Abstract
Glutathione S-transferases (GST) in pests plays a critical role in insecticide resistance. In this study, SfGSTe1 expression significant upregulation under chlorantraniliprole exposure, chlorantraniliprole susceptibility improvement after injected dsSfGSTe1 and miR-10-5p target located in the 3′ UTR of SfGSTe1 conducted by luciferase reporter assays were observed in Spodoptera frugiperda (Smith). Injected miR-10-5p agomir reduced significantly SfGSTe1 abundance, together with increased chlorantraniliprole susceptibility, while injected miR-10-5p antagomir improved significantly SfGSTe1 abundance and thus decreased chlorantraniliprole susceptibility in the larvae of S. frugiperda. We revealed that the mRNA stability of SfGSTe1 by preventing its degradation that could have been induced by miR-10-5p and thus increased the resistance of S. frugiperda to chlorantraniliprole. This finding would help to study the mechanism of miRNAs in mediating insecticide susceptibility in pests.
Similar content being viewed by others
Introduction
Agricultural insect pests could lead to great damage to crop production so severely that the plant produces abnormal crop or no crop at all1,2,3. Chemical insecticide application is the main strategy in insect pest management4,5. However, chemical insecticide resistance in insects is increasingly severe with a continuous increase use of these insecticides in order to achieve pest control, which reduce the negative effects in human health and the environmental problems caused by exposure to these chemicals6,7.
The fall armyworm Spodoptera frugiperda (Smith) was distributed worldwide and could damage many crops, including corn, rice, sorghumt and peanu8,9,10. Chlorantraniliprole is the primary insecticide showing significant control effect against several Lepidopteran pests and low toxicity to non-target organisms11,12,13. However, chlorantraniliprole-resistant pests from field populations have developed, such as Spodoptera litura, Chilo suppressalis, Plutella xylostella, and so on14,15,16.
The target mutation often causes a higher chlorantraniliprole resistance in pests6,17, the ryanodine receptor (RyR, the target of diamide insecticides), G4946E and I4790M/K have been known involved in the resistance to chlorantraniliprole18. Forthermore, metabolic detoxification is more frequent than mutations of target site in insecticide resistance, and the detoxication enzymes including cytochrome P450 (P450) and carboxylesterase by acting on the insecticide molecule directly19,20,21, glutathione S-transferases (GSTs) and UDP-glycosyltransferases (UGTs) by conjugating toxins with endogenous molecules20,22, and ATP-binding cassette transporters by facilitating the active transport of toxins23.
These detoxication enzymes may participate in the resistance to chlorantraniliprole. For instance, the GST activity in Bombyx mori could be induced significantly by chlorantraniliprole24. GST expression in P. xylostella were was significantly increased in chlorantraniliprole resistance25. GSTs could be related to chlorantraniliprole resistance in S. frugiperda26.
GST expression can be regulated by transcription and posttranscription20,27,28. microRNAs (miRNAs) could influence regulation of the target genes by combining to coding sequences (CDS), mRNA’s 3′ /5′untranslated region (3′/5′UTR) at the posttranscriptional level, ultimately causing translational inhibition, mRNA decay, or cleavage28,29,30. A lot of miRNAs are found in abundance in a variety of insects28,31,32 and many of them could participate in a lot of important biological processes, like insect metamorphosis33,34,35, reproduction36,37 and pest resistance to insecticides28,38.
It showed that regulation of GSTs could occur at posttranscriptional level when some miRNAs combined to their 3′/5′UTR or CDs. For instance, GSTu1 expression was regulated by miR-8525-5p in P. xylostella20, and TCGSTM4 expression was mediated by tci-miR-1-3p in Tetranychus cinnabarinus39. GSTs are crucial in the metabolic process of exogenous compounds including insecticides in insects, but the mechanism of GST response to insecticides has yet to be fully elucidated. Furthermore, miR-10-5p was predicted to be potentially involved in the upregulation of SfGSTe1 by competing for miR-10-5p binding and thus mediated chlorantraniliprole resistance in S. frugiperda based on our previous study.
Here, posttranscriptional regulation of SfGSTe1 by miR-10-5p was carried out. The target site of miR-10-5p isolated was combined to the 3’UTR of SfGSTe1, which was verified that the expression of SfGSTe1could be mediated by miR-10-5p and thus regulate chlorantraniliprole susceptibility. This highlights the importance of miR-10-5p in mediating SfGSTe1 expression. It is critical to explore the mechanism of GSTs in the tolerance of S. frugiperda to insecticides.
Materials and methods
Insects
The S. frugiperda strain used was collected from Jie Dong Village, Mengmao Town, Ruili City of Yunnan Province (23°58′35″ N, 97°48′51′ E) in 2019. And they have been reared in the greenhouse with a photoperiod of 16 h:8 h (L:D), a temperature of 25 ± 2 °C, and a relative humidity of 65 ± 5% without exposure to any insecticides. The artificial diet ingredient preparation, feeding and eggs collected as breeding the next generation were based on the method described previously40. The samples of developmental stages and the tissues in S. frugiperda were prepared as the method of Jiang et al.41 The artificial diet ingredients used to feed the larvae include soybean powder, propionic acid, and vitamin E. Three-day-old moths were fed an 8–10% sugar solution, and spawning was performed on fresh corn leaves.
Chlorantraniliprole and bioassay
The chlorantraniliprole (98% of the industrial-grade active compound) was supplied by Shenzhen Noposion Agricultural Chemical Co., LTD (Shenzhen, China), and Triton X-100 (X-100) was obtained by Sigma-Aldrich Co. (St. Louis, USA). The chlorantraniliprole toxicity to larvae and the samples treated were carried out by leaf-dipping method41. The 50% lethal concentration (LC50) was conducted by POLOPlus 2.0 software (LeOra Software Inc., Berkeley, CA, USA)42.
Cloning and RNA interference of SfGSTe1
For the first strand cDNAs of 3’/5' RACE, construction of the PCR system followed instructions from the SmartTM Race cDNA Amplification Kit (Clontech). Two gene-specific primers were used to amplify full-length cDNAs based on a fragment of the GST-like sequence SfGSTe1. Thermal cycling using touchdown PCR was carried out based on the procedure described by Zhang et al.43.
The cDNA fragments (PCR products) of SfGSTe1 and the green fluorescent protein (GFP) were expand by using reverse transcription PCR (RT-PCR) according to gene-specifific primers (Table S1). The PCR products served as templates for dsRNA synthesis according to the MEGAscriptRNAi kit based on manufacturer’s instructions from Ambion, Inc. The second instar larva each were injected about 300 ng of dsRNA via microinjector (Nanoliter 2000 Injector; WPI, Inc., Sarasota FL, USA). The 30 larvae post-injected dsRNA for 24 h were randomly selected for analysis of SfGSTe1 expression cunducted by quantitative real-time PCR (qPCR). The chlorantraniliprole toxicity against dsSfGSTe1- and dsGFP-injected larvae was carried out 24 h post-injection exposed to the LC50 chlorantraniliprole conducted by leaf-dipping method41. Corn leaves were cut into 20-mm-long pieces and immersed in TX-100 (0.1%, V/V) sterile water containing chlorantraniliprole for 15 s or in Triton X-100 (0.1%, V/V) sterile water without chlorantraniliprole as controls.
Homology modelling and molecular docking
The 3-dimensional structure of SfGSTe1 protein was carried out online by Swiss model server. The glutathione-S-transferase from Trichoplusia ni (Cabbage looper) (PDB ID: A0A7E5WJU8.1.A) with 74.89% sequence similarity was used standard model. ChemDraw 20.0 and Chem3D for the 3-dimensional structure of chlorantraniliprole, Autodock 4.2 for the default parameters of molecular docking and the combining free energy and interaction, and PLIP (Protein–Ligand Interaction Profiler) for the interaction of protein and ligand were conducted. The docking and modelling were presented by PyMOL.
Expression, purification, and metabolism assays of SfGSTe1
Sequencing-confirmed SfGSTe1 digested by BamHI and EcoRI, inserted into the expression vector pET-28a (+), and transferred into E. coli BL21 (DE3) cells for expression.
The recombinant protein expression of SfGSTe1 was induced, and identified by Luria–Bertani (LB) liquid medium including isopropyl β-d-thiogalactoside (IPTG), and SDS-PAGE, respectively. The recombinant SfGSTe1 protein was inhibited, and eluted by Ni–NTA His Bind Resin and imidazoles, respectively.
The purified SfGSTe1 protein dialyzed, and concentration determined by Tris–HCl buffer (10 mM, including 150 mM NaCl and 50% glycerol), and a Bicin-choninic Acid Protein Assay Kit20.
The specific recombinant SfGSTe1 activity was determined according to the method described by Zhang et al.44 The 200 μL of reaction volume contained 10 μL of recombinant SfGSTe1 protein, and the reaction mixture with 10.0 mM of GSH, 1.0 mM of CDNB and 50 mM PBS (phosphate-buffered saline, pH 6.5).
The chlorantraniliprole inhibition for recombinant SfGSTe1 activity was also conducted according to the method above, and the same volume with a 10 μM concentration of chlorantraniliprole was carried out based on our preliminary test. The same volume of PBS (50 mM, pH 6.5) and boiled recombinant SfGSTe1 protein was set as the negative control, and three biological replicates were carried out. Changes in absorbance values were recorded at 340 nm for 5 min using an Absorbance Microplate Reader.
The SfGSTe1 metabolism to chlorantraniliprole in vitro assays was carried out according to the method (high-performance liquid chromatography, HPLC grade) by Zhu et al.20 The 500 μL of the reaction volume including PBS (0.1 mM, pH 7.0), GSH (2.5 mM), chlorantraniliprole (0.50 mM), and recombinant SfGSTe1 protein (25 μL) was maintained at 34 °C for 2 h, and stopped by adding methanol (500 μL). A boiled recombinant SfGSTe1 protein was set as the negative control. The supernatant obtained by centrifuging was transferred into HPLC vials, and injected into an LC-18 reverse-phase analytical column. Chlorantraniliprole separation of was conducted by acetonitrile/water (70:30, v/v) as the mobile phase with flow rate at 0.4 ml min−1. Quantification by peak integration was determined by the change in absorbance at 260 nm and a standard curve (Fig. S1).
miRNA target studies of SfGSTe1
Three software programs of miRNA target prediction, the Target Scan, RNAhybrid, and Miranda programs45,46,47, were used as default arguments to speculate miRNAs that target the 3’UTRs of SfGSTe1 sequences. Standardized firefly luciferase activity assays was followed by Zhang et al.28 miR-10-5p agomir/antagomir along with their respective negative control (NC agomir/NC antagomir) was synthesized by Shanghai Genepharm Co., Ltd. The luciferase reporter plasmid construction was conducted by the manufacturer’s instructions.
Effect of miR-10-5p on chlorantraniliprole susceptibility
The 2nd instar larvae were collected and injected with microsyringes. Each larva received an injection of 150 nL of 40 μM miR-10-5p agomir/antagomir, while 2nd instar larvae injected with NC agomir/antagomir were as controls48. Furthermore, the sensitivity of injected larvae chlorantraniliprole was carried out by the leaf-dipping method after injection of agomir/antagomir of miRNA.
miR-10-5p and SfGSTe1 expression by qPCR
Total RNA extraction for SfGSTe1 expression, and miRNA expression according to the manufacturer’s instructions by using the TRIzol kit, and miRNeasy Mini Kit, respectively.
An Applied Biosystems 7500 Real-Time PCR system was used to determine Ct values of each sample in triplicate.
The miR-10-5p and SfGSTe1 expression was standardized to U6 small nuclear RNA (snRNA)28 and GAPDH mRNA40, respectively. The calculation and analysis of gene expression was according to the 2–ΔΔCt method49. Three biological replications was conducted for each sample. The primers tested were list in Table S1.
Statistical analysis
The data were analyzed by unpaired t tests using GraphPad InStat 3.0 software (GraphPad Software, San Diego, CA, USA). The relative luciferase activity of each transfection group and the expression in different developmental stages and tissues were compared by one-way analysis of variance (ANOVA) and Tukey’s multiple comparison test.
Results
Concentration mortality response
The chlorantraniliprole toxicity against 2nd instar larvae of S. frugiperda was conducted by by leaf-dipping method. The results showed that the LC25 and LC50 values were 0.968 and 1.586 μg mL−1, respectively. Concentration mortality and the effects of chlorantraniliprole on SfGSTe1 and miR-10-5p expression were determined based on these data (Table S2).
SfGSTe1 cloning
The GSTe1-like expression could be upregulated significantly by chlorantraniliprole indicating that it could be involved in chlorantraniliprole detoxication/metabolism in S. frugiperda. To study the function of GSTe1-like, we isolated a 687-bp open reading frame (ORF), coding for 228 amino acid residues, a 187-bp 3′UTR by the 3′/5′-RACE technique, and named SfGSTe1. The SfGSTe1 sequence in S. frugiperda was uploaded to NCBI (Accessions No: OR921092).
SfGSTe1 with chlorantraniliprole by molecular docking
The binding mode conducted by SWISS-MODELE indicated that SfGSTe1 can dock well with chlorantraniliprole (Fig. 1). The 3D structure of SfGSTe1 included 10 α-helixes, 4 β-sheets and linked by several loops. The hydrogen bond was formed by GLU148 with amide on the benzene ring, ARG100 with oxygen molecule on the amide, and TRP63 with pyridine ring or imidazole ring form hydrogen bond, with a length of 2.3 Å, 2.5 Å, 2.3 Å and 2.0 Å, respectively. Binding energy of SfGSTe1 with chlorantraniliprole is − 5.11 kcal/mol.
The interactions of SfGSTe1 with chlorantraniliprole. The yellow dotted lines indicate the hydrogen bond.
The metabolic activity of SfGSTe1 to chlorantraniliprole
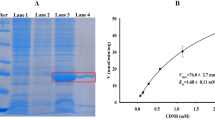
The recombinant SfGSTe1protein was soluble and expressed in Escherichia coli. The molecular mass was about 26 KD, and about 2 mg of the protein was collected by purification (Fig. 2A); The specific recombinant SfGSTe1 activity toward a substrate CDNB was 2.206 μmol/min/mg, and can be reduced by 49.3% under 10 μM chlorantraniliprole competition (Fig. 2B); The chlorantraniliprole metabolism by recombinant SfGSTe1 was determined by HPLC, indicating that about 43.2% of chlorantraniliprole could be metabolized by recombinant SfGSTe1 protein compared with the control (boiled SfGSTe1) (Fig. 2C).
The metabolic activity of recombinant protein SfGSTe1 to chlorantraniliprole. (A): Expression and purification of SfGSTe1 in the E. coli system (B): The specific activity of purified SfGSTe1 determined and inhibited by chlorantraniliprole. (C) Metabolic rate of chlorantraniliprole by recombinant SfGSTe1 (%). The results are shown as means ± S.E. for three independent biological replicates. Different letters indicate significant differences between the treatment and the control (Student’s t-test, P < 0.05).
miR-10-5p regulating SfGSTe1 expression
To comfirm if miR-10-5p could effect predicted combing site of the 3′ UTR and regulate SfGSTe1 expression in S. frugiperda. The wild-type sequence, 3′ UTR of SfGSTe1 cloned, was transfered into a pmirGLO-SfGSTe1-UTR (Fig. 3A). When miR-10-5p agomir and pmirGLO-SfGSTe1-UTR constructs were transfected into HEK293T cells, a significant decrease in luciferase activity by 40.9% was observed compared with the control (HEK293T cells transfected with NC agomir alone). In addition, the combining site of the 3 'UTR of SfGSTe1 sequence was mutated as a “seed” (complementary to miR-10-5p sequence) and transfected into the pmirGLO vector. When co-transfected with pmirGLO-SfGSTe1-Mut and miR-10-5p agomir into HEK293T cells, there was no significant changes compared with the control (HEK293T cells transfected with NC agomir alone) (Fig. 3B).
Interaction of miR-10-5p and SfGSTe1. (A) miR-10-5p predicted target sites within the 3′ UTR of SfGSTe1 in S. frugiperda. (B) A luciferase reporter assay was carried out by cotransfecting a luciferase reporter linked to the SfGSTe1 insert with miR-10-5p agomir. The bars represented by different letters (a, b) show significant difference (One-way ANOVA followed by Tukey’s multiple comparison tests, P < 0.05).
The effects of chlorantraniliprole on miR-10-5p and SfGSTe1 expression
To further explore miR-10-5p and SfGSTe1 expression under chlorantraniliprole exposure, the LC25 of chlorantraniliprole were used to treat 2nd instar larvae for 24 h. It showed that the expression of miR-10-5p was downregulated obviously by 46.5% under chlorantraniliprole treatment (Fig. 4A), whereas the expression of SfGSTe1 was upregulated obviously by 2.4-fold under chlorantraniliprole treatment (Fig. 4B).
SfGSTe1 and miR-10-5p expression under chlorantraniliprole treatment in S. frugiperda. (A) SfGSTe1 expression treated by the LC25 chlorantraniliprole for 24 h (B) miR-10-5p expression in S. frugiperda after the LC25 chlorantraniliprole exposure for 24 h. The results are shown as the means ± S. E. for three independent biological replicates. The significant differences between the treatment and control are represented by different letters (a, b) (Student’s t-test, P < 0.05).
The changes of chlorantraniliprole susceptibility after silencing SfGSTe1
To determine SfGSTe1 silencing efficiency, it showed suppression of RNAi (dsSfGSTe1) by qPCR. SfGSTe1 expression decreased significantly by 44.6% in 2nd instar larvae dsSfGSTe1-injected for 24 h compared to that in controls (2nd instar larvae dsGFP-injected for 24 h) (Fig. 5A). Forthermore, the mortality was also significantly increased in 2nd instar larvae dsSfGSTe1-injected for 24 h (46.7%) compared to that in controls (2nd instar larvae dsGFP-injected for 24 h) (24.5%) when dsRNA-injected 2nd instar larvae were treated by the LC25 of chlorantraniliprole for 24 h (Fig. 5B).
SfGSTe1 expression and the mortality under the LC25 chlorantraniliprole exposure. (A) SfGSTe1 expression (B) Mortality (%) of S. frugiperda under the LC25 chlorantraniliprole treatment. The results of three independent biological repetitions are expressed as the mean ± S. E. The bars represented by different letters (a, b) show significant difference (Student’s t-test, P < 0.05).
SfGSTe1 expression patterns
The expression patterns of SfGSTe1 highly expressed under insecticide treatments were further determined in different developmental stages, and tissue distribution, respectively. The SfGSTe1 expression was higher in 4th instar larvae, with 2.60-fold compared to the eggs; While SfGSTe1 expression was lower in 1st instar larvae, 3rd instar larvae and pupae, with 0.28-, 0.62-, and 0.79-fold compared to the eggs; No significant difference was observed among 3rd instar larvae, 5th instar larvae and adults, with 0.86-, 1.30-, and 1.25-fold compared to the eggs (Fig. 6A).
SfGSTe1 expression in developmental stages and tissues of S. frugiperda. (A) Different developmental stages. (B) Different tissues. The results of three independent biological repetitions are expressed as the mean ± S. E. The significant difference among different treatments are represented by different letters (One-way ANOVA followed by Tukey’s multiple comparison tests, P < 0.05).
For the analyses of SfGSTe1 expression pattern in different tissues from adults were explored. It showed that SfGSTe1 expression was higher in the thoraxes, wings, midguts, and fat bodies, with 1.64-, 1.45-, 3.35- and 3.75-fold compared to the heads, and the SfGSTe1 expression was highest in fat bodies (Fig. 6B).
The impacts of miR-10-5p abundance on the expression of SfGSTe1 and chlorantraniliprole susceptibility
miR-10-5p abundance was upregulated obviously by 4.56-fold in the larvae when the midgut was injected by miR-10-5p agomir for 48 h compared with the control (injection of NC agomir for 48 h) (Fig. 7A). Additionally, the expression of SfGSTe1 in the larvae when the midgut was injected by miR-10-5p agomir for 48 h was downregulated obviously by 42.1% compared to that in the control (injection of NC agomir for 48 h) (Fig. 7B). Forthermore, the mortality in 2nd instar larvae when the midgut was injected by miR-10-5p agomir for 24 h was increased obviously by 24.0% compared with the control (injection of NC agomir for 24 h) after the larvae miR-10-5p agomir/NC agomir injected were treated by the LC50 of chlorantraniliprole for 24 h (Fig. 7C). However, miR-10-5p abundance in 2nd instar larvae when the midgut was injected by miR-10-5p antagomir for 48 h was downregulated obviously by 44.2% compared with the control (injection of NC antagomir for 48 h) (Fig. 7D). Additionally, SfGSTe1 expression in 2nd instar larvae when the midgut was injected by miR-10-5p antagomir was upregulated obviously by 2.04-fold compared with the control (injection of NC antagomir for 48 h) (Fig. 7E). Forthermore, the mortality in 2nd instar larvae when the midgut was injected by miR-10-5p antagomir for 24 h was downregulated obviously by 19.33% compared wih the control (injection of NC antagomir for 24 h) after the larvae miR-10-5p antagomir/ NC antagomir injected were treated by the LC50 of chlorantraniliprole for 24 h (Fig. 7F).
The impacts of miR-10-5p abundance on SfGSTe1 expression and chlorantraniliprole susceptibility in S. frugiperda. (A, D): miR-10-5p expression in the midgut of 2nd larvae injected with miR-10-5p agomir (A) or antagomir (D) (48 h injection). (B, E): SfGSTe1 expression in the midgut of 2nd larvae injected with miR-10-5p agomir (B) or antagomir (E) (48 h injection). (C, F): Mortality in the midgut of 2nd larvae injected with miR-10-5p agomir (C) or antagomir (F) in S. frugiperda when exposed to the LC50 chlorantraniliprole. The results of three independent biological repetitions are expressed as the mean ± S. E. * indicated significant variations (Student’s t-test, * P < 0.05, **P < 0.01,***P < 0.001).
Discussion
GST as a critical detoxification enzyme could catalyze glutathione (GSH) binding to chemicals, improving solubility and promotes excretion in animal cells, and It has been reported that insecticide resistance development is from metabolism/detoxication by GSTs50. In our study, SfGSTe1 was verified to participate in the resistance to chlorantraniliprole by RNAi in S. frugiperda. Moreover, the recombinant SfGSTe1 protein activity was suppressed by chlorantraniliprole, indicating its contribution to insecticide resistance. Recombinant SfGSTe1 has showed high efficacy in metabolizing chlorantraniliprole. There were many studies that GSTs involved in metabolizing organochloride51,52, organophosphorus53, pyrethroid insecticides in insects54, and cyflumetofen in spider mites55.
miRNAs are found within organisms, playing a crucial role in mediating gene expression by combining to CDs or 3’/5' UTR of target genes and subsequently leading to cleavage, mRNA decay or translational repression. Insecticides are known to be significant environmental stressors for pests, and there is evidence suggesting that miRNAs could contribute to the resistance to insecticides by modulating gene expression of insecticide targets or detoxification enzymes28,48,56. For example, Aphis gossypii exhibits resistance to spirotetramat due to the regulation of acetyl-CoA carboxylase expression mediated by miR-3016 and miR-27657. miR-4133-3p could regulate CYP4CJ1 expression in A. gossypii involved in gossypol and tannic acid tolerance58. In Sitobion miscanthi, miR-316 may contribute to insecticides resistance by regulating CYP4CJ6 expression28. A similar study found that miR-316 can target three detoxication genes, a P450 gene and two GST genes in Manduca sexta59. In insects miRNAs can regulate gene expression exhibiting a negative relationship with their target gene expression60. Thus, miR-10-5p displayed downregulated while targeting SfGSTe1 expression upregulated when the 2nd instar larvae were treated by chlorantraniliprole. Forthermore, miR-10-5p identified to target SfGSTe1 was comfirmed to mediate SfGSTe1 expression and subsequently affect on chlorantraniliprole susceptibility in S. frugiperda conducted by in vitro injecting miR-10-5p antagomir/agomir and dual luciferase reporter assay methods. This indicated that miR-10-5p could change the susceptibility to insecticides by mediating the expression of SfGSTe1 in S. frugiperda.
There are reports that Hox gene expression regulated by miR-10 participated in embryonic development in Nile tilapia61. Furthermore, miR-10a-5p could promote chondrocyte apoptosis in osteoarthritis by mediating HOXA162; miR-10 could play an integral role in the evolution of numerous cancer types63. miRNAs could target specific gene clusters by multiple sites in the 3′UTR, and a lot of target sites could combine to the silencing complex sufficiently, leading to mRNA enrichment and thereby exhibiting more potent repressive effects64,65.
The luciferase activity analysis revealed that SfGSTe1 target sequences at these sites exhibiting strong combining affinity with miR-10-5p in S. frugiperda. Furthermore, we chose larvae to elucidate the role of SfGSTe1 in insecticide detoxification as they are more amenable for RNAi experiments due to their rapid acquisition of dsRNA through injection, as suggested by previous studies on dsRNA acquisition by larvae.
It has been reported that injection could induce RNAi in 2nd instar larvae found in Lepidoptera pests48,66,67. RNAi-based approaches offer the possibility for new perspectives on controlling pests. Knockdown of dsRNA by injecting indicated that dsRNA was efficient at silencing of target genes. Analysis of SfGSTe1 knockdown in the 2nd instar larvae impacts on insecticide sensitivity. For chlorantraniliprole exposure, the mortality was increased in dsSfGSTe1-injected 2nd instar larvae compared to that in controls (dsGFP-injected 2nd instar larvae for 24 h). This indicated that SfGSTe1 could be related to insecticide detoxification. Furthermore, developmental stages or tissue-specific expression patterns of GSTs can indicate their distinct functions in animals68,69,70. In our present study, SfGSTe1 was higher in the midguts and fat bodies compared with the heads. The SfGSTe1 overexpression in the midguts and fat bodies might reflect its important role in insecticide mechanisms/detoxication. This was somewhat similar with findings that SfGSTe1 may be involved in mechanisms/detoxication or insecticide regulation as the tissue-specific expression was related to detoxification, illustrating potentially their crucial functions in biology and physiology, and the midgut and fat body tissues in insects were the important detoxification organs. Additionally, the expression levels of these genes-related to insecticide resistance closely correlate with different developmental stages in insects71,72. Our results showed SfGSTe1 was highest in the 4th instar larvae and adults compared to the eggs. Additionally, our findings also support previous research indicating the possible role of these genes in insecticide mechanisms/detoxication, particularly in larvae, which are more susceptible to such processes. This strengthened the possibility that SfGSTe1 could be involved in insecticide mechanisms/detoxication in S. frugiperda. In addition, prolonged rearing of S. frugiperda on an artificial diet, followed by bioassays using host plant leaves, might influence gene expression.
In summary, miR-10-5p was confirmed to be potentially involved in the upregulation of SfGSTe1 to mediate chlorantraniliprole resistance. Injected miR-10-5p agomir was significantly downregulated SfGSTe1 expression to increase chlorantraniliprole susceptibility while injected miR-10-5p antagomir was significantly upregulated SfGSTe1 expression to decrease chlorantraniliprole susceptibility. This provided a crucial foundation for further study functions of miRNAs in the regulation of SfGSTe1 and chlorantraniliprole susceptibility in pests.
Data availability
The data presented in this study are available on request from the corresponding author.
References
Manosathiyadevan, M., Bhuvaneshwari, V. & Latha, R. Impact of insects and pests in loss of crop production: a review. In Sustainable Agriculture towards Food Security, 57–67 (2017)
Ali, M. A., Abdellah, I. M. & Eletmany, M. R. Towards sustainable management of insect pests: Protecting food security through ecological intensification. Int. J. Chem. Biochem. Sci. 24(4), 386–394 (2023).
Wang, L., Reddy, G. V., Wang, S. & Zhao, Z. The contrasting response of crop production and pest damage to ENSO cycles. Entomol. Gen. 43(2), 325–335 (2023).
van den Berg, H. et al. Recent trends in global insecticide use for disease vector control and potential implications for resistance management. Sci. Rep. 11(1), 23867 (2021).
Ran, X., Hadiatullah, H., Yuchi, Z., Yang, X. & Zhu, X. Sustainable use of pesticides. Agriculture 13(7), 1393 (2023).
Walsh, T. K. et al. Determinants of insecticide resistance evolution: comparative analysis among Heliothines. Annu. Rev. Entomol. 67, 387–406 (2022).
Chen, H. L. et al. Resistance monitoring and mechanism in the fall armyworm Spodoptera frugiperda (Lepidoptera: Noctuidae) for chlorantraniliprole from Sichuan Province, China. Front. Physiol. 14, 1180655 (2023).
Overton, K. et al. Global crop impacts, yield losses and action thresholds for fall armyworm (Spodoptera frugiperda): A review. Crop Prot. 145, 105641 (2021).
Kenis, M. et al. Invasiveness, biology, ecology, and management of the fall armyworm, Spodoptera frugiperda. Entomol. Gen. 43(2), 187–241 (2023).
Lv, S. et al. Low concentrations of cyantraniliprole negatively affects the development of Spodoptera frugiperda by disruption of ecdysteroid biosynthesis and carbohydrate and lipid metabolism. Pestic. Biochem. Physiol. 200, 105827 (2024).
Li, Y. P. et al. Current status of insecticide resistance in Spodoptera frugiperda and strategies for its chemical control. Plant Prot. 45, 14–19 (2019).
Wang, N. M., Li, J. J., Shang, Z. Y., Yu, Q. T. & Xue, C. B. Increased responses of phenoloxidase in chlorantraniliprole resistance of Plutella xylostella (Lepidoptera: Plutellidae). J. Insect Sci. 20(4), 2 (2020).
Li, X. et al. Drip application of chlorantraniliprole effectively controls invasive Spodoptera frugiperda (Lepidoptera: Noctuidae) and its distribution in maize in China. Crop Prot. 143, 105474 (2021).
Muthusamy, R., Vishnupriya, M. & Shivakumar, M. S. Biochemical mechanism of chlorantraniliprole resistance in Spodoptera litura (Fab)(Lepidoptera: Noctuidae). J. Asia-Pacific Entomol. 17(4), 865–869 (2014).
Xu, L. et al. Constitutive overexpression of cytochrome P450 monooxygenase genes contributes to chlorantraniliprole resistance in Chilo suppressalis (Walker). Pest Manag. Sci. 75(3), 718–725 (2019).
Shan, J. et al. Identification of ABCG transporter genes associated with chlorantraniliprole resistance in Plutella xylostella (L.). Pest Manag. Sci. 77(7), 3491–3499 (2021).
Ma, R. et al. Structural basis for diamide modulation of ryanodine receptor. Nat. Chem. Biol. 16(11), 1246–1254 (2020).
Teng, H. et al. High frequency of ryanodine receptor and cytochrome P450 CYP9A186 mutations in insecticide-resistant field populations of Spodoptera exigua from China. Pestic. Biochem. Physiol. 186, 105153 (2022).
Lu, K., Song, Y. & Zeng, R. The role of cytochrome P450-mediated detoxification in insect adaptation to xenobiotics. Curr. Opin. Insect Sci. 43, 103–107 (2021).
Zhu, B., Li, L., Wei, R., Liang, P. & Gao, X. Regulation of GSTu1-mediated insecticide resistance in Plutella xylostella by miRNA and lncRNA. PLoS Genet. 17(10), e1009888 (2021).
Zhang, Y., Li, J., Ma, Z., Shan, C. & Gao, X. Multiple mutations and overexpression of the MdaE7 carboxylesterase gene associated with male-linked malathion resistance in housefly, Musca domestica (Diptera: Muscidae). Sci. Rep. 8(1), 224 (2018).
Yan, M. W., Xing, X. R., Wu, F. A., Wang, J. & Sheng, S. UDP-glycosyltransferases contribute to the tolerance of parasitoid wasps towards insecticides. Pestic. Biochem. Physiol. 179, 104967 (2021).
Guan, D. et al. Identification and validation of ATP-binding cassette transporters involved in the detoxification of abamectin in rice stem borer, Chilo suppressalis. J. Agric. Food Chem. 70(15), 4611–4619 (2022).
Mao, T. et al. Effects of chlorantraniliprole exposure on detoxification enzyme activities and detoxification-related gene expression in the fat body of the silkworm, Bombyx mori. Ecotoxicol. Environ. Saf. 176, 58–63 (2019).
Yin, F. et al. The glutathione S-transferase (PxGST2L) may contribute to the detoxification metabolism of chlorantraniliprole in Plutella xylostella (L.). Ecotoxicology 30(6), 1007–1016t (2021).
Sun, Z. et al. Genome-wide scanning loci and differentially expressed gene analysis unveils the molecular mechanism of chlorantraniliprole resistance in Spodoptera frugiperda. J. Agric. Food Chem. 71(38), 14092–14107 (2023).
Pritchard, C. C., Cheng, H. H. & Tewari, M. MicroRNA profifiling: approaches and considerations. Nat. Rev. Genet. 13, 358–369 (2012).
Zhang, B. Z. et al. Identification of differentially expressed microRNAs under imidacloprid exposure in Sitobion miscanthi. Pestic. Biochem. Physiol. 177, 104885 (2021).
Gu, W. et al. The role of RNA structure at 5′ untranslated region in microRNA-mediated gene regulation. RNA 20(9), 1369–1375 (2014).
Yang, Y. et al. MicroRNA PC-5p-3991_515 mediates triflumezopyrim susceptibility in the small brown planthopper through regulating the post-transcriptional expression of P450 CYP417A2. Pest Manag. Sci. 80(4), 1761–1770 (2024).
Liang, P., Feng, B., Zhou, X. & Gao, X. Identification and developmental profiling of microRNAs in diamondback moth, Plutella xylostella (L.). PLoS ONE 8, e78787 (2013).
Zhu, B., Li, X., Liu, Y., Gao, X. & Liang, P. Global identification of microRNAs associated with chlorantraniliprole resistance in diamondback moth Plutella xylostella (L.). Sci. Rep. 7, 40713 (2017).
Yang, M. et al. miR-71 and miR-263 jointly regulate target genes chitin synthase and chitinase to control Locust molting. PLoS Genet 12, e1006257 (2016).
Belles, X. MicroRNAs and the evolution of insect metamorphosis. Annu. Rev. Entomol. 62(1), 111–125 (2017).
Ye, X. et al. miR-34 modulates wing polyphenism in planthopper. PLoS Genet. 15, e1008235 (2019).
Ling, L., Kokoza, V. A., Zhang, C., Aksoy, E. & Raikhel, A. S. MicroRNA-277 targets insulin-like peptides 7 and 8 to control lipid metabolism and reproducation in Aedes aegypti mosquitoes. Proc. Natl. Acad. Sci. U.S.A. 114, E8017–E8024 (2017).
Wang, N. et al. Characterization of microRNAs associated with reproduction in the brown planthopper, Nilaparvata lugens. Int. J. Mol. Sci. 23(14), 7808 (2022).
Peng, T. F. et al. Reduced abundance of the CYP6CY3-targeting let-7 and miR-100 miRNAs accounts for host adaptation of Myzus persicae nicotianae. Insect Biochem. Mol. Biol. 75, 89–97 (2016).
Zhang, Y. et al. A microRNA-1 gene, tci-miR-1-3p, is involved in cyflumetofen resistance by targeting a glutathione S-transferase gene, TCGSTM4, in Tetranychus cinnabarinus. Insect Mol. Biol. 27(3), 352–364 (2018).
Zhang, B. Z. et al. Silencing of cytochrome P450 in Spodoptera frugiperda (Lepidoptera: Noctuidae) by RNA interference enhances susceptibility to chlorantraniliprole. J. Insect Sci. 20(3), 12 (2020).
Jiang, Y. T. et al. MicroRNA-23a changes chlorantraniliprole susceptibility by targeting CYP9F1 in Spodoptera frugiperda. Entomologia Generalis 43(5), 1041–1049 (2023).
Desneux, N., Decourtye, A. & Delpuech, J. M. The sublethal effects of pesticides on beneficial arthropods. Annu. Rev. Entomol. 52, 81–106 (2007).
Zhang, B., Zhang, L., Cui, R., Zeng, X. & Gao, X. Cloning and expression of multiple cytochrome P450 genes: Induction by fipronil in workers of the red imported fire ant (Solenopsis invicta Buren). PLoS ONE 11(3), e0150915 (2016).
Zhang, L., Gao, X. W. & Liang, P. Beta-cypermethrin resistance associated with high carboxylesterase activities in a strain of house fly, Musca domestica (Diptera: Muscidae). Pestic. Biochem. Physiol. 89, 65–72 (2007).
Enright, A. J. et al. MicroRNA targets in Drosophila. Genome Biol. 5, R1 (2003).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004).
Betel, D., Wilson, M., Gabow, A., Marks, D. S. & Sander, C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 36, 149–153 (2008).
Zhu, B., Sun, X., Nie, X., Liang, P. & Gao, X. MicroRNA-998–3p contributes to Cry1Ac-resistance by targeting ABCC2 in Lepidopteran insects. Insect Biochem. Mol. Biol. 117, 103283 (2020).
Pfaffl, M. W. A new mathematical model for relative quantification in real–time RT–PCR. Nucleic Acids Res. 29, 45–45 (2001).
Pavlidi, N., Vontas, J. & Van Leeuwen, T. The role of glutathione S-transferases (GSTs) in insecticide resistance in crop pests and disease vectors. Curr. Opin. Insect Sci. 27, 97–102 (2018).
Low, W. Y. et al. Recognition and detoxification of the insecticide DDT by Drosophila melanogaster glutathione S-transferase D1. J. Mol. Biol. 399(3), 358–366 (2010).
Riveron, J. M. et al. A single mutation in the GSTe2 gene allows tracking of metabolically based insecticide resistance in a major malaria vector. Genome Biol. 15(2), R27 (2014).
Lu, X. P. et al. Divergent molecular evolution in glutathione S-transferase conferring malathion resistance in the oriental fruit fly, Bactrocera dorsalis (Hendel). Chemosphere 242, 125203 (2020).
Wang, W., Hu, C., Li, X. R., Wang, X. Q. & Yang, X. Q. CpGSTd3 is a lambda-cyhalothrin metabolizing glutathioneS-transferase from Cydia pomonella (L.). J. Agric. Food Chem. 67(4), 1165–1172 (2019).
Pavlidi, N. et al. A glutathione S-transferase (TuGSTd05) associated with acaricide resistance in Tetranychus urticae directly metabolizes the complex II inhibitor cyflumetofen. Insect Biochem. Mol. Biol. 80, 101–115 (2017).
Wu, M. et al. miR-317-3p and miR-283-5p play a crucial role in regulating the resistance to indoxacarb in Spodoptera frugiperda by targeting GSTs4. J. Agric. Food Chem. 72(13), 6889–6899 (2024).
Wei, P. et al. Characteristics of carboxylesterase genes and their expression-level between acaricide-susceptible and resistant Tetranychus cinnabarinus (Boisduval). Pestic. Biochem. Physiol. 131, 87–95 (2016).
Ma, Y. et al. miR-10a-5p promotes chondrocyte apoptosis in osteoarthritis by targeting HOXA1. Mol. Ther. Nucleic Acids 14, 398–409 (2019).
Zhang, X. et al. Identification and profiling of Manduca sexta microRNAs and their possible roles in regulating specific transcripts in fat body, hemocytes, and midgut. Insect Biochem. Mol. Biol. 62, 11–22 (2015).
Wang, A. et al. A microRNA, PC-5p-30_205949, regulates triflumezopyrim susceptibility in Laodelphax striatellus (Fallén) by targeting CYP419A1 and ABCG23. Pestic. Biochem. Physiol. 192, 105413 (2023).
Giusti, J. et al. MicroRNA-10 modulates Hox genes expression during Nile tilapia embryonic development. Mech. Dev. 140, 12–18 (2016).
Ma, K. et al. CYP4CJ1-mediated gossypol and tannic acid tolerance in Aphis gossypii glover. Chemosphere 219, 961–970 (2019).
Li, C. et al. Recent updates on the role of the MicroRNA-10 family in gynecological malignancies. J Oncol 2022, 1–8 (2022).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Schnall-Levin, M. et al. Unusually effective microRNA targeting within repeat-rich coding regions of mammalian mRNAs. Genome Res. 21, 1395–1403 (2011).
Wang, R. L. et al. Identification and functional analysis of a novel cytochrome P450 gene CYP9A105 associated with pyrethroid detoxification in Spodoptera exigua hübner. Int. J. Mol. Sci. 19(3), 737 (2018).
Zhu, B., Shan, J., Li, R., Liang, P. & Gao, X. Identification and RNAi-based function analysis of chitinase family genes in diamondback moth, Plutella xylostella. Pest Manag. Sci. 75(7), 1951–1961 (2019).
Knight, T. R., Choudhuri, S. & Klaassen, C. D. Constitutive mRNA expression of various glutathione S-transferase isoforms in different tissues of mice. Toxicol. Sci. 100(2), 513–524 (2007).
Mittapalli, O., Neal, J. J. & Shukle, R. H. Tissue and life stage specificity of glutathione S-transferase expression in the Hessian fly, Mayetiola destructor: implications for resistance to host allelochemicals. J. Insect Sci. 7(1), 20 (2007).
Tierbach, A., Groh, K. J., Schönenberger, R., Schirmer, K. & Suter, M. J. F. Glutathione S-transferase protein expression in different life stages of zebrafish (Danio rerio). Toxicol. Sci. 162(2), 702–712 (2018).
Long, C., Guo, J. & Wan, F. Molecular cloning, sequence analysis and developmental expression of cDNA fragment of juvenile hormone esterase from the whitefly Bemisia tabaci MED (Hemiptera: Aleyrodidae). Acta Entomol. Sin. 56, 234–242 (2013).
Mao, W. et al. Quercetin-metabolizing CYP6AS enzymes of the pollinator Apis mellifera (Hymenoptera: Apidae). Comp. Biochem. Physiol. Part B 154, 427–434 (2009).
Funding
This work is supported by Henan Provincial Natural Science Foundation of China (222300420440), the Project of Plant Protection Key Discipline of Henan Province (1070202190011005).
Author information
Authors and Affiliations
Contributions
F.K., L.C and B. Z. conceived and designed the experiments. YJ.F., P.Z., and M.Z. performed the experiments. Y.P. Y.X., M.Z., and B.Z. analyzed the data and wrote the manuscript. X.J, R.L., M.Z., and B. Z. participated in the data analysis. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kong, FB., Cui, LL., Zhang, BZ. et al. miR-10-5p mediated susceptibility to chlorantraniliprole by targeting for SfGSTe1 in Spodoptera frugiperda (Smith). Sci Rep 15, 19833 (2025). https://doi.org/10.1038/s41598-025-04684-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-04684-z