize the cross-entropy segmentation loss. For better training, we reduce the learning rate to half of the initial learning rate every 20 epochs.
Due to limitations of GPU memory, it is hard to input all points (\(10^5\) on average) into the network for training. We split all points into three sub-samples including 10240 points using FPS and set batch size as 2. During testing, due to the inconsistent number of points within each sample, in order to obtain their predictions, we also use multiple FPS to get multiple sub-samples with the size of 10240 until the number of the remaining points is less than 20480, and the batch size for testing is set as 1.
Experimental results
Comparison with state-of-the-art architectures
We make comparisons with state-of-the-art semantic segmentation methods on our dental dataset. The experimental results are shown in Table 1. Apart from the pioneering works PointNet32 and PointNet++20, we also choose representative innovative works from each period, including DGCNN50 and PCT16 involved in our model. PointNet and PointNet++, as early point cloud processing networks, exhibit shorter inference time and smaller parameter size, which may limit their ability to learn complex features and thus affect overall performance. Note that our PointRegion outperforms the other methods in terms of both OAcc and mIoU. Especially after boundary optimization using graph-cut algorithm72, we achieve 97.109% OAcc and 92.511% mIoU, respectively, surpassing PCT by 0.789% and 3.547%. In addition, compared to benchmark50 and Transformer-based methods14,16, PointRegion achieves significant improvement in inference time at the cost of a small increase in parameter size, demonstrating good efficiency. All the improvements are attributed to its partitioning strategy, which not only reduces input sequences, effectively reducing computational complexity, but also makes it easier for Transformer to learn differences between regions and get richer representations. Therefore, our RegionPartition module results in better performance.
Ablative analysis
Model hyperparameters
During training process, there are two model hyperparameters, i.e., number of regions, G, in the RegionPartition module and number of nearest neighbor regions, K, when establishing point-to-region association. In order to evaluate the impact of the two model hyperparameters on our model performance, we first fix K to 32 and vary G from 128 to 4096, and then fix G to 1024 while changing K. The results with different G or K are reported respectively in Table 2 and Table 3.
Results in Table 2 show that the performance of our method can be improved as G increases. When G is small, such as \(G=128\), the performance of segmentation is worse, with a gap of 2.39% mIoU from the best result when G is set to 1024. The fewer number of regions means the more diverse categories of points in each region, which makes it difficult to learn discriminative features. Furthermore, too large value of G can also lead to a decrease of OAcc and mIoU. The potential reason may be that excessive input sequences become a burden for modeling the correlation between regions. In the extreme case, where the number of regions is equal to the number of input point clouds (10240), we eliminate the branch of division of regions within the RegionPartition module and Point-to-Region Association mechanism, resulting in a significant increase in memory and computation compared to G=1024. Therefore, we set G to 1024 in the rest experiments.
Unlike regular pixels of 2D image, points of 3D point cloud are disordered, which means that one point may have a different number of neighbouring regions. In our method, we use KNN algorithm to find K regions nearest to each point. As shown in Table 3, our method performs best when \(K=32\). Note that the performance of our method drops significantly if \(K=G\). Therefore, we can get the same conclusion as in this work19 that there is no necessary to consider all regions as neighbouring regions.
Post-process hyperparameter
As well known, the tuning parameter \(\lambda\) in the graph-cut algorithm72 is used to balance the contribution of the data fitting term and the local smoothing term in the optimization objective function. We test the impact of different values of \(\lambda\) on the final results in Table 4. In order to achieve the best effect of boundary refinement, we adjust the value of \(\lambda\) with a step size of 5 within the range of 0 to 60. Note that \(\lambda =5\) can help mIoU and OAcc higher and make the boundary smoother.
Selection of region features
In Section Point level segmentation based on point and region association, we select the output of one of the cascaded L OA blocks as region features for building the point-to-region association. In order to show the impact of the output features at different OA layers on the performance of our method, we conduct a serial of experiments. Considering model size and referring to the setting in PCT16, we set L to 4. As shown in Table 5, for 4 cascaded OA blocks, selecting the output of the second OA block can help our model achieve optimal performance. This can be attributed to the trade-off between increasing the semantic richness of region features and reducing the gap between point and region features.
Partitioning methods
Partitioning methods can affect the efficiency and performance of our model. Inspired by this method19, we first naturally thought of using voxelization to partition the point cloud by analogy with images. However, the sparsity of point clouds makes non-empty voxels sparsely distributed and occupy a considerable portion in the voxel domain, thereby limiting the performance of this method. In comparison, NNC has the following advantages: 1) It can avoid empty regions, where at least one point exists in each region, saving time consumption like processing empty voxels. 2) This method is more flexible. It is not necessary to set the fixed number of points in each area, and clustering is completed completely according to the spatial semantic information of points. So in this experiment, we choose Voxelization and NNC for comparison. For sufficient fairness, we set the resolution of the voxelization method to 8, the number of regions for the NNC method to \(8^3=512\), and set the same number of nearest neighbors for both methods to \(3^3=27\). The experimental results in Table 6 demonstrate that NCC can improve performance without significantly reducing the computational efficiency of our model.
Qualitative results
Point-to-region association
As mentioned in Section Materials and methods, our PointRegion model is able to achieve accurately fine-grained point-level segmentation by establishing the association between each point and corresponding regions. In Fig. 5, we take one region from a source point cloud and visualize the probabilities of all the points associated with this region. Whether the region is located on the tooth or on the gingiva, we found that its association with the relevant points varied inversely with their distances. More specifically, with the distance between the point and the region increasing, the probability of the point belonging to the region becomes lower. This is in line with our expectations and confirms that it is reasonable to achieve point-level segmentation with low memory consumption and computational cost based on region logits and point-to-region probabilities.
Visualization of probabilities between specific region and associated points. For each set, Left: The yellow dots indicate the points located within a specific region, while the blue dots represent a set of region associated points outside the region; Right: Heatmap of the probabilities between a specific region and all its relevant points.
Visualization of segmentation results. Top to bottom: Case 1, 2, 3, 4 and 5. Left to right: Ground Truth, PCT, PointRegion, PointRegion with post-process.
Segmentation results in different dental cases
We visualize the segmentation results of our method and PCT16 in various dental cases, as shown in Fig. 6. Note that our PointRegion can accurately recognize gingiva and individual teeth, and performs well in situations such as the standard teeth, the crowded teeth, the small unerupted teeth, the missing teeth and the asymmetric arch, which demonstrates the effectiveness of our method. From the second and third columns, we can see that our method produces visually more continuous results by a per-region prediction fashion than PCT with dense per-point prediction. For ease of description, we divide maxillary (or mandible) into two quadrants by a midline passing through the teeth. Within the left quadrant, the teeth are numbered from right to left as L1 to L8. Similarly, within the right quadrant, the teeth are numbered from left to right as R1 to R8. Specifically, in Case 2, the crowded arrangement of teeth results in numerous discrete errors in boundary prediction by PCT in the L3-R3 region, with these errors predominantly concentrated around the gingiva adjacent to the tooth root. In the case of missing teeth, Case 4, PCT does not recognize the absence of tooth R2, erroneously assigning the R2 label to the gingiva at that location. In contrast, our PointRegion performs well in both scenarios, and exhibits nearly no erroneous segmentation in the R4-R8 and L4-L8 regions, particularly evident in Case 5 with the asymmetric dental arch. Besides, comparing the segmentation results of the last two columns, the tooth-tooth and teeth-gingiva boundaries are significantly improved and the discrete erroneous predictions are also corrected by the graph-cut algorithm72.
Conclusions
In this paper, we present PointRegion, a novel and efficient Transformer based 3D tooth segmentation model. In order to extend advanced method from 2D image segmentation to 3D point cloud segmentation, we design a RegionPartition module for region embedding and a RegionEncoder module for region prediction, as well as an innovative mechanism for establishing point-to-region association. By interpreting a point cloud as a set of learnable regions, we can apply global-based Transformer to large-scale point cloud dataset at a lower cost. Despite our method exhibits outstanding performance both qualitatively and quantitatively on our dental dataset, segmentation of boundary details remains a challenge. Unsmooth segmentation boundaries and segmentation errors tends to be more common in cases involving extremely crowded teeth, dental calculus and swollen ginigiva. The main reason could be that these case are quite rare, leading to insufficient learning opportunities for the model. The intricate dental arrangements, overlapping structures and noisy data present significant difficulties for accurate segmentation. Although the graph-cut post-processing algorithm can effectively improve the segmentation details at the edges of teeth, the introduction of additional complexity means that more computational resources and time costs are required, which to some extent affects the practicality of the method. Addressing these limitations would help improve the accuracy and efficiency of the dental treatment process. Next, we hope to further explore in terms of the boundary segmentation, and design dental software suitable for professional medical services based on our current work.
Data availability
Due to the involvement of patient privacy, the dental dataset analyzed during the current study is not publicly available. Upon a reasonable request, the corresponding author will provide access to the data supporting the conclusions of this study.
References
Fischler, M. A. & Bolles, R. C. Random sample consensus: A paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 24, 381–395 (1981).
Jiang, X. Y., Meier, U. & Bunke, H. Fast range image segmentation using high-level segmentation primitives. In Proceedings Third IEEE Workshop on Applications of Computer Vision. WACV’96, 83–88 (1996).
Chen, J. & Chen, B. Architectural modeling from sparsely scanned range data. Int. J. Comput. Vis. 78, 223–236 (2008).
Bahdanau, D., Cho, K. & Bengio, Y. Neural machine translation by jointly learning to align and translate. arXiv preprint arXiv:1409.0473 (2014).
Vaswani, A. et al. Attention is all you need. In Proceedings of the 31st International Conference on Neural Information Processing Systems, 6000–6010 (2017).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. Bert: Pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805 (2018).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929 (2020).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 9992–10002 (2021).
Zhao, H., Jiang, L., Jia, J., Torr, P. H. & Koltun, V. Point transformer. In 2021 IEEE/CVF International Conference on Computer Vision (ICCV), 16239–16248 (2021).
Pan, X., Xia, Z., Song, S., Li, L. E. & Huang, G. 3d object detection with pointformer. In 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 7459–7468 (2021).
Wu, L., Liu, X. & Liu, Q. Centroid transformers: Learning to abstract with attention. arXiv preprint arXiv:2102.08606 (2021).
Gao, Y. et al. Lft-net: Local feature transformer network for point clouds analysis. IEEE Trans. Intell. Transp. Syst. 24, 2158–2168 (2022).
Wang, Z., Wang, Y., An, L., Liu, J. & Liu, H. Local transformer network on 3d point cloud semantic segmentation. Information 13, 198 (2022).
Zhang, C., Wan, H., Shen, X. & Wu, Z. Pvt: Point-voxel transformer for point cloud learning. Int. J. Intell. Syst. 37, 11985–12008 (2022).
Lai, X. et al. Stratified transformer for 3d point cloud segmentation. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 8490–8499 (2022).
Guo, M.-H. et al. Pct: Point cloud transformer. Comput. Visual Media 7, 187–199 (2021).
Yu, X. et al. Point-bert: Pre-training 3d point cloud transformers with masked point modeling. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 19291–19300 (2022).
Pang, Y. et al. Masked autoencoders for point cloud self-supervised learning. arXiv preprint arXiv:2203.06604 (2022).
Zhang, Y., Pang, B. & Lu, C. Semantic segmentation by early region proxy. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 1248–1258 (2022).
Qi, C. R., Yi, L., Su, H. & Guibas, L. J. Pointnet++: Deep hierarchical feature learning on point sets in a metric space. In Proceedings of the 31st International Conference on Neural Information Processing Systems, 5105–5114 (2017).
Li, Y. et al. Pointcnn: Convolution on \({\cal X\it }\)-transformed points. arXiv preprint arXiv:1801.07791 (2018).
Wu, W., Qi, Z. & Fuxin, L. Pointconv: Deep convolutional networks on 3d point clouds. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 9613–9622 (2019).
Su, H., Maji, S., Kalogerakis, E. & Learned-Miller, E. Multi-view convolutional neural networks for 3d shape recognition. In 2015 IEEE International Conference on Computer Vision(ICCV), 945–953 (2015).
Maturana, D. & Scherer, S. Voxnet: A 3d convolutional neural network for real-time object recognition. In 2015 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), 922–928 (2015).
Tchapmi, L., Choy, C., Armeni, I., Gwak, J. & Savarese, S. Segcloud: Semantic segmentation of 3d point clouds. In 2017 International Conference on 3D Vision (3DV), 537–547 (2017).
Lawin, F. J. et al. Deep projective 3d semantic segmentation. In Computer Analysis of Images and Patterns: 17th International Conference, CAIP 2017, 95–107 (2017).
Boulch, A. et al. Unstructured point cloud semantic labeling using deep segmentation networks. Eurographics 3, 17–24 (2017).
Tatarchenko, M., Park, J., Koltun, V. & Zhou, Q.-Y. Tangent convolutions for dense prediction in 3d. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 3887–3896 (2018).
Wu, B., Wan, A., Yue, X. & Keutzer, K. Squeezeseg: Convolutional neural nets with recurrent crf for real-time road-object segmentation from 3d lidar point cloud. In 2018 IEEE International Conference on Robotics and Automation (ICRA), 1887–1893 (2018).
Rethage, D., Wald, J., Sturm, J., Navab, N. & Tombari, F. Fully-convolutional point networks for large-scale point clouds. In Proceedings of the European Conference on Computer Vision (ECCV), 596–611 (2018).
Wu, B., Zhou, X., Zhao, S., Yue, X. & Keutzer, K. Squeezesegv2: Improved model structure and unsupervised domain adaptation for road-object segmentation from a lidar point cloud. In 2019 International Conference on Robotics and Automation (ICRA), 4376–4382 (2019).
Qi, C. R., Su, H., Mo, K. & Guibas, L. J. Pointnet: Deep learning on point sets for 3d classification and segmentation. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 77–85 (2017).
Thomas, H. et al. Kpconv: Flexible and deformable convolution for point clouds. In 2019 IEEE/CVF International Conference on Computer Vision (ICCV), 6410–6419 (2019).
Yan, X., Zheng, C., Li, Z., Wang, S. & Cui, S. Pointasnl: Robust point clouds processing using nonlocal neural networks with adaptive sampling. In 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 5588–5597 (2020).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Lecture Notes in Computer Science, Medical Image Computing and Computer-Assisted Intervention - MICCAI Vol. 2015, 234–241 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 770–778 (2016).
Redmon, J., Divvala, S., Girshick, R. & Farhadi, A. You only look once: Unified, real-time object detection. In 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 779–788 (2016).
Ren, S., He, K., Girshick, R. & Sun, J. Faster r-cnn: Towards real-time object detection with region proposal networks. IEEE Trans. Pattern Anal. Mach. Intell. 39, 1137–1149 (2017).
Liu, Z. et al. A convnet for the 2020s. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 11966–11976 (2022).
Ando, A. et al. Rangevit: Towards vision transformers for 3d semantic segmentation in autonomous driving. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, 5240–5250 (2023).
Kong, L. et al. Rethinking range view representation for lidar segmentation. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 228–240 (2023).
Kweon, H., Kim, J. & Yoon, K.-J. Weakly supervised point cloud semantic segmentation via artificial oracle. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 3721–3731 (2024).
Peng, B. et al. Oa-cnns: Omni-adaptive sparse cnns for 3d semantic segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 21305–21315 (2024).
Graham, B., Engelcke, M. & Van Der Maaten, L. 3d semantic segmentation with submanifold sparse convolutional networks. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 9224–9232 (2018).
Choy, C., Gwak, J. & Savarese, S. 4d spatio-temporal convnets: Minkowski convolutional neural networks. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 3070–3079 (2019).
Wang, P.-S., Liu, Y., Guo, Y.-X., Sun, C.-Y. & Tong, X. O-cnn: Octree-based convolutional neural networks for 3d shape analysis. ACM Trans. Graph. 36, 1–11 (2017).
Riegler, G., Osman Ulusoy, A. & Geiger, A. Octnet: Learning deep 3d representations at high resolutions. In 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 6620–6629 (2017).
Klokov, R. & Lempitsky, V. Escape from cells: Deep kd-networks for the recognition of 3d point cloud models. In 2017 IEEE International Conference on Computer Vision (ICCV), 863–872 (2017).
Tian, S. et al. Automatic classification and segmentation of teeth on 3d dental model using hierarchical deep learning networks. IEEE Access 7, 84817–84828 (2019).
Wang, Y. et al. Dynamic graph CNN for learning on point clouds. ACM Trans. Graph. 38, 1–12 (2019).
Zhao, H., Jiang, L., Fu, C.-W. & Jia, J. Pointweb: Enhancing local neighborhood features for point cloud processing. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 5560–5568 (2019).
Ma, X., Qin, C., You, H., Ran, H. & Fu, Y. Rethinking network design and local geometry in point cloud: A simple residual mlp framework. arXiv preprint arXiv:2202.07123 (2022).
Hao, J. et al. Toward clinically applicable 3-dimensional tooth segmentation via deep learning. J. Dent. Res. 101, 304–311 (2022).
Deng, X., Zhang, W., Ding, Q. & Zhang, X. Pointvector: a vector representation in point cloud analysis. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 9455–9465 (2023).
Wu, X., Lao, Y., Jiang, L., Liu, X. & Zhao, H. Point transformer v2: Grouped vector attention and partition-based pooling. Adv. Neural. Inf. Process. Syst. 35, 33330–33342 (2022).
Wu, X. et al. Point transformer v3: Simpler faster stronger. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 4840–4851 (2024).
He, Y. et al. Full point encoding for local feature aggregation in 3-d point clouds. IEEE Transactions on Neural Networks and Learning Systems (2024).
Duan, L. et al. Condaformer: Disassembled transformer with local structure enhancement for 3d point cloud understanding. Adv. Neural Inf. Process. Syst. 36 (2024).
Lai, X., Chen, Y., Lu, F., Liu, J. & Jia, J. Spherical transformer for lidar-based 3d recognition. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 17545–17555 (2023).
Kolodiazhnyi, M., Vorontsova, A., Konushin, A. & Rukhovich, D. Oneformer3d: One transformer for unified point cloud segmentation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 20943–20953 (2024).
Lian, C. et al. Deep multi-scale mesh feature learning for automated labeling of raw dental surfaces from 3d intraoral scanners. IEEE Trans. Med. Imaging 39, 2440–2450 (2020).
Zhang, L. et al. Tsgcnet: Discriminative geometric feature learning with two-stream graph convolutional network for 3d dental model segmentation. In 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 6695–6704 (2021).
Zanjani, F. G. et al. Mask-mcnet: Tooth instance segmentation in 3d point clouds of intra-oral scans. Neurocomputing 453, 286–298 (2021).
Zheng, Y., Chen, B., Shen, Y. & Shen, K. Teethgnn: semantic 3d teeth segmentation with graph neural networks. IEEE Trans. Visual Comput. Graph. 29, 3158–3168 (2022).
Wu, T.-H. et al. Two-stage mesh deep learning for automated tooth segmentation and landmark localization on 3d intraoral scans. IEEE Trans. Med. Imaging 41, 3158–3166. https://doi.org/10.1109/tmi.2022.3180343 (2022).
Li, Z., Liu, T., Wang, J., Zhang, C. & Jia, X. Multi-scale bidirectional enhancement network for 3d dental model segmentation. In 2022 IEEE 19th International Symposium on Biomedical Imaging (ISBI), 1–5 (IEEE, 2022).
Wang, X. et al. Convolutional neural network for automated tooth segmentation on intraoral scans. BMC Oral Health 24, 804 (2024).
Qiu, L. et al. Darch: Dental arch prior-assisted 3d tooth instance segmentation with weak annotations. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 20752–20761 (2022).
Wang, H. et al. Weakly supervised tooth instance segmentation on 3d dental models with multi-label learning. In International Conference on Medical Image Computing and Computer-Assisted Intervention, 723–733 (Springer, 2024).
Almalki, A. & Latecki, L. J. Self-supervised learning with masked autoencoders for teeth segmentation from intra-oral 3d scans. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, 7820–7830 (2024).
Bruna, J., Zaremba, W., Szlam, A. & LeCun, Y. Spectral networks and locally connected networks on graphs. arXiv preprint arXiv:1312.6203 (2013).
Boykov, Y., Veksler, O. & Zabih, R. Fast approximate energy minimization via graph cuts. In Proceedings of the Seventh IEEE International Conference on Computer Vision, 377–384 (1999).
Wu, T.-H. et al.Machine (Deep) Learning for Orthodontic CAD/CAM Technologies, 117–129 (2021).
Author information
Authors and Affiliations
Contributions
Y.W., H.Y. and K.D. conceptualized the work. Y.W. performed experiments and analysed the results together with H.Y. and K.D. Y.W. wrote the first manuscript and revised by H.Y. and K.D. H.Y. and K.D. supervised the project and provided full guidance throughout the process. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Y., Yan, H. & Ding, K. Transformer based 3D tooth segmentation via point cloud region partition. Sci Rep 14, 28513 (2024). https://doi.org/10.1038/s41598-024-79485-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79485-x