Abstract
Pancreatic cancer exhibits a high degree of malignancy with a poor prognosis, lacking effective prognostic targets. Utilizing histopathological methodologies, this study endeavors to predict the expression of pathological features in pancreatic ductal adenocarcinoma (PAAD) and investigate their underlying molecular mechanisms. Pathological images, transcriptomic, and clinical data from TCGA-PAAD were collected for survival analysis. Image segmentation using unsupervised machine learning was employed to extract features, perform clustering, and establish models. The prognostic value of pathological features and associated clinical risk factors were evaluated; the correlation between pathological features and molecular mechanisms, gene mutations, and immune infiltration was analyzed. By clustering 45 effective pathological features, we divided PAAD patients into two groups: cluster 1 and cluster 2. Significant associations with poor prognosis were found for cluster 2 in both the training group (n = 113) and validation group (n = 75) (p = 0.006), with pathological stages II-IV identified as potential synergistic risk factors (HR = 2.421, 95% CI = 1.263–4.639, p = 0.008). Subsequently, through multi-omics correlation analysis, we further revealed a close association between cluster 2 and the oxidative phosphorylation mechanism. Within the cluster 2 group, 28 oxidative phosphorylation genes exhibited reduced expression, CDKN2A gene mutations were upregulated, and there was significant downregulation of Tregs infiltration and related immune gene expression. The pathomic model constructed using machine learning serves as a valuable prognostic target for PAAD. The histopathological features cluster 2 are closely associated with the downregulation of oxidative phosphorylation levels and Tregs immune infiltration.
Similar content being viewed by others
Introduction
Pancreatic cancer, arising from pancreatic duct and acini epithelial cells, is highly malignant1,2,3. It is notorious for its insidious onset, challenging early diagnosis, rapid progression, and poor prognosis, often dubbed the “king of cancers.“4,5,6 The most common subtype, pancreatic ductal adenocarcinoma (PAAD), accounts for 80–90% of cases according to WHO classification7,8. Options for Treating PAAD consist of surgery, radiation, drugs, targeted treatments, and immune system therapy9,10,11,12. Despite these treatments, PAAD is frequently diagnosed late, leading to high recurrence and metastasis rates. Patients face a grim prognosis, as the chances of five years are less than 10%13. Traditional prognostic indicators like tumor stage and invasion depth are insufficient for precision medicine’s needs14. Pancreatic cancer displays diverse histological subtypes and genomic profiles, resulting in varied challenges, outcomes, and responses to treatments. Hence, it is essential to develop a predictive algorithm that can detect patients at high-risk preoperatively by analyzing clinical and genomic features, paving the way for personalized precision treatment.
Oxidative phosphorylation (OXPHOS), as an important process of cellular energy metabolism, has complex interrelationships with various tumors15,16,17. Recent studies have shown that treatment with oxidative phosphorylation inhibitors can improve the anti-tumor response of certain cancers, such as melanoma18, lymphoma19, colorectal cancer20, leukemia21, and PAAD22, by regulating the “Warburg effect”. Abnormal expression of AMP-activated protein kinase (AMPK)23, nuclear factor erythroid 2-related factor 2 (Nrf2)24, mammalian target of rapamycin (mTOR)25, and sirtuin (SIRT) family genes26 directly or indirectly affects the oxidative phosphorylation process, thereby affecting the growth, metabolism, and metastasis of PAAD. Therefore, inhibition of oxidative phosphorylation might offer a safe and effective avenue for pancreatic cancer treatment27. Routine genetic testing may hinder the clinical application and promotion of these genes and molecules due to their high cost. Hence, it is essential to have efficient methods for categorizing the molecular characteristics of pancreatic cancer.
Analyzing PAAD specimens pathologically is essential for classifying patient prognosis. The characteristics displayed in the diseased tissue indicate the collective influence of the tumor surroundings on the behavior of cancer cells. The field of pathology has undergone a major transformation due to the increasing implementation of artificial intelligence28. Pathomics involves utilizing artificial intelligence to transform pathological images into detailed, easily analyzable data, including quantitative aspects like texture, shape, edge sharpness, and biological attributes. It is used to measure pathological diagnosis, molecular expression, and disease prognosis. The integration of histopathology and genomics has been attempted in some cancers, resulting in more precise patient prognostic stratification29,30. However, there have been limited reports on the utilization of computer-aided histopathological analysis in the diagnosis and prognosis prediction of PAAD.
Building upon the aforementioned factors, this study presents a novel approach utilizing pathomics technology and unsupervised machine learning to establish the relationship between clinical pathology, genomic information, and patient outcomes. Furthermore, it integrates bioinformatics analysis to investigate the underlying molecular mechanisms associated with pathomic subtyping.
Materials and methods
Ethical approval
We conducted a secondary analysis of publicly available and accessible data. The research ethics committee at Lihuili Hospital, affiliated with Ningbo University (Approval No. KY2024ML027), granted approval for the implementation of research procedures involving human subjects. All patients provided written informed consents before enrolling, in accordance with the Helsinki Declaration of 1964 and relevant ethical standards.
Image acquisition and processing
We obtained a PAAD patient dataset from The Cancer Genome Atlas (TCGA) database, which includes diverse clinical, H&E-stained histopathological images, and biological information, to be used as the training set (https://tcga-data.nci.nih.gov/tcga/). The clinical dataset from Li Hui Li Hospital, confirmed pathologically as pancreatic ductal adenocarcinoma between 2016 and 2022, was included as an external validation set. All pathological images are panoramic scans of formalin and paraffin-embedded tissue sections, in svs format, with a maximum magnification of 20x or 40x31,32.
The OTSU algorithm was utilized to acquire the tissue regions from the pathological slides. The OTSU algorithm, also referred to as the maximum inter-class variance method, is a threshold algorithm for image binarization that divides the image into two sections: the desired tissue region and the unwanted background for analysis33. The 40x images underwent segmentation into multiple sub-images measuring 1024 × 1024 pixels, while the 20x images were divided into numerous smaller images measuring 512 × 512 pixels and then enlarged to 1024 × 1024 pixels. Following review by pathologists, sub-images with poor quality (e.g., contamination, blurriness, or blank areas exceeding 50%) were excluded. For additional analysis, ten sub-images were chosen at random from each pathological slide31,32,34.
The PyRadiomics open-source package35 was used to standardize and extract original features from the sub-images. A total of 93 of first-order and second-order features were obtained. Additionally, higher-order features (original + wavelet (LL, LH, HL, HH)) were extracted. In total, 465 features were obtained. Subsequently, features were extracted from 10 smaller images within the pathological images of every patient. Each sample’s histopathological feature was determined by taking the corresponding average value. This information was utilized for subsequent analysis29,36,37.
Pathomic feature screening
After excluding four features with a variance of 0, we applied the interquartile range (IQR) normalization method to histopathological features (461 features extracted by the PyRadiomics package)38. The IQR normalization method was employed to eliminate the influence of variable dimensions and variation range. The calculation formula is as follows:
Subsequently, the univariate COX regression analysis method was employed to identify histopathological features associated with prognosis. Features with a p-value < 0.05 in the univariate COX regression analysis were considered to be associated with prognosis.
Unsupervised cluster analysis of pathomic features
NMF (non-negative matrix factorization)39 was used for unsupervised clustering of the aforementioned histopathological features associated with prognosis, resulting in different sample categories and histopathological subtypes. To ensure the stability of the results and the convergence of the algorithm, the clustering method of Brunet was employed during the clustering process, with a rank value ranging from 2 to 6. The optimal rank value was determined based upon the steepest decline of the cophenetic value, with nrun parameter set to 10. Based on the histopathological subtypes clustering, the baseline data of different clinical variables were summarized in both the training and validation datasets.
Survival analysis
We performed overall survival (OS) analysis on patients grouped by different histopathological subtypes. Survival rates among various groups of variables were illustrated through Kaplan-Meier (K-M) survival curves generated by the “survival” package in R. The median survival time represents the survival time at which the survival rate is 50%. The significance of survival rate differences among groups was evaluated using the Log-rank test.
Univariate and multivariate analysis
The Cox model is utilized to examine how one or multiple study variables are related to the likelihood of survival events happening40. Univariate Cox regression analyses were performed separately in the training set and validation set to explore independent factors influencing OS. Furthermore, we performed a multivariable Cox regression to simultaneously investigate the effects of multiple factors on OS. If the Hazard Ratio (HR) exceeds 1, the independent variable is seen as a risk factor; If the HR is less than 1, the independent variable is viewed as a protective factor.
Subgroup analysis and interaction test
We conducted univariate Cox regression to perform survival analysis on subgroups based on different pathological subtypes, aiming to explore the impact of various covariates on patient prognosis within different subgroups. Likelihood ratio test was employed to analyze the interaction effects between different pathological subtypes and other covariates, and the results were visualized by forest plots.
Differential expression genes (DEGs) analysis
Transcriptional data related to the PAAD project was obtained from the TCGA (https://portal.gdc.cancer.gov/) database. The RNA-sequence data in Fragments Per Kilobase per Million (FPKM) format was logarithmically transformed to base 2 (log2). DEGs analysis was conducted between different groups based on different pathological subtype clusters. Genes exhibiting fold change (FC) greater than 1.2 and p less than 0.05 were classified as differentially expressed.
Enrichment analysis
To elucidate the potential role of the target, a functional enrichment analysis of DEGs across distinct pathological subtypes was conducted. The Gene Ontology (GO) is a widely adopted resource for annotating gene functions, particularly encompassing molecular functions (MF), biological processes (BP), and cellular components (CC)41. The top 15 significantly enriched pathways in the enrichment analysis of BP, CC and MF were visualized, using a filtering criterion of corrected p < 0.05 with the Benjamini & Hochberg method.
Immune infiltration analysis
PAAD samples’ gene expression matrix was submitted to the CIBERSORTx database (https://cibersortx.stanford.edu/)42 in order to compute the immune cell infiltration level for each sample. Differences in levels of immune cell infiltration between various pathological subtypes were analyzed using the Wilcoxon rank-sum test. Furthermore, the same statistical method was used to examine the variation in expression of 37 immune-related genes43 among different pathological subtypes, with statistical significance defined as p < 0.05.
Differential expression analysis of OXPHOS-related genes
From the Kyoto Encyclopedia of Genes and Genomes (KEGG) website, we retrieved 121 genes linked to oxidative phosphorylation. The Wilcoxon rank sum test was used to evaluate their expression variances among distinct pathological groups. Genes with p < 0.05 were deemed statistically significant and visualized using a box plot.
Gene mutation analysis
The gene mutation data in the Mutation Annotation Format (MAF) for PAAD samples was downloaded from the TCGA database. The intersection of the downloaded mutation data and pathological data contained 101 samples. The analysis of mutation data and calculation of mutation frequency for each gene in the samples was performed using the maftools R package44. Fisher’s exact test was applied to examine potential variations in gene mutations across various pathological subtypes, with statistical significance defined as p < 0.05. The mutation profile shows the frequency of mutations in cancer-causing genes and typical carcinogenic pathways, with rows representing the reported genes and columns representing patients of each pathological subtype45,46,47,48,49,50,51. The pathway mutation frequency was calculated as follows:
Statistical methods
All statistical analyses and data calculations were carried out using the open-source language and environment for statistical computing and graphics, R software (version 4.1.0; available at https://www.r-project.org/). The threshold for statistical significance was set at a p < 0.05.
Results
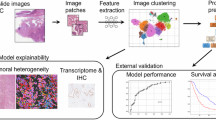
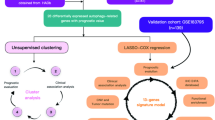
Figure 1 demonstrates the whole design and procedure of this study.
Study design flow chart. Pathological features of PAAD patients were processed for machine learning, survival analysis and bioinformatical analysis. PAAD pancreatic adenocarcinoma.
Data collection and processing
According to the inclusion and exclusion criteria, we obtained a combined total of 113 samples containing full pathological images, gene expression matrices, and clinical details from the TCGA-PAAD dataset, along with 75 samples containing complete pathological images and clinical data from an external dataset (Details were shown in Table S1-S2). The TCGA-PAAD dataset served as the training set, while the external dataset was utilized for validation. Baseline characteristics were summarized for each dataset separately using OS as the grouping variable. No notable variances were discovered among the groups for the eight co-variates in the training dataset (p > 0.05) (Table 1). A notable variation in pathologic stage (p = 0.006) was detected among the groups in the validation dataset (Table 2).
Pathological features clustering
Using univariate Cox regression analysis, we identified 45 pathological features associated with prognosis from a pool of 461 features (see Table S3 for details). Furthermore, during the unsupervised clustering process using NMF, we observed that the cophenetic value dropped most quickly when the number of clusters was two (Fig. 2A). Based on this observation, we divided the aforementioned pathological features into two main clusters, referred to as Cluster 1 and Cluster 2 (Fig. 2B-C).
Unsupervised cluster analysis of pathomic characteristics. (A) NMF rank survey on pathomic features of TCGA-PAAD; (B) 45 screened patholmic features clustering in Consensus matrix; (C) 45 screened patholmic features clustering in heatmap. NMF nonnegative matrix factorization, TCGA the cancer genome atlas.
Survival analysis and risk factor assessment
Patients in the training group were categorized into two clusters, Cluster1 (n = 57) and Cluster2 (n = 56), using NMF. Table 3 presented a summary of the baseline characteristics of clinical variables and significant differences were found in the distribution of age (p = 0.011), diabetes history (p < 0.001), and smoking history (p = 0.022) between the different pathological clusters (Table 3). Similarly, in the validation group, the patients were also categorized into Cluster1 (n = 25) and Cluster2 (n = 50) using NMF, but no notable differences were found in the distribution of clinical variables between the two pathological clusters (p > 0.05) (Table 4).
Survival analysis was conducted, and we found that the median OS for patients in Cluster1 was 24.6 months, while for those in Cluster2, it was 16.03 months in the training cohort, suggesting a significant correlation between Cluster2 and disease progression (p = 0.006) (Fig. 3A). On the other hand, the validation cohort showed no notable disparity in OS between the two groups, with a median OS of 18 months for Cluster1 and 12 months for Cluster2 (p = 0.102) (Fig. 3B).
Survival analysis and risk factor assessment on different pathological cluster groups. (A) Kaplan–Meier curves show a better OS in the cluster1 group than the cluster2 group in the training cohort; (B) Kaplan–Meier curves show no significant difference in OS between the cluster1 group and the cluster2 group in the validation cohort; (C) Univariate Cox regression analysis for the OS of training cohort; (D) multivariate Cox regression analysis for the OS of training cohort; (E) Univariate Cox regression analysis for the OS of validation cohort; (F) multivariate Cox regression analysis for the OS of validation cohort. OS, overall survival. *P < 0.05, **P < 0.01, ***P < 0.001.
Univariate Cox regression analysis revealed that Cluster2 (HR = 2.063, 95% confidence interval (CI) = 1.220–3.486, p = 0.007), pathologic stage of II-IV (HR = 3.703, 95% CI = 1.326–10.344, p = 0.013), and histologic grade of G3&G4 (HR = 1.808, 95% CI = 1.065–3.069, p = 0.028) were potential risk factors for OS in the training cohort (Fig. 3C). However, radiotherapy (HR = 0.447, 95% CI = 0.212–0.943, p = 0.035) was found to be a protective predictor for OS in the training cohort. The consistent findings were observed after multivariate Cox regression analysis. Cluster2 (HR = 2.593, 95% CI = 1.377–4.882, p = 0.003), pathologic stage of II-IV (HR = 3.438, 95% CI = 1.188–9.953, p = 0.023), and histologic grade of G3&G4 (HR = 1.973, 95% CI = 1.125–3.461, p = 0.018) were significant risk factors, while radiotherapy (HR = 0.383, 95% CI = 0.172–0.848, p = 0.018) was found to be a protective factor for OS in the training cohort (Fig. 3D).
In the validation cohort, only pathologic stage of II-IV (HR = 2.425, 95% CI = 1.430–4.112, p = 0.001) was observed to be a risk factor for OS using univariate Cox regression analysis (Fig. 3E). However, after conducting multivariate Cox regression analysis, Cluster2 (HR = 1.773, 95% CI = 1.029–3.057, p = 0.039) and pathologic stage of II-IV (HR = 3.113, 95% CI = 1.737–5.580, p < 0.001) were identified as potential risk factors for OS in the validation cohort (Fig. 3F).
Subgroup risk factor assessment
The subgroup analysis in the training cohort revealed that Cluster2 was a significant risk predictor for OS in the following subgroups: age < 65 (HR = 3.943, 95% CI = 1.789–8.693, p < 0.001), male gender (HR = 2.986, 95% CI = 1.222–7.294, p = 0.016), pathologic stage of II-IV (HR = 1.851, 95% CI = 1.097–3.121, p = 0.021), histologic grade of G3&G4 (HR = 2.550, 95% CI = 1.092–5.955, p = 0.031), absence of radiotherapy (HR = 2.107, 95% CI = 1.222–3.631, p = 0.007), and no smoking history (HR = 3.028, 95% CI = 1.298–7.067, p = 0.01) (Fig. 4A). However, in the validation cohort, Cluster2 was only observed to be a potential risk factor for OS in the subgroups of pathologic stage of II-IV (HR = 2.421, 95% CI = 1.263–4.639, p = 0.008) and perineural invasion (HR = 1.907, 95% CI = 1.024–3.553, p = 0.042) (Fig. 4B).
Subgroup risk factor assessment and co-variates interaction analysis. (A) Univariate Cox regression analysis for the OS of subgroups in training cohort; (B) Univariate Cox regression analysis for the OS of subgroups in validation cohort. *P < 0.05, **P < 0.01, ***P < 0.001.
DEGs and go analysis
Following the predefined threshold criterion, we identified 207 DEGs between the Cluster2 and Cluster1 groups. In the Cluster2 group, there were 64 genes exhibiting increased expression and 143 genes showing decreased expression compared to the Cluster1 group (Fig. 5A). To explore the potential functions of the DEGs further, we performed GO enrichment analysis. Our findings revealed a significant enrichment of DEGs in OXPHOS-related pathways, including fatty acid metabolic process, mitochondrial ATP synthesis coupled electron transport, mitochondrial inner membrane, respiratory chain complex, and oxidoreduction-driven active transmembrane transporter activity (all p < 0.001) (Fig. 5B).
DEGs screening and immune infiltration analysis. (A) Volcano plot of 207 DEGs; (B) GO enrichment analysis of genes in DEGs; (C) Infiltration abundance of 22 immune cell types between different pathological clusters; (D) Differential expression analysis of immune-related genes. DEGs, differential expressed genes; GO, gene ontology. *P < 0.05, **P < 0.01, ***P < 0.001.
Immune-related analysis
We analyzed the presence of 22 types of immune cell in various pathological subcategories. A significantly decreased presence of T cells regulatory (Tregs) was noted in the Cluster2 group of the TCGA-PAAD dataset (p < 0.01) (Fig. 5C). Further analysis of differential expression in 37 immune-related genes indicated a notable reduction in the expression of TMIGD2 (p < 0.05), TNFRSF4 (p < 0.01), and TNFRSF18 (p < 0.01) in the Cluster2 group (Fig. 5D).
OXPHOS-related genes analysis and gene mutation atlas
Through DEGs analysis, we identified significant expression differences in 28 out of 121 OXPHOS-related genes across different pathological subtypes. These genes exhibited down-regulated expression levels specifically in the Cluster2 group, including NDUFB2, ATP5F1D, NDUFB7, NDUFS7, NDUFA2, COX5B, NDUFS6, NDUFB11, ATP5ME, and UQCRFS1 (all p < 0.01) (Fig. 6A).
OXPHOS-related genes analysis and gene mutation atlas. (A) Differential expression analysis of OXPHOS-related genes; (B) Association between different pathological clusters and gene mutation features. OXPHOS oxidative phosphorylation. *P < 0.05, **P < 0.01, ***P < 0.001.
We further compared the gene mutation differences between the two groups. In the TCGA-PAAD dataset, both TP53 and Ras genes exhibited mutation rates exceeding 50% in both groups, with missense mutation being the predominant type. Additionally, PI3K-Akt, Wnt, and p53 signaling pathways also showed mutation rates higher than 50% in both groups. Comparatively, the mutation frequency of the CDKN2A gene was higher in Cluster2 (11/48, 23%) than in Cluster1 (4/53, 8%) (p = 0.048) (Fig. 6B).
Discussion
Despite recent advances in PAAD diagnosis and treatment outcomes driven by the flourishing development of multi-omics studies14,52,53, their translation into clinical practice remains limited, presenting an ongoing challenge to the global burden of PAAD. Molecular subtyping categorizes pancreatic cancer into distinct molecular-level prognoses54, yet it has not yielded adequate benefits for PAAD clinical management. This study leveraged histopathological image features and employed machine learning algorithms to cluster and establish pertinent Cox prognostic models. We identified the poorer prognosis and pertinent potential clinical risk factors for PAAD patients exhibiting cluster 2 features. Additionally, we unveiled differential gene expression, mechanistic pathways, immune infiltration, and gene mutations linked to this prognosis. To the best of our knowledge, the specific associations between pathological characteristics, prognosis, and potential mechanisms in PAAD, as identified through machine learning (NMF) and deep histopathological image analysis methods, have not been previously reported. Our results indicate a significant correlation between the histopathological features-based pathomic model and OS in PAAD patients, facilitating precise prognosis prediction and risk factor assessment. Furthermore, based on these histopathological features, we identified co-expressed modules of OXPHOS-related genes and pathways, gene mutation patterns, and immune infiltrating cells. Integrating pathomics and transcriptomics could enhance prognosis prediction and uncover interactive mechanisms for personalized treatment and risk stratification in clinical practice.
Due to tumor heterogeneity, molecular and genetic testing is now standard for characterizing cancer, particularly in precision oncology14,52,53,54. Variations in molecular expression can affect tissue and nuclear morphology, providing quantitative insights that enhance diagnostic and prognostic accuracy31. Artificial intelligence has bolstered image feature extraction, revealing valuable hidden information linked to tumor characteristics and survival outcomes55,56. Current research has found that histopathological images can predict the genetic mutation status of colorectal cancer57, liver cancer58, lung cancer59, and ovarian cancer60 through machine learning. In this study, we automated the extraction and identification of quantitative morphological features of histopathological sections, including Zernike shape features, using whole slide imaging (WSI) technology and the computer OTSU algorithm. This efficient and accurate method not only confirms the potential practical value of histopathological image analysis in predicting the prognosis of PAAD patients but also significantly reduces human resource costs, making it suitable for routine practice. Therefore, we believe that, in situations where molecular genomic data is limited, pathomics may be an excellent alternative strategy for predicting the prognosis and common mutations of PAAD patients.
Although histopathological images have been successfully utilized in the establishment of prognostic models for various tumors, the application of this submicroscopic information in stratified prognosis prediction for PAAD is unprecedented. In this study, we innovatively classified PAAD patients into two subtypes based on submicroscopic image features, revealing that patients with cluster 2 pathological characteristics had significantly lower OS than those with cluster 1 pathological characteristics (p < 0.01) (Fig. 3A). Similar results were also observed in an external validation set, although the difference did not reach statistical significance possibly due to insufficient clinical sample size. Furthermore, through univariate and multivariate Cox regression models, we identified cluster 2 and pathologic stage II/III/IV as key risk factors for poor prognosis in PAAD patients, both in the training and validation sets. Subgroup analysis of these patients further demonstrated that pathologic stage II/III/IV was a synergistic risk factor with cluster 2, leading to worse prognosis for PAAD patients (p < 0.05) (Fig. 4). However, the correlation between clinical prognosis and histopathological submicroscopic features still requires further validation in larger cohort studies.
The close association between pathological and radiological features and molecular genetics carries a wealth of microscopic-to-clinical intermediary information. Perez-Johnston R et al. also studied the CT-based radiogenomic and pathomic information of clinical stage I lung cancer patients, demonstrating the existence of associations among these three types of information61. In our study, GO enrichment analysis was employed to reveal a strong correlation between cluster 2 group and oxidative phosphorylation processes, such as ATP synthesis coupled electron transport, respiratory chain complex, electron transfer activity, etc.(Fig. 5B). Previous research has indicated the downregulation of OXPHOS in pancreatic cancer15, which is consistent with our findings. Additionally, we found that PAAD patients with cluster 2 subtype exhibited lower OXPHOS levels compared to those with cluster 1 subtype, especially in the expression of 28 OXPHOS-related genes (NDUF subunits, ATP5 subunits, COX subunits, etc.), which were significantly downregulated (all p < 0.05) (Fig. 6A). These findings imply a direct link between the extent of OXPHOS downregulation and poorer OS in PAAD, which probably implies that intervening in the OXPHOS pathway represents a promising therapeutic strategy for pancreatic cancer. Presently, efforts are being made to formulate targeted therapeutic agents for PAAD targeting this pathway22,62.
CDKN2A mutation is considered to be associated with various primary tumors, particularly one of the most common gene mutations in pancreatic cancer63,64. Baek et al.64 found that CDKN2A mutations were associated with poor prognosis in PAAD through gene mutation survival analysis. However, a study analyzing genomic alterations in a cohort of 608 Chinese PAAD patients did not support the prognostic value of CDKN2A mutations65. Nevertheless, our research found that CDKN2A mutations were not only associated with OS in PAAD patients but also closely linked to the poor OS of patients with cluster 2 histopathological subtyping, suggesting a dose-response relationship between the mutation of this gene and PAAD prognosis. However, further in-depth research is needed to investigate the prognostic value of this gene in pancreatic cancer due to limitations in sample sizes and cohorts.
On the other hand, Tregs have long been considered closely associated with the progression of PAAD, playing a crucial role in the tumor microenvironment (TME). However, whether Tregs promote or inhibit pancreatic cancer is complex and contentious66. Previous studies have indicated that high levels of FOXP3+ Treg infiltration may exacerbate the development of pancreatic cancer by inhibiting the function of effector T cells and modulating the immunosuppressive state within the tumor microenvironment66,67,68. However, there is also evidence suggesting that in certain cases, reducing Tregs levels may lead to worsening conditions, as this can trigger the activation of other immunosuppressive mechanisms, such as increased recruitment of myeloid-derived suppressor cells (MDSCs), thereby accelerating tumor progression69,70, or blocking the differentiation of specific types of cancer-associated fibroblasts, thereby promoting tumor growth71. Our findings are consistent with these, as analysis of 22 immune cell infiltrations revealed a significant downregulation of Tregs expression in PAAD patients with cluster 2 subtype (p < 0.05) (Fig. 5C), suggesting its association with poor prognosis. However, the potential reasons for this contradictory phenomenon may lie in the varying impacts of different Treg subpopulations on tumor prognosis72. This necessitates further foundational experiments to identify and validate the previously undisclosed Treg subpopulations associated with pancreatic cancer prognosis. Furthermore, we also observed a significant downregulation of TNFRSF14, TNFRSF18, and TMIGD2, which are closely associated with Tregs infiltration, in the cluster 2 group (p < 0.05) (Fig. 5D). These mutually supportive results indicate that the degree of Tregs infiltration in the tumor microenvironment of PAAD is significantly correlated with prognosis, and targeting this pathway may be an effective approach for anti-tumor immunotherapy73,74.
Although we have pioneered the use of pathomics in pancreatic cancer research, there are still some limitations. Currently, research on the correlation between histopathology and related molecular mechanism is still in its infancy. Data from public databases showed considerable inter-individual image variability. Enhancing the alignment between TCGA and clinical datasets and refining prognostic accuracy are imperative. Achieving this necessitates enhanced submicroscopic image interpretation skills and iterative machine learning models. Secondly, this is a single-center retrospective study, inevitably affected by confounding factors and limited sample size. We plan to collaborate with multiple pathology centers to expand the sample size, further validate the prognostic value of pathomics in PAAD, and develop relevant prognostic models. Finally, only representative sub-images were selected or identified in lesion delineation, which cannot reflect the overall characteristics of WSI, resulting in selection bias. This requires artificial intelligence with stronger computing power to screen and analyze effective information.
In summary, the acquisition of submicroscopic image features of histopathological slides through deep learning is an important biomarker for predicting OS, molecular subtypes, gene mutations, and potential mechanisms in PAAD patients. Our study defined, for the first time, the association between cluster 2 histopathological features and worse OS in PAAD patients, as well as revealed the potential biological mechanisms of the OXPHOS pathway, Tregs depletion, and CDKN2A mutations. The rise of big data, advancements in histopathology, and the growing need for precision medicine will lead to a shift towards integrating pathomics with genomics and proteomics in future research.
Data availability
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding author/s.
Change history
03 December 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41598-024-81889-8
References
Connor, A. A. & Gallinger, S. Pancreatic cancer evolution and heterogeneity: integrating omics and clinical data. Nat. Rev. Cancer 22(3), 131–142 (2022).
Flowers, B. M. et al. Cell of Origin influences Pancreatic Cancer Subtype. Cancer Discov 11(3), 660–677 (2021).
Backx, E. et al. On the origin of pancreatic Cancer: Molecular Tumor subtypes in Perspective of Exocrine Cell plasticity. Cell. Mol. Gastroenterol. Hepatol. 13(4), 1243–1253 (2022).
Park, W., Chawla, A. & O’Reilly, E. M. Pancreat. Cancer: Rev. JAMA 326(9):851–862. (2021).
Klein, A. P. Pancreatic cancer epidemiology: understanding the role of lifestyle and inherited risk factors. Nat. Rev. Gastroenterol. Hepatol. 18(7), 493–502 (2021).
Zhao, Z. & Liu, W. Pancreatic Cancer: a review of risk factors, diagnosis, and treatment. Technol. Cancer Res. Treat. 19, 1533033820962117 (2020).
McGuigan, A. et al. Pancreatic cancer: a review of clinical diagnosis, epidemiology, treatment and outcomes. World J. Gastroenterol. 24(43), 4846–4861 (2018).
Tempero, M. A. NCCN guidelines updates: pancreatic Cancer. J. Natl. Compr. Canc Netw. 17(5.5), 603–605 (2019).
Neoptolemos, J. P. et al. Therapeutic developments in pancreatic cancer: current and future perspectives. Nat. Rev. Gastroenterol. Hepatol. 15(6), 333–348 (2018).
Kolbeinsson, H. M. et al. Pancreatic Cancer: a review of current treatment and Novel therapies. J. Invest. Surg. 36(1), 2129884 (2023).
Gugenheim, J., Crovetto, A. & Petrucciani, N. Neoadjuvant therapy for pancreatic cancer. Updates Surg. 74(1), 35–42 (2022).
Mukherji, R. et al. The role of Immunotherapy in Pancreatic Cancer. Curr. Oncol. 29(10), 6864–6892 (2022).
Humpton, T. J. et al. Oncogenic KRAS induces NIX-Mediated Mitophagy to promote pancreatic Cancer. Cancer Discov 9(9), 1268–1287 (2019).
Dai, L. et al. Pancreatic adenocarcinoma associated immune-gene signature as a novo risk factor for clinical prognosis prediction in hepatocellular carcinoma. Sci. Rep. 12(1), 11944 (2022).
Ashton, T. M. et al. Oxidative phosphorylation as an emerging target in Cancer Therapy. Clin. Cancer Res. 24(11), 2482–2490 (2018).
Evans, K. W. et al. Oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast Cancer. Cancer Res. 81(21), 5572–5581 (2021).
Sica, V. et al. Oxidative phosphorylation as a potential therapeutic target for cancer therapy. Int. J. Cancer 146(1), 10–17 (2020).
Kumar, P. R. et al. Mitochondrial oxidative phosphorylation in cutaneous melanoma. Br. J. Cancer 124(1), 115–123 (2021).
Nie, Y. et al. Targeting metabolic reprogramming in chronic lymphocytic leukemia. Exp. Hematol. Oncol. 11(1), 39 (2022).
Ren, L. et al. PHB2 promotes colorectal cancer cell proliferation and tumorigenesis through NDUFS1-mediated oxidative phosphorylation. Cell. Death Dis. 14(1), 44 (2023).
Amaya, M. L. et al. The STAT3-MYC axis promotes survival of leukemia stem cells by regulating SLC1A5 and oxidative phosphorylation. Blood 139(4), 584–596 (2022).
Xue, D. et al. Multiparameter optimization of oxidative phosphorylation inhibitors for the treatment of pancreatic Cancer. J. Med. Chem. 65(4), 3404–3419 (2022).
Xu, H. et al. AMPactivated protein kinase family member 5 is an independent prognostic indicator of pancreatic adenocarcinoma: a study based on the Cancer Genome Atlas. Mol. Med. Rep. 22(5), 4329–4339 (2020).
Cykowiak, M. & Krajka-Kuźniak, V. Role of Nrf2 in pancreatic Cancer. Antioxid. (Basel) 11(1), 98 (2021).
Cheng, C. S. et al. Functional inhibition of lactate dehydrogenase suppresses pancreatic adenocarcinoma progression. Clin. Transl Med. 11(6), e467 (2021).
Hu, T. et al. Metabolic rewiring by loss of Sirt5 promotes Kras-Induced Pancreatic Cancer Progression. Gastroenterology 161(5), 1584–1600 (2021).
Zhao, Z. et al. The effect of oxidative phosphorylation on Cancer Drug Resistance. Cancers (Basel) 15(1), 62 (2022).
Liu, K. & Hu, J. Classification of acute myeloid leukemia M1 and M2 subtypes using machine learning. Comput. Biol. Med. 147, 105741 (2022).
Nishio, M. et al. Homology-based image Processing for Automatic classification of histopathological images of lung tissue. Cancers (Basel) 13(6), 1192 (2021).
Banna, G. L. et al. The Promise of Digital Biopsy for the Prediction of Tumor Molecular Features and clinical outcomes Associated with Immunotherapy. Front. Med. (Lausanne) 6, 172 (2019).
Chen, L. et al. Histopathological image and gene expression pattern analysis for predicting molecular features and prognosis of head and neck squamous cell carcinoma. Cancer Med. 10(13), 4615–4628 (2021).
Zeng, H. et al. Integration of histopathological images and multi-dimensional omics analyses predicts molecular features and prognosis in high-grade serous ovarian cancer. Gynecol. Oncol. 163(1), 171–180 (2021).
Wang, X. et al. Weakly supervised deep learning for whole Slide Lung Cancer Image Analysis. IEEE Trans. Cybern 50(9), 3950–3962 (2020).
Prasad, R. R. & Udupa, K. BCHisto-Net: breast histopathological image classification by global and local feature aggregation. Artif. Intell. Med. 121, 102191 (2021).
Wang, T. et al. Radiomics for Survival Risk Stratification of Clinical and Pathologic Stage IA pure-solid Non-small Cell Lung Cancer. Radiology 302(2), 425–434 (2022).
Saednia, K. et al. Quantitative digital histopathology and machine learning to predict pathological complete response to chemotherapy in breast cancer patients using pre-treatment tumor biopsies. Sci. Rep. 12(1), 9690 (2022).
Li, H. et al. Integrative Analysis of Histopathological Images and genomic data in Colon adenocarcinoma. Front. Oncol. 11, 636451 (2021).
Habibzadeh, F. Statistical Data Editing in Scientific Articles. J. Korean Med. Sci. 32(7), 1072–1076 (2017).
Nagayama, M. et al. Detecting cell assemblies by NMF-based clustering from calcium imaging data. Neural Netw. 149, 29–39 (2022).
McLernon, D. J. et al. Assessing performance and clinical usefulness in Prediction models with Survival outcomes: practical Guidance for Cox Proportional hazards models. Ann. Intern. Med. 176(1), 105–114 (2023).
Dai, L. et al. A pan-cancer bioinformatic analysis of the carcinogenic role of SMARCA1 in human carcinomas. PLoS One 17(9), e0274823 (2022).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37(7), 773–782 (2019).
Xie, J. et al. A necroptosis-related Prognostic Model of Uveal Melanoma was constructed by Single-Cell Sequencing Analysis and Weighted Co-expression Network Analysis Based on Public Databases. Front. Immunol. 13, 847624 (2022).
Mayakonda, A. et al. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28(11), 1747–1756 (2018).
Morris, J. P. 4th, Wang, S. C., Hebrok, M. K. R. A. S. & Hedgehog Wnt and the twisted developmental biology of pancreatic ductal adenocarcinoma. Nat. Rev. Cancer 10(10), 683–695 (2010).
Neuzillet, C. et al. Targeting the Ras-ERK pathway in pancreatic adenocarcinoma. Cancer Metastasis Rev. 32(1–2), 147–162 (2013).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518(7540), 495–501 (2015).
Zhao, M., Mishra, L. & Deng, C. X. The role of TGF-β/SMAD4 signaling in cancer. Int. J. Biol. Sci. 14(2), 111–123 (2018).
Hu, C. et al. Association between inherited germline mutations in Cancer Predisposition genes and risk of pancreatic Cancer. JAMA 319(23), 2401–2409 (2018).
Hayashi, H. et al. Biological significance of YAP/TAZ in pancreatic ductal adenocarcinoma. Front. Oncol. 11, 700315 (2021).
Stanciu, S. et al. Targeting PI3K/AKT/mTOR signaling pathway in pancreatic Cancer: from Molecular to clinical aspects. Int. J. Mol. Sci. 23(17), 10132 (2022).
Yan, C. et al. System analysis based on the pyroptosis-related genes identifies GSDMC as a novel therapy target for pancreatic adenocarcinoma. J. Transl Med. 20(1), 455 (2022).
Zhang, B. et al. Integrated multi-omics identified the novel intratumor microbiome-derived subtypes and signature to predict the outcome, tumor microenvironment heterogeneity, and immunotherapy response for pancreatic cancer patients. Front. Pharmacol. 14, 1244752 (2023).
Huang, X. et al. Identification of tumor antigens and immune subtypes of pancreatic adenocarcinoma for mRNA vaccine development. Mol. Cancer 20(1), 44 (2021).
Romo-Bucheli, D. et al. Automated tubule nuclei quantification and correlation with Oncotype DX risk categories in ER + breast Cancer whole slide images. Sci. Rep. 6, 32706 (2016).
Moon, W. K. et al. Computer-aided prediction of axillary lymph node status in breast cancer using tumor surrounding tissue features in ultrasound images. Comput. Methods Programs Biomed. 146, 143–150 (2017).
Kather, J. N. et al. Predicting survival from colorectal cancer histology slides using deep learning: a retrospective multicenter study. PLoS Med. 16(1), e1002730 (2019).
Liao, H. et al. Classification and prognosis prediction from Histopathological Images of Hepatocellular Carcinoma by a fully automated Pipeline based on machine learning. Ann. Surg. Oncol. 27(7), 2359–2369 (2020).
Niu, Y. et al. Predicting Tumor Mutational Burden from Lung Adenocarcinoma histopathological images using deep learning. Front. Oncol. 12, 927426 (2022).
Miyagawa, C. et al. Histopathological subtyping of high-grade serous ovarian cancer using whole slide imaging. J. Gynecol. Oncol. 34(4), e47 (2023).
Perez-Johnston, R. et al. CT-based Radiogenomic Analysis of Clinical Stage I Lung Adenocarcinoma with histopathologic features and oncologic outcomes. Radiology 303(3), 664–672 (2022).
Xu, Y. et al. First-in-Class NADH/Ubiquinone Oxidoreductase Core Subunit S7 (NDUFS7) antagonist for the treatment of pancreatic Cancer. ACS Pharmacol. Transl Sci. 6(8), 1164–1181 (2023).
Wang, J. Z. et al. Increased mRNA expression of CDKN2A is a transcriptomic marker of clinically aggressive meningiomas. Acta Neuropathol. 146(1), 145–162 (2023).
Baek, B. & Lee, H. Prediction of survival and recurrence in patients with pancreatic cancer by integrating multi-omics data. Sci. Rep. 10(1), 18951 (2020).
Zhan, Q. et al. Identification of copy number variation-driven molecular subtypes informative for prognosis and treatment in pancreatic adenocarcinoma of a Chinese cohort. EBioMedicine 74, 103716 (2021).
Saleh, R. & Elkord, E. FoxP3 + T regulatory cells in cancer: prognostic biomarkers and therapeutic targets. Cancer Lett. 490, 174–185 (2020).
Liu, M. F. et al. Helios serves as a suppression marker to reduce regulatory T cell function in pancreatic cancer patients. Immunol. Res. 69(3), 275–284 (2021).
Su, K. F., Peng, Y. & Yu, H. C. Prognostic value of regulator T cells in patients with pancreatic cancer: a systematic review and meta-analysis. Eur. Rev. Med. Pharmacol. Sci. 26(8), 2906–2917 (2022).
Zhang, Y. et al. Regulatory T-cell depletion alters the Tumor Microenvironment and accelerates pancreatic carcinogenesis. Cancer Discov 10(3), 422–439 (2020).
Aykut, B., Chen, R. & Miller, G. Regulatory T cells keep pancreatic Cancer at Bay. Cancer Discov 10(3), 345–347 (2020).
Ohlund, D. et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 214(3), 579–596 (2017).
Saito, T. et al. Two FOXP3(+)CD4(+) T cell subpopulations distinctly control the prognosis of colorectal cancers. Nat. Med. 22(6), 679–684 (2016).
Knee, D. A., Hewes, B. & Brogdon, J. L. Rationale for anti-GITR cancer immunotherapy. Eur. J. Cancer 67, 1–10 (2016).
Chen, Q. et al. B7-H5/CD28H is a co-stimulatory pathway and correlates with improved prognosis in pancreatic ductal adenocarcinoma. Cancer Sci. 110(2), 530–539 (2019).
Funding
This study was supported by Ningbo Top Medical and Health Research Program (2024020818), Municipal Key Technical Research and Development Program of Ningbo (2023Z160), Natural Science Foundation of Ningbo (2023J221) and Medical and Health Research Project of Zhejiang Province (2024KY1480).
Author information
Authors and Affiliations
Contributions
Conception and writing-original draft preparation, L.D and L.J.P; Charting and methodology, G.H.L; data analysis and visualization, L.D; reference acquisition, Y.L and H.L; comments and suggestions, J.H; manuscript revision and funding acquisition, L.D. All the authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article Yang Li and Lujuan Pan were omitted as equally contributing authors. Full information regarding the corrections made can be found in the correction for this Article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Pan, L., Mugaanyi, J. et al. Pathomic and bioinformatics analysis of clinical-pathological and genomic factors for pancreatic cancer prognosis. Sci Rep 14, 27769 (2024). https://doi.org/10.1038/s41598-024-79619-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-79619-1
Keywords
This article is cited by
-
Dysregulated expression of cell cycle regulators CDC20, PLK1, BUB1, CDC45, CDCA5 in pancreatic ductal adenocarcinoma
Scientific Reports (2026)
-
Research progress in diagnosis and treatment of pancreatic cancer with mismatch repair and microsatellite instability
Clinical and Translational Oncology (2026)