Abstract
Osteosarcoma (OS) is a prevalent form of bone cancer among younger people, particularly children and adolescents. Ferroptosis is a non-apoptotic cell death identified by increased levels of iron-dependent lipid peroxidation. This study was designed to develop a prognostic model based on differentially expressed genes (DEGs) associated with ferroptosis and examined the functions of ferroptosis-related genes (FRGs) in OS cells. Gene expression profiles in OS were retrieved from TARGET and GEO databases, while GTEx provided data for healthy tissues. Prognostic genes were identified through bioinformatics analysis and data integration. In vitro experiments, cell cultures, qRT-PCR, immunohistochemistry (IHC), cell transfection, Edu assays, DHE assays, migration, and invasion assays validated the prognostic model and explored the functional role of FRGs in OS cells. Univariate Cox regression analysis demonstrated that 12 DEGs were differentially expressed. Based on four FRGs in OS constructed a risk-scoring model. The high-risk (HR) group showed a considerably lower OS rate than the low-risk (LR) group (p < 0.001 in the TARGET and p < 0.05 in the GSE21257 cohorts). A risk score was validated as an independent predictive factor for OS via multivariate Cox regression. Functional analysis shows that these FRGs affect the occurrence of ferroptosis by influencing the intracellular ROS levels and play a regulatory role in the proliferation, migration, and infiltration of OS cells. The findings suggested that four FRGs demonstrate significant prognostic value in OS, offering potential insights into novel therapeutic targets for OS treatment.
Similar content being viewed by others
Introduction
Osteosarcoma is a prevalent and aggressive form of bone cancer, particularly affecting children and adolescents1,2. The current standard treatment involves a combination of systemic chemotherapy and extensive surgical resection. Recent advancements in limb-salvage surgery and neoadjuvant chemotherapy have substantially improved the 5-year survival rate to approximately 65 to 70% in patients with non-metastatic disease3. However, distant metastasis is also found in around 20% of patients, with lung metastasis accounting for 90% of them4, which dramatically decreases the 5-year survival rate (15 to 30%)5,6,7,8. This necessitates the development of novel prognostic models to enhance the survival rates of patients with OS.
Ferroptosis is an iron-dependent, non-apoptotic cell death and is distinguished by the accumulation of lipid reactive oxygen species (ROS)9. Ferroptosis has recently attracted substantial interest and attention as a possible therapeutic option for cancer treatment. Reported studies show that ferroptosis remarkably impacts the destruction of tumor cells and the prevention of tumor growth9,10. In the OS cell line D-27, Isani et al. initially described the ferroptosis mechanism. They demonstrated that the cells initiate ferroptosis by restricting the signaling pathway that involves glutathione peroxidase 4 (GPX4), signal transducer and activator of transcription 3 (STAT3), and nuclear factor erythroid 2-related factor 2 (Nrf2)11. Meanwhile, ferroptosis is regulated by multiple genes12. Current studies have revealed the significant role of ferroptosis in OS, with particular genes like STAT3, Nrf2, GPX4, FPN, FTH1, DMT1, and IRP2 known for their negative regulation of ferroptosis11,13. Genes such as FTH1 and SLC40A1 have been found to positively regulate ferroptosis14. As ferroptosis continues to be investigated, FRGsg has been identified as novel therapeutic targets with prognostic effects in OS15,16. In previous studies, the etiology, advancement, and prognosis of OS have been substantially correlated with ferroptosis17. Building upon this premise, oncologists have validated the activation of ferroptosis as an innovative and potentially efficacious therapeutic strategy for OS18. Bioinformatics analysis serves as a valuable tool for identifying possible drug targets and diagnostic markers, enabling a more potent therapeutic strategy. However, the prognostic significance of ferroptosis and FRGs in OS has not been fully elucidated. This study developed a novel prognostic model for OS by integrating FRGs (confirmed in an external patient cohort), OS cell lines, and tissue samples. This model promotes evaluating treatment outcomes and prognosis while advancing the understanding of the molecular mechanisms underlying ferroptosis in OS development and progression.
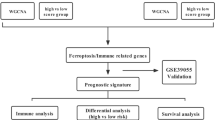
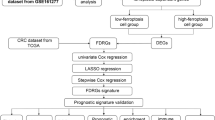
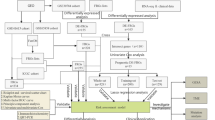
In this study, RNA-seq and clinical information for OS patients were sourced from publicly accessible databases. A prognostic multigene signature was then developed via ferroptosis-associated DEGs identified in the TARGET cohort. These signatures were verified in various cohorts, such as GSE21257, OS cell lines, and OS tissue samples. A nomogram model was developed to further validate its clinical application value.
Materials and methods
Data source and FRGs
The training cohort comprised 84 OS patients, with data retrieved from the TARGET database (https://portal.gdc.cancer.gov/). The validation cohort comprised data from 53 OS samples, which were obtained from the GEO database (http://www.ncbi.nlm.nih.gov/geo/) (accesserum-freeE21257). The clinical characteristics of the patients are summarized in Table 1. Data for RNA expression from healthy tissues, including 396 musculoskeletal samples, were provided from the GTEx database (https://gtexportal.org/). Moreover, a total of 247 FRGs were retrieved from the FerrDb database (http://www.zhounan.org/ferrdb/), a comprehensive resource that contains information on regulators and markers of ferroptosis, as well as the relationship between ferroptosis and multiple diseases. The associated R programming codes used in this manuscript were uploaded in the repository of GitHub (https://github.com/Bob111111-spec/R.git).
Differentially expressed FRGs analysis and functional annotation.
To minimize batch variations between the TARGET and GTEx databases, gene transcript expression data for each sample was logarithmically modified by adding 1 to the FPKM value, followed by a log2 transformation. Differentially expressed FRGs between tumor and healthy tissues were detected via the “limma” R package, with the selection criteria set at FDR < 0.05 and |logFC|≥ 2. Next, a univariate Cox analysis of OS was performed to identify FRGs with prognostic significance. The study also conducted GO and KEGG19,20,21 pathways analyses to examine the biological roles and pathways correlated with the differentially expressed FRGs. To visualize the interaction among these genes, the STRING website (https://cn.string-db.org/) was used to develop an interaction network22.
Constructing and validating the prognostic FRG signature
Univariate Cox regression analysis of 58 differentially expressed FRGs in the TARGET cohort was carried out via the “survival” R package. Prognostic relevance was determined as per the significance threshold of p < 0.05. After that, a prognostic model was developed via multiCox regression analysis via the same “survival” R package23. Based on the gene expression signature, the gene normalized expression, as well as their respective regression coefficients, have been processed to determine the patient’s risk scores by using the given formula:
Here \(Coef_{i}\) is the gene regression coefficient and the gene expression level.
The “stats” R package’s “prcomp” function was used for Principal Component Analysis (PCA), while t-SNE analysis was conducted using the “Rtsne” R package to evaluate group distributions. The “survminer” R package’s “surv cutpoint” function was used to conclude suitable cut-off expression values for gene survival analysis. Moreover, the “survivalROC” R package assessed the predictive potency of the gene signature via time-dependent ROC curve studies.
Nomogram model development and validation
A nomogram model was developed and validated using the “rms” R package. In line with the results of multivariate Cox regression, key parameters, such as patient age, sex, metastatic status, and risk scores, were added to the nomogram to provide a comprehensive prognostic evaluation. The model’s performance was assessed by estimating the C-index, and a calibration curve was plotted to evaluate the concordance between actual and predicted clinical outcomes.
Cell cultures and transfection
MG63, Saos-2, 143B, U2OS, and hFOB 1.19 were obtained from the Shanghai Institute of Cell Biology in China. Cells were cultured in DMEM (ATCC, USA) with 10% FBS with 1% penicillin and streptomycin at 37◦C with 5% CO2 in a humidified atmosphere. To reduce the expression of key genes in SAOS-2 and MG-63 cells, siRNA was transfected into the cells as per the manufacturer’s guidelines. Essential gene-targeting siRNAs and a negative control siRNA (siRNA-NC) were sourced from Guangzhou Ruibo Biotechnology Co., Ltd. Transfection of Si-NC and specific siRNAs was performed using Lipofectamine 3000 (Invitrogen) in 6-well plates. The impact of siRNA knockdown was evaluated 48 h post-transfection via qRT-PCR, in line with the manufacturer’s procedure. All sequences of the siRNAs are listed in Table S1.
Gene expression via qRT-PCR
Total RNA was isolated via TRIzol reagent (Invitrogen), followed by cDNA synthesis via the PrimeScriptTMRT reagent Kit (Takara). SYBR Green qPCR Master Mix was used for qRT-PCR analysis (Thermo Fisher Scientific). The expression of mRNA was normalized with GAPDH. Table 2 displays the details of the primer.
Immunohistochemistry (IHC) assay
From December 2019 to January 2021, 5 patients with OS underwent resection at Nanchang University’s Second Affiliated Hospital.The clinical characteristics of these individuals was summarized by Table S2. The approval for the existing research was provided by the Second Affiliated Hospital of Nanchang University of Ethics Committee [2019] No. (088). Furthermore, each patient signed informed consent in written form.
The fixing of OS and surrounding tissues was carried out, followed by embedding in paraffin. Next, these tissues were cut into sections, and deparaffinized. Serum-free protein block buffer (Dako, Glostrup, Denmark) was used to block these sections for 1.5 h, followed by incubating them with BNIP3, G6PD, PGD (1:200, Abcam, Cambridge), and TGFBR1 (1:100, Boster Biological Technology co.ltd, China) antibodies. After taking pictures under the microscope, H-score was used for analysis.
Cell migration assay
Cell migration potential was observed via wound healing assay. Initially, 5 × 105 cells in logarithmic growth were seeded in 3.5 cm culture plates and grown at 37 °C for 24 h with 5% CO₂. Once cells reached 90% confluence and covered dish bottoms, a single scratch was induced via a 200 μL pipette tip. After PBS washing, cells were transfected and cultured at 37 °C with 5% CO₂ for 24 and 48 h. Cell migration was observed under an Olympus microscope, and images were acquired.
Transwell assay
This assay was used to evaluate cell invasion. Approximately 5 × 104 cells were allowed to grow with serum-free medium (8 μm pore size, Corning, New York, USA) for migration assays using transwell chambers. An enriched medium (10% FBS) was added to the lower chamber. The procedure for invasion experiments with transwell chambers was similar to migration assay, except for pre-coating the upper chamber with 10% Matrigel (Corning). After 36 h incubation, the cells that remained in the upper chamber were removed using swabs. However, cells that crossed the membrane were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet.
5-ethynyl-2’-deoxyuridine (Edu) assay
For this assay, cells were incubated with 50 µM Edu at 37 °C for 4 h. Next, cells were fixed in 4% formaldehyde at 25 °C for 15 min, then permeabilized with 5% Triton X-100. They were treated with 100 µL of 1X Apollo®567 reaction cocktail for 30 min. Afterward, they were stained with 100 µL of Hoechst 33,342 for 30 min in the dark and examined under a light microscope (Olympus, Tokyo, Japan).
Assessment of oxidative stress
In order to assess the levels of ROS in tissues, we carried out a staining procedure on kidney slices using dihydroethidium (DHE). The DHE powder was first dissolved in dimethyl sulfoxide. Subsequently, the cryosections were incubated with the prepared DHE solution at room temperature for a duration of 30 min. After that, the samples were observed with the aid of confocal microscopy. To measure the levels of mitochondrial ROS, MitoSOX Red was employed. The cells were incubated with the MitoSOX Red solution at 37 ℃ for 30 min. Following the incubation, images were captured under a microscope. Finally, the integrated optical density of the ratios DHE/DAPI and MitoSOX/DAPI was calculated using the Image-Pro Plus (IPP) software.
Statistical analysis
Data was statistically analyzed via R (v3.5.3) or SPSS (v25.0) software. The sample size was determined through power analysis, taking into account the effect size, desired power (typically 80%), and significance level, and different formulas were used depending on the type of statistical test. The normality of the data was evaluated using the Shapiro–Wilk test in R or the Explore function in SPSS. Quantitative data were presented as SD ± mean. For all quantitative analyses, each analysis condition was performed in triplicate. Student’s t-test was used to compare differences between two groups. One-way ANOVA was used for comparisons among multiple groups, and statistical differences were analyzed using the Bonferroni post hoc test in GraphPad Prism 5.0. Proportion changes were analyzed via Chi-squared tests. Overall survival was evaluated with log-rank and Kaplan–Meier tests. Uni- and multivariate Cox regression analyses were carried out to predict OS independently. p-values < 0.05 denoted significance threshold, and “ns” indicated no significant difference.
Results
Evaluation of differentially expressed FRGs
A total of differentially expressed 58 FRGs were detected (logFC > 1, FDR < 0.05). The expression patterns of these 58 RGs were illustrated in Fig.1 A, B, revealing that 37 were decreased and 21 were increased. Univariate Cox regression analysis was conducted on the 58 differentially expressed FRGs, which resulted in the identification of 12 FRGs that were related to the prognosis of OS patients, as shown in Fig. 1C. The network of protein–protein interaction (PPI) revealed how the underlined genes interacted with each other (Fig. 1D). The relationship between these genes is displayed in Fig. 1E, where the blue line indicates a negative association and the red line represents a positive association. Furthermore, the GO and KEGG enrichment pathway analyses showed significant enrichment of iron ion binding and ferroptosis pathways in DEGs.
Identification of the key FRGs: The 58 differentially-expressed FRGs (37 downregulated and 21 upregulated) were identified (A, B); Forest plots depicting the outcomes of univariate Cox regression analysis regarding gene expression and OS were presented (C); The PPI network, sourced from the STRING database, illustrated the interactions among the candidate genes (D); The correlation network of the candidate genes was analyzed and the results were presented (E).
Further, they were related to pathways involving ROS and hypoxia. These correlations were statistically significant with a p-value adjustment of < 0.05, as shown in Fig. 2A, B. It is widely observed that ferroptosis is characterized by lipid ROS accumulation. The relationship between hypoxia and ferroptosis has also been confirmed in the literature. Studies have shown that hypoxia-inducible factors.
GO enrichment and KEGG pathway analyses: For the differentially expressed FRGs, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses were conducted (A, B).
(HIF) can adjust iron regulatory proteins in tumors or during inflammation. They also observed that HIF regulates the transferrin receptor (TfR) and divalent metal transporter 1 (DMT1) to control iron import, and it also modulates ferroportin to control iron export24,25,26,27,28.
Prognostic model development
A prognostic model was developed via a multivariate Cox regression test. The optimal value of λ was used to identify a four-gene signature (Table S3). Patient groups were distributed into HR and LR groups as per the median cutoff values. The survival curves indicated that the HR group had a considerably lower OS than the LR group (Fig. 3A). The ROC curves revealed that with time-dependent AUCs, the inferential results of the risk score for OS were 0.857 for 1 year, 0.840 for 3 years, and 0.818 for five years (Fig. 3B). Both HR and LR groups included 42 patients, resulting in a total of 84 patients (Fig. 3C). As shown in Fig. 3D, most LR patients survived longer than HR patients. It was observed from PCA and t-SNE that patients could be conveniently separated into various risk groups (Fig. 3E,F).
Prognostic analysis of the 4 FRGs signature model in the TARGET cohort: Kaplan–Meier curves depicting the OS of patients were generated for both the HR and LR groups (A); Time-dependent ROC curves were used to calculate the AUC in the TARGET cohort (B); The distribution and median value of the risk scores within the TARGET cohort were also analyzed (C); The distributions of OS, OS, and risk score (D); Principal component analysis (PCA) was utilized to plot the TARGET cohort (E); T-distributed stochastic neighbor embedding (T-SNE) analysis was performed for further data exploration (F).
Validation of the four FRG signatures in the GEO cohort
The accuracy of the developed model was measured via both the TARGET and GSE21257 cohorts. A comparison between both groups revealed that the LR group showed substantially enhanced OS, aligning with the findings from the TARGET cohort (Fig. 4A) (p < 0.05). The ROC curve analysis demonstrated that the AUC values for the OS risk score at one, three, and five years were 0.745, 0.617, and 0.632, respectively (Fig. 4B). The HR group comprised 29 patients, while the LR group consisted of 24 patients (Fig. 4C). Patients in the HR group tended to experience earlier mortality (Fig. 4D). The patient classification into two subgroups was further confirmed via t-SNE and PCA analyses (Fig. 4E. F).
Validation of the four FRGs signatures in the GSE21257 cohort: Kaplan–Meier curves were utilized to compare the OS rates between patients classified into the HR and LR groups (A); The AUC values from time-dependent ROC curves were calculated for the GSE21257 cohort (B); The risk scores’ distribution and median value were analyzed within the GSE21257 cohort(C); The distributions of OS, OS and risk score (D); Principal component analysis (PCA) was conducted to visualize the data distribution within the GSE21257 cohort (E); T-distributed stochastic neighbor Embedding (T-SNE) analysis was carried out for further data exploration (F).
Independent predictive value of the FRG signature
Univariate and multivariate Cox regression analyses evaluated the risk score’s potential to predict OS independently in the provided variables. A substantial association between the risk score and OS was observed in the TARGET cohort via a univariate Cox regression (HR = 1.302; 95% CI = 1.193–1.421) with p < 0.001 (Fig. 5A). Further, the risk score was detected as a distinct predictor of OS (p < 0.001) in the multivariate Cox regression analysis (HR = 1.324, 95% CI = 1.203–1.458), as mentioned in Fig. 5B.
Cox regression analyses (uni- and multivariate) and the interrelation of risk scores and clinic-pathological features: These analyses assessed the impact of OS in the TARGET cohort (A, B); The association between the expression profiles of four prognostic FRGs, stratified by risk score, and the clinicopathological features of OS were investigated in both the TARGET and GSE21257 cohorts (C, D).
The function of the underlined 4-FRGs in OS was determined by the correlation between the expression patterns of the four genes and comparable biological and pathological features of OS, such as age, sex metastasis, risk score, and longevity status, as displayed in Fig. 5C and Fig. 6D. The results revealed that BNIP3 mRNA expression was reduced, and TGFBR1, G6PD, and PGD mRNA increased in the LR group.
Nomogram for the prediction of the survival rate of patients with OS: Nomogram was developed using patient age, sex, metastasis status, and risk score in the TARGET cohort (A); Calibration plots were then used to demonstrate the effectiveness of the nomogram in predicting the 3 and 5-years OS rates of glioma patients in respective cohorts (B-E).
Development and confirmation of a nomogram model
The prognostic mode’s potential clinical applications were evaluated by constructing the nomogram model in both the TARGET and GSE21257 cohorts using multivariate Cox regression analysis. The model included sex, age, metastasis, and the risk score to evaluate its predictive ability (Fig. 6A). Based on the calibration curves, the nomogram had substantially high predictive accuracy in identifying the 3- and 5-year OS of patients in the TARGET and GSE21257 cohort (Fig. 6B-E). Nomogram model C index was 0.812 in the TARGET cohorts and 0.741 in the GSE21257 cohorts. The underlined results suggested that the nomogram model derived from a prognostic model could be a possible prognostic model.
In vitro experimental validation of the four FRGs signature
To assess the crucial genes for developing the risk score signature, qRT-PCR was conducted on OS and normal cells. Further, immunohistochemistry (IHC) was performed to compare protein levels between adjacent normal and OS tissues. As illustrated in Fig. 7A, TGFBR1, G6PD, and PGD levels were substantially upregulated in MG-63 and SAOS-2 cells relative to normal cells, whereas BNIP3 expression was markedly downregulated in SAOS-2 and MG-63 cells. In addition, the four FRGs were also validated in 143B and U2OS cells (Fig. S1). IHC revealed that in OS tissues, compared to adjacent normal tissues, the protein staining for BNIP3 was dramatically reduced, whereas the staining for TGFBR1, G6PD, and PGD was significantly enhanced (Fig. 7B). Fig S2 shows the quantitative analysis of the protein expression levels of these four prognostic FRGs in OS tissues and adjacent normal tissues in IHC. Importantly, the expression trends of G6PD29, PGD30, and BNIP331 in OS were consistent with findings from previous studies.
Both mRNA and protein levels of four prognostic FRGs signature: The mRNA expression profiles of four FRGs with prognostic value were assessed in both OS cells and normal cells (n = 3) (A); Immunohistochemical analysis compared the protein expression levels of these four prognostic FRGs in OS specimens (n = 3) (B). Significance threshold was denoted as *p < 0.05, ** p < 0.01, and ***p < 0.001.
The expression pattern of the target genes was inhibited via siRNA to conduct comprehensive research into the function of four FRGs in OS cells. In order to ensure the specificity of the results, we have already added the si-NC group to the gene validation process. The qRT-PCR results confirmed a substantial reduction in BNIP3, G6PD, PGD, and TGFBR1 in MG-63 and SAOS-2 cells (Fig. 8A, B). Wound healing (Fig. 8C, D,) and Transwell (Fig. 8E, F) assays demonstrated that the knockdown of TGFBR1 and G6PD genes considerably reduced the migration and invasion potential of MG-63 and SAOS-2 cells. Interestingly, the knockdown of BNIP3 resulted in enhanced migration and invasion of cancer cells, which aligns with previous research indicating a negative correlation of BNIP3 with survival risk and risk scores31. The current experiments also demonstrated that the suppression of PGD in OS cells yielded similar results to those observed in normal cells without statistically significant variances. Fig S3A shows the quantitative analysis of the wound healing assay results. Next, an Edu incorporation assay was carried out to investigate the impact of silencing these four genes on OS cell proliferation. The results indicated that cells transfected with si-G6PD and si-TGFBR1 revealed a significant decrease in Edu-positive cells, suggesting that the knockdown of G6PD and PGD effectively inhibited cell growth (Fig. 8G, Fig. S3B). Conversely, cells transfected with si-BNIP3 showed a significant increase in Edu-positive cells, indicating that knockdown of BNIP3 promoted cell growth. In si-PGD-transfected cells, no substantial change was observed in Edu-positive cells.
Impact of the knockdown of four prognostic FRGs on the migration and invasion of OS cells was investigated: The mRNA expression levels of BNIP3, G6PD, PGD, and TGFBR1 in MG-63 and SAOS-2 cells were quantified using qRT-PCR following siRNA-mediated knockdown of the four prognostic FRGs(n = 3) (A, B). Healing assays in MG-63 and SAOS-2 cells with siRNA knockdown of the four prognostic FRGs (100 μm) (n = 3) (C, D). Evaluation of invasion capabilities of MG-63 and SAOS-2 cells with siRNA knockdown of the four prognostic FRGs using Transwell assays (100 μm) (n = 3) (E, F). Comparison of the proportion of Edu-positive cells in MG-63 and SAOS-2 cells with siRNA knockdown of the four prognostic FRGs via Edu staining (100 μm) (n = 3) (G). Significance threshold was denoted as *p < 0.05, ** p < 0.01, *** p < 0.001; ns, not significant.
To measure the ROS levels in OS cells after silencing these four genes, we conducted a DHE assay (4). The results indicated that G6PD and TGFBR1 could reduce the ROS levels in OS cells. Their downregulation increased intracellular ROS levels, facilitating ferroptosis in these cells. Conversely, knocking down BNIP3 decreased the intracellular ROS levels and inhibited iron-induced cell death in OS cells. The downregulation of PGD had a minor effect on increasing the intracellular ROS levels of OS cells.
Discussion
Osteosarcoma (OS), a malignant bone tumor predominantly affecting adolescents, is marked by an unclear etiology and generally poor prognosis32. Despite advancements in management strategies, the survival rate of OS patients has remained largely unchanged over the past 30 years33. Thus, it is crucial to recognize and evaluate key molecular markers that determine the prognosis rate to improve the clinical efficacy of therapeutic interventions and optimize diagnosis and treatment.
Recently, it has been revealed that ferroptosis considerably contributes to destroying tumor cells and the prognosis of tumor patients. Several research has confirmed that 14 FRGs are related to a poor prognosis of hepatocellular carcinoma34,35. In another study, the gene expression analysis predicted the survival rate of glioma patients. The results of this study indicated that 45 FRGs were substantially correlated with the patient survival rate36,37. Recently, the role of ferroptosis in the formation and treatment of OS has attracted substantial scientific interest38. Multiple studies have indicated significant differential expression of various FRGs in OS patients, which is closely associated with adverse prognoses39,40,41,42.
The model proposed in this study includes four FRGs: BNIP3, G6PD, PGD, and TGFBR1. The crucial role of these four FRGs in determining OS prognosis was demonstrated by the considerable increase in both risk scores and mortality rates over time in the time-dependent model.
BNIP3 belongs to the B-cell lymphoma family and can induce cell death and autophagy. BNIP3 silencing has been shown to increase intracellular iron levels, thereby promoting Ferroptosis43. Early research has identified a strong association between BNIP3 expression and poor prognosis in different cancers, such as breast, bladder, and renal cell carcinoma44,45,46. Another study by Ye et al. showed evidence of BNIP3-mediated cell death through ROS in OS cells, highlighting BNIP3’s potential role in OS prognosis47.
In cancer cells, glucose-6-phosphate dehydrogenase (G6PD) is crucial for producing NADPH and glycolysis48. NADPH is a crucial antioxidant molecule within cells and serves the function of reducing oxidized glutathione (GSH), thereby maintaining the redox balance within cells49. In cases where the activity of G6PD is diminished, the production of NADPH is reduced, which in turn leads to insufficient synthesis of GSH. As a result, the antioxidant capacity of the cell decreases, causing the accumulation of ROS. This accumulation subsequently triggers lipid peroxidation and facilitates the occurrence of ferroptosis50. Current studies have demonstrated that the level of G6PD is enhanced in different cancer types, and this expression level is related to a poor prognosis for cancer patients48,51. Cao et al. observed that iron-induced cell death in HCC cells is inhibited by the upregulation of G6PD52.
Phosphogluconate dehydrogenase (PGD) is the third enzyme in the pentose phosphate pathway. It functions similarly to G6PD in generating NADPH and is involved in the regulation of energy metabolism and iron-induced cell death53. Previous investigations have demonstrated that PGD exhibits an upregulated expression pattern in various types of cancers54, examples of such cancers include gastrointestinal stromal tumors55, liver cancer56, and pancreatic cancer57,58, and this upregulation of PGD contributes to the proliferation of cancer cells. Additionally, PGD is associated with multiple tumor signaling pathways, including the epidermal growth factor signaling pathway and the nuclear factor E2-related factor signaling pathway, etc.59.Further, a study found that the propagation of OS cells is inhibited by the downregulation of G6PD 60, PGD also regulates energy metabolism and iron-induced cell death. Erastin-induced ferroptosis can be blocked by silencing G6PD and PGD in NSCLC cells9,61,62.
Transforming Growth Factor β Receptor 1 (TGFBR1) is a crucial mediator of the TGF-β signaling pathway. It functions primarily by binding to TGF-β ligands, which in turn activates downstream Smad proteins, thereby regulating multiple biological processes including cell proliferation, differentiation, and apoptosis63,64. TGFBR1 enhances the expression of glutathione peroxidase 4 (GPX4) through interaction with the NRF2. GPX4 clears ROS by converting lipid hydroperoxides into non-toxic lipid alcohols, maintaining intracellular redox balance and alleviating etanercept-induced ferroptosis65. Growing evidence indicates that TGFBR1 is related to survival rates in different cancer types, such as gastric66, colorectal67, and head and neck47 cancers.
Overall, the evidence indicates that the investigated FRGs may play a crucial role in cancer, necessitating further research to explore their roles in OS cells specifically. Moreover, qRT-PCR determined the characteristics of the four FRGs by detecting the relative expression of BNIP3, G6PD, PGD, and TGFBR1 mRNA in normal osteoblasts and two human OS cell lines, MG-63, and SAOS-2 cells. The results showed significantly increased G6PD, PGD, and TGFBR1 expression in OS cells, while BNIP3 expression was downregulated. Further, the study evaluated the protein expression of these genes in OS and adjacent tissues using IHC, resulting in consistent findings. Interestingly, consistent with the current findings, previous research has also demonstrated an inverse relationship between BNIP3 expression and survival risk and risk scores31. Lei et al.'s research showed an upward trend in G6PD and PGD expression in OS15. G6PD, PGD, and TGFBR1 upregulation has also been confirmed in other cancers51,52,56,60,68. This study also designed a siRNA knockdown experiment to inhibit the expression of these four essential genes and confirm their impact on malignant biological behaviors, i.e., cell propagation and migration in OS cells. The propagation and migration potentials of tumor cells were inhibited by the knockdown of the G6PD and TGFBR1 genes, as demonstrated by wound healing, Transwell, and Edu incorporation assays. Both G6PD and TGFBR1 may function as positive risk factors related to a poor prognosis in OS. The expression of BNIP 3 is also consistent with its function as a negative risk factor. Furthermore, the DHE assay shows that the downregulation of G6PD and TGFBR1 increases the intracellular ROS levels and promotes ferroptosis in OS cells. Conversely, the knockout of BNIP3 can reduce the intracellular ROS levels in OS cells and inhibit iron-induced cell death. This experiment demonstrates that these four FRGs influence the occurrence of ferroptosis by affecting the intracellular ROS levels and play a regulatory role in the proliferation, invasion, and migration of OS cells.
The current research clearly reveals the crucial role of FRGs in the proliferation and migration processes of OS cells, bringing new ideas and potential intervention directions for the clinical treatment of OS. Among them, the design of small interfering RNA (siRNA) drugs based on RNA interference (RNAi) technology has brought new hope for osteosarcoma treatment69. Research shows that G6PD and TGFBR1 are highly expressed in OS cells, and their expression levels are closely related to the poor prognosis of OS. By designing specific siRNA drugs targeting G6PD and TGFBR1, the expression of these two genes can be effectively silenced at the mRNA level, thereby increasing the accumulation of ROS, promoting ferroptosis, and ultimately inhibiting the proliferation of OS cells. On the other hand, BNIP3 is downregulated in OS cells and is a negative risk factor associated with a better prognosis. In this regard, siRNA drugs can play a role in an indirect way, that is, by designing siRNA drugs targeting upstream regulatory factors that inhibit the expression of BNIP3, the inhibition of BNIP3 can be relieved, thereby increasing its expression level in cells, inducing cell death and autophagy, and simultaneously promoting ferroptosis, effectively inhibiting tumor growth. Although siRNA drugs have great potential in the OS treatment strategy based on FRGs, there are still many technical challenges to achieve clinical application, requiring continuous optimization of drug design, improvement of delivery systems, and verification of their safety and effectiveness through large-scale clinical trials. In addition, in clinical practice, detecting the expression levels of BNIP3, G6PD, PGD, and TGFBR1 in patients’ tumor tissues can serve as an effective prognostic evaluation tool, helping doctors to more accurately assess patients’ prognosis and providing an important reference for formulating personalized treatment plans.
This study has identified four FRGs, which may play important roles in the development of ferroptosis in OS, and was the first to integrate clinical samples into validating FRG signaling mechanisms in OS cells. However, it is necessary to address the limitations of the current study. Firstly, there are currently few public gene expression databases containing the prognostic information of OS patients, resulting in a small number of tumor samples in our study. In the future, a larger sample size should be used to establish a more accurate prognostic model. Secondly, the FRGs are sourced from the continuously updated website FerrDb, and more relevant genes remain to be discovered. Finally, further validation of the prognostic significance of these four ferroptosis-related signatures in OS through large-scale clinical research is required. Additionally, in future research, it is necessary to conduct further functional experimental studies to explore the specific mechanism of action of these four FRGs in OS cells.
Conclusion
Four FRG signatures were also observed in OS cells and tissues. In vitro experiments demonstrated their functional significance in OS cell propagation and migration. Moreover, additional studies are needed to verify the current findings and obtain a more comprehensive understanding of the fundamental processes of ferroptosis in OS.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
Abbreviations
- DEGs:
-
Differentially expressed genes
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- IHC:
-
Immunohistochemistry
- HR:
-
High-risk
- LR:
-
Low-risk
- FRGs:
-
Ferroptosis-related genes
- DHE:
-
Dihydroethidium
- OS:
-
Osteosarcoma
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GO:
-
Gene Ontology; PCA: principal component analysis
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- BNIP3:
-
BCL2 interacting protein 3
- G6PD:
-
Glucose-6-phosphate dehydrogenase
- TGFBR1:
-
Transforming growth factor beta receptor 1
- PGD:
-
Phosphogluconate dehydrogenase
- STAT3:
-
Transcription 3
- GPX4:
-
Glutathione peroxidase 4
- GSH:
-
Glutathione
- Nrf2:
-
NF-E2-related factor
- AUC:
-
Area under the curve
- ROS:
-
Reactive oxygen species
- HIF:
-
Hypoxia-inducible factors
- TFR:
-
The transferrin receptor
- DMT1:
-
Divalent metal transporter 1
References
Hameed, M. & Mandelker, D. Tumor Syndromes Predisposing to Osteosarcoma. Adv. Anat. Pathol. 25, 217–222. https://doi.org/10.1097/pap.0000000000000190 (2018).
Ying, H., Li, Z. Q., Li, M. P. & Liu, W. C. Metabolism and senescence in the immune microenvironment of osteosarcoma: focus on new therapeutic strategies. Front Endocrinol (Lausanne) 14, 1217669, https://doi.org/10.3389/fendo.2023.1217669 (2023).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 68, 7–30. https://doi.org/10.3322/caac.21442 (2018).
Anninga, J. K. et al. Chemotherapeutic adjuvant treatment for osteosarcoma: where do we stand?. Eur. J. Cancer 47, 2431–2445. https://doi.org/10.1016/j.ejca.2011.05.030 (2011).
Luetke, A., Meyers, P. A., Lewis, I. & Juergens, H. Osteosarcoma treatment - where do we stand? A state of the art review. Cancer Treat Rev 40, 523–532. https://doi.org/10.1016/j.ctrv.2013.11.006 (2014).
Gianferante, D. M., Mirabello, L. & Savage, S. A. Germline and somatic genetics of osteosarcoma - connecting aetiology, biology and therapy. Nat. Rev. Endocrinol. 13, 480–491. https://doi.org/10.1038/nrendo.2017.16 (2017).
Lin, Y. H. et al. Osteosarcoma: Molecular Pathogenesis and iPSC Modeling. Trends Mol. Med. 23, 737–755. https://doi.org/10.1016/j.molmed.2017.06.004 (2017).
Whelan, J. S. & Davis, L. E. Osteosarcoma, Chondrosarcoma, and Chordoma. J. Clin. Oncol. 36, 188–193. https://doi.org/10.1200/jco.2017.75.1743 (2018).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072. https://doi.org/10.1016/j.cell.2012.03.042 (2012).
Ooko, E. et al. Artemisinin derivatives induce iron-dependent cell death (ferroptosis) in tumor cells. Phytomedicine 22, 1045–1054. https://doi.org/10.1016/j.phymed.2015.08.002 (2015).
Liu, Q. & Wang, K. The induction of ferroptosis by impairing STAT3/Nrf2/GPx4 signaling enhances the sensitivity of osteosarcoma cells to cisplatin. Cell Biol. Int. 43, 1245–1256. https://doi.org/10.1002/cbin.11121 (2019).
Yamaguchi, H. et al. Caspase-independent cell death is involved in the negative effect of EGF receptor inhibitors on cisplatin in non-small cell lung cancer cells. Clin. Cancer Res. 19, 845–854. https://doi.org/10.1158/1078-0432.Ccr-12-2621 (2013).
Li, J. et al. Ferroptosis: past, present and future. Cell Death Dis. 11, 88. https://doi.org/10.1038/s41419-020-2298-2 (2020).
Lv, H., Zhen, C., Liu, J. & Shang, P. β-Phenethyl Isothiocyanate Induces Cell Death in Human Osteosarcoma through Altering Iron Metabolism, Disturbing the Redox Balance, and Activating the MAPK Signaling Pathway. Oxid. Med. Cell Longev. 2020, 5021983. https://doi.org/10.1155/2020/5021983 (2020).
Lei, T., Qian, H., Lei, P. & Hu, Y. Ferroptosis-related gene signature associates with immunity and predicts prognosis accurately in patients with osteosarcoma. Cancer Sci. 112, 4785–4798. https://doi.org/10.1111/cas.15131 (2021).
Zhao, J., Zhao, Y., Ma, X., Feng, H. & Jia, L. Outstanding prognostic value of novel ferroptosis-related genes in chemoresistance osteosarcoma patients. Sci. Rep. 12, 5029. https://doi.org/10.1038/s41598-022-09080-5 (2022).
Zhang, F. et al. Oridonin-induced ferroptosis and apoptosis: a dual approach to suppress the growth of osteosarcoma cells. BMC Cancer 24, 198. https://doi.org/10.1186/s12885-024-11951-1 (2024).
Fan, L., Ru, J., Liu, T. & Ma, C. Identification of a Novel Prognostic Gene Signature From the Immune Cell Infiltration Landscape of Osteosarcoma. Front. Cell Dev. Biol. 9, 718624. https://doi.org/10.3389/fcell.2021.718624 (2021).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672-d677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 39, D561-568. https://doi.org/10.1093/nar/gkq973 (2011).
Simon, N., Friedman, J., Hastie, T. & Tibshirani, R. Regularization Paths for Cox’s Proportional Hazards Model via Coordinate Descent. J Stat. Softw. 39, 1–13. https://doi.org/10.18637/jss.v039.i05 (2011).
Fuhrmann, D. C., Mondorf, A., Beifuß, J., Jung, M. & Brüne, B. Hypoxia inhibits ferritinophagy, increases mitochondrial ferritin, and protects from ferroptosis. Redox Biol. 36, 101670. https://doi.org/10.1016/j.redox.2020.101670 (2020).
Taylor, M. et al. Hypoxia-inducible factor-2α mediates the adaptive increase of intestinal ferroportin during iron deficiency in mice. Gastroenterology 140, 2044–2055. https://doi.org/10.1053/j.gastro.2011.03.007 (2011).
Tacchini, L., Gammella, E., De Ponti, C., Recalcati, S. & Cairo, G. Role of HIF-1 and NF-kappaB transcription factors in the modulation of transferrin receptor by inflammatory and anti-inflammatory signals. J. Biol. Chem. 283, 20674–20686. https://doi.org/10.1074/jbc.M800365200 (2008).
Matak, P. et al. Copper deficiency leads to anemia, duodenal hypoxia, upregulation of HIF-2α and altered expression of iron absorption genes in mice. PLoS ONE https://doi.org/10.1371/journal.pone.0059538 (2013).
Ma, X. et al. SMAD family member 3 (SMAD3) and SMAD4 repress HIF2α-dependent iron-regulatory genes. J. Biol. Chem. 294, 3974–3986. https://doi.org/10.1074/jbc.RA118.005549 (2019).
Li, J. et al. A novel ferroptosis-related gene signature to predict overall survival in patients with osteosarcoma. Am. J. Transl. Res. 14, 6082–6094 (2022).
Jiang, M. et al. A Risk-Scoring Model Based on Evaluation of Ferroptosis-Related Genes in Osteosarcoma. J. Oncol. 2022, 4221756. https://doi.org/10.1155/2022/4221756 (2022).
Yang, L., Liu, J. & Liu, S. Clinical significance and immune landscape of a novel ferroptosis-related prognosis signature in osteosarcoma. BMC Cancer 23, 229. https://doi.org/10.1186/s12885-023-10688-7 (2023).
Bone sarcomas: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 23 Suppl 7, vii100–109, https://doi.org/10.1093/annonc/mds254 (2012).
Fathizadeh, H., Mirzaei, H. & Asemi, Z. Melatonin: an anti-tumor agent for osteosarcoma. Cancer Cell Int. 19, 319. https://doi.org/10.1186/s12935-019-1044-2 (2019).
Liang, J. Y. et al. A Novel Ferroptosis-related Gene Signature for Overall Survival Prediction in Patients with Hepatocellular Carcinoma. Int. J. Biol. Sci. 16, 2430–2441. https://doi.org/10.7150/ijbs.45050 (2020).
Tang, B. et al. The ferroptosis and iron-metabolism signature robustly predicts clinical diagnosis, prognosis and immune microenvironment for hepatocellular carcinoma. Cell Commun. Signal 18, 174. https://doi.org/10.1186/s12964-020-00663-1 (2020).
Zhuo, S. et al. Clinical and Biological Significances of a Ferroptosis-Related Gene Signature in Glioma. Front Oncol. https://doi.org/10.3389/fonc.2020.590861 (2020).
Liu, H. J. et al. Ferroptosis-Related Gene Signature Predicts Glioma Cell Death and Glioma Patient Progression. Front Cell Dev. Biol. 8, 538. https://doi.org/10.3389/fcell.2020.00538 (2020).
Li, G. et al. Integrative analyses of ferroptosis and immune related biomarkers and the osteosarcoma associated mechanisms. Sci. Rep. 13, 5770. https://doi.org/10.1038/s41598-023-33009-1 (2023).
Zhao, J., Zhao, Y., Ma, X., Zhang, B. & Feng, H. Targeting ferroptosis in osteosarcoma. J. Bone Oncol. 30, 100380. https://doi.org/10.1016/j.jbo.2021.100380 (2021).
Xu, Z., Chen, L., Wang, C., Zhang, L. & Xu, W. MicroRNA-1287-5p promotes ferroptosis of osteosarcoma cells through inhibiting GPX4. Free Radic. Res. 55, 1119–1129. https://doi.org/10.1080/10715762.2021.2024816 (2021).
Chen, M., Jiang, Y. & Sun, Y. KDM4A-mediated histone demethylation of SLC7A11 inhibits cell ferroptosis in osteosarcoma. Biochem. Biophys. Res. Commun. 550, 77–83. https://doi.org/10.1016/j.bbrc.2021.02.137 (2021).
Lin, H. et al. EF24 induces ferroptosis in osteosarcoma cells through HMOX1. Biomed. Pharmacother. 136, 111202. https://doi.org/10.1016/j.biopha.2020.111202 (2021).
Vara-Pérez, M. et al. BNIP3 promotes HIF-1α-driven melanoma growth by curbing intracellular iron homeostasis. Embo J. 40, e106214. https://doi.org/10.15252/embj.2020106214 (2021).
Macher-Goeppinger, S. et al. Expression and Functional Characterization of the BNIP3 Protein in Renal Cell Carcinomas. Transl. Oncol. 10, 869–875. https://doi.org/10.1016/j.tranon.2017.08.008 (2017).
Chourasia, A. H. et al. Mitophagy defects arising from BNip3 loss promote mammary tumor progression to metastasis. EMBO Rep. 16, 1145–1163. https://doi.org/10.15252/embr.201540759 (2015).
García-Baquero, R. et al. Methylation of a novel panel of tumor suppressor genes in urine moves forward noninvasive diagnosis and prognosis of bladder cancer: a 2-center prospective study. J. Urol. 190, 723–730. https://doi.org/10.1016/j.juro.2013.01.105 (2013).
Ye, F. et al. Baicalein induces human osteosarcoma cell line MG-63 apoptosis via ROS-induced BNIP3 expression. Tumour Biol. 36, 4731–4740. https://doi.org/10.1007/s13277-015-3122-y (2015).
Wang, X., Chen, K. & Zhao, Z. LncRNA OR3A4 Regulated the Growth of Osteosarcoma Cells by Modulating the miR-1207-5p/G6PD Signaling. Onco. Targets Ther. 13, 3117–3128. https://doi.org/10.2147/ott.S234514 (2020).
Bjørklund, G. et al. The impact of glutathione metabolism in autism spectrum disorder. Pharmacol. Res. 166, 105437. https://doi.org/10.1016/j.phrs.2021.105437 (2021).
Lei, G., Mao, C., Yan, Y., Zhuang, L. & Gan, B. Ferroptosis, radiotherapy, and combination therapeutic strategies. Protein Cell. 12, 836–857. https://doi.org/10.1007/s13238-021-00841-y (2021).
Wang, J. et al. Overexpression of G6PD is associated with poor clinical outcome in gastric cancer. Tumour. Biol. 33, 95–101. https://doi.org/10.1007/s13277-011-0251-9 (2012).
Cao, F., Luo, A. & Yang, C. G6PD inhibits ferroptosis in hepatocellular carcinoma by targeting cytochrome P450 oxidoreductase. Cell Signal 87, 110098. https://doi.org/10.1016/j.cellsig.2021.110098 (2021).
Aydemir, D., Hashemkhani, M., Acar, H. Y. & Ulusu, N. N. In vitro interaction of glutathione S-transferase-pi enzyme with glutathione-coated silver sulfide quantum dots: A novel method for biodetection of glutathione S-transferase enzyme. Chem. Biol. Drug. Des. 94, 2094–2102. https://doi.org/10.1111/cbdd.13614 (2019).
Liu, R. et al. Tyrosine phosphorylation activates 6-phosphogluconate dehydrogenase and promotes tumor growth and radiation resistance. Nat. Commun. 10, 991. https://doi.org/10.1038/s41467-019-08921-8 (2019).
Xu, K. et al. HIF-1α regulates cellular metabolism, and Imatinib resistance by targeting phosphogluconate dehydrogenase in gastrointestinal stromal tumors. Cell Death Dis. 11, 586. https://doi.org/10.1038/s41419-020-02768-4 (2020).
Chen, H. et al. 6PGD inhibition sensitizes hepatocellular carcinoma to chemotherapy via AMPK activation and metabolic reprogramming. Biomed. Pharmacother. 111, 1353–1358. https://doi.org/10.1016/j.biopha.2019.01.028 (2019).
Bechard, M. E. et al. Pancreatic cancers suppress negative feedback of glucose transport to reprogram chromatin for metastasis. Nat. Commun. 11, 4055. https://doi.org/10.1038/s41467-020-17839-5 (2020).
Bechard, M. E. et al. Pentose conversions support the tumorigenesis of pancreatic cancer distant metastases. Oncogene 37, 5248–5256. https://doi.org/10.1038/s41388-018-0346-5 (2018).
Sarfraz, I. et al. 6-Phosphogluconate dehydrogenase fuels multiple aspects of cancer cells: From cancer initiation to metastasis and chemoresistance. BioFactors 46, 550–562. https://doi.org/10.1002/biof.1624 (2020).
Zheng, W. et al. Inhibition of 6-phosphogluconate Dehydrogenase Reverses Cisplatin Resistance in Ovarian and Lung Cancer. Front Pharmacol. 8, 421. https://doi.org/10.3389/fphar.2017.00421 (2017).
Shan, C. et al. Lysine acetylation activates 6-phosphogluconate dehydrogenase to promote tumor growth. Mol. Cell 55, 552–565. https://doi.org/10.1016/j.molcel.2014.06.020 (2014).
Fujiki, K., Inamura, H., Sugaya, T. & Matsuoka, M. Blockade of ALK4/5 signaling suppresses cadmium- and erastin-induced cell death in renal proximal tubular epithelial cells via distinct signaling mechanisms. Cell Death Differ. 26, 2371–2385. https://doi.org/10.1038/s41418-019-0307-8 (2019).
Wrana, J. L., Attisano, L., Wieser, R., Ventura, F. & Massagué, J. Mechanism of activation of the TGF-beta receptor. Nature 370, 341–347. https://doi.org/10.1038/370341a0 (1994).
Lebrin, F., Deckers, M., Bertolino, P. & Ten Dijke, P. TGF-beta receptor function in the endothelium. Cardiovasc Res. 65, 599–608. https://doi.org/10.1016/j.cardiores.2004.10.036 (2005).
He, B. et al. Polymorphisms of TGFBR1, TLR4 are associated with prognosis of gastric cancer in a Chinese population. Cancer Cell. Int. 18, 191. https://doi.org/10.1186/s12935-018-0682-0 (2018).
Lampropoulos, P. et al. Prognostic significance of transforming growth factor beta (TGF-β) signaling axis molecules and E-cadherin in colorectal cancer. Tumour Biol. 33, 1005–1014. https://doi.org/10.1007/s13277-012-0333-3 (2012).
Fanelli, M. F. et al. Evaluation of incidence, significance, and prognostic role of circulating tumor microemboli and transforming growth factor-β receptor I in head and neck cancer. Head Neck 39, 2283–2292. https://doi.org/10.1002/hed.24899 (2017).
Zhang, L. et al. Circular RNA CircCACTIN Promotes Gastric Cancer Progression by Sponging MiR-331-3p and Regulating TGFBR1 Expression. Int. J. Biol. Sci. 15, 1091–1103. https://doi.org/10.7150/ijbs.31533 (2019).
Tang, Q. & Khvorova, A. RNAi-based drug design: considerations and future directions. Nat. Rev. Drug Discov. 23, 341–364. https://doi.org/10.1038/s41573-024-00912-9 (2024).
Acknowledgements
We thank all the participants in this study and gratefully acknowledge contributions from the TARGET cohort, GSE21257 cohort, and GTEx cohort.
Funding
The Hospital Incubation Project Of The Second Affiliated Hospital Of Nanchang University (Grant No. 2022YNFY12003).
Author information
Authors and Affiliations
Contributions
The experimental work was designed by XC, JD, RD, and LL. RD, TL, JC, JZ and XM collected the data, analyzed figures and tables. The experimental procedures were conducted by LL, QC, JZ, and SC. The main text was written by LL, XC, JD and RD. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The Ethics Committee of Nanchang University’s Second Affiliated Hospital granted ethical approval for this study (Approval No. [2019]088). Each patient involved in this study provided written informed consent. We confirmed that all experiments in this study were performed in accordance with the relevant guidelines and regulations.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, R., Liao, L., Chen, J. et al. Downregulation of ferroptosis-related Genes can regulate the invasion and migration of osteosarcoma cells. Sci Rep 15, 17582 (2025). https://doi.org/10.1038/s41598-025-02319-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02319-x