Abstract
Antiphospholipid syndrome (APS) is associated with recurrent pregnancy morbidity, yet the underlying mechanisms remain elusive. We performed multifaceted characterization of the biological and transcriptomic signatures of mouse placenta and uterine natural killer (uNK) cells in APS. Histological analysis of APS placentas unveiled placental abnormalities, including disturbed angiogenesis, occasional necrotic areas, fibrin deposition, and nucleated red blood cell enrichment. Analyses of APS placentas showed a reduced cell proliferation, lower protein content and thinning of endothelial cells. Disturbances in APS trophoblast cells were linked to a cell cycle shift in cytotrophoblast cells, and a reduced number of spiral artery-associated trophoblast giant cells (SpA-TGC). Transcriptomic profiling of placental tissue highlighted disruptions in cell cycle regulation with notable downregulation of genes involved in developmental or signaling processes. Cellular senescence, metabolic and p53-related pathways were also enriched, suggesting potential mechanisms underlying placental dysfunction in APS. Thrombotic events, though occasionally detected, appeared to have no significant impact on the overall pathological changes. The increased number of dysfunctional uNK cells was not associated with enhanced cytotoxic capabilities. Transcriptomic data corroborated these findings, showing prominent suppression of NK cell secretory capacity and cytokine signaling pathways. Our study highlights the multifactorial nature of APS-associated placental pathologies, which involve disrupted angiogenesis, cell cycle regulation, and NK cell functionality.
Similar content being viewed by others
Introduction
Reproductive health is crucial to women’s physical and emotional well-being. The presence of antiphospholipid antibodies (aPL), the serological hallmark of antiphospholipid syndrome (APS), is a well-recognized factor associated with reproductive failure1. Antibodies targeting β2GPI are increasingly accepted as the most, if not the only, clinically relevant in APS2. aPL can bind to various cells within placental tissue, including trophoblasts, loose connective tissue cells, Hofbauer and NK cells, utilizing a range of receptors such as TLR2, TLR4, apolipoprotein E receptor 23. Furthermore, β2-Glycoprotein I (β2GPI) is constitutively expressed on the cell surface by all placental trophoblast subpopulations and maternal endothelial cells4, rendering the placenta a major target for β2GPI-dependent pathogenic aPL binding.
In accordance with recent estimates, adverse pregnancy outcomes reported to occur at a rate of 5.6% in the group of aPL-negative women, as high as 41.9% in women with defined APS, 25% in non-criteria APS, and 28.3% in aPL-carriers5. Pregnant women with aPL face elevated risks of thrombosis, recurrent pregnancy loss, and various pregnancy related complications6. Despite significant advancements in the understanding of obstetrical APS in recent decades, many questions remain unanswered to date. Numerous theories and hypotheses exist for placenta-related pathophysiology of adverse outcomes in APS pregnancy, encompassing inflammatory processes in the decidua, deposition of active components of complement, an increase in syncytial knots, hypercoagulation and thrombosis7. Inadequate vascular remodeling resulting from inflammation-induced trophoblast or endothelial dysfunction may cause insufficient delivery of oxygen and nutrients to the developing placental tissue and eventually to the fetus8. In APS, poor placental angiogenesis and vascular pathological events such as impaired spiral artery remodeling, intimal thickening, and fibrinoid necrosis are believed to be secondary to aPL-mediated inflammation or thrombosis7. Another evidence suggests that direct aPL-mediated targeting of the placenta via β2GPI abundantly expressed on the surface of trophoblast cells may reduce decidual and vascular trophoblast invasion4.
Healthy pregnancy poses a unique immunological challenge where the maternal immune system ensures a tolerance towards the semi-allogeneic fetus while preserving the resistance against infection9. Over the last ten years, different types of immune cells have been considered to be involved in the establishment of maternal-fetal immune tolerance. In this context, natural killer (NK) cells play an intriguing role during pregnancy. Uterine NK (uNK) cells form the most abundant immune cell type, comprising approximately 70% of the total lymphocytes that occupies the maternal-fetal interface and facilitates implantation and maintenance of pregnancy both in human and mice pregnancy10. In contrast to peripheral blood NK cells (pNK), uNK cells are poorly cytotoxic, expressing killer cell immunoglobulin-like receptors and releasing numerous soluble factors that promote trophoblast invasion, tissue remodeling, embryonic development, and placentation11. Our recent study has shown that NK cells might be considered as candidate cells for APS-related obstetric complications12, however detailed investigation of uNK cells is required to elucidate their potential role in APS pregnancy.
Though there has been progress in obstetrical APS research, additional studies are required to cover open issues. Research studies on the human placenta encounter a number of ethical and technical issues, which can be effectively addressed with the use of animal models. The mouse is an excellent model organism to study mammalian biology and widely utilized for investigations of pregnancy progression. Thus, we established a mouse model of APS to explore potential disruptions in placental tissue development during gestation using a multi-pronged study of endothelial and trophoblast cells. We also aimed to generate a detailed map of transcriptomic, phenotypic, and functional diversity of uNK cells, shedding light on immune mechanisms causing pregnancy complications in APS.
Results
Serological, coagulatory and clinical criteria of APS animals
Before conducting the experiments (Fig. 1A), initial observations in mice revealed a decrease in the placenta weight (p < 0.05), size and decreased live birth rate in a group of immunized (APS) mice compared with non-immunized (Ctl) (Fig. 1E, F,H). Rates of maternal (28%) and neonatal (49.8%, p < 0.05) survival in the APS group were lower than in the control group (Fig. 1I, J). Resorption rate in the APS group was tended to be higher than in the control group, albeit not significantly (Fig. 1G). These characteristics may potentially serve as surrogate markers of maternal malperfusion and a predictor for abortion or stillbirth, and additionally serve as a criterion for a successful APS model.
Assessment of APS pregnancy outcomes. (A) Schematic representation of study design. Applied immunization, mating and termination time-point are indicated with arrows. Blood sampling for aPL titers monitoring (asterisk) was performed before any other manipulation. (B) Time-dependent anti-β2GPI response in immunized mice. Data presented as normalization to values in non-immunized animals, multiplied by a dilution factor. (C) Tissue factor (TF, CD142) surface expression on THP-1 cells was analyzed using flow cytometry after exposure to aPL− and aPL + IgG at 50 ng/ml for 4 h. (D) Relative mRNA levels of TF gene in PBMCs isolated from peripheral blood of mice. Data was calculated with 2−ΔΔCT method. Dotted line represents the control value of 1. (E) Representative macroscopic appearances of control and APS placentas. Infarct areas and hemorrhages are observed widely in APS. (F) Clinical assessment of the APS model was performed on embryonic days 12.5 by weighting a single placenta. (G) Total resorption rate determined from the difference between the number of conceptus and viable embryos. (H) Postpartum assessment included calculating number of pups per litter. (I) The percentage of litters survival. (J) The Kaplan–Meier survival estimates 96-day maternal postpartum survival.
To further validate the clinical relevance of APS model animals, we assessed the potential of aPL + IgG, generated in response to β2GPI immunization, to induce procoagulatory responses. Increased expression of TF (tissue factor, CD142) by monocytes or endothelial cells is one of main procoagulatory effects precipitated by aPL autoantibodies13. We observed a significant increase of TF expression on THP-1 cells upon exposure to aPL + IgG compared to aPL- treated or control unstimulated cells (Fig. 1C). In parallel, we measured TF expression in peripheral blood mononuclear cells (PBMCs) isolated from peripheral blood of experimental mice. As appeared, PBMCs from immunized mice exhibited higher mRNA levels of TF (p < 0.01) as compared with the control group (Fig. 1D).
Monitoring of anti-β2GPI titres in immunized mice was performed weekly with the use of serum samples obtained by tail vein puncture. Steady rise of antibodies titers was observed from 7th day post first immunization and reached its maximum level at 35th day following first immunization. Steady high titers of aPL were observed for up to 47 days (Fig. 1B). Animals which did not demonstrate signs of pregnancy during this period were excluded from the experiment.
Histological evidences of placental pathology in APS mice
Histological analyses uncovered inter-individual variability in the placentas of APS mice, indicating heterogeneity in their characteristics and pathologies. Histologic patterns of placental vascular malperfusion have been detected. Hematoxylin and eosin (H&E) staining of APS placenta sections at day 12.5 of pregnancy demonstrated a fibrin deposition in the lateral edge and decidua and small areas of disintegration and necrosis in the junctional zone close to decidua (Fig. 2B). Fibrin depositions were also present in the border between junctional and trophoblast giant cell (TGC) areas. The number of vessels was comparable in both studied experimental groups. Thin vessels of the junctional zone and signs of emerging fibrin thrombus were infrequently visualized in the APS group. Occasional thrombotic events in vessels of APS placenta were also detected (Fig. 2A).
Microscopic appearances of H&E stained control and APS placentas. (A) Fibrin depositions in APS blood vessels marked with (*). (B) Diffuse trophoblast necrosis in the APS junctional zone is outlined with a dotted-line. Spiral artery-associated trophoblast giant cells (SpA-TGC) are shown in white arrows. (C) Nucleated erythrocytes (black arrows) in the labyrinth zone. (D) Uterine NK cell infiltrations in decidua. APS uNK cells (white arrows) showed low granularity and partial signs of degranulation.
In APS placenta, histological analysis revealed particular enrichment of nucleated (immature) red blood cells in the vessels within the labyrinth (Fig. 2C), a feature characteristic for throughout early stages of gestation. Elevated number of nucleated red blood cells at late stages of gestation can be associated with numerous complications including hypoxia, prenatal brain damage, preterm birth, respiratory distress syndrome and preeclampsia14. In sections from control animals, occasional presence of nucleated red blood cells was also observed.
Organ-related changes in NK cells
The absolute number of total NK cells derived from tissues (blood, spleen, placenta) and their subpopulational distribution were analyzed using flow cytometry. The gating strategy for NK cell identification is shown in Fig. 3A. The analysis showed an increased absolute number of NK cells in the APS mice group across all studied organs compared with the control group (Fig. 3B, C). Importantly, the count of uNK cells was validated through histological section analysis, which also demonstrated the equal higher distribution of NK cells in different compartments of placenta, including maternal gland, junctional zone, and decidua (Figs. 2D, 3B). uNK cells were identified based on their specific morphology, featuring large, round nuclei, abundant cytoplasm, and characteristic dark pink cytoplasmic granules. Subpopulation analysis revealed an increased percentage of cytotoxic CD11highCD27low circulating NK cells in APS mice compared with the control group (Fig. 3D). However, there was no significant difference in distribution of cytotoxic NK cells in the spleen and placenta, as well as CD11lowCD27high IFNγ-producing NK in all investigated tissues (Fig. 3D, E).
Distribution and characteristics of mouse NK cells from blood, spleen and placenta. (A) Gating strategy for flow cytometric analysis to identify NK cell subsets. Following doublets exclusion and lymphocyte gate selection by forward (FSC) and side scatter (SSC) properties, NK cells were determined as CD3-CD49b+. NK cell subsets were selected as follows: CD11lowCD27high cytotoxic NK cells and CD11highCD27low IFNγ-secreting NK cells. Analysis of pNK cells was performed with the use of peripheral blood isolated PBMC. sNK and uNK cell were isolated with EasySep™ Mouse NK Cell Isolation Kit. (B) Quantification of uNK cells performed by delineating placental zones using hematoxylin and eosin (H&E) stained histological sections. For each section, the cells were counted in 10–15 random fields. (C) Quantification of pNK and sNK cells using cytometry data with absolute cell counts. (D) Distribution of cytotoxic (CD11lowCD27high) NK cells in studied organs. (E) Distribution of IFNγ-producing (CD11highCD27low) NK cells in studied organs. (F) Cytotoxic abilities of placenta-derived NK cells directed towards THP-1 target cells at ratio 5:1 measured by the direct killing assay, employing THP-1 cells as the target cells and LDH assay. (G) IFNγ production assessed by measuring its levels in the supernatants of sNK and uNK cells that were pretreated with IL-2 for 24 h.
Next, we aimed to analyze whether the changes in the number of NK cells impact their functionality. We assessed cytotoxic activity using two complementary methods: the LDH assay and the direct killing assay, employing THP-1 cells as the target cells. As appeared, the increased number of uNK cells (Fig. 3B) does not correlate with the alterations in cytotoxic activity (Fig. 3F). The absence of differences in the number of IFNγ-producing NK cells paralleled the lack of disparity observed in the supernatants of cultured NK cells and plasma samples (Fig. 3E, G).
Fluorescent staining
A summary of Ki-67, VEGF-A, and DAPI expression in the control and APS placenta sections are presented in Fig. 4A, B. We evaluated the immunoreactivity of placental sections for Ki-67, a marker expressed in all phases of the cell cycle except the resting phase G0. In general, placentas from both studied groups exhibited low levels Ki-67-defined cytotrophoblast proliferation, which correspond to the gestational age. In APS placentas, there was no evidence of a significantly altered number of trophoblasts expressing Ki-67 compared to control placentas. Angiogenesis-related VEGF-A expression was observed in different compartments of the maternal and fetal placenta. VEGF-A expression was found to be lower in placentas from the APS group in comparison with control ones, suggesting compromised angiogenic processes in APS.
Immunohistochemical and immunofluorescent staining of placental sections from control and APS mice. (A) Sections were stained with Ki67 (green) and counterstained with DAPI (blue). (B) Sections were stained with VEGF-A (red) and counterstained with DAPI (blue). Three placental sections from each experimental group were examined. Magnification x40. Scale bar = 100 μm. (C) Sections of placental tissues stained with combined Feulgen–Naphthol Yellow. Arrows point to the endothelial cells. Magnification x100. (D) Cell cycle phases of endothelial and trophoblast cells measured using micro-spectrophotometry. The histograms represent the percentage of cells in each phase of the cell cycle: G0/G1 (blue), S (yellow) and G2/M (red). (E) Violin plots displaying protein levels in endothelial and trophoblast cells of control and APS placentas, measured with micro-spectrophotometry.
Cytophotometric analysis of DNA and protein cellular content in placental endothelial cells and trophoblasts
Spectrophotometric analysis of DNA content in Feulgen-Naphthol Yellow stained sections revealed distinct patterns in endothelial cells from control and APS animal groups (Fig. 4C). In control placentas, the majority of endothelial cells exhibited diploid DNA content, corresponding to the G1 stage of the cell cycle. 34% of the total cell population showed intermediate values indicative of the S stage, while about 2% of cells were tetraploid (G2 stage). In the APS group, the number of cells in the S phase of the cell cycle was decreased and constituted 26% of total cells. The number of tetraploid cells was higher (6%) than in the control group, which may suggest potential arrest of the cells in the G2/M phase (Fig. 4D). Generally, endothelial cells in both groups exhibited heterogeneity in terms of protein content, ranging from 28 to 190 pg/cell. In the APS group, endothelial cells were mainly with lower protein content compared with the control group (81.27 vs. 97.31 pg/cell, correspondingly) (Fig. 4E). A shortage of proteins may potentially result in defective DNA replication dynamics, impairing the accuracy and consistency of DNA replication15.
Regarding trophoblast cells, there was a slight difference in protein content. Compared with the control group, APS cytotrophoblast (81.45 vs. 75.86 pg/cell, correspondingly) contained lower amounts of protein (Fig. 4E). There were also detected differences in trophoblast cell ploidy between APS and control groups. Namely, a proportion of APS cytotrophoblasts in S phase was found to be reduced in comparison with the control group (9% vs. 27%, correspondingly) (Fig. 4D).
Transcriptome profiles of placentas from control and APS mice
With the goal of determining the changes in expression profiles in APS placentas, we collected placental tissues from APS (n = 7) and control (n = 7) animals and conducted bulk RNA-Seq analysis. Abundant up- and down-regulated differentially expressed genes (DEGs) were discovered between APS and control groups (Fig. 5). Analysis of DEGs up-regulated in APS placenta revealed that top enriched GO biological processes (GO: BP) terms almost exclusively include processes related to regulation of cell cycle, including cell cycle, chromosome segregation, mitotic cell cycle, nuclear division and sister chromatid segregation. The analysis of down-regulated genes showed enrichment of terms mainly related to developmental processes, multicellular organism development, anatomical structure development and animal organ development (Supplementary Fig. S1). Molecular functions (GO: MF) of DEGs up-regulated in APS were primarily related to nucleotide and purinergic metabolisms, including heterocyclic compound binding, nucleotide binding, ATP and anion binding (Online Supplementary Table S2). GO: MF terms of DEGs down-regulated in APS revealed predominantly metabolism related terms such as protein binding, hormone activity, signaling receptor regulator activity and prolactin receptor binding (Online Supplemental Fig. S1).
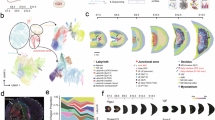
Transcriptomic analysis of placenta and uNK cells in APS and control mice. (A) The volcano plot analysis displaying a notable alteration in the expression of 1072 protein-coding genes (476 were up-regulated and 596 down-regulated) in placenta and 1696 genes (440 up-regulated and 1256 down-regulated) in uNK samples. For both groups, meeting criteria was log2 fold change (log2FC) ± 1 and adjusted p < 0.05. (B) Principal component analysis (PCA) plot illustrating distinctions based on gene expression in experimental groups. PCA identified two clusters in the data separated along the first and second principal components. (C) Heatmaps depicting hierarchical clustering of all differentially expressed genes (DEGs) and variability between expression profiles of mice from control and APS groups.
Transcriptome profiles of uNK cells from control and APS mice
Next, to test whether transcriptome alterations in the placenta between APS and control animals could be driven by abnormalities in uNK cells, we conducted RNA-Seq analysis of uNK cells from APS and control placentas. Analysis of biological function of up- and down- regulated genes in uNK cells revealed substantial similarities with significant GO: MF and GO: BP terms found in placental samples, particularly up-regulation of cell cycle machinery and down-regulation of developmental processes (Fig. 5). List of top deregulated terms presented in Online Supplementary Fig. S2. It is notable that the number of down-regulated genes notably exceeded that of the up-regulated genes in APS uNK. The common DEGs displayed significant enrichments in various processes associated with NK cell effector function. Particularly, 100% of deregulated uNK cells genes enriched in KEGG Lysosome pathway (mmu04142) and 92% of genes enriched in Protein processing in endoplasmic reticulum (mmu04141) pathway were down-regulated in APS (Online Supplementary Fig. S3). Similarly, total suppression was observed for genes enriched in KEGG Cytokine-cytokine receptor interaction pathway (mmu04060) (Online Supplementary Fig. S3).
DEGs associated with blood clotting in APS placenta were not significantly enriched
Considering the procoagulatory nature of APS, we expected enrichment in coagulation related pathways. However, according to functional enrichment analysis, as assessed by g: Profiler tool, “homeostatic processes” (GO:0042592) was the only related term found among the significantly involved GO: BP terms when the total list of up- and down-regulated placental DEG being analyzed. Among genes comprising KEGG Complement and coagulation cascades (mmu04610) pathway, only three genes (F11, Bdkrb2 and F2rl2) were found deregulated in APS placentas compared to control placentas.
Expression pattern of accelerated cell senescence in APS
One of the most enriched KEGG pathways identified in APS placentas and uNK was Cellular senescence (mmu04218). Cellular senescence is a state of irreversible proliferation arrest, accompanied with morphological, metabolic and transcriptional changes. Trophoblast senescence is considered to be physiological and gradually progresses as pregnancy advances to term. However, placental stress-driven premature or accelerated cell senescence may lead to several adverse pregnancy outcomes, such as preeclampsia and restricted fetal growth16. In APS placentas, a total of 17 significant senescence related DEGs were detected. In uNK from APS animals a totally 20 genes from genes involved in KEGG Cellular senescence were found to be deregulated (Online Supplementary Fig. S1, S2) The enrichment of this pathway was expected, given the deregulation of genes involved in signaling pathways known to induce cell cycle arrest and senescence. These include p53, p38/MAPK, PI3K/AKT, JAK2/STAT3 which expressions were found highly variable in the APS group17.
Expression pattern of motor proteins in APS placenta
Among a total of 32 kinesin proteins encompassing the KEGG Motor proteins (mmu04814) pathway, expression of 9 kinesins was up-regulated and 3 down- regulated in both the APS placentas and uNK cells (Online Supplementary Fig. S4). An increased expression pattern of kinesins on trophoblasts or villi endothelium was found in patients with preeclampsia or diabetes, suggesting that these proteins may be involved in placental pathologies18. Members of the kinesin motors family play a central role in controlling cell cycle and maintenance of cell growth, the processes that are disrupted in the APS placenta19.
Expression pattern of pathways related to metabolism and hypoxia in APS
KEGG pathway analysis of the total list of APS placental DEGs identified Metabolic pathway (mmu01100) as the most enriched term, with 74 genes comprising this pathway being differentially expressed (Online Supplementary Table S2). Particularly, “Carbon metabolism” and “Pyrimidine metabolism” were found mainly affected. Metabolic reprogramming of the placenta might partially appear as a compensatory mechanism for the chronic reduction in maternal oxygen supply20.
Among significantly enriched terms in APS placenta, a few respiration and oxygen carrying/transporting pathways, namely GO: CC hemoglobin complex (GO:0005833), haptoglobin-hemoglobin complex (GO:0031838); GO: MF oxygen carrier activity (GO:0005344) and oxygen binding (GO:0019825) were identified. DEGs related to these processes included genes of the α globin (Hba-x, Hba-a1, and Hba-a2) and the β globin (Hbb-y, Hbb-bh1, Hbb-bt) loci. Also, albeit hypoxia-related HIF-1 KEGG pathway was not among significantly enriched, still eight genes (Angpt2, Crebbp, Flt1, Ldhb, Nos3, Slc2a1, Vhl, Eno1b) involved in the regulation of this signaling were differentially expressed in APS placentas.
Regulation of cell cycle and outcome of REAC (biological pathways reactome) analysis
According to GO analysis of APS placentas and uNK cells, the cell cycle (mmu04110) was found as the most significantly enriched KEGG pathway (Online Supplementary Fig. S5A). In scope of GO analysis of up-regulated in APS placenta genes, output of REAC (Biological pathways (Reactome)) significantly enriched terms was also saturated with cell cycle processes, such as Cell Cycle Checkpoints, G2/M Checkpoints, G1/S Transition, S Phase, Mitotic G1 phase and G1/S transition and Mitotic G2-G2/M phases (Online Supplemental Fig. S5B).
Deconvolution of bulk RNAseq data revealed reduced number of SpA-TGC in APS placentas
We applied the single-cell deconvolution reference to estimate cell type proportions in a bulk RNAseq data between APS and control placentas. Our analysis showed a significantly lower representation of spiral artery-associated trophoblast giant cells (SpA-TGC), a type of trophoblast cells, in APS placentas in comparison to the controls (1.9 fold). SpA-TGC are primarily located at the junction of the maternal spiral arteries and the developing placenta and ensure adequate blood supply to the growing fetus21. Thus, a reduction in the number of SpA-TGC may contribute to an inflammatory environment in APS and hinder blood supply to the fetus. To confirm the findings, we examined the spatial distribution of SpA-TGC within the placenta using histology (Fig. 2B). In line with deconvolution data, the number SpA-TGC in APS placentas was reduced in comparison with control ones. Moreover, SpA-TGCs in APS exhibited a more granular chromatin structure and an irregularly shaped nucleus that is less lobulated compared to the control group. Worth mentioning, our study didn’t confirm an increased number of macrophages in APS placentas, reported by others22.
Discussion
Here, we performed an in-depth analysis of the placenta and uNK cells resulting from immunization of mice with β2GPI, to gain better insight into placenta pathologies associated with APS. The placenta maintains a delicate balance in regulating cell cycle progression and apoptosis, ensuring that growth and differentiation occur at optimal gestational stages, thereby supporting the overall health of the pregnancy23,24. In our study, biological functions of genes up-regulated in APS are extensively linked to the cell cycle related processes. The findings revealed a significant enrichment of p53 signaling pathway, as evidenced by gene ontology analysis of up-regulated DEGs in APS placenta and uNK cells. In early pregnancy, p53 plays an important role in regulation of cell cycle, angiogenesis, trophoblast differentiation, including fusion of cytotrophoblast into syncytiotrophoblast and autophagic activity in villous trophoblasts25,26. When up-regulated, p53 inhibits cell proliferation via regulating the progression of the G1/S phase of the cell cycle, and thereby irreversibly blocking cell cycle entry16. Consistent with these roles, our study revealed a reduction in cytotrophoblast proliferation, with fewer cells in S phase in APS. In line with our data, increased apoptosis of ectoplacental cone giant cells27, and aberrant cell death in the syncytiotrophoblast28 were reported in APS. Apoptotic signaling can interfere with the cell cycle machinery, causing cells to arrest in a particular phase or preventing them from progressing through the cycle29; this disruption can halt the proliferation of trophoblast cells. Ability of aPL antibodies to affect the regulation of the cell cycle in trophoblastic cells was recently addressed in in vitro studies, albeit the conclusions have been controversial. Some authors have reported the arrest of trophoblast cell cycle in the G2/M phase and accelerated apoptotic rates mediated by aPL30, while others have observed increased cell proliferation, migration and invasion31. The variability in reported cell proliferation rates could stem from the use of different trophoblastic cell lines. Evidence from our RNA-Seq analysis indicated another deregulated mechanism interfering with the regulation of the cell cycle within the placenta, namely a motor and kinesin proteins. An increased expression pattern of kinesins on trophoblasts or villi endothelium was found in patients with preeclampsia or diabetes, suggesting that these proteins may be involved in placental pathologies18,32. Dysregulation of kinesin expression can potentially lead to cell cycle arrest, causing abnormal trophoblast proliferation. In turn, this dysregulation could impede normal trophoblast invasion and placental vascularization, exacerbating the risk of adverse pregnancy outcomes.
There is growing evidence that differences in trophoblast proliferation and differentiation are closely linked to the oxygen homeostasis and VEGF-A expression33,34. VEGF-A was identified as a downstream mediator in the MALAT1-dependent inhibition of trophoblast proliferation and endovascular differentiation35. Supporting this mechanistic pathway, our study showed reduced VEGF-A expression, cell proliferation, and low total protein content in endothelial cells from APS placentas compared to the control placentas. The disturbances in the placental vessel network corresponded well with the fetuses showing high lethality, and placentas being small in size at E12.5. As a potent inducer of placental endothelial cell proliferation and migration34, reduced levels of VEGF-A observed in our study could contribute to impaired endothelial cell proliferation and function. Histological examination also showed the thinning of vessel walls of the junctional zone, where we found a reduced number of SpA-TGC. Reduced expression of VEGF-A and the consequent loss of proliferative abilities of endothelial cells may have been the primary cause of disturbed cytotrophoblast proliferation, syncytiotrophoblast differentiation or impairments in APS placentas. The decreased efficiency of oxygen transport and the exchange of other substances in the APS placental labyrinth was reflected by an increases in the number of nucleated erythrocytes, and a decrease in the number of giant trophoblast cells.
Reduced placental growth and its abnormalities, along with endothelial dysfunction, point to the presence of preeclampsia in our mouse APS model, reflecting key features of the disease observed in humans. Moreover, despite being smaller in size, junctional zones of APS placentas were larger compared to control, as already reported in mice with uteroplacental malperfusion36. The importance of preeclampsia for obstetrical complications in APS has already been demonstrated in a mouse model4 and several observations in humans37. Preeclampsia is a pregnancy-specific condition which stands as one of the primary contributors to neonatal and fetal mortality or preterm birth38. Numerous conditions are recognized as risk factors for the development of preeclampsia, including but not limited to placental hypoxia, maternal infection and inflammation, fetal distress, placental aging, maternal metabolic disturbances, the so-called “dangerous father” phenomenon, endocrine or autoimmune disorders including APS38,39. According to the recent estimations, preeclampsia affects around 10–17% of APS pregnancies and progresses with severe features37. Preeclampsia onset with placental dysfunction triggering syncytiotrophoblast stress manifests as oxidative stress, mitochondrial damage, dysregulated metabolism and apoptosis resulting in clinical manifestation of preeclampsia40. Defective trophoblast invasion and decidual transformation at early gestation were suggested as the main underlying mechanisms of APS that are associated with preeclampsia37.
Our study demonstrated that anti-β2GPI antibodies reshape the uNK cells into a dysfunctional state by transforming their transcriptional profile, which to some extent may be influenced by an altered intrauterine environment shaped by the aPL. uNK cells have received considerable attention in relation to their role in implantation and placental development and in terms of pregnancy pathology41. Alterations of uNK cell number or function were linked with various pregnancy complications, such as recurrent miscarriage, preeclampsia, and potentially infertility41,42. Previous studies on uNK cells in APS pointed out their possible role in the development of pregnancy morbidity. However, those assumptions were based mainly on activated status or abnormal number of NK cells present in peripheral blood43,44, high count in placental specimens45 or in vitro activation of peripheral blood NK cells by aPL antibodies12. Thereby, until now, our understanding of the phenotypical and activation status of uNK cells during APS pregnancy has been vague. Current study, in line with others46 and our previous reports12,13, once again pointed to the potential of aPL antibodies to promote cytotoxic activation of NK cells in circulation. Taking into account that the rapid increase in uNK cell numbers during early pregnancy primarily arises from the in situ differentiation of decidual precursor cells47,48, we suggest that activated phenotype of peripheral NK cells would not necessarily reflect with activatory status of uNK cells. Even assuming that some portion of uNK cells are recruited from circulation47, it is well documented that upon recruitment, NK cells rapidly alter their function and adopt unique tissue-residency specific phenotypic characteristics49.
Secretory capacity of uNK cells is one of the important mechanisms regulating tissue homeostasis and vascularisation to ensure optimal fetal development50. The transcriptomic signature of NK cells derived from APS mice placentas clearly showed prominent suppression of cytokine signaling pathways, which may potentially disrupt immune tolerance and pro-angiogenic processes crucial for placental function. We propose that the diminished expression pattern of key genes in uNK cells render these cells partially dysfunctional, which eventually contributes to the pathological alterations observed in placental tissue upon histological and RNA-Seq examinations.
Conclusion
Summing up, our findings from the animal model of APS indicate that aPL antibodies significantly impact uNK cell function, leading to pathological alterations in the placenta. The observed changes in placental transcriptome profiling and morphology suggest a link between APS pregnancy abnormalities and the mechanisms underlying the development of preeclampsia. Importantly, our study underscores the role of NK cells in APS. The diminished expression of key genes in NK cells leads to their partial dysfunction, affecting their cytotoxic capabilities and cytokine signaling pathways. This dysfunction likely contributes to the pathological alterations in the placenta, including disrupted immune tolerance and impaired angiogenesis. The findings highlight the need for further investigation into the mechanisms underlying placental pathology to develop targeted therapeutic strategies. Addressing these pathways could potentially mitigate the adverse effects of APS on pregnancy outcomes, improving maternal and fetal health.
Materials and methods
Study animals
For this study, 8 weeks old BALB/c female mice (weighing 20 ± 2 g) were used. The mice were housed in standard cages (1/cage) in a controlled environment, with a relative humidity of 60 ± 5%, 12 h/12 h light-dark cycle and a constant temperature of 25 ± 2 °C. Animals had ad libitum access to tap water and rodent chow throughout the whole study. All methods are reported in accordance with ARRIVE guidelines. Euthanasia methods comply with AVMA standards. Mice were kept under controlled conditions according to protocol #05072021/1; 07/05/2021, issued by the Committee of the Institute of Molecular Biology of the National Academy of Sciences of the Republic of Armenia for the Care and Maintenance of Animals. All methods were carried out in accordance with relevant guidelines and regulations.
Establishing and validating an animal model for APS
For the establishment of the APS animal model, we selected a model of immunization with β2GPI. In contrast to other existing mouse models51, which used mainly total IgG passive antibody transfer to trigger autoimmune reactions, we aimed to induce the production exclusively anti-β2GPI antibodies, avoiding the involvement of other autoantibodies of unknown or undefined origin. The APS animal model was established in BALB/c mice via three-step immunization 10 days apart. Mice were immunized with solubilized Human Apolipoprotein H Native Protein (β2GPI) (10 µg/per injection, Invitrogen) emulsified in equal amounts with complete Freund’s adjuvant (Sigma). Mice from the control group were receiving injections with adjuvant. aPL-positive and negative status was confirmed with in-house ELISA. Shortly, high absorbance 96-well plate with hydrophilic/hydrophobic mix surface was coated overnight with 3 µg/mL β2GPI, washed and blocked with 0.1% BSA/PBS solution for 1 h and incubated with 50 µl of 1:500 diluted mouse sera for 1 h. Next, the plate was incubated with streptavidin conjugated goat anti-mouse secondary IgG (R&D Systems). Colorimetric reaction was achieved by addition of TMB substrate (R&D Systems). Following 15 min incubation, the reaction was stopped and absorbance was measured with ELISA plate reader at 450 nm. Determination of antibody levels was performed with freshly obtained sera samples.
Clinical relevance of the APS animal model was assessed in a pre-experimental group of animals. This trial group was immunized with the same scheme and immunization-induced antibody response was monitored weekly, during 7 weeks post-first immunization (Fig. 1A). On day 28, animals were mated and pregnancy outcome was assessed on embryonic day 12.5 (n = 5) or during 48 h post-partum (n = 4). Particularly, placental weight, resorption rate, maternal and pups survival were estimated.
Following confirmation of aPL status, aPL-positive (aPL+) and negative (aPL−) IgG were isolated from the APS (day 47) and healthy mice, respectively. Plasma samples were used for total IgG isolation with the Protein G SpinTrap (Cytiva) according to the manufacturer instructions. IgG concentrations were measured using a NanoQ (Mecasys). To estimate the procoagulatory potential of isolated IgG, THP-1 cells (ATCC TIB-202) were cultured in absence or presence of 50 µg/ml of aPL- and aPL + IgG for 4 h at 37 °C and 5% CO2 in a humid atmosphere. Following this, surface expression of tissue factor (TF, CD142-PE) was analyzed with flow cytometry. Isolated aPL− and aPL + IgG were tested for their procoagulatory potential individually and pooled.
Mouse pregnancy and placenta collection procedure
Immunized and non-immunized animals were routinely examined for aPL status. Monitoring of aPL titres revealed that by the 5th week post last immunization, the antibody response reached pike and remained stable for next few weeks (Fig. 1B). Selection of animals for both APS and control groups for the mating was based on anti-β2GPI titers. On day 28 post last immunization, aPL+ (APS) and aPL− (control) animals were paired and allowed to mate for 24 h. The presence of a vaginal plug confirmed the onset of pregnancy. On gestation day 12–13, mice were sacrificed by cervical dislocation and rapid decapitation for truncal blood, placental tissue and spleen collection. Micro-scissors were used for opening the uterus, detaching the conceptus and separating the placentas from the embryo and uterus wall. From each selected animal, one placenta was placed in RNAlater immediately after collection, and one was embedded in paraffin for immunohistochemistry and histology.
Uterine and spleen NK cells isolation
uNK and spleen NK (sNK) cells were isolated for downstream analyses of phenotype, cytotoxicity evaluation and RNAseq. For this, whole placentas or spleens were separately and aseptically rinsed with cold PBS immediately after collection, placed on pre-wet 70 μm cell strainer and gently dissociated with sterile rubber plunger with circular motion applying with exceptionally light pressure. The obtained single cell suspension was washed and further enriched with Histopaque gradient centrifugation. uNK and sNK cells were further isolated with the use of EasySep™ Mouse NK Cell Isolation Kit (STEMCELL, USA). pNK cells were isolated from peripheral blood mononuclear cells (PBMCs) fraction with the same kit. The purity of NK cells following isolation with the negative selection kit was determined with flow cytometry, based on the percentage of CD3-CD49b + cells within the viable lymphocyte population. The purity of NK cells was 90–95%. Immediately following isolation, the cells were used for phenotype analysis, cytotoxicity assay and cytokine production. The rest of the uNK cells were placed in RNAlater and stored for RNAseq analysis.
qPCR analysis
To detect the expression of TF in mouse PBMC, total RNA was extracted with TRIzol™ Reagent (ThermoFisher Scientific, MA, USA). One step probe-based real-time RT-PCR kit qPCRBIO Probe 1-Step Go (PCR Biosystems, USA) was used for relative quantification. The expression level of the target gene was normalized to GAPDH level using the 2−ΔΔCt method. Primer sequences are provided in online Supplemental Table S1.
NK cells phenotyping and cytotoxicity assay
Aliquots of freshly isolated PBMCs, sNK and uNK cells were stained with anti-mouse fluorochrome conjugated following antibodies: CD3-APC, CD49b-FITC, CD27-PE-Cy7, CD11b-PE and analyzed with Novocyte flow cytometer (Agilent Technologies). After exclusion of doublets and dead cells, NK cells were determined as CD3− and CD45 + cells. Subpopulational distribution of NK cells was determined according to CD27 and CD11b expression levels. It is generally accepted that mouse CD11blowCD27high and CD11bhighCD27high NK cells demonstrate superior cytokine secretion abilities, while CD11bhighCD27low NK cells exhibit heightened cytolytic function. NK cells lacking both CD11b and CD27 display an immature phenotype.
With the aim to estimate the cytotoxicity of mouse sNK and uNK, isolated cells were pretreated with 2000 U/ml of IL-2 for 24 h. Afterwards, pretreated cells were co-cultured with CFSE-labeled THP-1 cells at effector to target (E: T) ratio 5:1 for 4 h. Following the culture period, the percentage of killed target cells was assessed as PI positive CFSE-labeled THP-1 cells, calculated using the following equation: [(experiment target cell death-spontaneous death)/(100-spontaneous death)×100]. Acquisition was performed with BD FACSCalibur™ Flow Cytometer (Becton Dickinson).
Cytotoxicity was additionally assessed by lactate dehydrogenase (LDH) release from target cells into the culture medium. The activity of LDH in the medium was determined using a commercially available CytoTox 96® Non-Radioactive Cytotoxicity Assay kit (Promega Corporation, Madison, WI, USA). Aliquots of media and reagents were mixed in a 96-well plate according to protocol instructions and the optical absorbance was measured using a HiPo MPP-96, Microplate Photometer (Biosan) at 490 nm. The following equation was used to calculate the percentage of cytotoxicity: Cytotoxicity (%) = (Experimental Value-Effector Cells Spontaneous Control-Target Cells Spontaneous Control)/(Target Cell Maximum Control-Target Cells Spontaneous Control) ×100.
Cytokine measurement
IFN-γ was quantified in plasma samples from animals and cultured media of NK cells isolated from spleen and placenta after 24 h of incubation in the presence of IL-2 (2000 U/ml) with the use of ELISA MAX™ Deluxe Set Mouse IFN-γ kit (BioLegend; 430804).
Fluorescent microscopy
Formalin fixed, paraffin-embedded tissue Sect. (5-µm) were mounted on positively charged glass slides (Superfrost plus microscope slides, Fisher Scientific). The sections were heated at 60 °C for 1 h and deparaffinized by serial immersions in xylene and ethanol dilutions. Antigen retrieval was achieved by placing slides in pre-warmed to 60 °C citrate buffer (pH 6.0) and heated up to 95 °C for 20 min, using a water bath. Cooled tissue sections were blocked and permeabilized for 1 h at room temperature (0.3% Triton X-100, 5% Bovine Serum Albumin). Stainings were carried out consecutively with the primary anti-VEGF-A (vascular endothelial growth factor A) (ab185238) and anti-Ki-67 rabbit monoclonal primary antibody (Roche; 05278384001) being added first and incubated for 1 h at RT, followed by staining with the secondary anti-rabbit IgG (Cy5) (Abcam) and for 2 h at RT. Following this, the sections were stained with Hoechst 33,342 (Thermo Scientific) for nucleus identification and mounted with Fluoromount G (Invitrogen) and analyzed with Cytation 10 (Agilent Technologies).
Histopathological examination
Placental sections from control and APS mice were deparaffinized as described above and rehydrated with deionized H2O. Slides were stained with Hematoxylin Solution, rinsed and counterstained with Eosin solution. Alternatively, slides were treated with the combined Feulgen-Naphthol Yellow staining (FNYS) procedure. Stained slides were layered with the Permount™ histological mounting medium (Fisher Scientific), followed by a glass coverslip.
Micro-spectrophotometric analysis of DNA and protein content in endothelial and trophoblast cells
Placental sections, stained with the combined Feulgen-Naphthol Yellow staining (FNYS), were subjected to the micro-spectrophotometric determination of the quantities of DNA and protein at the same time in each individual cell. Samples were analyzed with microscope-photometer SMP 05 (OPTON), on a 575 nm wave. For each group a total of at least 400 cells in at least 40 fields were observed and analyzed. Quantity of DNA was defined in conventional units (C.U.) Following quantitative DNA-staining, the nuclear Integrated Optical Density (IOD), the cytometric equivalent of its DNA content, was determined. For the quantification of nuclear DNA IOD values were rescaled by comparison with those from cells with the known DNA content, particularly, unstimulated lymphocytes were used as standards. Therefore, the DNA content is expressed in a “c” scale in which 1c is half (haploid) of the nuclear DNA content of cells from a normal (non-pathological) diploid population in G0/G1 cell cycle phase. The DNA cytometric measurements identified cell nuclei as aneuploid if they deviate more than 10% from 2c, 4c, 8c i.e. if they are outside of 2c ± 0.2, 4c ± 0.4, 8c ± 0.8. The number of all cells in euploid regions of the DNA histogram rescaled by the mean corrective factor of the tissue type under the investigation (1.8c–2.2c; 3.6c–4.4c; 7.2c–8.8c) was also calculated. The ploidy status of cells was determined by the amount of DNA, expressed as a DNA index (DI), i.e., the ratio between the peak channel number of the cells sample in G0/G1 and the peak channel number of the diploid standard population in G0/G1. Mouse red blood cells were used as a reference standard to determine the protein content of mouse placenta endothelial cells. The recalculation of optical density into picograms per cell (pg/cell) was carried out similarly to the recalculation of DNA.
RNA sequencing
Total RNA was isolated from placenta tissues (7 APS and 7 controls) and isolated uNK cells (6 APS and 4 controls) using RNAqueous™-Micro Total RNA Isolation Kit (Thermo Fisher Sci Inc., Waltham, MA, USA) according to the manufacturer’s protocol. Total RNA was quantified using the Qubit™ RNA HS Assay Kit (Thermo Fisher Sci Inc., Waltham, MA, USA). Sample quality assessment was performed using the Agilent RNA 6000 Pico Chip on the Agilent Bioanalyzer 2100 according to the manufacturer’s instructions (Agilent Technologies, Palo Alto, CA, USA). The RNA Integrity Number (RIN) and DNA contamination was evaluated using the Agilent 2100 Expert software (Software Version 2100 Expert B.02.10.SI764). Double-Strand specific DNase pretreatment (ArcticZymes Technologies, Tromsø, Norway) was performed to remove residual DNA according to the manufacturer’s instructionsSequencing libraries were constructed using the SMARTer (for NK cells) Stranded Total RNA-Seq Kit - Pico input Mammalian v3 (Takara Bio USA, Mountain View, CA, USA) and NEBNext Ultra II Directional RNA Library prep kit after enrichment for mRNA using the NEBNext poly(A) mRNA Isolation (NEB) (for placenta samples) kit according to the manufacturer’s instructions. Quality control of the libraries was performed using an Agilent DNA HS Chip on an Agilent Bioanalyzer 2100 (Agilent Technologies, Palo Alto, CA, USA) according to the manufacturer’s protocol. Libraries were sequenced by next-generation sequencing on a NovaSeq 6000 (Illumina Inc., CA, USA) using paired-end sequencing (2 × 151 bp) with a minimum depth of 50 million reads per sample.
Bioinformatic analysis of the differentially expressed genes
RNA-Seq analyses for placentas and NK cells were performed using the NextFlow nf-core/rnaseq pipeline (v.3.14.0; https://doi.org/10.5281/zenodo.1400710, https://doi.org/10.1038/s41587-020-0439-x, https://doi.org/10.1038/nbt.3820) using GRCm39 mouse (Mus musculus) genome reference with default parameters (in case of SMARTer specific setting handling UMIs was used). Tools STAR (https://doi.org/10.1093/bioinformatics/bts635) was used for alignment and Salmon (https://doi.org/10.1038/nmeth.4197) for quantification. Principal component analysis (PCA), volcano plots, transcripts per million (TPM) generation, normalization of data were carried out using 3D shiny RNA-Seq App and for DEG analysis the limma R package was used (https://doi.org/10.1080/15476286.2020.1858253, R Core Team (2024). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/). In the case of placentas, batch effect removal using the RUVr function from the RUVSeq R package was used (https://doi.org/10.1038/nbt.2931). To identify DEGs, a false discovery rate (FDR) threshold of < 0.05 and a log2 fold change (Log2FC) cut-off of ± 1 were used.
To comprehensively analyze the DEGs obtained from our study, the functional enrichment analysis of these DEGs was carried out by using the g: Profiler gene annotation tool, an online platform for the assessment of Gene Ontology (GO) terms and KEGG pathways (https://biit.cs.ut.ee/gprofiler/gost) tool that covered three GO categories: biological processes (BPs), molecular functions (MFs) and cellular compartment (CCs). Separate analysis of up- and down-regulated DEGs in placenta resulted in a notable enrichment of a large number of GO terms, Kegg and REAC pathways. Furthermore, to emphasize the extensive impact of aPL antibodies on placental function, we highlighted GO terms with the highest significance in each category (Online Supplementary Fig. S1).
This analysis aimed at elucidating crucial biological processes and pathways affected in APS (Online Supplementary Figs. S1, S2). KEGG pathway diagrams were generated to visualize the interaction of total up- and down- regulated DEGs, with the use of KEGG Mapper color tool (https://www.genome.jp/kegg/mapper/color.html). The results of these analyses highlighted enriched terms and provided valuable insights into the functional roles and interactions of the identified genes (Online Supplementary Figs. S1, S2, S3, S4).
Deconvolution of bulk RNAseq data from whole placentas with scRNAseq as a reference
For the deconvolution of bulk RNA-seq from the placenta cells, thus providing insights into the cellular composition of our samples, we employed the MuSiC tool (https://doi.org/10.1038/s41467-018-08023-x), which leverages single-cell RNA sequencing (scRNA-seq) data to estimate the cell-type proportions. As reference, scRNA-seq dataset from the study “A differentiation roadmap of murine placentation at single-cell resolution” by Jiang et al. was used21.
Statistics
The statistical analysis of obtained data was performed using Graph Pad Prism 9 (GraphPad Software, San Diego, CA). Data is presented as the mean ± the standard deviation (SD). The Shapiro–Wilk test was used to assess the normality of data distributions. Data comparison was performed by using one-way repeated measures ANOVA with Tukey’s multiple comparison post hoc test. Analysis between independent data was done with unpaired t test or Mann–Whitney test, based on data distribution. A p-value < 0.05 was considered significant.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Galarza-Maldonado, C. et al. Obstetric antiphospholipid syndrome. Autoimmun. Rev. 11(4), 288–295. https://doi.org/10.1016/j.autrev.2011.10.006 (2012).
de Groot, P. G. & Urbanus, R. T. The significance of autoantibodies against β2-glycoprotein I. Blood 120(2), 266–274. https://doi.org/10.1182/blood-2012-03-378646 (2012).
Tong, M., Viall, C. A. & Chamley, L. W. Antiphospholipid antibodies and the placenta: a systematic review of their in vitro effects and modulation by treatment. Hum. Reprod. Update 21(1), 97–118. https://doi.org/10.1093/humupd/dmu049 (2015).
Chu, H. et al. Protein phosphatase 2A activation via ApoER2 in trophoblasts drives preeclampsia in a mouse model of the antiphospholipid syndrome. Circ. Res. 129(7), 735–750. https://doi.org/10.1161/CIRCRESAHA.120.318941 (2021).
Yang, J. & Liang, M. Risk factors for pregnancy morbidity in women with antiphospholipid syndrome. J. Reprod. Immunol. 145, 103315. https://doi.org/10.1016/j.jri.2021.103315 (2021).
Schreiber, K. & Hunt, B. J. Managing antiphospholipid syndrome in pregnancy. Thromb. Res. 181, S41–S46. https://doi.org/10.1016/S0049-3848(19)30366-4 (2019).
Meroni, P. L. et al. Obstetric and vascular antiphospholipid syndrome: same antibodies but different diseases? Nat. Rev. Rheumatol. 14(7), 433–440. https://doi.org/10.1038/s41584-018-0032-6 (2018).
Schiffer, V. et al. Spiral artery blood flow during pregnancy: a systematic review and meta-analysis. BMC Pregn. Childbirth 20(1), 680. https://doi.org/10.1186/s12884-020-03150-0 (2020).
Cornish, E. F. et al. Innate immune responses to acute viral infection during pregnancy. Front. Immunol. 11, 572567. https://doi.org/10.3389/fimmu.2020.572567 (2020).
Sfakianoudis, K. et al. The role of uterine natural killer cells on recurrent miscarriage and recurrent implantation failure: from pathophysiology to treatment. Biomedicines 9(10), 1425. https://doi.org/10.3390/biomedicines9101425 (2021).
Zhang, X. & Wei, H. Role of decidual natural killer cells in human pregnancy and related pregnancy complications. Front. Immunol. 12, 728291. https://doi.org/10.3389/fimmu.2021.728291 (2021).
Manukyan, G. et al. Antiphospholipid antibody-mediated NK cell cytotoxicity. J. Reprod. Immunol. 155, 103791. https://doi.org/10.1016/j.jri.2022.103791 (2023).
Manukyan, G. et al. Anti-domain 1 β2 glycoprotein antibodies increase expression of tissue factor on monocytes and activate NK cells and CD8 + cells in vitro. Auto Immun. Highlights 11(1), 5. https://doi.org/10.1186/s13317-020-00128-y (2020).
Bayram, F. et al. Perinatal asphyxia is associated with the umbilical cord nucleated red blood cell count in pre-eclamptic pregnancies. J. Obstet. Gynaecol. 30(4), 383–386. https://doi.org/10.3109/01443611003706928 (2010).
Gemble, S. et al. Genetic instability from a single S phase after whole-genome duplication. Nature 604(7904), 146–151. https://doi.org/10.1038/s41586-022-04578-4 (2022).
Sultana, Z., Maiti, K., Dedman, L. & Smith, R. Is there a role for placental senescence in the genesis of obstetric complications and fetal growth restriction? Am. J. Obstet. Gynecol. 218(2S), S762–S773. https://doi.org/10.1016/j.ajog.2017.11.567 (2018).
Bai, H. et al. Progressive senescence programs induce intrinsic vulnerability to aging-related female breast cancer. Nat. Commun. 15(1), 5154. https://doi.org/10.1038/s41467-024-49106-2 (2024).
Wong-Riley, M. T. & Besharse, J. C. The kinesin superfamily protein KIF17: one protein with many functions. Biomol. Concepts 3(3), 267–282. https://doi.org/10.1515/bmc-2011-0064 (2012).
Mann, B. J. & Wadsworth, P. Kinesin-5 regulation and function in mitosis. Trends Cell. Biol. 29(1), 66–79. https://doi.org/10.1016/j.tcb.2018.08.004 (2019).
Giussani, D. A. Breath of life: heart disease link to developmental hypoxia. Circulation 144(17), 1429–1443. https://doi.org/10.1161/CIRCULATIONAHA.121.054689 (2021).
Jiang, X. et al. A differentiation roadmap of murine placentation at single-cell resolution. Cell. Discov. 9(1), 30. https://doi.org/10.1038/s41421-022-00513-z (2023).
Lu, C. et al. Single-cell transcriptome analyses reveal disturbed decidual homoeostasis in obstetric antiphospholipid syndrome. Ann. Rheum. Dis. 83(5), 624–637. https://doi.org/10.1136/ard-2023-224930 (2024).
Kar, M., Ghosh, D. & Sengupta, J. Histochemical and morphological examination of proliferation and apoptosis in human first trimester villous trophoblast. Hum. Reprod. 22(11), 2814–2823. https://doi.org/10.1093/humrep/dem284 (2007).
Knöfler, M. et al. Human placenta and trophoblast development: key molecular mechanisms and model systems. Cell. Mol. Life Sci. 76(18), 3479–3496. https://doi.org/10.1007/s00018-019-03104-6 (2019).
Levy, R. et al. Trophoblast apoptosis from pregnancies complicated by fetal growth restriction is associated with enhanced p53 expression. Am. J. Obstet. Gynecol. 186(5), 1056–1061. https://doi.org/10.1067/mob.2002.122250 (2002).
Liu, T. et al. The role of p53-MDM2 signaling in missed abortion and possible pathogenesis. J. Obstet. Gynaecol. Res. 48(11), 2686–2696. https://doi.org/10.1111/jog.15385 (2022).
Ornoy, A. et al. The effects of antiphospholipid antibodies obtained from women with SLE/APS and associated pregnancy loss on rat embryos and placental explants in culture. Lupus 12(7), 573–578. https://doi.org/10.1191/0961203303lu405oa (2003).
Pantham, P. et al. Antiphospholipid antibodies alter cell-death-regulating lipid metabolites in first and third trimester human placentae. Am. J. Reprod. Immunol. 74(2), 181–199. https://doi.org/10.1111/aji.12387 (2015).
Kumari, R. & Jat, P. Mechanisms of cellular senescence: cell cycle arrest and senescence associated secretory phenotype. Front. Cell. Dev. Biol. 9, 645593. https://doi.org/10.3389/fcell.2021.645593 (2021).
Huang, Z. et al. Inactivation of yes-associated protein mediates trophoblast dysfunction: a new mechanism of pregnancy loss associated with anti-phospholipid antibodies?. Biomedicines 10(12), 3296. https://doi.org/10.3390/biomedicines10123296 (2022).
Lu, X. et al. Effect and mechanism of the aβ2–GP I/rhβ2–GP I complex on JEG–3 cell proliferation, migration and invasion. Mol. Med. Rep. 17(6), 7505–7512. https://doi.org/10.3892/mmr.2018.8822 (2018).
Sati, L. et al. The presence of kinesin superfamily motor proteins KIFC1 and KIF17 in normal and pathological human placenta. Placenta 30(10), 848–854. https://doi.org/10.1016/j.placenta.2009.07.005 (2009).
Natale, B. V. et al. Reduced uteroplacental perfusion pressure (RUPP) causes altered trophoblast differentiation and pericyte reduction in the mouse placenta labyrinth. Sci. Rep. 8(1), 17162. https://doi.org/10.1038/s41598-018-35606-x (2018).
Fan, X. et al. VEGF maintains maternal vascular space homeostasis in the mouse placenta through modulation of trophoblast giant cell functions. Biomolecules 11(7), 1062. https://doi.org/10.3390/biom11071062 (2021).
Li, Q. et al. MALAT1 modulates trophoblast phenotype via miR-101-3p/VEGFA axis. Arch. Biochem. Biophys. 744, 109692. https://doi.org/10.1016/j.abb.2023.109692 (2023).
Bobek, G. et al. Quantification of placental change in mouse models of preeclampsia using magnetic resonance microscopy. Eur. J. Histochem. 62(2), 2868. https://doi.org/10.4081/ejh.2018.2868 (2018).
Mayer-Pickel, K. et al. Preeclampsia and the antiphospholipid syndrome. Biomedicines 11(8), 2298. https://doi.org/10.3390/biomedicines11082298 (2023).
Committee on Practice Bulletins-Obstetrics. Gestational hypertension and preeclampsia: ACOG practice bulletin summary, number 222. Obstet. Gynecol. 135(6), 1492–1495. https://doi.org/10.1097/AOG.0000000000003892 (2020).
Opichka, M. A. et al. Vascular dysfunction in preeclampsia. Cells 10(11), 3055. https://doi.org/10.3390/cells10113055 (2021).
Dimitriadis, E. et al. Pre-eclampsia. Nat. Rev. Dis. Primers 9(1), 8. https://doi.org/10.1038/s41572-023-00417-6 (2013). Erratum in: Nat. Rev. Dis. Primers 9(1), 35. https://doi.org/10.1038/s41572-023-00451-4 (2013).
Von Woon, E. et al. Number and function of uterine natural killer cells in recurrent miscarriage and implantation failure: a systematic review and meta-analysis. Hum. Reprod. Update 28(4), 548–582 https://doi.org/10.1093/humupd/dmac006 (2022).
Moffett, A., Regan, L. & Braude, P. Natural killer cells, miscarriage, and infertility. BMJ 329(7477), 1283–1285. https://doi.org/10.1136/bmj.329.7477.1283 (2004).
Perricone, C. et al. High levels of NK cells in the peripheral blood of patients affected with anti-phospholipid syndrome and recurrent spontaneous abortion: a potential new hypothesis. Rheumatology (Oxford) 46(10), 1574–1578. https://doi.org/10.1093/rheumatology/kem197 (2007).
Carbone, J. et al. Quantitative abnormalities of peripheral blood distinct T, B, and natural killer cell subsets and clinical findings in obstetric antiphospholipid syndrome. J. Rheumatol. 36(6), 1217–1225. https://doi.org/10.3899/jrheum.081079 (2009).
Gomaa, M. F. et al. Uterine CD56dim and CD16 + cells in refractory antiphospholipid antibody-related pregnancy loss and chromosomally intact abortuses: a case-control study. J. Hum. Reprod. Sci. 10(1), 18–23. https://doi.org/10.4103/jhrs.JHRS_65_16 (2017).
Zhang, Y. et al. Increased peripheral NKG2A-NKG2D + CD3-CD16 + CD56dim NK cell subset was positively correlated with antiphospholipid antibodies in patients of obstetric antiphospholipid syndrome. Immunol. Investig. 51(2), 425–437. https://doi.org/10.1080/08820139.2020.1835949 (2022).
Sojka, D. K., Yang, L. & Yokoyama, W. M. Uterine natural killer cells. Front. Immunol. 10, 960. https://doi.org/10.3389/fimmu.2019.00960 (2019).
Xie, M. et al. Uterine natural killer cells: a rising star in human pregnancy regulation. Front. Immunol. 1, 13:918550. https://doi.org/10.3389/fimmu.2022.918550 (2022).
Dean, I. et al. Rapid functional impairment of natural killer cells following tumor entry limits anti-tumor immunity. Nat. Commun. 15(1), 683. https://doi.org/10.1038/s41467-024-44789-z (2024).
Yockey, L. J. & Iwasaki, A. Interferons and proinflammatory cytokines in pregnancy and fetal development. Immunity 49(3), 397–412. https://doi.org/10.1016/j.immuni.2018.07.017 (2018).
Tincani, A. et al. Immunization of naive BALB/c mice with human beta2-glycoprotein I breaks tolerance to the murine molecule. Arthritis Rheum. 46(5), 1399–1404. https://doi.org/10.1002/art.10304 (2002).
Funding
This work was supported by a State Committee Science MES RA, in the frame of the research Project No. SCS 21AG-1F072 and in part by MH CZ-DRO (FNOl, 00098892) and IGA LF UP_2024_013.
Author information
Authors and Affiliations
Contributions
Martirosyan A: Writing - original draft; Investigation; Methodology; Formal analysis; Visualization; Validation. Kriegova E: Funding acquisition; Writing - review & editing; Resources; Investigation; Project administration. Savara J: Validation; Visualization; Software; Formal analysis. Abroyan L: Formal analysis; Investigation. Ghonyan S: Formal analysis; Investigation. Slobodova Z: Validation; Formal analysis; Investigation; Visualization. Nesnadna R: Formal analysis; Investigation. Manukyan G: Conceptualization; Writing - original draft; Writing - review & editing; Supervision; Data curation; Project administration; Methodology; Resources; Investigation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal procedures were approved by the Institutional Animal Care and Use Committee (IACUC) at the Institute of Molecular Biology, NAS RA, and performed under the authority of the project license (#04072022/1).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Martirosyan, A., Kriegova, E., Savara, J. et al. Impact of antiphospholipid syndrome on placenta and uterine NK cell function: insights from a mouse model. Sci Rep 14, 31163 (2024). https://doi.org/10.1038/s41598-024-82451-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-82451-2