Abstract
Scapular morphological attributes show promise as prognostic indicators of retear following rotator cuff repair. Current evaluation techniques using single-slice magnetic-resonance imaging (MRI) are, however, prone to error, while more accurate computed tomography (CT)-based three-dimensional techniques, are limited by cost and radiation exposure. In this study we propose deep learning-based methods that enable automatic scapular morphological analysis from diagnostic MRI despite the anisotropic resolution and reduced field of view, compared to CT. A deep learning-based segmentation network was trained with paired CT derived scapula segmentations. An algorithm to fuse multi-plane segmentations was developed to generated high-resolution 3D models of the scapula on which morphological landmark- and axes were predicted using a second deep learning network for morphological analysis. Using the proposed methods, the critical shoulder angle, glenoid inclination and version were measured from MRI with accuracies of -1.3 ± 1.7 degrees, 1.3 ± 2.1 degree, and − 1.4 ± 3.4 degrees respectively, compared to CT. Inter-class correlation between MRI and CT derived metrics were substantial for the glenoid version and almost perfect for the other metrics. This study demonstrates how deep learning can overcome reduced resolution, bone border contrast and field of view, to enable 3D scapular morphology analysis on MRI.
Similar content being viewed by others
Introduction
Several morphological attributes of the scapula, such as the glenoid inclination1, the extension of the acromion2, and the critical shoulder angle (CSA)3,4 are associated with the risk of a retear of rotator cuff (RC) repair surgery. Scapular morphological metrics are typically evaluated manually in two-dimensions (2D) on radiographic images. As such, they are prone to inaccuracies due to projection direction and slice orientation5,6. Improved assessment methods on computed tomography (CT) data have been proposed, allowing the user to define the precise slice position and orientations for the measurements, resulting in higher accuracy measurements6,7,8. Three-dimensional (3D) morphology assessment approaches, in which anatomical landmarks are depicted on a 3D surface model of the scapula, generated from segmentation of the scapula from CT, have also been proposed9. However, as magnetic resonance imaging (MRI) is the state-of-the art imaging modality for RC tear diagnosis10,11, the associated cost and irradiation of an additional CT acquisition, prevents use of such techniques in clinical routine. To date no method, allowing for 3D analysis of scapular morphology from MRI with comparable accuracies to CT have been described. For effective 3D-measurement on MRI, accurate segmentation of the scapula to create a 3D surface model, as well as the depiction of bony landmarks is required. MRI analysis is limited, however, compared to CT, by; (1) decreased bone contrast, (2) considerably lower resolution, especially in the out-of-plane direction; and (3) a restricted field-of-view (FOV) often failing to include the entire scapular body and thus also landmarks required for morphological analysis.
To date, semantic segmentation of the shoulder from MRI for 3D surface modelling has mostly been performed manually12,13,14 or with semi-automatic methods15,16. Recently, deep learning algorithms have facilitated automatic segmentation of the scapula, humerus as well as the rotator cuff muscles on MRI with excellent accuracies compared to manual segmentation17,18,19. However, with slice thicknesses of diagnostic MRI typically three to four millimetres, the accuracy of generated surface models remains limited compared to CT. The increased surface model error, expressed as a step-artifact between slices in generated 3D surface models, results in increased error in landmark detection on the bone surface and thus increased error in scapular morphological analysis. 3D surface models generated from isotropic 3D-MRI such as Dixon15,16 or zero echo time MRI which allows for excellent bone imaging20,21,22 have been reported, however, such sequences are not standard in clinical RC tear diagnosis. For analysis of the brain and prostate, methods to fuse the information of multiple non-isotropic MRI planes to generate high-resolution images or 3D models, have been described23,24,25,26. However, presented methods are yet to be applied to musculoskeletal analysis and require the same MRI weighting along all planes or a perfect overlapping FOV of all MRI planes.
To reduce the effect of user variability on morphological measure calculations, deep learning approaches, enabling automatic 3D landmark localisation have been described for the knee27 and hip28 on MRI and for the shoulder on CT29. In case of diagnostic shoulder MRI with restricted FOV, however, the medial and inferior scapular borders, which are necessary to define measurement planes, are often cropped, preventing accurate manual or automatic analysis. We propose, that alternatively, deep learning could be used to predict the scapular axes directly from incomplete scapular surface models, eliminating the reliance on landmarks outside of the image FOV.
The goal of this study was to develop deep learning-based methods for accurate, automatic scapular morphology analysis on non-isotropic MRI without the need for additional CT scans. We hypothesised that the proposed methods would enable automated scapular morphology analysis on MRI that is equivalent to CT based analysis for the diagnosis of RCT patients.
Methods
Data description
In this study, two datasets (Dataset 1, Dataset 2) from patients with posterosuperior RC tears without bone defects or arthritic changes were used. With approval of the local ethical commission (CCER 2020–02670), data collected during a prospective study was used in this research, which was conducted in accordance with the Declaration of Helsinki. Informed consent was obtained from all participants. Dataset 1 comprised paired CT-MRI data of the same shoulder from 30 patients with a mean age of 58.6 ± 15.1 years (range 28–87 years), comprising 16 females (53%) and 14 males (47%). CT data was anisotropic and had a mean in-plane resolutions 0.88 ± 0.18 mm (range 0.47–0.98 mm) in the transversal plane and mean slice-thickness of 0.43 ± 0.16 mm (range 0.30–0.90 mm), while MR arthrograms had in-plane resolutions of 0.2–0.6 mm and slice-thicknesses of 2.8–5.5 mm. The MRI weightings and planes are depicted in Table 1. Though the patients were all seen at the same clinic, the diagnostic MRI was performed at seven different institutions following their standard clinical protocols on different MRI machines from different vendors with field strengths ranging from 1 to 3 Tesla. Dataset 1 was randomly split into a training- (N = 20) and a test-dataset (N = 10). A second dataset (Dataset 2), consisting of CT data only, from an additional 57 RC tear patients with a mean age of 58.0 ± 9.4 year (range 33–77 years), comprising 29 females (51%) and 28 males (49%) (non-isotropic resolution, mean in-plane resolutions 0.87 ± 0.19 mm (range 0.43–0.98 mm) in the transversal plane and mean slice-thickness of 0.43 ± 0.16 mm (range 0.30–0.90 mm)) was also collected.
Manual CT data labelling
The scapula was segmented from the CT data of Dataset 1and Dataset 2 using an intensity threshold approach with manual checking and correction by an expert. A 3D surface models of the scapula were generated from the resulting segmentation masks in Mimics 16.0 (Materialise, Leuven, Belgium).
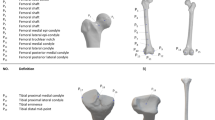
For calculation of morphological parameters, seven anatomical landmarks were manually depicted by a clinical expert (J.T.R (fellowship trained)) on the 3D surface models of the scapulae generated from the CT data of Dataset 1 and Dataset 230. To define the glenoid centre, three anatomical landmarks on the lower glenoid rim (anterior, posterior, and inferior glenoid rim) were manually depicted (Fig. 1). The centre of a circle fitted to these three points was defined as the glenoid centre and the origin of the scapular coordinate system (Fig. 1). These three points also defined the glenoid plane as proposed in31. The two points describing the vertex of the inferior scapular angle and the centre of the trigonum spinae scapulae were also manually depicted (Fig. 1). The z-axis (transverse scapular axis) was defined by the line crossing the centre of the glenoid, and the centre of the trigonum spinae scapulae, pointing in the lateral direction. The coronal scapular plane (CSP) was defined by three points, the centre of the glenoid, the vertex of the inferior scapular angle and the centre of the trigonum spinae scapulae, as defined by Suter et al.6. The x-axis was defined as the vector normal to the CSP pointing in the anterior direction with the origin at glenoid centre and the y-axis was defined perpendicular to the x- and -z axis. Additionally, the superior glenoid tubercule, and at the anterior lateral corner of the acromion were manually depicted (Fig. 1).
Anatomical landmarks of the scapula: (1) Anterior- (2) Posterior-. (3) Inferior- points on the lower glenoid rim. (4) Glenoid centre, origin of the scapular coordinates system (circle fit from points 1,2, and 3). (5) Inferior scapular angle. (6) Centre of the trigonum spinae scapulae. (7) Superior glenoid tubercule. (8) anterior lateral corner of the acromion. The glenoid plane is defined using points 1,2, and 3. The coronal scapular plane (CSP) is defined using points 4, 5, and 6. The transverse scapular axis runs through points 4) and 6), corresponding to the z-axis of the scapular coordinate system.
Automatic scapular morphology analysis on MRI
For automatic, accurate 3D morphology analysis of the scapula from diagnostic MRI our proposed methods comprise the subsequent four steps (Fig. 2): (1) Accurate segmentation of the bony border of the scapula on MRI, performed in the coronal, sagittal and axial directions. (2) Generation of a high-resolution surface model of the scapula by the fusion of the low-resolution masks along the different planes. (3) Automatic, accurate landmark depiction and scapular axes prediction on the generated high-resolution scapular model and (4) scapular morphology analysis using the predicted landmarks and axes.
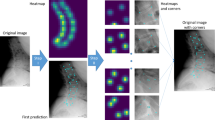
Overview of the pipeline for automatic morphology analysis. (a) From diagnostic magnetic resonance image (MRI) data, automatic segmentation is performed on multiple image orientations for the same patient. This information is fused to create isotropic, high-resolution 3D masks of the scapula. (b) From these high-resolution 3D masks, landmarks and axes are automatically detected. The landmark coordinates and axes directions are used for morphological analysis.
Automatic scapula segmentation on MRI
For accurate segmentation of the scapula in the MRI, the paired CT-MRI image training data from Dataset 1 was used to train a deep learning network. The CT and MRI were rigidly aligned using a mutual information algorithm32 (Python 3.10, Elastix extension for the SimpleITK-library33) to derive the transformation needed to transfer the high-resolution segmentation masks from the CT to the MRI. After registration, the CT scapula mask was interpolated to the spacing of the aligned MRI without manual correction of the transferred mask.
The MRI and corresponding transferred CT-based scapula masks were then used to train a deep learning network with U-net architecture to segment the scapula from the MRI (all orientations and weightings). The network was trained with CT segmentations rather than direct segmentations of the MRI to allow the network to exploit the more accurate depiction of skeletal structures in CT as compared with MRI. The 3D- full-resolution network was trained for 100 epochs using the nnU-Net framework34. The ensemble of the results from the five different networks trained during 5-fold cross validation training procedure was applied to maximize the network performance.
Multi-planar high-resolution shape reconstruction
Because of the low out-of-plane resolution, 3D surface masks generated from segmented coronal, sagittal and axial MRIs exhibit distinct inaccuracies. To generate an isotropic high-resolution 3D model from MRI, a shape-based fusion of the coronal, sagittal and axial MRI masks was performed (Fig. 3). The proposed method automatically consolidates the regions of highest accuracy from the different low-resolution masks. These regions are characterized by a consistent mask periphery across the adjacent mask slices in the low-resolution direction (Fig. 3).
Methods for multi-planar, high-resolution shape reconstruction. (a) Fusion of separately segmented masks from coronal sagittal and axial magnetic resonance imaging (MRI). (b) Example of a scapula segmentation in different sagittal slices, indicated with different colours showing different regions of 1) High consistency; where the mask border remains consistent over the adjacent image slices, 2) medium consistency; where the mask border differs between adjacent image slices, 3) low consistency, where the mask border is highly inconsistent over adjacent image slices.
The algorithm generates a high-resolution mask, henceforth referred to as HR-MRI-mask, by integrating three binary segmentations (coronal, sagittal, and axial masks). If multiple MRI sequences along a particular orientation are available, the sequence with the maximum FOV is selected. Employing a slice-wise distance map for each image orientation, the mask periphery is delineated, with pixel values representing the distance to the scapula border, up until a threshold of ± 15 pixels (Fig. 4). Mask consistency is quantified as the absolute difference between consecutive slices’ distance maps, normalized to values between 0 (indicating a zone of low consistency) and 1 (indicating a zone of high consistency) (Fig. 4). The initial binary masks are re-labelled with values of 1 inside and − 1 outside the scapula segmentation. Subsequently, for each orientation, the new binary mask and the consistency map are interpolated to an isotropic high-resolution space (isotropic resolution 0.5 mm) and consolidated by voxel-wise multiplication to generate a weighted mask. The weighted masks from all three orientations are summed, and the HR-MRI-mask is generated by classifying voxels with values greater than zero as scapula and those with values of zero or lower as background.
Visualization of intermediate steps in generating the high-resolution scapula model from multiple MRI planes. The blue crosshair indicates the same glenoid position across all views. (a) In-plane distance map of the scapula segmentation on coronal, transverse and sagittal MRI planes, with distances ranging from − 15 and 15. Bright areas represent high segmentation border distances within the scapula mask, while dark areas indicate large distances outside the mask. (b) Mask border consistency along the out-of-plane direction in the coronal, transverse and sagittal MRI planes, ranging from 0 and 1. White areas denote regions with low border consistency (assigned lower weighting for plane-combination), while dark areas indicate regions with higher consistency and weighting.
Automatic landmark and axes detection
For automatic landmark depiction, a deep learning-based segmentation approach was developed using the manually segmented masks from Dataset 2 (CT data) with landmarks depicted manually by clinical experts. Each manual landmark was encoded as a separate spherical mask with a radius of 7 mm, with the centre of the sphere defining the 3D landmark position. An exception was made for the centre of the trigonum spinae scapulae, and the vertex of the inferior scapular angle. These two points were not encoded as landmarks because they are expected to be outside of the FOV of the MRI medially and inferiorly. Instead, the axes which are defined by these two landmarks, the transverse scapular axis, passing through the glenoid centre in the direction of the centre of the trigonum spinae scapulae; and the axis passing through the glenoid centre in the direction of the vertex of the inferior scapular angle, were encoded. With these axes, the scapular coordinate system is established without requiring the entire scapula. The axes were encoded as separate rod-shaped masks with a radius of 7 mm. A best fit line to the voxels of the mask defined the axis. The 3D-full resolution deep learning network was trained for 200 epochs using the nnU-Net framework34. The network was trained to predict the sphere and rod segmentations depicting the scapula landmarks and axes from the binary masks of the scapula.
Automatic glenoid detection
For automatic scapular morphology analysis, a method for extracting the glenoid from the scapula surface model, based on the defined landmarks on the glenoid and scapular body, was developed. A sub-volume containing the glenoid is defined by a rectangular cuboid bounding box with a 5 mm margin around the anterior, inferior, posterior, and superior glenoid landmarks. In the glenoid plane, surface points of the 3D scapula model, contained within the bounding box, are split into 90 angular sectors with a central angle of 4° about the glenoid centre point. In each section, the glenoid surface points are identified by increasing the radius until the points at the arc became more medial (in relation to the glenoid plane), than those with a smaller radius, effectively defining the rim. We defined the glenoid axis as a line passing through the inferior- and superior-most points of the glenoid for further morphology analysis.
Automatic scapular morphology analysis
Algorithms enabling the automatic analysis of scapular morphology from the calculated landmarks and axes were developed. The glenoid circle radius is calculated as the radius of the circle fitted to the three anatomical landmarks on the lower glenoid rim. The glenoid height is defined as the distance between the superior glenoidal tubercule landmark and the inferior landmark on the lower glenoid rim.
The glenoid version (GV) is evaluated as the complementary of the angle between the glenoid plane, and the transverse scapular axis as defined by Friedman et al.35. The glenoid inclination (GI) is similarly assessed as the angle between the glenoid axis and the transverse scapular axis, projected on the CSP as defined by Serrano et al.36.
The critical shoulder angle (CSA) is automatically measured, as defined by Moor et al.37, as the angle between the glenoid axis and the CSA-axis; defined as the line passing through the inferior-most extrema point of the lower glenoid rim and the inferolateral acromion extrema point projected in the true anterior-posterior direction. The CSA-projection plane is defined from the CSP rotated by the scapula’s respective GV, as described by Suter et al.6. The inferolateral acromion extrema point is evaluated using an optimization procedure, initialized using the anatomical landmark located on the anterior lateral corner of the acromion. The surface points within a 5 mm radius of this landmark are evaluated, and the point which yields the highest CSA angle is selected.
Evaluation
The evaluation of our pipeline including the accuracy of the automatic MRI segmentation, the HR-MRI-mask reconstruction, the landmark and axis prediction and the morphological analysis was performed on the test dataset of Dataset 1 (N = 10).
The performance of the proposed automatic scapula segmentation network was evaluated by comparing the masks predicted from MRI to the transferred manual masks from CT of the same shoulder using the Dice similarity coefficient38.
To evaluate the performance of the HR-MRI-mask reconstruction algorithm, the predicted HR-MRI-mask and CT masks of the same shoulder were compared. To account for the restricted FOV of the MRI and only compare the area inside the FOV of the MRIs, the masks were cropped 50 mm medially and 40 mm inferiorly from the centre of the glenoid. The cropped models were aligned using the iterative closest point (ICP) optimisation algorithm and the root mean square error (RMSE) of the models was calculated.
The accuracy of the proposed automatic landmark detection method was quantified using the Euclidian distance between the predicted landmark centre position on the HR-MRI-masks and the ground-truth manual landmark positions on the manual CT masks. The angular errors of the scapula axis (z-axis) and the CSP of the HR-MRI-masks and those defined by the manual landmarks were also measured.
Ground-truth scapula morphology parameters, including the glenoid radius, the glenoid height, the GV, the GI, and the CSA, were calculated by applying the algorithms described in Chaps. 2.3.4 and 2.3.5 to the CT manual segmentation and manually depicted landmarks. The mean absolute errors between the automatically evaluated scapula morphology parameters on the HR-MRI-mask and the ground-truth morphology analysis on CT, were calculated. The Wilcoxon signed-rank test was performed to compare the differences between the values calculated from MRI and CT39 with a level of significance of p < 0.05. The intraclass correlation coefficient (ICC)40, was used to measure the absolute agreement between the methods. ICC estimates and their 95% confident intervals (95%CI) were calculated using the Pingouin41 Python statistical package, version 0.5.4, based on a single rater, absolute-agreement, 2-way mixed-effects model. The ICC results were further categorized according to Koo & Li et al.40.
Results
Segmentation accuracy
The mean Dice similarity coefficient score between the transferred manual segmentation from CT, and the automated segmentation of the scapula from MRI was 0.87 ± 0.05.
High-resolution-MR image mask precision
In the test dataset, the MRI with the FOV per plane was used to generate the high-resolution MRI mask for each of the 10 patients. Among these, 6 coronal, 8 sagittal and 6 transversal MRI were T1-weighted and the rest T2- (4) or pd- (6) weighted. Despite using MRIs with the largest FOV, no scapula was fully contained within the MRI dataset FOV.
The mean and standard deviation RMSE between the aligned cropped HR-MRI masks and the manually segmented CT masks was 1.34 mm ± 0.26 mm. Figure 5 depicts the 3D model from a HR-MRI mask, the CT mask and the color-coded absolute error surface distance map on the CT mask between the HR-MRI and CT masks of a patient with an RMSE of 1.54 mm.
3D models from a patient with RMSE of 1.54. (a) 3D model of the HR-MRI-mask, (b) 3D model from CT, (c) color-coded map of absolute error surface distance between HR-MRI and CT 3D-models (blue = 0 mm, to yellow ≥ 5 mm).
Landmark and axis prediction accuracy
The mean and standard deviation Euclidian distance between the automatically detected scapular landmarks on HR-MRI and CT masks was 2.0 mm ± 1.0 mm, 2.4 mm ± 1.5 mm and 2.0 mm ± 0.8 mm, for the anterior-, inferior- and posterior glenoid rim point, respectively. The superior glenoid tubercule was detected with a Euclidian distance of 2.1 mm ± 0.7 mm to the manual landmarks on the CT mask and the acromion lateral corner with a Euclidian distance of 3.3 mm ± 1.6 mm. The predicted transverse scapular axis had a mean angular error of 3.6° ± 1.8° and the predicted CSP normal a mean error of 4.7° ± 4.3°.
The CSA projection plane was predicted with a mean and standard deviation error of 0.2° ± 3.1° around the y-axis (corresponding to ante- and retro- version error) and 1.0° ± 5.5° around the transverse scapular axis (corresponding to an error of extension).
The mean and standard deviation Euclidian distance between the extrema points on the HR-MRI and CT masks were 2.7 mm ± 1.2 mm and 3.3 mm ± 1.1 mm for the inferior-most and superior-most glenoid points, respectively, and 5.9 mm ± 3.1 mm for the CSA point on the acromion.
Morphology parameters
The results of the comparison between the morphological measures taken from manual landmarks and segmentations on CT images, and our automatic pipeline with automatic segmentation and automatic landmark detection on MRI are shown in Table 2. For the glenoid anatomic dimensions, the mean difference was less than one millimetre between the two methods (-0.7 ± 0.8 mm for the glenoid radius and − 0.3 ± 1.9 mm for the glenoid height respectively). For the morphological metrics, the CSA had a mean error of -1.3 ± 1.7 degrees, the GI, a mean error of 1.3 ± 2.1 degrees, and the GV, a mean error of -1.4 ± 3.4 degrees. For the glenoid anatomic dimensions, the absolute agreement between the CT and MRI measurements was good, with ICC values of 0.81 (95%CI 0.40–0.95; poor to excellent) for the glenoid radius, 0.89 (95%CI 0.62–0.97; moderate to excellent) for the glenoid height. The absolute agreement between the and the GI and CSA was excellent 0.93 (95%CI 0.74–0.93; moderate to excellent) for the GI and 0.91 (95%CI 0.69–0.98; moderate to excellent) for the CSA respectively. For the GV, the absolute agreement was moderate, with an ICC of 0.73 (95%CI 0.23–0.93; poor to excellent). CT and MRI based measures were only statistically significantly different in case of the GR and the CSA.
Discussion
We described a deep learning-based method that enabled the automatic calculation of scapular morphological metrics on diagnostic MRI with anisotropic resolution and reduced FOV. For all morphological measurements the ICC was substantial or almost perfect, highlighting the feasibility of 3D MRI based analysis for clinical diagnosis.
The proposed pipeline ensures compatibility with common clinical practices, by requiring a single transverse, coronal, and sagittal MRI, in any of the commonly available diagnostic T1-, T2 or pd-weighted sequence. The pipeline starts with deep learning-based scapula segmentation from the MRI in these planes. Subsequently, high-resolution 3D models are generated from these segmentations. Finally, we present a deep learning-based landmark and axes detection on the high-resolution mask for morphology calculation.
With the proposed network, the scapula was segmented with a Dice similarity coefficient of 0.87 ± 0.05 compared to CT. This Dice coefficient value is lower than our previously presented value for automatic deep learning-based segmentation of the scapula from T1-weighted MRI, which was 0.92 ± 0.0517. However, in this work the segmentation accuracy was evaluated relative to CT-derived segmentations, while previous evaluations were performed relative to manual segmentation from MRI only. When the network presented in17 was applied to the T1-weighted MRI images from our test dataset, an average Dice similarity coefficient of 0.79 ± 0.08 was achieved compared to the CT-derived ground-truth segmentation. This indicates a significant improvement in segmentation accuracy with our proposed approach. Manual segmentation borders of the scapula in MRI are placed considering the visualized structures in the image, potentially leading to misalignments compared to the true bone borders. Due to improved imaging quality of skeletal structures in CT, the accuracy of manual segmentation of the scapula in CT is significantly higher than that of MRI. Using CT segmentation for network training thus allows the network to predict the scapula borders more accurately than can be visualised in MRI alone. The lower Dice similarity coefficient of the network, when compared to CT-derived scapula segmentations, might be due to the border of the transferred mask from CT not aligning with any clearly visible structure in an MRI, making it harder for the network to predict accurately. Compared to alternative approaches for automatic segmentation of the scapula in anisotropic MRI using anatomical priors, our algorithm achieved superior precision42,43.
The RMSE of the test-patients’ 3D high resolution models of the scapula (N = 10) compared to CT was 1.34 mm ± 0.26 mm. Figure 4 shows a typical distance map of an example patient with average RMSE. Visually, large surface differences between the HR-MRI models to the CT models were located in the periphery and mainly at the medial parts of the scapula. In regions where the morphology measurements are performed, such as the glenoid and the acromion, these errors were significantly lower. This might be due to the fact, that these regions are visualized in the MRI along all orientations whereas the structures in the periphery are often outside the FOV of one or multiple planes.
The GI was measured automatically on the MRI data with a mean error of -1.4° ± 3.4° and an ICC of 0.93 (95%CI 0.74–0.98) which is superior to published interobserver variations of − 3.0° ± 3.6° for CT44 and the ICC of 0.88 (95%CI 0.78–0.94) between manual MRI and CT measurements45. The GV was measured with a mean absolute error of 2.6° ± 2.5° and an ICC of 0.73 (95%CI 0.23–0.93) on MRI data compared to the measurement on CT. These errors are comparable or lower to discrepancies observed between the GV and GI measurements of two commercially available shoulder arthroplasty planning systems, where GI measurement differences were less than 5° in only 54% of cases, and GV measurement differences were less than 5° in 70% of cases46. This ICC is in the range of a previously reported interobserver ICC of 0.68 (95%CI 0.45–0.83) on CT and 0.76 (95%CI 0.57–0.88) on MRI47. The Pearson correlation of GV measured on MRI compared to CT was r = 0.75, superior to that found by Parada et al., who documented a correlation of r = 0.63 between manual MRI and CT GV measurements47. Additionally, in47 measurements were performed on MRI-images with at least 75% of the scapular width. The differences of the CSA measurements from MRI and CT were statistically significant, however the mean absolute error was low with 1.8° ± 1.1° and the ICC was high with 0.91 (95%CI 0.69–0.98). This mean absolute error is higher than previously reported absolute interobserver agreement of 1.0° ± 1.0° if measured manually on CT6 but is lower than the accepted errors if measured on radiographic images6. Additionally, our ICC is higher than the ICC reported between raters of 0.75 (95%CI not reported) if manually measured on MRI48, where clinicians used T1- and T2-weighted MRIs to perform three-dimensional CSA measurements in the sagittal MRI plane, marking the lateral-most point of the acromion and scrolling the images to the glenoid midline for measurements. The glenoid radius measurement from MRI and CT were significant different. However, the mean absolute error of the glenoid height and radius between those automatically measured on MRI and those from CT were comparable to those reported between manual and semi-automatic measurements on the same CT data49.
The main scapula axes are significant for accurate morphology analysis as they impact GI5,50, the CSA12 and GV51,52 measurements. Until now, the need for accurate imaging of the full scapula for definition of the scapular axes has limited morphology analysis on MRI. We have demonstrated the feasibility of alternatively predicting the required axes directly from a partial scapular model, eliminating reliance on landmarks on the often cropped medial and proximal borders. This method allows for greater variation in imaging protocols and relaxes segmentation accuracy requirements for the thin medial scapular border. While the mean orientation error of our predicted measurement plane for the CSA was 0.24° ± 3.07° is in an acceptable projection error range12, the mean angular error of the predicted transverse scapular axis of 3.60° ± 1.82° could have negatively influenced the accuracy of the automatic GV measurements52.
Our CSA and GI measurement error on MRI compared to CT falls below the range of reported clinically relevant differences. Studies on patients with isolated supraspinatus tear reported CSA differences of 2° to 4.3° between patients with retear and without retear after surgical repair3,53,54. Another study on RC tear patients found GI differences of 5° between patients without retear and full retear after surgical repair3. However, considering that previous morphology studies of RC tear patients were performed in 2D on MRI or radiography, which are known to suffer from large measurement errors, we believe that our proposed method has the potential to enable the discovery of more clinically significant morphological differences of various RC tear patient groups.
One limitation of our study is the limited size of the test dataset, which restricts the ability to draw definitive conclusions regarding the significant difference of the accuracy of landmark measurement between MRI and CT imaging. The mean differences between MRI and CT morphology measurements also indicate a tendency of our methods to underestimate the CSA and GV and overestimate the GI. However, testing on larger datasets would be required to determine the significance of such trends and to identify if these underestimations are due to errors of the automatic segmentation, or due to the different resolution of the MRI and CTs as observed by Neubert et al.55, or if the proposed combination algorithm lead to a slightly smaller scapula model. Furthermore, in this study, the performance of the proposed algorithms was not tested on shoulders with bone defects or with arthritic changes. Accurate shoulder morphology analysis is expected to be especially challenging on arthritic glenoids with osteophytes due to their fine structures. Future studies will investigate the accuracy of these methods in such patients, and the influence of MRI resolution and weighting.
To further improve the accuracy of segmentation obtained from conventional diagnostic MRI, our method could be refined to use a regional weighting function when performing the multivariate interpolation of the three anatomical planes. This refinement could emphasize the information contained in a specific plane in certain regions of the scapula, enhancing segmentation precision. In the future, scapular shape completion using statistical shape model fitting or deep learning-based shape completion could alternatively be explored to address the challenge of medial and inferior landmarks not available within the current FOV. This enhancement may improve accuracy in detecting the CSP and transverse scapular axis, thereby improving morphological analysis precision. More broadly, the methods presented herein for 3D scapula modelling from MRI hold potential to eliminate the reliance on CT in applications such as shoulder arthroplasty planning.
Conclusions
In conclusion, our study demonstrates the feasibility and accuracy of a fully automatic 3D morphology analysis pipeline for diagnostic MR arthrogram data. Despite the challenges posed by the reduced resolution and restricted FOV of diagnostic MRI our method enabled robust segmentation and landmark detection, providing reliable scapular morphology measurements with clinically acceptable precision even in diverse imaging settings. Furthermore, our approach facilitates the application of 3D scapula morphology analysis to RC tear patients without the reliance on CT imaging, enabling the standardised investigation of large cohorts, and versatility for extension to include both established and novel morphological parameters.
Data availability
The data that support the findings of this study are available on request from M.Z. The data is not publicly available due to the contained information that could compromise the privacy of the research participants.
References
Zaid, M. B. et al. Radiographic shoulder parameters and their relationship to outcomes following rotator cuff repair: a systematic review. Shoulder Elb. 13, 371–379 (2021).
Zumstein, M. A., Jost, B., Hempel, J., Hodler, J. & Gerber, C. The clinical and structural long-term results of open repair of massive tears of the rotator cuff. J. Bone Joint Surgery-American Volume. 90, 2423–2431 (2008).
Garcia, G. H. et al. Higher critical shoulder angle increases the risk of retear after rotator cuff repair. J. Shoulder Elb. Surg. 26, 241–245 (2017).
Sheean, A. J. et al. Does an increased critical shoulder angle affect re-tear rates and clinical outcomes following primary rotator cuff repair? A systematic review. Arthroscopy: J. Arthroscopic Relat. Surg. 35, 2938–2947e1 (2019).
Chalmers, P. N., Salazar, D., Chamberlain, A. & Keener, J. D. Radiographic characterization of the B2 glenoid: the effect of computed tomographic axis orientation. J. Shoulder Elb. Surg. 26, 258–264 (2017).
Suter, T. et al. The influence of radiographic viewing perspective and demographics on the critical shoulder angle. J. Shoulder Elbow Surg. 24, e149–e158 (2015).
Hoenecke, H. R., Hermida, J. C., Flores-Hernandez, C. & D’Lima, D. D. Accuracy of CT-based measurements of glenoid version for total shoulder arthroplasty. J. Shoulder Elb. Surg. 19, 166–171 (2010).
Chalmers, P. N. et al. Influence of radiographic viewing perspective on glenoid inclination measurement. J. Shoulder Elb. Arthroplasty. 3, 2471549218824986 (2019).
Welsch, G. et al. CT-based preoperative analysis of scapula morphology and glenohumeral joint geometry. Comput. Aided Surg. 8, 264–268 (2003).
Yoo, J. C. et al. Correlation of arthroscopic repairability of large to massive rotator cuff tears with preoperative magnetic resonance imaging scans. Arthroscopy: J. Arthroscopic Relat. Surg. 25, 573–582 (2009).
Brunner, U. et al. S2e-Leitlinie „Rotatorenmanschette AWMF-Leitlinien-Register Nr. 033/041 (2017).
Yanke, A. B. et al. Three-dimensional magnetic resonance imaging quantification of glenoid bone loss is equivalent to 3-dimensional computed tomography quantification: cadaveric study. Arthroscopy: J. Arthroscopic Relat. Surg. 33, 709–715 (2017).
Gyftopoulos, S. et al. 3DMR osseous reconstructions of the shoulder using a gradient-echo based two-point Dixon reconstruction: a feasibility study. Skeletal Radiol. 42, 347–352 (2013).
Akbari-Shandiz, M. et al. MRI vs CT-based 2D-3D auto-registration accuracy for quantifying shoulder motion using biplane video-radiography. J. Biomech. 82, 375–380 (2019).
Vopat, B. G. et al. Measurement of glenoid bone loss with 3-dimensional magnetic resonance imaging: a matched computed tomography analysis. Arthroscopy: J. Arthroscopic Relat. Surg. 34, 3141–3147 (2018).
Lansdown, D. A. et al. Automated 3-dimensional magnetic resonance imaging allows for accurate evaluation of glenoid bone loss compared with 3-dimensional computed tomography. Arthroscopy: J. Arthroscopic Relat. Surg. 35, 734–740 (2019).
Hess, H. et al. Deep-learning-based segmentation of the shoulder from mri with inference accuracy prediction. Diagnostics (Basel Switzerland) 13 (2023).
Medina, G., Buckless, C. G., Thomasson, E., Oh, L. S. & Torriani, M. Deep learning method for segmentation of rotator cuff muscles on MR images. Skeletal Radiol. 50, 683–692 (2021).
Shim, E. et al. Automated rotator cuff tear classification using 3D convolutional neural network. Sci. Rep. 10, 15632 (2020).
Breighner, R. E. et al. Technical developments: zero echo time imaging of the shoulder: enhanced osseous detail by using MR imaging. Radiology 286, 960–966 (2018).
Mello, R. A. F. et al. Three-dimensional zero echo time magnetic resonance imaging versus 3-dimensional computed tomography for glenoid bone assessment. Arthroscopy: J. Arthroscopic Relat. Surg. : Official Publication Arthrosc. Association North. Am. Int. Arthrosc. Association. 36, 2391–2400 (2020).
Carretero-Gómez, L. et al. Deep learning-enhanced zero echo time MRI for glenohumeral assessment in shoulder instability: a comparative study with CT. Skeletal Radiol. https://doi.org/10.1007/s00256-024-04830-0 (2024).
Yuan, X. & Yuan, X. Fusion of multi-planar images for improved three-dimensional object reconstruction. Comput. Med. Imaging Graph. 35, 373–382 (2011).
Meyer, A. et al. 177–181 (2018).
Meyer, A. et al. Anisotropic 3D multi-stream CNN for accurate prostate segmentation from multi-planar MRI. Comput. Methods Programs Biomed. 200, 105821 (2021).
Shanmugalingam, K. et al. (eds) 217–226 (2024).
Xue, N., Doellinger, M., Ho, C. P., Surowiec, R. K. & Schwarz, R. Automatic detection of anatomical landmarks on the knee joint using MRI data. J. Magn. Reson. Imaging. 41, 183–192 (2015).
Fischer, M. et al. Automated morphometric analysis of the hip joint on MRI from the German national cohort study. Radiology: Artif. Intell. 3, e200213 (2021).
Liu, S., He, J. L. & Liao, S. H. 1–6 (2020).
Rojas, J. T., Jost, B., Zipeto, C., Budassi, P. & Zumstein, M. A. Glenoid component placement in reverse shoulder arthroplasty assisted with augmented reality through a head-mounted display leads to low deviation between planned and postoperative parameters. J. Shoulder Elb. Surg. 32, e587–e596 (2023).
Verstraeten, T., de Wilde, L. & Victor, J. The normal 3D gleno-humeral relationship and anatomy of the glenoid planes. J. Belg. Soc. Radiol. 102, 18 (2018).
Pluim, J., Maintz, J. & Viergever, M. Mutual information matching in multiresolution contexts. Image Vis. Comput. 19, 45–52 (2001).
Marstal, K., Berendsen, F., Staring, M. & Klein, S. pp. 134–142. (2016).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods. 18, 203–211 (2021).
Friedman, R. J., Hawthorne, K. B. & Genez, B. M. The use of computerized tomography in the measurement of glenoid version. JBJS 74, 1032 (1992).
Serrano, N. et al. CT-based and morphological comparison of glenoid inclination and version angles and mineralisation distribution in human body donors. BMC Musculoskelet. Disord. 22, 849 (2021).
Moor, B. K., Bouaicha, S., Rothenfluh, D. A., Sukthankar, A. & Gerber, C. Is there an association between the individual anatomy of the scapula and the development of rotator cuff tears or osteoarthritis of the glenohumeral joint? Bone Joint J. 95-B, 935–941 (2013).
Dice, L. R. Measures of the amount of ecologic association between species. Ecology 26, 297–302 (1945).
Wilcoxon, F. Individual comparisons by ranking methods. Biometrics Bull. 1, 80–83 (1945).
Koo, T. K. & Li, M. Y. A guideline of selecting and reporting intraclass correlation coefficients for reliability research. J. Chiropr. Med. 15, 155–163 (2016).
Vallat, R. Pingouin: statistics in Python. JOSS 3, 1026 (2018).
Boutillon, A., Borotikar, B., Burdin, V. & Conze, P. H. 1164–1167 (2020).
Yang, Z. et al. Automatic bone segmentation and bone-cartilage interface extraction for the shoulder joint from magnetic resonance images. Phys. Med. Biol. 60, 1441 (2015).
Maurer, A. et al. Assessment of glenoid inclination on routine clinical radiographs and computed tomography examinations of the shoulder. J. Shoulder Elb. Surg. 21, 1096–1103 (2012).
Chalmers, P. N., Beck, L., Granger, E., Henninger, H. & Tashjian, R. Z. Superior glenoid inclination and rotator cuff tears. J. Shoulder Elb. Surg. 27, 1444–1450 (2018).
Denard, P. J. et al. Version and inclination obtained with 3-dimensional planning in total shoulder arthroplasty: do different programs produce the same results? JSES open. Access. 2, 200–204 (2018).
Parada, S. A. et al. Magnetic resonance imaging correlates with computed tomography for glenoid version calculation despite lack of visibility of medial scapula. Arthroscopy: J. Arthroscopic Relat. Surg. 36, 99–105 (2020).
Schiefer, M. et al. MRI is a reliable method for measurement of critical shoulder angle and acromial index. Rev. bras. Ortop. 58, 719–726 (2023).
Rodrigues, C. Three-dimensional MRI bone models of the glenohumeral joint using deep learning: evaluation of normal anatomy and glenoid bone loss. Radiol. Artif. Intell. 2, e190116 (2020).
Daggett, M., Werner, B., Gauci, M. O., Chaoui, J. & Walch, G. Comparison of glenoid inclination angle using different clinical imaging modalities. J. Shoulder Elb. Surg. 25, 180–185 (2016).
Budge, M. D. et al. Comparison of standard two-dimensional and three-dimensional corrected glenoid version measurements. J. Shoulder Elb. Surg. 20, 577–583 (2011).
Bryce, C. D. et al. Two-dimensional glenoid version measurements vary with coronal and sagittal scapular rotation. JBJS 92, 692 (2010).
Gerber, C., Catanzaro, S., Betz, M. & Ernstbrunner, L. Arthroscopic correction of the critical shoulder angle through lateral acromioplasty: a safe adjunct to rotator cuff repair. Arthroscopy: J. Arthroscopic Relat. Surg. 34, 771–780 (2018).
Scheiderer, B. et al. Higher critical shoulder angle and acromion index are associated with increased retear risk after isolated supraspinatus tendon repair at short-term follow up. Arthroscopy: J. Arthroscopic Relat. Surg. 34, 2748–2754 (2018).
Neubert, A. et al. Comparison of 3D bone models of the knee joint derived from CT and 3T MR imaging. Eur. J. Radiol. 93, 178–184 (2017).
Author information
Authors and Affiliations
Contributions
H.H and K.G. conceived of the presented idea. T.J.R., A.L. and M.Z. processed and analysed the clinical data. H.H. and A.O designed the computational framework. H.H. and A.O performed the computations and the comparative analysis. K.G. helped supervise the project. H.H., A.O. and K.G. wrote the manuscript with consultation from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Alexandra Oswald: The author’s work is funded by the Innosuise Grant 35656.1 IP-LS “Computer Assisted Planning for Rotator Cuff Repair” which includes partial funding by Synthes GmbH. J. Tomas Rojas: The author, their immediate family, and any research foundation with which they are affiliated have not received any financial payments or other benefits from any commercial entity related to the subject of this article. Alexandre Lädermann: This author is a paid consultant for Arthrex, Stryker, Medacta, and Enovis. He received royalties from Stryker and Medacta. He is the (co-)founder of FORE, Med4Cast, and BeeMed. He owns stock options in Medacta and Follow Health. He is on the board of the French Arthroscopic Society. Matthias A. Zumstein: This author reports grants from Medacta and non-financial support from Angiocrine Biosciences, outside the submitted work. Kate Gerber: The author’s work is funded by the Innosuise Grant 35656.1 IP-LS “Computer Assisted Planning for Rotator Cuff Repair” which includes partial funding by Synthes GmbH.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hess, H., Oswald, A., Rojas, J.T. et al. Deep learning algorithms enable MRI-based scapular morphology analysis with values comparable to CT-based assessments. Sci Rep 15, 1591 (2025). https://doi.org/10.1038/s41598-024-84107-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-84107-7