Abstract
In European seabass (Dicentrarchus labrax), dietary tryptophan (TRP) surplus has a notable modulatory effect on the hypothalamic–pituitary–interrenal axis under chronic stress and acute inflammation, affecting cortisol levels and neuroendocrine- and immune-related gene expression. A transcriptomic approach (RNA-seq) was applied to head-kidney samples of fish submitted to confinement stress and/or acute inflammation to uncover the biological mechanisms behind these effects. Undisturbed seabass fed dietary TRP supplementation showed an up-regulation of various innate immune functions, contrasting previous studies which indicated mainly a TRP regulatory role. Upon bacterial injection, TRP-fed fish showed a transcriptomic profile similar to their counterparts fed on control diet, indicating TRP’s inability to modulate immune mechanisms under bacterial challenge. Under confinement stress, TRP-fed fish exhibited a molecular profile similar to unstressed control fish, highlighting TRP’s role in mitigating stress. However, combining dietary TRP supplementation with confinement stress and immune stimulation by bacterial inoculation resulted in a unique molecular profile. Stressed fish fed TRP did not show the restorative effect of immune stimulation on carbohydrate metabolism and showed downregulated genes related to glycolysis and glycogenolysis. Additionally, transcription upregulation in these fish after bacterial injection included terms related to serine and steroid metabolism (carboxyl ester lipase 2), indicating tryptophan-induced changes in lipid mobilization in the head-kidney, potentially affecting cortisol synthesis and other hormones.
Similar content being viewed by others
Introduction
Immune-enhancing nutrition is currently used to maintain and improve animal health by including functional ingredients. In aquafeeds, the incorporation of functional ingredients, such as amino acids, stimulates or modulates fish immune system. The activation of the immune system, an inflammatory or a stress response, increases the metabolic energy needs, resulting in a depletion of essential nutrient provisions1,2. The supply of certain nutrients in the diets could be a strategy to be applied in specific situations when requirements are higher, such as anthropogenic stressful and immune-threatening situations3, inherent to aquaculture production4,5,6,7,8. Tryptophan is an essential amino acid (AA), only assimilated through diet since fish are not able to synthesize it endogenously4,9. This AA has recognized modulatory effects in the neuroendocrine system by being the precursor of the neurotransmitter serotonin (5-hydroxytryptamine; 5-HT) affecting both stress and behavioural responses10,11,12,13.
Previous studies with European seabass (Dicentrarchus labrax), Senegalese sole (Solea senegalensis), mrigal carp (Cirrhinus mrigala), common carp (Cyprinus carpio), Atlantic salmon (Salmo salar) and rainbow trout (Oncorhynchus mykiss) have consistently highlighted the modulatory role of tryptophan in fish stress mitigation14,15,16,17,18,19,20,21,22,23,24. Nevertheless, tryptophan modulation can yield both beneficial and adverse effects contingent upon the neuroendocrine condition of the fish14,15,16,20. For instance, a dietary tryptophan surplus enhanced Senegalese sole survival rates against bacterial infection if fish were reared under stressful conditions (high rearing density), whereas it worsened fish performance against disease when fish were kept in a lower, non-stressful rearing density16. Additionally, Lepage et al.20 demonstrated an increase in plasma cortisol levels of undisturbed rainbow trout triggered by graded tryptophan inclusion levels; in contrast, the same diets hampered cortisol secretion in fish subjected to an acute handling stressor. Immunosuppressive effects and a concomitant decrease in disease resistance were reported in European seabass-fed tryptophan-supplemented diets14,25. In particular, these fish presented lower gut superoxide activity and down-regulation of head-kidney key immune-related genes. However, if fish were in a previous state of stress, the tested diet improved inflammatory response, reduced plasma cortisol levels and up-regulated several immune-related genes following infection. Other studies using stressed European seabass-fed tryptophan-supplemented diets15, described, the modulatory role of tryptophan surplus in hypothalamus-pituitary-interrenal (HPI) axis and hepatic metabolism. In unstressed fish, a tryptophan surplus suppresses the up-regulation of pro-inflammatory cytokines (interleukin 6 and 1β) in response to intraperitoneal injection (i.p.) and increased expression of the serotonin-producing enzyme, tryptophan hydroxylase (TPHα), suggesting higher levels of this neurotransmitter15. Conversely, in stressed fish, after i.p. injection of Photobacterium damselae subspecies piscicida (Phdp) fed tryptophan supplemented diet counteracted the neuroendocrine response by decreasing pro-opiomelanocortin b-like levels and stress-induced cortisol, as well as by orchestrating the energy consumption for homeostatic condition15,24.
Considering all these findings, a comprehensive understanding of the modulatory effects of tryptophan dietary supplementation on the physiological panorama of European seabass becomes imperative. The present study aimed to unveil the intricate biological mechanisms associated with tryptophan supplementation in European seabass juveniles under stressful rearing conditions (i.e. crowding for 15 days) followed by immune stimulation, applying a transcriptomic approach.
Materials and methods
Diets composition
The experimental diets were formulated and manufactured by Sparos Lda. located in Olhão, Portugal. The control diet (CTRL) was specifically formulated to meet the indispensable AA profile recommended for European seabass, as outlined by Kaushik26. Additionally, a variant of the CTRL-based diet was developed by supplementing it with 0.3% L-tryptophan (dry matter; TRP), at the expense of wheat meal. Ingredients composition and AA profile of experimental diets were described by Machado et al.14 and Peixoto et al.15.
Phdp inoculum preparation
The inflammatory bacterial challenge was performed using Phdp strain PP3, isolated from yellowtail (Seriola quinqueradiata; Japan) by Doctor Andrew C. Barnes (Marine Laboratory, Aberdeen, UK), following the methodology described by Machado et al.14.
Experimental design
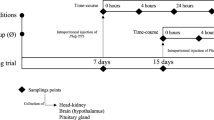
European seabass juveniles (12.02 ± 2.77 g) were obtained from Sonrı́onansa S.L. (Santander, Spain) and transported to the experimental facilities of CIIMAR (Portugal). Fish were randomly distributed in two independent recirculating seawater systems (8 tanks each, n = 22 per tank) with a density of 5 kg m−3 (temperature 20 ± 0.5 °C; salinity 32; photoperiod 10:14 h dark: light). In a complete randomized design, the two dietary treatments were evaluated in quadruplicate tanks of each system. Fish were fed these diets twice a day for 15 days, with a daily average ration of 2% of body weight. By lowering the water level in one of the systems, fish were kept within a low water column (≈ 10 cm height; 10 kg m−3) as a stressful condition. Groups will henceforth be named CTRL_Ø, CTRL_stress, TRP_Ø and TRP_stress. At the end of the feeding period, 12 fish per treatment were euthanized by an overdose of 2-phenoxyethanol (1 mL L−1, Merck KGaA) and head-kidney tissues were collected. These first sampled groups were considered non-injected (0 h, Fig. 1). The remaining fish were i.p. injected with 100 µL of Phdp (5 × 107 cfu mL−1) and moved to a new system with the same dimensions to avoid horizontal infection with other groups (temperature 24 ± 0.5 °C; salinity 32‰; photoperiod 10:14 h dark: light) and similarly sampled at 24-hours post-infection (12 fish per treatment). No mortality was observed during the trial.
Experimental setup of dietary tryptophan supplementation under confinement conditions followed by an acute inflammation. RAS – recirculating aquaculture system.
RNA extraction and sequencing
Head-kidney samples were individually processed for total RNA extraction using the NZY Total RNA Isolation kit (NZYTech) following the manufacturer’s instructions. The RNA was measured with an Invitrogen Qubit 3.0 Fluorometer using the Qubit RNA HS Assay Kit (ThermoFisher Scientific), following the manufacturer’s instructions. Pools of 3 samples were made with an RNA concentration of 100 ng µL−1 in a final volume of 30 µL, rendering 4 pools for each treatment (96 fish in total). Samples were barcoded and prepared for sequencing by Macrogen, Inc. (Republic of Korea) using TruSeq Stranded mRNA LT Sample Prep Kit on an Illumina NovaSeq 6000 as 101 bp paired-end reads with an average of 25 M reads (minimum 23 M) and RNA sequenced using 3’mRNA tag libraries.
RNA-seq analysis
The sequencing output quality was assessed using FastQC v0.12.0 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and low-quality reads (Phred quality score < 15 and read length < 30 bp) were discarded using Fastp v0.23.427 (Table S1 and Fig. S1). Transcript-level expression was estimated using Kallisto v.0.44.028 using the seabass transcriptome as reference (annotation and gene names extracted from Ensembl using Biomart, dlabrax2021 (GCA_905237075.1). Transcript level expression was imported into R v4.0.2 and summarised to the gene level using the R/tximport v1.10.1. Principal component analysis (PCA) was performed using R/plotPCA29 to visualize the spatial distribution of the dietary groups at each sampling time. Differential expression analysis was performed using R/Deseq2 v1.28.129 (Table S2), and genes with False Discovery Rate (FDR) adjusted p-values < 0.05 were considered differentially expressed. The experiment, rearing conditions and dietary treatments, were carefully designed to allow the evaluation of effects induced by diet, inflammation and stress independently, but also of their interactions. For the analysis of differentially expressed genes (DEGs), transcriptomes of all groups (CTRL_Ø, CTRL_stress, TRP_Ø and TRP_stress sampled at either 0–24 h) were compared to that of CTRL_ Ø sampled at 0 h.
Gene identification and their associated pathways
To improve the annotation of the European seabass transcriptome, canonical protein sequences for European seabass genes were obtained from Ensembl and used for protein-protein BLAST (BLASTp, NCBI) searches against the NCBI standard database. European seabass orthologues for literature-annotated sets of zebrafish (Danio rerio) genes were identified using g: Profiler (version e109_eg56_p17_1 d3191 d) with g: Orth for mapping orthologous genes across species. After that, gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG)30,31,32 enrichment analyses were performed using g: GOSt for functional enrichment analysis33 with Benjamini-Hochberg FDR for multiple testing correction.
Results
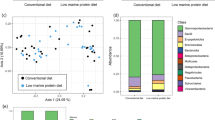
A total of 32 head-kidney pools were RNA sequenced, producing an average of 25 M reads per pool. A principal component analyses (PCA) showed two well-defined clusters distanced by the first principal component, and the main factor underlying this separation was sampling time (0 and 24 h-post-injection; Fig. 2A). On the cluster formed by fish sampled before injection (0 h), there was a secondary, less conspicuous separation between two groups, with dietary treatment being the main factor (second principal component), regardless of rearing condition. As opposed to the pre-injection scenario, at 24 h, a single cluster was observed, with no clear separation between elements (i.e. no clear influence of dietary treatment or rearing condition, Fig. 2A).
Transcriptome modulations in the head-kidney of European seabass-fed experimental diets (CTRL and TRP) during 15 days under stressful crowding conditions (Stressed) or not (Ø), followed by a bacterial challenge. (A) Principal Components Analysis showing the clustering of RNA-seq data. (B) Number of differentially expressed genes (DEGs). For fish sampled 24 h post-injection a log2 (Fold Change) ≥ 5 was applied.
Differential gene expression analysis
Differential expression analyses between the different groups revealed between 7 and 126 DEGS per comparison, where the complete differential expression results are available in Supplementary Tables S1 and S2.
Before immune stimulation (0 h, left panel of Fig. 2B), and in the absence of stressful conditions, the effect of tryptophan dietary supplementation was mild (TRP_Ø - a total of 32 DEGs). In control diet fish, a stronger effect of stressful conditions was observed (CTRL_stress – a total of 78 DEGs), but TRP supplementation seemed to overturn the impact of stress, as almost no DEGs were observed (TRP_stress – a total of 7 DEGs).
Regarding transcriptomes of fish held in similar experimental conditions but subsequently immune-stimulated by i.p. bacterial injection (24 h, right panel of Fig. 2B), due to the very high number of DEGs obtained in the analysis, only those with log2 (Fold Change) ≥ 5 will be considered for subsequent analytical approaches. In what the effect of diet is concerned, CTRL_Ø and TRP_Ø displayed a similar number of DEGs (a total of 59 and 58 DEGs, respectively). Stressful conditions significantly affected the transcriptome of infected fish (CTRL_stress – a total of 103 DEGs) while this number was slightly increased when TRP was provided instead of CTRL (TRP_stress – a total of 126 DEGs).
Transcriptomic changes induced by stress during infection
To assess stress-induced transcriptomic changes in fish before and after a bacterial challenge, gene expression of CTRL_stress fish (sampled at 0–24 h) was compared to that of CTRL_Ø_0 h fish, and enrichment analyses were performed (Supplementary Table S5 and S6).
The up-regulation of genes in non-infected CTRL_stress fish led to the enrichment of GO terms and KEGG pathways mostly associated with regulatory processes, immune response and energy consumption, such as “ATP adenylyltransferase activity” (GO:0003877), “glutathione transferase activity” (GO:0004364) and “methionine adenosyltransferase complex” (GO:0048269) (Figs. 3 and 4 and Table S5). The DEGs involved in these enriched terms were microsomal Glutathione S-Transferase 1 (MGST1.2), methionine adenosyltransferase 2 A (MAT2 AL) and lysyl-tRNA synthetase 1 (KARS1; Tables S3 and S5).
The enriched terms and KEGG pathways related with genes that were found downregulated in non-infected, stressed fish were mostly associated with lipids, carbohydrates and amino acids metabolism, as well as homeostasis (Fig. 5 and Table S5): “carbohydrate metabolic process” (GO:0005975), “Glycolysis/Gluconeogenesis” (KEGG:00010) and “Biosynthesis of amino acids” (KEGG:01230; Fig. 5 and Table S5). Some of the DEGs fuelling the GO terms enrichment analysis were phosphofructokinase (PFKma), aldolase a, fructose-bisphosphate aldolase b (ALDOAb), glycerol-3-phosphate dehydrogenase 1b (GPD1b) and phosphorylase, glycogen, muscle A (PYGMA; Fig. 6A, Tables S3 and S5). The same genes (PFKma and ALDOAb) encoding phosphofructokinase-1 (EC 2.7.1.11) and fructose-bisphosphate aldolase (EC 4.1.2.13) enzymes (Fig. 4A-C) contributed to the different KEGG pathways identified, with a fold (Log2) increase of 5 (Fig. 6A, Tables S3 and S5).
When CTRL_stress fish were exposed to infection, genes related to immune and endocrine processes, amino acids biosynthesis, energy consumption and cell metabolism were significantly upregulated (Fig. 3–4 and Table S6). These changes in the upregulated genes were responsible for the enrichment of GO terms and KEGG pathways associated with “lysozyme activity” (GO:0003796), “defence response” (GO:0006952) and response to stress” (GO:0006950; Figs. 3 and 4 and Table S6). For the enrichment of these terms and pathways, the following DEGs were involved: interleukin 4 induced 1 (IL4il1), cysteine dioxygenase 1 (CDO1), serum amyloid A (SAA), lysozyme g-like (LYGL1), CC Motif Chemokine Ligand 19 (CCL19a) and fibrinogen gamma gene (FGG; Fig. 6B and Table S6). No immune or endocrine-related terms were found among the down-regulated genes in immune-stimulated fish following infection.
Bubble plot illustrating the enrichment analysis results of up-regulated differentially expressed genes involved in immune and cell responses of European seabass fed a TRP-supplemented diet during 15 days under crowding conditions (stress) or not (Ø), followed by a bacterial challenge. Benjamini-Hochberg False Discovery Rate (FDR as p-value) for simple multiple testing correction, measuring the expected proportion of false significant matches (Type I errors) within results. BP biologic process, MF molecular function, CC cellular component.
Bubble plot illustrating the enrichment analysis results of up-regulated differentially expressed genes involved in the metabolic response of European seabass fed a TRP-supplemented diet during 15 days under crowding conditions (stress) or not (Ø), followed by a bacterial challenge. Benjamini-Hochberg False Discovery Rate (FDR as p-value) for simple multiple testing correction, measuring the expected proportion of false significant matches (Type I errors) within results. KEGG Kyoto Encyclopedia of Genes and Genomes, BP biologic process, MF molecular function, CC cellular component.
Bubble plot illustrating the enrichment analysis results of down-regulated differentially expressed genes of European seabass fed a TRP-supplemented diet during 15 days under crowding conditions (stress) or not (Ø), followed by a bacterial challenge. Benjamini-Hochberg False Discovery Rate (FDR as p-value) for simple multiple testing correction, measuring the expected proportion of false significant matches (Type I errors) within results. KEGG Kyoto Encyclopedia of Genes and Genomes, BP biologic process, MF molecular function.
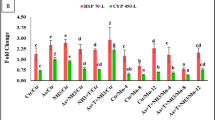
Expression profiles for selected key metabolic- (A) and immune-related genes (B). Log2 fold change values for selected genes of unstressed and crowding stressed fish for 15 days and fed either with CTRL or TRP-supplemented diets, and thereafter submitted or not to a bacterial challenge. A - PRSS1 – serine protease; ALDOAb – aldolase a, fructose-bisphosphate, b; PYGMma – phosphorylase, glycogen, muscle a; PFkma – phosphofructokinase, muscle a; ATP2a1 – ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1; TFa – transferrin-a; CEL.2 – carboxyl ester lipase, tandem duplicate 2; CHST7 – carbohydrate (N-acetylglucosamine 6-O) sulfotransferase 7; GPD1b – glycerol-3-phosphate dehydrogenase 1b. B - C9 – complement component 9; SAA – serum amyloid A; IL1β – interleukin 1β; MAP1a – microtubule-associated protein 1 Aa; PLP1b – proteolipid protein 1b isoform X1; LYGL1 – lysozyme g-like protein 1 precursor; CDO1 – cysteine dioxygenase type 1; CCL19 – C-C motif chemokine 19-like isoform x1; IL4i1 – L-amino acid oxidase 1 precursor. The complete dataset is available in Tables S3 and S4 of the Supplementary File.
Transcriptomic changes induced by tryptophan dietary supplementation following immune stimulation by bacterial infection
To evaluate TRP-induced transcriptomic changes in non-stressed fish before and after an immune challenge with virulent bacteria, gene expression of CTRL_Ø fish sampled at 24 h and that of TRP_Ø fish sampled at 0 and 24 h were all compared to that of CTRL_Ø_0 h fish, and enrichment analyses performed (Supplementary Tables S7–S9).
In non-infected fish, the transcriptomic up-regulation triggered by the TRP-supplemented diet contributed to the enrichment of terms associated to the immune response, such as “acute-phase response” (GO:0006953), “neutrophil mediated immunity” (GO:0002446), “hemoglobin metabolic process” (GO:0020027; Fig. 3 and Table S7). Contributing to these terms were genes such as serum amyloid A gene (SAA) and transferrin-a (TFA) (Fig. 6B and Tables S2 and S7). The enrichment analysis performed to downregulated DEGs of non-infected TRP_Ø fish revealed no major GO terms or KEGG pathways associated with immune and neuroendocrine response, or with metabolic mechanisms.
Upon i.p. bacterial infection, gene expression up-regulation of unstressed fish fed a control diet was mostly related to defence mechanisms, inflammation and metabolic pathways and some of the associated enriched terms and KEGG pathways were “defence response” (GO:0006952), “inflammatory response” (GO:0006954), “carbohydrate derivative binding” (GO:0097367, Figs. 3 and 4 and Tables S3 and S8). Contributing to these terms and KEGG pathways were genes such as SAA, LYGL1, CCL19 and interleukin 1β (IL1β) (Fig. 6 and Tab. S8).
In parallel, down-regulated DEGs contributed to the enrichment of terms such as “cell development” (GO:0048468), “neurogenesis” (GO:0022008) and “gliogenesis” (GO:0042063; Fig. 5 and Tables S1 and S6). The down-regulated genes that mostly contributed to the listed terms were microtubule-associated protein 1 Aa (MAP1aa) and proteolipid protein 1b isoform X1 (PLP1b; Fig. 6B and Table S8).
When immune-stimulated fish by bacterial infection were previously fed TRP for 15 days, a similar immune response was observed compared to those fed CTRL, regarding gene up-regulation (Figs. 2 and 3 and Table S8-S9). Regarding the metabolic response, a lower number of GO terms was observed compared to fish-fed CTRL after infection. No major terms and KEGG pathways associated with immune or neuroendocrine response were observed amongst the down-regulated genes, nor those related to metabolic mechanisms.
Transcriptomic changes induced by tryptophan dietary supplementation after crowding stressful conditions in combination with immune stimulation by bacterial infection
To evaluate TRP-induced transcriptomic changes in stressed fish before and after an immune challenge, gene expression of TRP_stress fish (sampled at 0 and 24 h) was compared to that of CTRL_Ø_0 h fish, and enrichment analyses performed (Supplementary Table S10 and S11).
The enrichment analysis performed on DEGs of non-infected TRP_stress fish rendered very few significant results, and no enriched terms or KEGG pathways related to immune, endocrine or metabolic processes (Figs. 3, 4, 5 and 6 and Table S10).
Concerning the enrichment analysis performed to up-regulated DEGs of TRP_stress fish submitted to a bacterial infection, a similar pattern of enriched GO terms and KEGG pathways to immune-stimulated CTRL-stress fish was found. These were mostly associated with metabolism and immunity: α-amino acid serine metabolism (serine-type endopeptidase activity - GO:0004252), immune response (lysozyme activity - GO:0003796, ID:3), and with the cholesterol metabolism (cholinesterase activity – GO:0004104; Steroid biosynthesis – KEGG:00100) (Figs. 3 and 4 and Table S10). The DEGs contributing to the enrichment of the abovementioned GO terms and KEGG pathways were serine protease (PRSS), IL1β, LYGL1, CDO1, SAA, IL4il, CCL19 and carboxyl ester lipase 2 (CEL.2, EC 3.1.1.; Figs. 4 and 6, Tables S4 and S8).
A higher number of enriched GO terms and KEGG pathways was found as a result of transcriptomic downregulation in infected TRP_stress fish than in their CTRL counterparts (Fig. 5). Enriched terms of downregulated genes in fish fed TRP were mostly associated with metabolism and AA biosynthesis: “generation of precursor metabolites and energy” (GO:0006091), “Biosynthesis of amino acids” (KEGG:01230) and Glycolysis/Gluconeogenesis (KEGG:00010; Fig. 5 and Table S11). The DEGs contributing to the enrichment of these GO terms and KEGG pathways included ALDOAb, PYGMa, PFKma and glycerol-3-phosphate dehydrogenase 1b (GPD1b; Fig. 6 and Tables S4 and S11).
Discussion
Different approaches to the study of tryptophan’s modulatory role in immune and neuroendocrine mechanisms of European seabass have shed light on its effects on the HPI axis response, central serotonergic activity, plasma cortisol and immune parameters, as well as fish metabolic responses14,15. However, an upstream overview of tryptophan-driven changes in the molecular response of European seabass to stressful conditions (prolonged crowding stress followed by immune stimulation) is here presented as a more in-depth approach. This will hopefully enlighten on tryptophan modes of action when offered above its nutritional requirement. For a clearer discussion of the results, this section will follow a similar structure to that used in the Results section.
Molecular patterns of stressed European Seabass and their response to immune stimulation by bacterial infection
It has been well documented that chronic stress dysregulates immune response by inducing enduring alterations in energetic metabolism, persistent low-level inflammation, and the suppression of immune cell function and cytokine production8,34. While acute stressors such as fish handling have been proven to enhance catabolic pathways that sustain very high energy demands35, the persistent presence of a chronic stressor might culminate in the establishment of inhibitory processes, such as the shutdown of energy production10. Results from the present study showed that chronic stress per se (in the absence of immune stimulation by bacterial challenge) led to a pronounced downregulation of genes encoding key enzymes of carbohydrates metabolism such as phosphofructokinase-1, fructose-bisphosphate aldolase and glycerol-3-phosphate dehydrogenase (glycolysis and glycolipids metabolism) but also enzymes related to mobilization of energy reserves and glycogenolysis (e.g. glycogen phosphorylase). In accordance, it can be suggested that crowding stress conditions for 15 days induced considerable suppressing effects in energy dynamics. The abovementioned enzymes have been described as tools for decreasing glycolysis rate during the stress response of fish and humans, increasing cytotoxicity, oxidative stress and decreasing ATP availability9,36,37,38. Given the important roles played by these molecules in the energy metabolism, their downregulation in chronically stressed fish might reflect metabolic changes induced by stressful conditions that eventually lead to a maladaptive response7,39,40. On the other hand, transcriptomes from chronically crowded fish contrasted previous studies and revealed an enhancement of immune-related and regulatory mechanisms, favoured by the increased expression of genes that translate to glutathione S-transferase 1 and methionine adenosyltransferase 2α, among others. These particular enzymes are respectively related to the control of oxidative stress and methionine metabolism, the latter being the source of S-adenosyl methionine, used in DNA methylation41. Chronic stress is described as affecting the immune system and the energetic-metabolic machinery that results in an impairment of metabolism, disease resistance, growth and behaviour, combined in what is known as the tertiary response to stress39.
At 24 h post-injection of virulent bacteria, the number of DEGs increased significantly in seabass that were both chronically stressed and infected (CTRL_stress sampled at 24 h). This experimental group was setup to assess the effects of a previous ongoing neuroendocrine response (to chronic crowding) on the development of an immune response following a trigger and for that, a comparison of their transcriptome to that observed in infected, non-stressed fish (CTRL_Ø sampled at 24 h) is more useful. The total number of DEGs in crowded stressed fish was almost twice the number observed in non-injected stressed fish following infection, suggesting a modulatory and synergistic effect of crowding stress in fish exposed to immune stimulation by virulent bacteria. And despite there being a more conspicuous increase in down-regulated genes of crowded stressed fish, both groups lacked any enrichment of immune, endocrine or metabolic-related terms. In turn, gene ontology analysis on upregulated DEGs of crowded stressed fish revealed a reduced number of immune-related GO terms, with a marked absence of others significantly enriched in non-crowded fish (e.g., neutrophil-mediated immunity, leukocyte chemotaxis, neutrophil migration). The most relevant genes contributing to this particular enrichment were those encoding serum amyloid A and CC motif chemokine ligand 19 (SAA, CCL19). SAA is known as a major acute-phase protein and exhibits increased expression in response to various inflammatory conditions42, and so does the cytokine CCL19, also known for its role in facilitating lymphocyte migration43,44. In addition to this apparent stress-mediated immunosuppression, the transcriptome of crowded stressed seabass following infection showed comparatively poorer amino acid metabolism than in immune-stimulated control fish. A previous approach in European seabass regarding the study of crowding conditions similar to those in the present work, provided evidence of treatment-induced alterations (tumor necrosis factor, interleukin 1 beta, 6 and 10) in both pre-inflammatory and post-inflammatory physiological status15. Asian seabass, Lates calcarifer, reared at high density, not only presented elevated plasma cortisol and glucose levels but also a decreased activity of immune-related enzymes, reflecting a compromised health status45. The present results corroborate those described above, suggesting that space confinement for 15 days before an inflammatory insult significantly affects both immune and metabolic responses in the European seabass.
Tryptophan-induced changes in the molecular patterns of European Seabass following infection
In the quest to unveil the interactions of tryptophan’s higher availability with the immune response after infection (TRP_Ø sampled at 24 h), the present study first looked into possible modulatory effects of tryptophan alone (TRP_Ø sampled at 0 h). Seabass juveniles fed TRP for 15 days showed a modest profile of DEGs compared to CTRL-fed counterparts. Amongst the upregulated genes, these fish presented increased gene expression of complement components, SAA and transferrin-a (TFA), implicated in immune defences and iron metabolism. This upregulation and the consequent enrichment of several innate immune-related GO terms was somewhat unexpected, as previous experiments with tryptophan dietary supplementation in European seabass in particular, and in mammals too, frequently pictured an anti-inflammatory role14,46,47,48.
In what the immune response is concerned, dietary tryptophan supplementation for 15 days (TRP_Ø sampled at 24 h) produced a modest number of changes in the transcriptomic profile compared to that of infected fish provided a control diet. This was particularly true amongst upregulated DEGs and their related enriched GO terms/KEGG pathways, in which fewer or less enriched immune-related terms were obtained. This outcome is somewhat expected, given the generally regulatory character attributed to tryptophan as a functional ingredient to modulate immunity15,25,49. Such regulatory mechanisms, though not yet extensively investigated, are believed to be partly mediated by metabolites borne from the kynurenine pathway50. Given that the gene expression of the first catalytic enzyme of the kynurenine pathway in immune cells, indoleamine 2,3-dyoxygenase, was not amongst any DEGs listed in this study, its importance during fish immune responses remains unclear, and so does the reasoning behind tryptophan-mediated immunosuppression previously witnessed.
Also noteworthy is the absence of enriched terms related to neuroendocrine processes since firstly, it is widely accepted that an ongoing immune response is centrally perceived as an acute stress and activates neuroendocrine mechanisms51; and secondly, there is evidence of tryptophan-induced alterations in serotonergic dynamics13. In teleosts and mammals, the increase in the tryptophan intake can increase its uptake in the brain where it is converted to serotonin52. This neurotransmitter can stimulate or inhibit adrenocorticotropic hormone (ACTH) production and thereby increase or inhibit cortisol production, depending on the fish stress condition20. Machado et al.25 and co-workers, reinforced the theory of Winberg and Lepage53 about increased plasma cortisol levels fed a tryptophan surplus resulting in inhibited innate immune defences. Tryptophan dietary supplementation has even been reported to compromise non-stressed Senegalese sole disease resistance16. In the present study, the lack of genes related to serotonin activity or any neuroendocrine-related mechanisms should therefore be subject of additional investigation. Still, the fact that a threshold of 5 was established as minimum fold change in injected fish gene expression, might have prevented the observation of neuroendocrine- and serotonin-related genes.
Unlike CTRL_Ø fish, TRP_Ø fish did not demonstrate, based on their gene ontology enrichment analysis, any important alterations in the development and proliferation of head-kidney peripheral nervous system (PNS) cells. GO terms associated with downregulated genes of CTRL_Ø_24 h point to a negative impact of the inflammatory response on the PNS, impairing macrophages’ glial cell development and proliferation, central system gliogenesis and neurogenesis. Under stressful conditions, cortisol elevated levels lead to a decrease in neurogenesis, cell proliferation, neuronal differentiation and cell survival as observed in the results described by Alfonso et al., Best et al. and Tea et al.54,55,56. In this study, the GO terms associated with cell development at the PNS were given by the expression of MAP1aa and PLP1b. Proteolipid protein (PLP1b) is described as being recruited into myelin from cellular functions, like neuronal process formation57,58,59, and MAP1aa as an essential step in neurogenesis60.
When tryptophan was provided before immune stimulation, such changes in the development and proliferation of head-kidney PNS cells were not observed. As abovementioned, these fish presented a less intense general response to the bacteria injection, as indicated by their FDR (p-value). This less intense response might be linked with less damage in the development and proliferation of head-kidney PNS.
Tryptophan-induced changes in molecular patterns of stressed European seabass and their response to immune stimulation by bacterial challenge
As discussed before in Sect. 4.1., crowding stress by itself (i.e., without infection) induced regulatory effects on energy metabolism and the homeostatic condition (a total of 78 DEGs). In contrast, TRP_stress fish showed no major transcriptome alterations, thus presenting a molecular profile similar to that observed in CTRL_Ø fish sampled at 0 h (a total of 7 DEGs). This outcome is in agreement with insights from previous research highlighting the role of tryptophan in mitigating stress in fish15,16,20,22. The present approach suggests that tryptophan dietary supplementation in chronically crowded-stressed fish may effectively alleviate the negative impacts of stress. Also of particular interest was the fact that the few immune-related genes whose expression was induced by TRP alone were unaffected under crowding conditions. Even though the stress response was attenuated by dietary treatment, crowding stress was still able to prevent dietary tryptophan changes. Thereby, these results seem to pinpoint a bidirectional, mutually inhibitory effect of this interaction.
In this study, the transcriptomic analysis in the head-kidney of European seabass revealed that tryptophan supplementation did not significantly alter fish response to immune stimulation by infection, but effectively alleviated the negative impacts of crowding stress. When all three conditions (dietary treatment, space confinement and immune stimulation) were simultaneously tested, fish presented a singular molecular profile not at all foreseen.
Firstly, the restorative effect of immune stimulation on carbohydrate metabolism was not observed when crowded stressed fish were fed TRP for 15 days, and TRP_stress fish sampled 24 h post-injection presented the same downregulated genes related to glycolysis and glycogenolysis (PFKma, ALDOAb, GPD1b and PYGMa) identified in crowded stressed fish fed CTRL. Indeed, infection with virulent bacteria seemed to have restored stress-impaired carbohydrate metabolism, as no inhibition of glucose metabolic enzymes was observed in immune-stimulated CTRL_stress fish. This restorative effect seems to be a compensatory mechanism of the inflammatory response that being an acute stressor itself, requires extra energy of fish. In crowded fish subsequently immune-stimulated by Phdp injection, and in other studies, higher activities of glycolytic enzymes were reported after exposure to an acute stressor in fish, reactivating glycogen breakdown into glucose24,61,62. However, when the dietary treatment was TRP, this restorative effect on impaired carbohydrate metabolism was not observed, and these fish presented the same metabolic inhibition. These results corroborate those previously obtained in a parallel approach by Peixoto et al.15, in which it is hypothesized that the dietary treatment TRP acted as a buffer against the acute stress induced by immune stimulation, inhibiting further neuroendocrine activation.
Secondly, amongst the enriched terms and KEGG pathways associated with transcription up-regulation there were terms related to serine and steroid metabolism. One particular gene of interest observed to be upregulated in this group was that of carboxyl ester lipase 2 (cel.2). All classes of steroid hormones including reproductive hormones and cortisol are synthesized de novo from the common precursor cholesterol63,64, frequently stored in the inert form of cholesteryl ester in lipid droplets within cells65,66. Carboxyl ester lipase (CEL) stands as a lipolytic enzyme, capable of hydrolysing cholesteryl esters67,68. In teleost fish, steroid biosynthesis is regulated by the HPI axis69. As mentioned above, the overall transcripts profile of crowded stressed fish fed a tryptophan-supplemented diet and sampled 24 h following infection resembled that of unstressed fish fed CTRL, unveiling a homeostatic trend conferred by tryptophan. Moreover, Machado et al.14 observed a decrease in plasma cortisol levels in stressed fish provided with a dietary tryptophan surplus compared to fish fed a control diet in the course of a similar inflammatory response. Having in mind that cholesterol, the substrate of CEL, is the precursor not only of cortisol but of several other hormones, the fact cel.2 expression was found upregulated in the present study shall not be necessarily associated with an increase in cortisol synthesis. Still, it is an indication of tryptophan-induced changes in head-kidney lipids mobilization that might in turn affect cortisol dynamics.
Conclusion
This study employed a transcriptomic approach to elucidate on the physiological mechanisms underlying chronic crowding stress and immune responses following infection in the head-kidney of European seabass, while also investigating the modulatory impact of dietary tryptophan supplementation on such responses.
Dietary tryptophan supplementation alone showed enrichment of several innate immune-related GO terms, which was somewhat unexpected, as previous experiments with tryptophan dietary supplementation in European seabass in particular, and in mammals too, frequently pictured a more inhibitory role.
Upon bacterial injection, fish fed TRP presented a similar transcriptomic profile to that of infected fish fed CTRL, highlighting the inability of TRP to modulate the activation of immune mechanisms upon a bacterial challenge.
When fish fed TRP were under crowding stress conditions, no major transcriptome alterations were observed, and these fish presented a molecular profile similar to that of unstressed and non-infected fish fed CTRL. This highlights the modulatory role of tryptophan in mitigating stress in fish.
Nevertheless, when dietary tryptophan supplementation was combined with stressful crowding rearing conditions and immune stimulation by virulent bacteria, the restorative effect of immune stimulation on carbohydrate metabolism observed in those fed CTRL was not observed, with genes related to glycolysis and glycogenolysis found to be downregulated. Moreover, steroid metabolism-related genes were upregulated (carboxyl ester lipase 2 - cel.2) shedding light on tryptophan-induced changes in head-kidney lipids mobilization that might in turn affect hormone synthesis such as cortisol. Nonetheless, further research is needed to comprehensively understand all the mechanisms involved in metabolic and neuroendocrine-immune responses.
Data availability
Data is provided within the supplementary information files and in the online OSF repository, DOI 10.17605/OSF.IO/UAFMW, https://osf.io/uafmw/?view_only=de703918908843f59ff6f299 d66aa67a.
References
Kiron, V. Fish immune system and its nutritional modulation for preventive health care. Anim. Feed Sci. Technol. 173, 111–133. https://doi.org/10.1016/j.anifeedsci.2011.12.015 (2012).
Andersen, S. M. et al. Dietary arginine affects energy metabolism through polyamine turnover in juvenile Atlantic salmon (Salmo salar). Br. J. Nutr. 110, 1968–1977. https://doi.org/10.1017/S0007114513001402 (2013).
F Grimble, R. Basics in clinical nutrition: Immunonutrition – Nutrients which influence immunity: effect and mechanism of action. e-SPEN Eur. E-J. Clin. Nutr. Metabolism. 4, e10–e13. https://doi.org/10.1016/j.eclnm.2008.07.015 (2009).
Conceição, L. E. et al. Dietary nitrogen and fish welfare. Fish. Physiol. Biochem. 38, 119–141. https://doi.org/10.1007/s10695-011-9592-y (2012).
Salamanca, N., Giraldez, I., Morales, E., de La Rosa, I. & Herrera, M. Phenylalanine and tyrosine as feed additives for reducing stress and enhancing welfare in Gilthead seabream and meagre. Animals (Basel). https://doi.org/10.3390/ani11010045 (2020).
Verburg-van Kemenade, B. M. L., Stolte, E. H., Metz, J. R. & Chadzinska, M. Academic Press, in Fish Neuroendocrinology Vol. 28 Fish Physiology (eds N Bernier, G Van Der Kraak, A Farrel, & C Brauner) Ch. 7, 313–364 (2009).
Tort, L. Stress and immune modulation in fish. Dev. Comp. Immunol. 35, 1366–1375. https://doi.org/10.1016/j.dci.2011.07.002 (2011).
Uren Webster, T. M. et al. Contrasting effects of acute and chronic stress on the transcriptome, epigenome, and immune response of Atlantic salmon. Epigenetics 13, 1191–1207. https://doi.org/10.1080/15592294.2018.1554520 (2018).
Herrera, M. et al. Metabolic and stress responses in Senegalese soles (Solea senegalensis Kaup) fed Tryptophan supplements: effects of concentration and feeding period. Animals (Basel). 9. https://doi.org/10.3390/ani9060320 (2019).
Schreck, C. & Tort, L. Fish Physiology (eds Carl, S. et al.) 2–34 (Academic Press, 2016).
Azeredo, R. Amino acids as novel nutraceutics to modulate immune mechanisms and increase disease resistance in fish PhD thesis, Faculty of Sciences of University of Porto (2017).
Li, P., Yin, Y. L., Li, D., Kim, S. W. & Wu, G. Amino acids and immune function. Br. J. Nutr. 98, 237–252. https://doi.org/10.1017/S000711450769936X (2007).
Höglund, E., Øverli, Ø. & Winberg, S. Tryptophan metabolic pathways and brain serotonergic activity: A comparative review. Front. Endocrinol. (Lausanne). 10, 158. https://doi.org/10.3389/fendo.2019.00158 (2019).
Machado, M. et al. Tryptophan modulatory role in European Seabass (Dicentrarchus labrax) immune response to acute inflammation under stressful conditions. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms232012475 (2022).
Peixoto, D. et al. Dietary Tryptophan intervention counteracts stress-induced transcriptional changes in a teleost fish HPI axis during inflammation. Sci. Rep. 14 https://doi.org/10.1038/s41598-024-57761-0 (2024).
Azeredo, R. et al. Dietary Tryptophan induces opposite Health-Related responses in the Senegalese sole (Solea senegalensis) reared at low or high stocking densities with implications in disease resistance. Front. Physiol. 10, 508. https://doi.org/10.3389/fphys.2019.00508 (2019).
Basic, D. et al. Short- and long-term effects of dietary l-tryptophan supplementation on the neuroendocrine stress response in seawater-reared Atlantic salmon (Salmo salar). Aquaculture 388–391, 8–13. https://doi.org/10.1016/j.aquaculture.2013.01.014 (2013).
Basic, D. et al. Changes in regional brain monoaminergic activity and temporary down-regulation in stress response from dietary supplementation with l-tryptophan in Atlantic Cod (Gadus morhua). Br. J. Nutr. 109, 2166–2174. https://doi.org/10.1017/S0007114512004345 (2013).
Hoseini, S. M. & Hosseini, S. A. Effect of dietary L-tryptophan on osmotic stress tolerance in common carp, Cyprinus carpio, juveniles. Fish. Physiol. Biochem. 36, 1061–1067. https://doi.org/10.1007/s10695-010-9383-x (2010).
Lepage, O., Tottmar, O. & Winberg, S. Elevated dietary intake of L-tryptophan counteracts the stress-induced elevation of plasma cortisol in rainbow trout (Oncorhynchus mykiss). J. Exp. Biol. 205, 3679–3687. https://doi.org/10.1242/jeb.205.23.3679 (2002).
Lepage, O., Vilchez, I. M., Pottinger, T. G. & Winberg, S. Time-course of the effect of dietary L-tryptophan on plasma cortisol levels in rainbow trout Oncorhynchus mykiss. J. Exp. Biol. 206, 3589–3599. https://doi.org/10.1242/jeb.00614 (2003).
Tejpal, C. S. et al. Dietary supplementation of L-tryptophan mitigates crowding stress and augments the growth in Cirrhinus mrigala fingerlings. Aquaculture 293, 272–277. https://doi.org/10.1016/j.aquaculture.2008.09.014 (2009).
Peixoto, D. et al. Synergistic effects of dietary Tryptophan and dip vaccination in the immune response of European Seabass juveniles. Int. J. Mol. Sci. 25 https://doi.org/10.3390/ijms252212200 (2024).
Peixoto, D., Martos-Sitcha, J. A., Costas, B., Azeredo, R. & Mancera, J. M. Tryptophan-supplemented diet modulates the metabolic response of European Seabass (Dicentrarchus labrax) juveniles reared under space-confined conditions and submitted to acute inflammation. Fish. Physiol. Biochem. 51, 10. https://doi.org/10.1007/s10695-024-01427-1 (2025).
Machado, M. et al. Dietary Tryptophan deficiency and its supplementation compromises inflammatory mechanisms and disease resistance in a teleost fish. Sci. Rep. 9, 7689. https://doi.org/10.1038/s41598-019-44205-3 (2019).
Kaushik, S. Whole body amino acid composition of European Seabass (Dicentrarchus labrax), Gilthead seabream (Sparus aurata) and turbot (Psetta maxima) with an Estimation of their LAA requirement profiles. Aquat. Living Resour. 355–358. https://doi.org/10.1016/S0990-7440(98)80007-7 (1998).
Carvalho, I. et al. Exploring the effects of dietary methionine supplementation on European Seabass mucosal immune responses against Tenacibaculum maritimum. Front. Immunol. 16, 1513516. https://doi.org/10.3389/fimmu.2025.1513516 (2025).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527. https://doi.org/10.1038/nbt.3519 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated Estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. & Goto, S. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/gkae909 (2000).
Raudvere, U. et al. G:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198. https://doi.org/10.1093/nar/gkz369 (2019).
Dhabhar, F. S. Effects of stress on immune function: the good, the bad, and the beautiful. Immunol. Res. 58, 193–210. https://doi.org/10.1007/s12026-014-8517-0 (2014).
Krasnov, A. et al. Transcriptome responses to carbon tetrachloride and pyrene in the kidney and liver of juvenile rainbow trout (Oncorhynchus mykiss). Aquat. Toxicol. 74, 70–81. https://doi.org/10.1016/j.aquatox.2005.04.009 (2005).
Magalhães, C. R. et al. Gilthead seabream liver integrative proteomics and metabolomics analysis reveals regulation by different prosurvival pathways in the metabolic adaptation to stress. Int. J. Mol. Sci. 23 https://doi.org/10.3390/ijms232315395 (2022).
Li, L., Fath, M. A., Scarbrough, P. M., Watson, W. H. & Spitz, D. R. Combined Inhibition of glycolysis, the Pentose cycle, and thioredoxin metabolism selectively increases cytotoxicity and oxidative stress in human breast and prostate cancer. Redox Biol. 4, 127–135. https://doi.org/10.1016/j.redox.2014.12.001 (2015).
Abdel-Tawwab, M., Monier, M. N., Hoseinifar, S. H. & Faggio, C. Fish response to hypoxia stress: growth, physiological, and immunological biomarkers. Fish. Physiol. Biochem. 45, 997–1013. https://doi.org/10.1007/s10695-019-00614-9 (2019).
Van Weerd, J. & Komen, J. The effects of chronic stress on growth in fish: a critical appraisal. Comp. Biochem. Physiol. A: Mol. Integr. Physiol. 120, 107–112. https://doi.org/10.1016/S1095-6433(98)10017-X (1998).
Samaras, A. et al. Cortisol concentration in scales is a valid indicator for the assessment of chronic stress in European sea bass. Dicentrarchus labrax L Aquaculture 545 https://doi.org/10.1016/j.aquaculture.2021.737257 (2021).
Zhang, N. Role of methionine on epigenetic modification of DNA methylation and gene expression in animals. Anim. Nutr. 4, 11–16. https://doi.org/10.1016/j.aninu.2017.08.009 (2018).
Castellano, M. et al. Serum amyloid A is a positive acute phase protein in Russian sturgeon challenged with Aeromonas hydrophila. Sci. Rep. 10, 22162. https://doi.org/10.1038/s41598-020-79065-9 (2020).
Chen, F., Lu, X. J., Nie, L., Ning, Y. J. & Chen, J. Molecular characterization of a CC motif chemokine 19-like gene in Ayu (Plecoglossus altivelis) and its role in leukocyte trafficking. Fish. Shellfish Immunol. 72, 301–308. https://doi.org/10.1016/j.fsi.2017.11.012 (2018).
Romagnani, S. IL4I1: key immunoregulator at a crossroads of divergent T-cell functions. Eur. J. Immunol. 46, 2302–2305. https://doi.org/10.1002/eji.201646617 (2016).
Sadhu, N., Sharma, S., Dube, P., Joseph, S. & Philipose, K. First results of culture of Asian Seabass (Lates calcarifer, Bloch) in open sea floating net cages in India: effect of stocking density on survival and growth. Indian J. Geo-Mar. Sci. 44, 1540–1544 (2015).
Azeredo, R. et al. Dietary Tryptophan plays a role as an Anti-Inflammatory agent in European Seabass (Dicentrarchus labrax) juveniles during chronic inflammation. Biology 13, 309. https://doi.org/10.3390/biology13050309 (2024).
Larkin, P. B. et al. Tryptophan 2,3-dioxygenase and indoleamine 2,3-dioxygenase 1 make separate, tissue-specific contributions to basal and inflammation-induced kynurenine pathway metabolism in mice. Biochim. Biophys. Acta. 1860, 2345–2354. https://doi.org/10.1016/j.bbagen.2016.07.002 (2016).
Mbongue, J. C. et al. The role of indoleamine 2, 3-Dioxygenase in immune suppression and autoimmunity. Vaccines (Basel). 3, 703–729. https://doi.org/10.3390/vaccines3030703 (2015).
Machado, M. et al. Dietary Tryptophan and methionine as modulators of European Seabass (Dicentrarchus labrax) immune status and inflammatory response. Fish Shellfish Immunol. 42, 353–362. https://doi.org/10.1016/j.fsi.2014.11.024 (2015).
Badawy, A. Kynurenine pathway of Tryptophan metabolism: regulatory and functional aspects. Int. J. Tryptophan Res. 10, 1–20. https://doi.org/10.1177/1178646917691938 (2017).
Verburg-van Kemenade, B. M. L., Cohen, N. & Chadzinska, M. Neuroendocrine-immune interaction: evolutionarily conserved mechanisms that maintain allostasis in an ever-changing environment. Dev. Comp. Immunol. 66, 2–23. https://doi.org/10.1016/j.dci.2016.05.015 (2017).
Johnston, W. L., Atkinson, J., Hilton, J. W. & Were, K. E. Effect of dietary Tryptophan on plasma and brain Tryptophan, brain serotonin, and brain 5-hydroxyindoleacetic acid in rainbow trout. J. Nutr. Biochem. 1, 49–54. https://doi.org/10.1016/0955-2863(90)90100-Y (1989).
Winberg, S. & Lepage, O. Elevation of brain 5-HT activity POMC expression and plasma cortisol in socially subordinate. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 274 https://doi.org/10.1152/ajpregu.1998.274.3.R645 (1998).
Alfonso, S. et al. Coping styles in European sea bass: the link between boldness, stress response and neurogenesis. Physiol. Behav. 207, 76–85. https://doi.org/10.1016/j.physbeh.2019.04.020 (2019).
Best, C., Kurrasch, D. M. & Vijayan, M. M. Maternal cortisol stimulates neurogenesis and affects larval behaviour in zebrafish. Sci. Rep. 7, 40905. https://doi.org/10.1038/srep40905 (2017).
Tea, J., Alderman, S. L. & Gilmour, K. M. Social stress increases plasma cortisol and reduces forebrain cell proliferation in subordinate male zebrafish (Danio rerio). J. Exp. Biol. 222 https://doi.org/10.1242/jeb.194894 (2019).
Möbius, W., Patzig, J., Nave, K. A. & Werner, H. B. Phylogeny of proteolipid proteins: divergence, constraints, and the evolution of novel functions in myelination and neuroprotection. Neuron Glia Biol. 4, 111–127. https://doi.org/10.1017/S1740925X0900009X (2009).
Brösamle, C. The Myelin proteolipid dmalpha in fishes. Neuron Glia Biol. 6, 109–112. https://doi.org/10.1017/S1740925X09000131 (2009).
Hernandez-Linares, Y. et al. 3,5-T2 and 3,3’,5-T3 regulate cerebellar thyroid hormone signalling and Myelin molecular dynamics in Tilapia. Sci. Rep. 9, 7359. https://doi.org/10.1038/s41598-019-43701-w (2019).
Halpain, S. & Dehmelt, L. The MAP1 family of microtubule-associated proteins. Genome Biol. 7, 224. https://doi.org/10.1186/gb-2006-7-6-224 (2006).
Mommsen, T., Vijayan, M. M. & Moon, W. Cortisol in teleosts: dynamics, mechanisms of action, and metabolic regulation. Rev. Fish Biol. Fish. 9 https://doi.org/10.1023/A:1008924418720 (1999).
Iwama, G., Afonso, L. & Vijayan, M. in The Fhysiology of Fishes (eds Evans, D. H. & Claiborne, J. B) 319–335 (CRC Press, 2006).
Miller, W. L. Steroid hormone synthesis in mitochondria. Mol. Cell. Endocrinol. 379, 62–73. https://doi.org/10.1016/j.mce.2013.04.014 (2013).
Miller, W. L. & Auchus, R. J. The molecular biology, biochemistry, and physiology of human steroidogenesis and its disorders. Endocr. Rev. 32, 81–151. https://doi.org/10.1210/er.2010-0013 (2011).
Tabas, I. Consequences of cellular cholesterol accumulation: basic concepts and physiological implications. J. Clin. Invest. 110, 905–911. https://doi.org/10.1172/jci0216452 (2002).
Kandutsch, A. A., Chen, W. & heiniger, H. J. Biological activity of some oxygenated sterols. Science 201 https://doi.org/10.1126/science.663671 (1978).
Hui, D. Y. & Howles, P. N. Carboxyl ester lipase: structure-function relationship and physiological role in lipoprotein metabolism and atherosclerosis. J. Lipid Res. 43, 2017–2030. https://doi.org/10.1194/jlr.r200013-jlr200 (2002).
Edwards, M. & Mohiuddin, S. S. Biochemistry Lipolysis (StatPearls Publishing, 2024).
Tokarz, J., Moller, G., Hrabe de Angelis, M. & Adamski, J. Steroids in teleost fishes: A functional point of view. Steroids 103, 123–144. https://doi.org/10.1016/j.steroids.2015.06.011 (2015).
Acknowledgements
The present work was supported by the IMMUNAA (reference PTDC/CVT-CVT/7741/2020) project, financed by national funds through Fundação para a Ciência e a Tecnologia (FCT, Portugal), and within the scope of UIDB/04423/2020 and UIDP/04423/2020. DP, IC, BC and RA were supported by FCT (UI/BD/150900/2021, 2021.04867.BD, 2020.00290.CEECIND and 2022.03248.CEECIND, respectively). DR was supported by BBSRC Institute Strategic Grants to the Roslin Institute (BBS/E/20002172, BBS/E/D/30002275, BBS/E/D/10002070 and BBS/E/RL/230002 A), and by the Oportunius programme of the Axencia Galega the Innovación (GAIN, Xunta de Galicia).
Funding
The present work was supported by the IMMUNAA (reference PTDC/CVT-CVT/7741/2020) project, financed by national funds through Fundação para a Ciência e a Tecnologia (FCT, Portugal), and within the scope of UIDB/04423/2020 and UIDP/04423/2020. DP, IC, BC and RA were supported by FCT (UI/BD/150900/2021, 2021.04867.BD, 2020.00290.CEECIND and 2022.03248.CEECIND, respectively). DR was supported by BBSRC Institute Strategic Grants to the Roslin Institute (BBS/E/20002172, BBS/E/D/30002275, BBS/E/D/10002070 and BBS/E/RL/230002 A), and by the Oportunius programme of the Axencia Galega the Innovación (GAIN, Xunta de Galicia).
Author information
Authors and Affiliations
Contributions
Conceptualization, R.A., M.M. and B.C.; methodology, B.C., D.P., M.M. and R.A.; validation, D.R., B.C. and R.A.; investigation, D.P., I.C., D.R. and R.A.; data curation, D.P., I.C., D.R., P.P., A.F. and R.A.; writing original draft preparation, D.P.; writing review and editing, D.R., R.A. and B.C.; supervision, B.C. and R.A.; funding acquisition, B.C. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The experiments were approved by the Animal Welfare Committee of the Interdisciplinary Centre of Marine and Environmental Research and carried out in a registered installation (N16091.UDER). All experiments and methodologies were performed by trained scientists (following FELASA category C recommendations) in full compliance with national rules, following both the European Directive 2010/63/EU of the European Parliament and the European Union Council on the protection of animals used for scientific purposes, and the relevant ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peixoto, D., Carvalho, I., Robledo, D. et al. Tryptophan-induced transcriptomic changes in the European Seabass are highly dependent on neuroendocrine-immune conditions. Sci Rep 15, 16340 (2025). https://doi.org/10.1038/s41598-025-01079-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-01079-y