Abstract
The molecular pathophysiology of nephrotic syndrome remains largely elusive in pediatric patients. While most children with minimal change disease (MCD) show favorable responses to immunosuppressive therapy, those with focal segmental glomerulosclerosis (FSGS) often exhibit poorer treatment responses, with many experiencing either partial remission or no remission of proteinuria. The need for reliable glomerular disease biomarkers to predict treatment response and understand molecular pathways governing responsiveness and resistance is a critical unmet need in pediatric nephrology. In this study, we sought to characterize urine proteomes in children with MCD and FSGS to identify biomarkers distinguishing disease activity and associated molecular pathways. Using quantitative proteomics, urine proteins from children with MCD and FSGS in the CureGN Study were identified and correlated with disease onset and activity. Unbiased cluster analyses of nephrotic urine proteomes demonstrated a cluster with relatively increased immune response and complement proteins, suggesting important distinctions in disease characteristics within the nephrotic subgroups. These analyses yielded patient subpopulations with proteinuria and distinct urine proteome differences associated with 116 proteins exerting cluster separation in the multivariate analyses. These findings highlight the potential of unsupervised clustering to identify disease subgroups and provide insights into the underlying molecular heterogeneity within nephrotic syndrome, paving the way for more tailored therapeutic strategies and improved patient management.
Similar content being viewed by others
Introduction
Mechanisms of nephrotic syndrome remain to be characterized in pediatric patients with minimal change disease (MCD) and focal segmental glomerulosclerosis (FSGS). Therapeutic response in pediatric patients with FSGS is variable as compared to MCD where positive intervention is more likely and outcomes are more positive. Development of glomerular disease biomarkers able to reliably differentiate subgroups could be useful towards identifying molecular patterns in disease activity and potential response to therapy. Proteomics analyses of biofluids in these patient groups could lead to population level stratification tools to better understand treatment outcomes and sensitivity. Identification and characterization of molecular signatures of nephrotic syndrome and changes in patient grouping based on quantitative cluster analysis of the proteome promise more productive implementation of biomarkers in the clinic.
Characterization of the urine proteome has the potential to identify biomarkers of pediatric kidney diseases. Identifying minimally invasive sources of diagnosis and response to therapy could decrease the need for invasive renal biopsies and improve treatment regimens. High-throughput quantitative mass spectrometry approaches have been successful in identifying renal disease biomarkers such as PLA2R1, HTRA12 and THSD7A3 in membranous nephropathy. Translation of such high throughput methods to the study of large numbers of patient urine samples should enable urine biomarker identification to distinguish disease activity, offering diagnostic and prognostic markers of disease progression and therapeutic efficacy. In this context, the current studies were designed to test the hypothesis that characterization of urine proteomes in children with MCD and FSGS could enable identification of urinary biomarkers able to distinguish disease activity levels and identify molecular pathways regulating these differences. These studies utilized urine samples collected from children enrolled in the Cure Glomerulopathy (CureGN) Study, a large prospective multi-center observational study of ~ 2400 adults and children with biopsy-proven glomerular disease in whom detailed clinical data, as well as serial blood and urine biosamples, had been collected. Our current study utilizes quantitative proteomics approaches to characterize the urine proteomes in pediatric MCD and FSGS patients enrolled in the prospective CureGN cohort. We determined differences in patient populations with the same disease activity by performing clustering analysis on urine specimens to visualize differences in the nephrotic urine proteome, offering potential patient stratification and insights into disease heterogeneity.
Results
Patient characteristics and demographics
Urine proteomic analysis was used to cluster patients by unsupervised K-means clustering approaches as detailed below and in Fig. 1. Table 1 describes clinical characteristics and patient demographics in each cluster (CL1 and CL2) at the time of enrollment and biopsy collection. Included are age, sex, race, patient diagnosis, eGFR (estimated glomerular filtration rate), UPCR (urinary protein to creatinine ratio) and kidney failure rate at study follow-up. Table 2 shows clinical characteristics and patient demographics at the time of cluster assignment. UPCR was determined at University of Louisville (UofL) on each analyzed urine specimen at the time of proteomic sample preparation.
Proteomic data processing and bioinformatic analysis. Flowchart illustrating proteomic data handling and processing. Each enclosed box indicates specific data acquisition and processing approaches used to determine patient stratification and clustering as a function of the urine proteome. Box 1) shows data acquisition by mass spectrometry and initial data handling post proteome search by MaxQuant. High stringency identifications below 1% false discovery rate (FDR < 1%) are the initial list cut parameter followed by removal of single peptide identifications, thus increasing stringency. Box 2) shows data filtering and missing value imputations calculated in Metaboanalyst from the comma separated value (CSV) converted files from the initial proteome identification lists and removal of any patients deemed as CKD5 to diminish data skewing. Box 3) shows the re-clustering approach from initial groups of patients based on UPCR > 2 that would normally be grouped together based on clinical data (UPCR, GFR etc.) K-means clustering was used to identify patients with UPCR > 2 and determine if sub-groups existed based on urine proteome differences. Box 4) candidate selection by supervised multivariate analysis of patient clusters to determine proteins with the most influence and drivers of group separation. Box 5) shows pathways and ontological network analyses to define changes in patient groups.
Urine proteome and clustering analysis
Quantitative differences in the pediatric urine proteome were measured by isobaric reporter ion tagging followed by 2D-LC–MS/MS. Overall, a total of over 1,100 proteins were identified with 2 peptides or greater at less than 2.3% false discovery rate (see data processing scheme in Fig. 1). Following data filtering, log transformation and median normalization to remove highly variable values and data imputation, 226 proteins from the filtered and normalized dataset were used for univariate and multivariate analyses in Metaboanalyst. Unsupervised k-means clustering (Fig. 2A) identified two patient groups within the nephrotic and sub-nephrotic patient groups (UPCR > 2) (CL1-NP n = 71, CL2-NP = 17). Multivariate analysis of the clusters (Fig. 2B) highlighted differences in the proteome contents between these patient groups, red is CL1 and green is CL2. Figure 2C shows VIP (variable of importance in projection) scores for proteins that markedly influence the partial least-squares discriminant analysis (PLSDA) separation of the clusters and the relative importance of each protein in the stratification each protein. (Supplemental Table 1 sheets contain TMT normalized protein list (table S1A), k-means cluster list (table S1B), processed data (table S1C) and VIP scored proteins list table S1D).
Unsupervised clustering analyses of the nephrotic urine proteome. (A) K-means clustering of nephrotic/subnephrotic patient (UPCR > 2) urine proteome. Dark boundary line is the median intensity of each cluster, score plot shows the PCA (principle component analysis) of the normalized data. (B) Partial least squares discriminant analysis (PLSDA) plot shows separation of patients into two distinct clusters, CL1-NP (n = 71) and CL2-NP (n = 17). Inset histogram showing PLSDA model statistics: R2 = 0.95, Q2 = 0.73 for 5 components. (C) Plot of VIP (variable of importance in projection) for the most influential proteins in each patient cluster from PLSDA analyses.
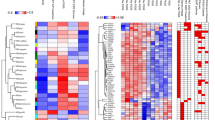
Hierarchical heatmapping (Fig. 3A) illustrates the distribution of proteins for each cluster with the top 40 most enriched proteins shown in Fig. 3B. These analyses indicate potential biological differences in the proteome contents in patients otherwise categorized as nephrotic (UPCR > 3 mg/mg) or “subnephrotic” (UPCR 2–3 mg/mg).
Hierarchical heatmapping of the urine proteome in cluster 1 and 2 patients from PLSDA analysis. (A) Overall heatmap of total proteome post-filtering and normalization shows robust clustering and enrichment of cluster 2 proteins. (B) Heatmap of the top 40 enriched proteins.
Tissue specific network analysis
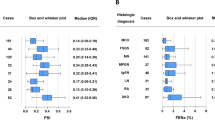
Determining tissue origin of urine proteins in nephrotic syndrome could be helpful in understanding the mechanism of the disease. To this end, we utilized tissue specific network analysis of the VIP proteins for mapping gene/protein networks4. Kidney and podocyte proteins were mapped to modules for protein functional predictions, Fig. 4 illustrates the interactions between specific groups of proteins and connectivity with other modules in the renal tubules and podocyte tissue sources. The tables show functional characteristics associated with protein groups within each module and the statistical value (q-val) for ontology enrichment. Modules enriched in the renal tubules (Fig. 4A) have predicted functions including neutrophil aggregation, complement activation, and JAK-STAT signaling. When networks are mapped to the podocyte (Fig. 4B), the functional characteristics map to endocytosis, immune responses, cytokine secretion and complement activation.
Enriched proteins (VIP > 0.9 in PLSDA) were analyzed for tissue network enrichment to determine biological processes and functions. (A) Glomerular networked proteins are separated into modules with lines between proteins and modules showing interconnectivity. Tables show functional characteristics mapped to each module and statistical evidence for ontology enrichment (q-val). (B) Podocyte networked proteins grouped into 6 modules, the table shows ontological enrichment for each module.
Protein interaction network analysis
To further investigate functional characteristics of the proteins eliciting group stratification, proteins with a VIP score greater than 0.9 were submitted for String-DB analysis to determine ontology and biological significance of the urine proteins in these two clusters (Fig. 5). Urine proteins in cluster 2 were elevated compared to cluster 1, enriched terms for this urine proteome included immune response, complement activation and leukocyte mediated immunity (Fig. 5B). Clustering analysis demonstrates the potential biological differences in the urine proteome contents of the two patient clusters, offering insights into patient stratification based on proteome profiles.
STRING analysis of protein interactions for proteins from cluster 2 nephrotic patient urine with VIP > 0.9. Biological processes by gene ontology were sorted by false discovery rate, included are observed genes from the total and strength of association calculations for ontology and false-discovery rate.
Discussion
The current studies tested the hypothesis that comparison of urine proteomes in children with MCD and FSGS could enable identication of urinary biomarkers able to identify molecular pathways regulating differences in disease activity and to stratify patient populations based on differences in the urine proteome by unbiased cluster analysis. Quantitative proteomics was used to identify and quantify urine proteins in children with MCD and FSGS in the CureGN Study associated with disease activity. Unsupervised clustering differentiated the urine proteome into two groups of patients with UPCR > 2. The patients showed enrichment in groups of proteins involved in immune responses, complement and neutrophil infiltration.
Our data suggest a major role for complement cascade in cluster 2 patients. Complement is a component of the innate immune system that is activated to promote a cascade of protein cleavages that allow for attachment to glycosylated proteins and further activation of downstream complement proteins to form an immune attack complex and initiate inflammation5. Overactivation of complement has been implicated in many kidney pathologies including C3 Glomerulopathy, Dense Deposit Disease, IgA Nephropathy and Membranous Nephropathy where C3 accumulation occurs in the mesangium and glomerular basement membrane and can lead to damage to these structures6,7,8. Other studies have shown that complement components are increased in the urine of FSGS patients9,10.
Urine proteins enriched in nephrotic patients were cross-referenced with the plasma proteome from the protein abundance database (PaxDb) to determine relative contribution of kidney cell injury and plasma protein leakage11. Proteins we identified in urine such as DNAAF1 , HMCN1, SLURP1 and RNASE2 are less prominent or absent in plasma and serum proteome studies and would suggest kidney cell injury is contributing to the large protein abundance changes in our dataset instead of only resulting from plasma protein leakage across the damaged glomerular basement membrane (see Supplemental Table 2). Additionally, comparison of the urine proteome dataset with single nuclear transcripts of the kidney indicated that ~ 43% of the identified urine proteins have cognate mRNA in kidney cells (see Supplemental Fig. 1)12. This suggests that a significant proportion of the urine proteome in our dataset could be derived from kidney cell sources.
Proteomic profiling of urine has been used to propose candidate biomarkers of chronic kidney diseases over the past few decades13,14. Early determination of disease onset may be achieved with a better understanding and characterization of the disease urinary proteome. Improvements in mass spectrometry and quantitative proteomics methods are beginning to provide tools for better interogation and analysis of biofluids towards the goal of identifying useful biomarkers15. The lack of biomarkers for pediatric kidney diseases presents the opportunity to apply cutting-edge proteomics methods and approaches towards bridging the gap in this knowledge on well-defined urine specimens. Recent studies have shown that circulating anti-nephrin antibodies are commonly elevated in pediatric MCD patients and in idiopathic nephrotic syndrome16. Our study did not account for anti-nephrin in the patient cohort and is a limitation of the findings that need to be considered in future proteomic analyses of nephrotic syndrome pediatric patients.
In our current work, we investigated the urine proteome of MCD and FSGS in nephrotic syndrome patients with proteinuria. Urine samples were obtained from the CureGN biorepository from a pediatric cohort of patients under age 18 at time of biopsy, urine was collected at initial and follow-up visits for analyses. We performed multivariate analysis of urine proteome quantitation to determine differences in patient populations. Changes in the urine proteome in response to disease progression and remission would be useful towards determining biomarkers of therapeutic efficacy or resistance. The urine proteome is accessible, noninvasive, and can be repeatedly probed to determine changes in specific proteins.
Cluster analysis has been useful to distinguish within groups of patients based on biomarker characteristics17,18. Clustering of the nephrotic/subnephrotic urine proteomes indicated the potential for stratifying patient groups based on differences in the quantitative profile of the urine proteome. Although we did not find correlation with clinical features such as eGFR, slope of eGFR or time to remission (see Supplemental Fig. 2), we do see major differences in the urine proteome within the nephrotic/subnephrotic groups when we used unsupervised k-means clustering to differentiate patient groups with the same clinical diagnosis. This suggests the potential for identifying meaningful differences in patient urine proteomes that could lead to a stratification approach for disease severity and potential for relapse.
Methods
Study approval, ethics statement, patients and urine collections
All research protocols and consent documents were approved by the Institutional Review Board of Nationwide Children’s Hospital as the coordinating center (approval numbers IRB07-00400, IRB12-00039 and IRB05-00544), as well as by each of the participating centers. See Tables 1 and 2 for patient demographics and Supplementary Methods for clinical data for patients in the proteomics clusters. All research was conducted in adherence with the Declaration of Helsinki. All enrolled participants provided either informed consent or assent obtained from a parent or legal guardian, as appropriate.
Pediatric patient selection criteria
CureGN (Cure Glomerulonephropathy Consortium) is a multi-center prospective observational cohort study of children and adults with biopsy-proven glomerular disease19. Eligible patients have a diagnostic kidney biopsy within 5 years of enrollment with MCD, FSGS, Membranous Nephropathy (MN) or IgA nephropathy. Patients are excluded if they have end stage kidney disease at the time of enrollment or any of the following before kidney biopsy: solid organ or bone marrow transplant, active HIV infection, hepatitis B or C infection, diabetes mellitus, lupus or active malignancy. Participants are followed prospectively 2–3 times per year with collection of demographics, comorbid health conditions, medications, local laboratory measurements and biospecimens at least yearly. For this study, CureGN participants were included if they were < 18 years at the time of biopsy and had one visit at the time of disease activity as determined by locally reported labs and a minimum of one additional follow-up visit. Bioposy confirmed cases of MCD have poorer outcomes and could be a interpreted as biased towards analysis of patients with worse proteinuria. Active disease based on locally reported labs was defined for MCD, FSGS as UPCR > 3 mg/mg. Informatics analyses on the urine proteome indicated that many patients in the sub 3 UPCR range had proteome signatures indicating nephrotic syndrome flare, leading us to further analyze these patient subgroups as “subnephrotic” and potentially distinguishable when considering the urine proteome content and quantification.
Urine sample collection and determination of creatinine and protein values
Spot urine samples (preserved with a protease inhibitor) were collected from 216 participants at two study visits, initial active disease visit and a follow-up study visit. For patients without a subsequent 12 months visit, the closest visit to 12 months was selected. For the proteomics analysis we included only the MCD and FSGS specimens, though the raw data includes IgAN and MN participant data. Urine creatinine and protein are reported as mg/dL. Urine creatinine was measured by the Jaffe rate method in the Synchron DXC system by reacting urine creatinine with picric acid and measuring the production of creatinine-picrate absorbance at 520 nm. Urine protein was measured in a separate assay by reacting protein with pyrogallol red and molybdate and determining absorbance at 600 nm for PYR-Mb-Protein complex. Both tests were performed using FDA approved calibrators, controls, reagents and instrumentation in a CLIA-certified hospital laboratory. Ratio of urine protein to urine creatinine (UPCR) was used to determine disease activity with > 3.0 mg/mg defined as nephrotic and 2–3.0 mg/mg defined as subnephrotic.
Protein concentration for proteomic analyses
Protein was concentrated in 5000 MWCO centrifugation filters (Sartorius). Urine was concentrated to ~ 200 μL and buffered in 50 mM HEPES supplemented with 1X HALT™ (Thermo) protease and phosphatase inhibitors, and 0.1 M EDTA. Samples were brought from 1 volume to 3 mL with deionized HPLC grade water. Final protein concentration was estimated by Bradford assay in triplicate. Equal amounts of urinary protein were then processed by S-trap proteolysis and TMT labeling as described in the supplemental methods and reference20. Mass spectrometry analysis is described in Supplemental Methods.
Bioinformatics analysis
Determining different cluster groups within NP patients was applied to distinguish patient groups based on the urine proteome. K-means clustering was used to determine if NP groups with clinical phenotype overlap could be shown to cluster as a function of the urine proteome. Metaboanalyst 5.0 was used to perform k-means clustering analyses. NP groups were clustered into two clusters and then regrouped based on proteome profile similarity for multivariate analyses. These groups were then analyzed by PLSDA (partial least squares discriminant analysis) to find the proteins most important in discriminating clusters. Proteins with a VIP score > 0.9 were then analyzed by String-DB to determine functional characteristics of the clusters and groups based on the PLSDA. Tissue network analysis was performed using the Human Base tool (https://hb.flatironinstitute.org/module) to indicate contributions of specific network modules of proteins within the renal tubules and podocyte4.
Data availability
All proteomics raw, processed files and tables will be made available through supplemental files associated with this manuscript or through partial submission of raw files to massive.ucsd.edu in accordance with HUPO (Human Proteome Organization) recommendations for publication and data sharing of proteomics experiments and data.
References
Beck, L. H. et al. M-type phospholipase A2 receptor as target antigen in idiopathic membranous nephropathy. N. Engl. J. Med. 361, 11–21 (2009).
Al-Rabadi, L. F. et al. Serine protease HTRA1 as a novel target antigen in primary membranous nephropathy. J. Am. Soc. Nephrol. 32, 1666–1681 (2021).
Tomas, N. M. et al. Thrombospondin type-1 domain-containing 7A in idiopathic membranous nephropathy. N. Engl. J. Med. 371, 2277–2287 (2014).
Greene, C. S. et al. Understanding multicellular function and disease with human tissue-specific networks. Nat. Genet. 47, 569–576 (2015).
Ricklin, D., Reis, E. S., Mastellos, D. C., Gros, P. & Lambris, J. D. Complement component C3 - The ‘Swiss Army Knife’ of innate immunity and host defense. Immunol. Rev. 274, 33–58 (2016).
Servais, A., Noël, L.-H., Frémeaux-Bacchi, V. & Lesavre, P. C3 glomerulopathy. Contrib. Nephrol. 181, 185–193 (2013).
Tortajada, A., Gutierrez, E., Pickering, M. C., Praga Terente, M. & Medjeral-Thomas, N. The role of complement in IgA nephropathy. Mol. Immunol. 114, 123–132 (2019).
Zhang, M.-F. et al. Complement activation products in the circulation and urine of primary membranous nephropathy. BMC Nephrol. 20, 313 (2019).
Huang, J. et al. Complement activation profile of patients with primary focal segmental glomerulosclerosis. PLoS ONE 15, e0234934 (2020).
Thurman, J. M. et al. Complement activation in patients with focal segmental glomerulosclerosis. PLoS ONE 10, e0136558 (2015).
Wang, M., Herrmann, C. J., Simonovic, M., Szklarczyk, D. & von Mering, C. Version 4.0 of PaxDb: Protein abundance data, integrated across model organisms, tissues, and cell-lines. Proteomics 15, 3163–3168 (2015).
Mariani, L. H. et al. Precision nephrology identified tumor necrosis factor activation variability in minimal change disease and focal segmental glomerulosclerosis. Kidney Int. S0085–2538(22), 00966–00968. https://doi.org/10.1016/j.kint.2022.10.023 (2022).
Greenberg, J. H. et al. Urine biomarkers of kidney tubule health, injury, and inflammation are associated with progression of CKD in children. J. Am. Soc. Nephrol. JASN 32, 2664–2677 (2021).
Pontillo, C. & Mischak, H. Urinary biomarkers to predict CKD: Is the future in multi-marker panels?. Nephrol. Dial. Transplant. Off. Publ. Eur. Dial. Transpl. Assoc. Eur. Ren. Assoc. 31, 1373–1375 (2016).
Cummins, T. D. et al. Advances in proteomic profiling of pediatric kidney diseases. Pediatr. Nephrol. Berl. Ger. https://doi.org/10.1007/s00467-022-05497-2 (2022).
Hengel, F. E. et al. Autoantibodies targeting nephrin in podocytopathies. N. Engl. J. Med. 391, 422–433 (2024).
Whelan, S. A. et al. Assessment of a 60-biomarker health surveillance panel (HSP) on whole blood from remote sampling devices by targeted LC/MRM-MS and discovery DIA-MS analysis. Anal. Chem. https://doi.org/10.1021/acs.analchem.3c01189 (2023).
Bhatraju, P. K. et al. Integrated analysis of blood and urine biomarkers to identify acute kidney injury subphenotypes and associations with long-term outcomes. Am. J. Kidney Dis. Off. J. Natl. Kidney Found. https://doi.org/10.1053/j.ajkd.2023.01.449 (2023).
Mariani, L. H. et al. CureGN study rationale, design, and methods: Establishing a large prospective observational study of glomerular disease. Am. J. Kidney Dis Off. J. Natl. Kidney Found. 73, 218–229 (2019).
Cummins, T. D. et al. Quantitative mass spectrometry normalization in urine biomarker analysis in nephrotic syndrome. Glomerular Dis. https://doi.org/10.1159/000522217 (2022).
Consortium Name: CureGN Consortium Contributors: CureGN CollaboratorsThe CureGN Consortium members listed below, from within the four Participating Clinical Center networks and Data Coordinating Center, are acknowledged by the authors as Collaborators. **CureGN Principal Investigators: Ali Gharavi, Larry Greenbaum, William Smoyer, Jon Klein, Ronald Falk, Lawrence Holzman, Bruce Robinson, Brenda Gillespie, Debbie Gipson, Matthias Kretzler, Laura Mariani. *CureGN Site Principal Investigators: Krzysztof Mucha, Gian Marco Ghiggeri, Diego Aviles, Tarak Srivastava, Sun-Young Ahn, Amrish Jain, Elif Erkan, Sherene Mason, Rasheed Gbadegesin, Liliana Gomez-Mendez, Yi Cai, Donald Weaver, Jerome Lane, Carl Cramer, Rajasree Sreedharan, Katherine Twombley, Mahmoud Kallash, Amira Al-Uzri, Myda Khalid, Craig Belsha, Joseph Alge, Scott Wenderfer, Tetyana Vasylyeva, Daniel Feig, Melisha Hannah, Carla Nester, Aftab Chishti, Wacharee Seeherunvong, Michelle Rheault, Craig Wong, Nisha Mathews, John Barcia, Sharon Bartosh, Tracy Hunley, Vikas Dharnidharka, Milos Budisavljevic, Louis-Philippe Laurin, Shikha Wadhwani, Brad Rovin, Dana Rizk, Amy Mottl, Neil Sanghani, Jason Kidd, Jeffrey Schelling, Kevin Lemley, Kevin Meyers, Raed Bou Matar, John O’Toole, John Sedor, Christine Sethna, John Sperati, Sharon Adler, Fernando Fervenza, Kimberly Reidy, Jeffrey Kopp, Richard Lafayette, Katherine Tuttle, Michelle Hladunewich, Rulan Parekh, Alessia Fornoni, Daniel Cattran, Patrick Gipson, Blaise Abromovitz, Michael Mortiz, J. Ashley Jefferson, Kamal Sambandam. #CureGN Lead Coordinators: Maddalena Marasa, Corinna Bowers, Mary Dreher, Samantha Sharpe, Caroline Poulton, Suzanne Vento.
Funding
This work was supported by NIDDK-NIH (DK110077 to JBK, WES), (DK091584 to JBK, MLM), internal funds from Nationwide Children’s Hospital (NCH) to WES; and clinical research funding from the University of Louisville Kidney Disease Program. Funding for the CureGN consortium is provided by U24DK100845 (formerly UM1DK100845), U01DK100846 (formerly UM1DK100846), U01DK100876 (formerly UM1DK100876), U01DK100866 (formerly UM1DK100866), and U01DK100867 (formerly UM1DK100867) from the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK). Patient recruitment is supported by NephCure Kidney International. Dates of funding for first phase of CureGN was 9/16/2013–5/31/2019.
Author information
Authors and Affiliations
Consortia
Contributions
Study Design and Conceptualization: JBK, MLM, WES, MK, SS Methodology and Investigation: TDC, DWW, MLM, MJR, SAJ, WJ, MH, CT Figure Preparation: TDC, DWW, JAD, PM, MH Funding: JBK, WES Manuscript Preparation and Drafting: TDC, MLM, WES, JBK, YK, JAD Manuscript Editing: TDC, WES, LHM, JBK, SM, WP, MLM.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cummins, T.D., Mariani, L.H., Wilkey, D.W. et al. Defining subgroups of pediatric nephrotic patients with urine proteomics. Sci Rep 15, 25064 (2025). https://doi.org/10.1038/s41598-025-05150-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05150-6