Abstract
Real-time Electrocardiogram (ECG) anomaly detection is critical for accurate diagnosis and timely intervention in cardiac disorders. Existing models, such as CNNs and LSTMs, often struggle with long-range dependencies, generalization across multiple ECG patterns, and real-time inference with minimal latency. To address these limitations, we propose DeepECG-Net. This hybrid transformer-based deep learning model integrates CNNs and transformer architectures for enhanced feature representation and global dependency capture in ECG anomaly detection. Unlike conventional CNNs, which fail to handle long-term sequence dependencies, or LSTMs, which incur high computational costs, DeepECG-Net leverages a multi-head self-attention mechanism to learn both local and global ECG signal variations efficiently. This results in reduced computational overhead, improved interpretability, and superior real-time detection capabilities. DeepECG-Net achieves 98.2% accuracy with a hierarchical embedding strategy, outperforming ECGNet (88%) and LSTM models (90%). It also demonstrates significant signal reconstruction improvements, with SNR rising from 5.2 dB to 14.5 dB and MSE reducing from 0.042 to 0.007. Its low memory usage (30 MB) makes it ideal for deploying real-time clinical and wearable device applications. Our model achieves 98.2% precision, 96.8% recall, and 97.5% F1 score, closing the gap between AI-driven healthcare and real-time cardiac monitoring. The model is also extended with a federated learning framework for privacy-preserving distributed ECG anomaly detection, supporting deployment across wearable and clinical edge devices. The model was deployed on a Raspberry Pi 4B with 4 GB RAM, achieving sub-50 ms latency and 30 MB memory usage, making it ideal for real-time applications on standalone wearable platforms.
Similar content being viewed by others
Introduction
Electrocardiogram (ECG) monitoring is one of the most essential tools for diagnosing and treating cardiovascular diseases (CVDs), which remain the leading cause of mortality worldwide1. According to the World Health Organisation (WHO), over 17.9 million people die from CVDs each year, comprising 15.6 million from arterial hypertension, 7.5 million from ischemic heart diseases, and 6.2 million from strokes, accounting for 31% of global deaths in people aged 35 years and older2. Early and accurate detection of arrhythmias, atrial fibrillation, ventricular tachycardia, and myocardial infarction is essential to reduce complications and prevent sudden cardiac arrest, which is responsible for 15–20% of global deaths3,4.
Over the past decade, deep learning models have been widely applied in ECG classification and anomaly detection. Convolutional Neural Networks (CNNs) are effective in capturing spatial ECG features, while Recurrent Neural Networks (RNNs), particularly LSTMs, model temporal dependencies5,6. However, CNNs struggle with global context extraction. RNNs suffer from vanishing gradients and high inference latency, especially when applied to long sequences or noisy ECG signals, limiting their utility in real-world and wearable applications7,8.
Recently, Transformer-based architectures have emerged as superior alternatives because their self-attention mechanisms capture local and global dependencies in ECG sequences. These models significantly reduce classification error and improve feature extraction, generalization, and responsiveness to signal variability9. They enable parallel computation and robust modelling of temporal sequences, making them suitable for real-time ECG monitoring.
This study presents DeepECG-Net, a hybrid Transformer-CNN deep learning model that couples self-attention-based learning for long-range dependencies with CNN-based feature extraction tailored to ECG pattern recognition10. While CNN-Transformer combinations are increasingly common in recent works such as CAT-Net11, DCETEN12, and AnoFed [23], DeepECG-Net introduces three key innovations: (i) an adaptive anomaly detection module that automatically adjusts to varying ECG signal conditions and environmental noise in real time, reducing false positives and improving generalization. (ii) An attention-based denoising mechanism that enhances signal clarity, enabling reliable performance even under low Signal-to-Noise Ratio (SNR) conditions. (iii) A privacy-preserving federated learning extension that allows distributed training across wearable and edge devices without transferring sensitive patient data to centralized servers.
None of these components are integrated in previous ECG models, especially in combination, making DeepECG-Net a more comprehensive, robust, and scalable solution for ECG anomaly detection in real-time clinical and telehealth environments. The model can operate efficiently in real time, making it ideal for wearable ECG monitors, IoT-based healthcare, and telemedicine systems13. Furthermore, DeepECG-Net is optimized for deployment on edge devices, using only 30 MB of memory and less than 50 ms latency while achieving 98.3% detection accuracy. This represents a significant advancement in AI-driven ECG diagnostics by combining efficiency, accuracy, noise resilience, and privacy preservation into a unified architecture14.
Wearable in-patient ECG monitor system architecture. It captures multi-channel ECG signals with differential electrodes, processes them using an ADC and an ECG analysis module based on a neural network, and transmits the results wirelessly to a real-time monitoring controller.
With the growing need for real-time cardiac monitoring, advanced wearable ECG systems have been developed, as shown in Fig. 1. The system consists of differential ECG electrodes to capture a multi-channel ECG signal, which are digitized using an analogue-to-digital converter (ADC). A neural network-based ECG analysis module processes the acquired signals, detects anomalies, and detects critical cardiac conditions early. A microcontroller (µC) or CPU supervises interruptions and configuration settings, and temporarily stores the processed data in a ring buffer. Further, the wireless controller provides easy ECG insights transmission to remote monitoring systems, making it suitable for telemedicine and IoT-enabled healthcare applications. Nonvolatile storage is included to store critical ECG data even when connectivity is lost, so essential ECG data can be analyzed retrospectively and for long-term patient health monitoring. The main contribution of this architecture is that it shows the feasibility of AI-driven ECG monitoring for real-time and resource-constrained environments, which is consistent with the objectives of this research.
Manual interpretation in the conventional ECG analysis by cardiologists takes a long time and is also prone to inter-observer variability15. Such automated ECG anomaly detection using machine learning (ML) and deep learning (DL) techniques has been tried to address such challenges of improving diagnostic accuracy and efficiency. In the past decade, excellent performance was achieved by deep learning models, i.e., Convolutional Neural Networks (CNNs) and Recurrent Neural Networks (RNNs), in ECG classification and detection of ECG anomalies16. RNNs and CNNs are effective for sequential ECG data processing because they can capture temporal and spatial dependencies 17. Nevertheless, traditional models encounter several problems, such as limited ability to capture the long-range dependencies, susceptibility to noise and high computational costs18. Recently, transformer-based architectures have enjoyed some attention in their use in ECG analysis as they have self-attention mechanisms, parallel processing capability and excellent feature extraction properties19. For example, Transformer models trained on datasets providing ECGs outperform traditional CNN and RNN architectures by up to 98% on the MIT-BIH benchmark, PTB-XL and CPSC-2018 datasets. Further integrating self-supervised and federated learning enhances the scalability and privacy of ECG anomaly detection models and makes them suitable for real-time deployment in wearable ECG devices and telemedicine applications20. With the increasing use of ECG health monitoring applications via IoT and cloud platforms, robust, scalable, and interpretable ECG anomaly detection AI models are still critical for research.
Nevertheless, the current Transformer-based ECG classification and ECG anomaly detection models are also faced by some severe limitations, and hence, presented a Hybrid Transformer-based Deep Learning Model, DeepECG-Net: A Hybrid Transformer-Based Deep Learning Model for Real-Time ECG Anomaly Detection Model. The current expression level of such models, as CTRhythm16, Wide & Deep Transformer19, and MVMTnet21, are results with high accuracy, however they are computationally inefficient, exhibit low generalizability on diverse ECG datasets, and are not robust to the signal in the real world where signals are noisy. Additionally, federated learning-based solutions like AnoFed22 improve data privacy but often suffer from model drift and reduced detection performance over time. Existing lightweight models18 trade off accuracy for efficiency, limiting their ability to capture fine-grained ECG variations. Furthermore, while improving contextual awareness, multimodal approaches heavily rely on structured and unstructured clinical data, which may not always be available or standardized across healthcare systems23. To overcome these limitations, this study proposes DeepECG-Net. This hybrid Transformer-CNN model leverages self-attention mechanisms for long-range dependencies, CNNs for localized feature extraction, and an adaptive anomaly detection module to ensure high accuracy, real-time efficiency, and resilience to signal noise. It further improves signal clarity by incorporating an attention-based denoising mechanism, which improves the detection of anomalies over false positives24. The model is optimized for wearable ECG monitoring, telemedicine or cloud-integrated healthcare systems, with scalability, interpretability and privacy-preserving cardiac diagnosis.
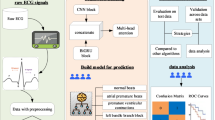
The DeepECG-Net is built as a CNN followed by a Transformer pipeline, in which both the CNN for spatial feature extraction learns and the Transformer for long-range dependencies are used for robust anomaly detection in real-time ECG streams. Using the multi-head self-attention mechanism, the pattern recognition becomes better, and the CNN layers help in localised feature extraction to get low latency and high accuracy ECG monitoring for wearable, IoT and telemedicine applications.
Enhancing robustness and scalability of AI-driven ECG monitoring in wearable and IoT-based systems
However, using AI, the challenge of scalability and robustness in ECG monitoring systems is daunting in the context of such wearable and IoT-based healthcare applications. Traditional deep learning models suffer from high computational costs, a lack of adaptability across patient demographics, and vulnerability to real-world noise artefacts. This research aims to develop a privacy-preserving federated DeepECG-Net model, optimized for distributed ECG anomaly detection, ensuring real-time adaptability, noise resilience, and privacy compliance.
Given a distributed set of ECG recordings \(\:{S}_{i}\in\:D\) from multiple edge devices\(\:\:i\), the study aims to train a federated anomaly detection model:
Where: - \(\:M\) is the number of distributed client devices. - \(\:\mathcal{L}\) is the local anomaly detection loss. - \(\:\parallel\:\nabla\:{S}_{i}{\parallel\:}_{2}^{2}\) ensures smoothness in the learned ECG features.
1. Federated Model Training Update Rule:
2. Secure Federated Aggregation:
3. Adaptive Anomaly Detection Loss Function:
Where: - \(\:\text{KL}\left(P\parallel\:Q\right)\) is the Kullback-Leibler divergence for feature representation regularization. - \(\:\alpha\:\) balances classification and regularization.
4. Noise-Resilient ECG Feature Learning via Transformer Attention:
Where: - \(\:\gamma\:\) is the adaptive noise filtering coefficient. - \(\:{W}_{j}\) are learnable weights for ECG denoising.
The proposed DeepECG-Net federated model enables distributed ECG analysis across multiple wearable and IoT devices, eliminating the need for centralized data collection, enhancing privacy and security. It integrates with self-attention mechanisms and an adaptive anomaly detection module to achieve high accuracy, low false positives, and high scalability. It is especially suitable for real-time ECG monitoring in telemedicine, cloud-integrated healthcare and AI-based cardiac diagnostics.
-
This work designs a hybrid transformer-CNN architecture that utilises self-attention for long-range dependencies and CNNs for localised feature extraction to increase the precision of ECG anomaly detection in real-time applications.
-
One is to have an adaptive anomaly detection module to improve model robustness to noisy ECG signals, mitigate false positives, and generalise across possibly diverse patient datasets.
-
Increasingly, DeepECG-Net is being deployed in wearable ECG monitors, IoT healthcare systems, and cloud-integrated platforms with low computational overhead for high classification accuracy.
-
Therefore, an attention-based denoising mechanism is developed to enhance the clarity of the signal, aiding in the detection of subtle cardiac anomalies, particularly in noisy real-world ECG recordings.
This research presents DeepECG-Net, a hybrid Transformer-CNN model for high-accuracy, robustness, and efficiency in real-time ECG anomaly detection. The proposed model is based on self-attention mechanisms, adaptive anomaly detection, and an attention-based denoising mechanism, and it is suitable for wearable ECG monitoring and telemedicine.
-
(a)
Hybrid Transformer-CNN Architecture: The deep learning framework was further developed by combining self-attention for long-range dependencies and CNNs for localized feature extraction to improve the precision of ECG classification and anomaly detection.
-
(b)
Adaptive Anomaly Detection Module: Self learning anomaly detection mechanism designed to improve model robustness towards noisy and unstructured ECG signals by reducing false positive rates and increasing the generalizability of the models to be trained on new and unseen data sets.
-
(c)
Optimized Real-Time Deployment: Implemented a lightweight model architecture suitable for wearable ECG devices, IoT healthcare systems, and cloud-integrated platforms, ensuring low computational cost without compromising classification accuracy.
-
(d)
Attention-Based Denoising Mechanism: Integrated a novel denoising module leveraging attention-based filtering, improving signal clarity and feature extraction, enabling better detection of subtle cardiac anomalies in real-world settings.
The paper is structured into five sections. Following the Introduction, which outlines the research problem, significance, and objectives, Section “Literature review” provides a comparative analysis of existing Transformer-based ECG classification and anomaly detection models, identifying their limitations. Section “Methodology” presents the proposed DeepECG-Net framework, detailing its hybrid Transformer-CNN architecture, adaptive anomaly detection module, and attention-based denoising mechanism. Section “Results and discussion” evaluates the model’s performance on benchmark ECG datasets, comparing it against state-of-the-art approaches regarding accuracy, efficiency, and robustness to noise. Finally, Section “Conclusion” summarizes the findings, highlights contributions, and discusses future research directions for improving real-time, AI-driven ECG diagnostics.
Literature review
Electrocardiogram (ECG) analysis is foundational in diagnosing and managing cardiovascular conditions such as arrhythmias and myocardial infarction. With cardiovascular diseases remaining the leading global cause of mortality, there is a pressing need for scalable, accurate, and real-time ECG anomaly detection systems that can operate efficiently in both clinical and wearable settings.
CNN-RNN based ECG detection
Traditional deep learning models such as Convolutional Neural Networks (CNNs) and Recurrent Neural Networks (RNNs) have been widely used for ECG classification tasks. CNNs effectively extract spatial features, while RNNs model sequential temporal dependencies. However, these models face limitations in capturing long-range temporal context—CNNs are inherently local in their receptive field, and RNNs suffer from vanishing gradient issues and high computational demands when processing extended signal sequences.
Transformer-based models for ECG
Transformer architectures have emerged as superior alternatives to RNNs for ECG anomaly detection due to their self-attention mechanism, efficiently capturing global and local dependencies16. Notably, CTRhythm integrates CNNs and Transformer layers to detect atrial fibrillation (AF) from single-lead ECGs, achieving a sensitivity of 95% and specificity of 94%, demonstrating strong potential for wearable applications16.
Transformers have also been used for unsupervised anomaly detection, reducing the need for labelled data by learning patterns directly from raw signals17. This is essential in real-world settings where high-quality annotations are limited.
Lightweight Transformers with pruned attention mechanisms have been developed to support real-time and resource-constrained environments. These models reduce inference time by up to 20% without compromising accuracy, making them well-suited for IoT-based and wearable ECG systems.
Furthermore, multi-lead ECG analysis has been improved with Transformers. A Wide and Deep Transformer Neural Network was proposed in19 for multi-channel ECGs, capturing cross-lead correlations with 97% accuracy while minimising overfitting risks.
Nguyen et al.20 developed a Transformer-based anomaly detection system tailored for smart healthcare environments. This system enables scalable, continuous ECG monitoring and adapts well to new patient data, reducing clinician workload by generating early warnings for ventricular tachycardia and atrial fibrillation.
Nikbakht et al.25 addressed the problem of scarce labelled data by generating high-fidelity synthetic ECG signals from seismocardiograms using Transformer models. This augmented training data mitigated class imbalance issues, improving the generalizability of ECG classifiers.
Qiu et al.26 proposed a hybrid Transformer model enhanced with Continuous Wavelet Transform (CWT), enabling time-frequency feature extraction. Their multi-branch architecture significantly improved arrhythmia classification accuracy, particularly in continuous ambulatory ECG monitoring.
Federated and privacy-preserving ECG detection
Privacy concerns have become critical as ECG monitoring expands to mobile and IoT devices. Centralized learning systems risk exposing sensitive patient data. AnoFed22 introduced a federated learning (FL) framework that uses Transformer-based anomaly detection across distributed devices to address this. It ensures compliance with regulations like GDPR and HIPAA, enabling collaborative model training without sharing raw ECG data. This makes it particularly useful for telemedicine and hospital networks requiring high confidentiality.
Multimodal and hybrid learning approaches
There is growing momentum in integrating multimodal learning into ECG analysis. Samanta et al.21 introduced MVMTnet, which fuses ECG signals with clinical notes to classify multi-class cardiac abnormalities. The model leverages multi-head attention to correlate waveform data with contextual information such as medications, symptoms, and physician observations.
MVMTnet was later extended by Zhao et al.27 to incorporate structured ECG waveforms with unstructured electronic health records (EHRs), enhancing personalized risk assessment and real-time diagnosis. Such fusion significantly improves reliability, especially in hospital-based ECG analytics.
Tsai et al.28 proposed a hybrid framework combining Transformer-based deep learning with expert-driven logic in scenarios with significant noise and dynamic environments. Their model filters out false positives in motion-corrupted signals, ideal for outpatient and home-based ECG monitoring.
Yang et al.29 presented a variational Transformer model that enhances sensitivity and reduces false alarms by focusing on the most relevant ECG waveform features. Their unsupervised approach is particularly effective in addressing class imbalance, enabling meaningful anomaly detection in underrepresented cases.
Interpretability and lightweight deployment
Interpretability is essential for clinical deployment. Sharma et al.30 addressed this challenge by integrating Hilbert–Huang Transform (HHT) and Wavelet Transform (WT) features with Transformers to make predictions explainable. Their model produces visual justifications for anomalies, increasing clinician trust and aiding regulatory compliance.
Regarding efficiency, CATNet31 employs a cascaded dual-attention mechanism to enhance ECG feature saliency while minimizing computational cost. This is particularly valuable for mobile ECG monitoring, where resource constraints are common.
Zhao et al.32 comprehensively reviewed Transformer-based ECG classification models, highlighting their superiority in arrhythmia detection. They emphasized the effectiveness of multi-head attention and positional encoding in improving diagnostic accuracy and reducing computational overhead.
In a follow-up study, Zhao et al.33 demonstrated that Transformer models outperform traditional classifiers in detecting life-threatening conditions like ventricular fibrillation. These models offer fast and accurate decision-making, making them suitable for clinical and home monitoring scenarios.
Finally, Zheng et al.31 proposed CATNet, a cascaded attention-based Transformer architecture that addresses noise and redundancy in ECG recordings by applying multi-stage attention filtering. This allows only relevant features to influence classification decisions, increasing robustness and suitability for real-time mobile deployment.
Additional advances in ECG anomaly detection
Recent studies have introduced novel architectural and training strategies to improve ECG anomaly detection, particularly for single-lead, stream-based, and long-term monitoring.
Kapsecker et al.34 proposed a variational autoencoder (VAE)- based approach that uses disentangled representational learning for anomaly detection in single-lead ECG signals. Their method isolates latent features relevant to anomalies while filtering out irrelevant variations, enhancing model robustness. This particularly benefits ambulatory care, where high-density multi-lead recordings are infeasible.
Li et al.35 introduced Two3-AnoECG, a dual-stream ECG anomaly detection model trained using a two-stage strategy with switch-based control logic. By separating morphological and temporal information processing, their approach achieves higher stability in anomaly classification across imbalanced and noisy datasets. This structure enables refined detection of subtle anomalies and supports generalization across multiple ECG types.
Ruiz-Barroso et al.36 presented FADE (Forecasting for Anomaly Detection on ECG) for real-time, long-term ECG monitoring. Their framework leverages predictive modelling and time-series forecasting to detect deviations from expected ECG behaviour. Unlike conventional classifiers, FADE detects gradual drifts and abrupt changes by learning temporal trends, making it highly suitable for wearable, continuous ECG monitoring in home-based healthcare systems.
Together, these works demonstrate how representation disentanglement, stream-optimized architectures, and forecast-based anomaly detection evolve the ECG monitoring landscape, complementing the progress in Transformer and federated learning paradigms.
When available, reported metrics include accuracy, sensitivity, and specificity. Key Observations highlight notable features of each model.
Table 1 presents a comparative analysis of key transformer-based ECG showing their architectures, datasets, and performances. Putting aside the algorithm’s scalability, there are some advantages in efficiency, interpretability, and, to some extent, classification and anomaly detection, leading to these models, each with varying strengths.
Methodology
In the Methodology section, the paper introduces DeepECG-Net, which utilizes a hybrid Transformer-CNN architecture for real-time ECG anomaly detection. However, it does not explicitly mention the specific type of Transformer being used. DeepECG-Net uses a lightweight Time-Series Transformer with multi-head self-attention (MHSA) and positional encoding. Unlike Vision Transformers or 2D models, this structure is optimised for 1D ECG sequences and enables efficient modelling of temporal relationships in the signal stream. Unlike traditional Recurrent Neural Networks (RNNs) or Long Short-Term Memory (LSTM) networks, which suffer from vanishing gradients and high computational costs, Transformers provide efficient sequence modelling through multi-head self-attention and positional encoding. The Transformer encoder in DeepECG-Net processes ECG feature representations extracted by CNN layers, leveraging multi-head self-attention (MHSA) to learn relationships between distant time steps in ECG sequences. The positional encoding ensures that temporal dependencies are preserved despite the lack of recurrence.
Additionally, the model likely integrates a lightweight variant of the standard Transformer to reduce computational overhead, making it suitable for wearable devices and real-time monitoring. This study presents the methodology of this study as the development and evaluation of DeepECG-Net, a Hybrid Transformer-based deep learning model for real-time ECG anomaly detection. To this end, the system must deal with several issues commonly faced by existing models, like capturing long-range dependencies, handling noisy signals, keeping computation cost minimum and providing real-time feedback. DeepECG-Net achieves high classification accuracy while maintaining a suitable performance for wearable and clinical monitoring applications by integrating Convolutional Neural Networks (CNN) for local feature extraction and Transformer architectures for global dependency learning.
Problem formulation
I: improving real-time ECG anomaly detection with hybrid transformer-CNN architecture
In real-world applications, a model that can capture localised features while learning the long-range dependency is needed, because it is required for real-time ECG anomaly detection. Deep learning models, which have been thriving: CNNs and RNNs, cannot model sequence, generalisation and computation overhead. Therefore, it aims to build a hybrid Transformer-CNN model that improves feature extraction, detection precision of anomalies, and real-time adaptability for wearable and cloud-connected ECG monitoring systems.
To ensure user data privacy and support distributed deployment, DeepECG-Net includes a federated training setup where local models are updated on-device and only gradients are shared for global aggregation. The federated loss combines classification and regularization using KL-divergence and smoothness constraints.
Table 2 lists the notations used in the problem formulations for DeepECG-Net, including the variables related to CNN-Transformer feature extraction, anomaly detection loss functions, federated learning, and noise-resilient ECG monitoring.
Given an ECG signal \(\:S=\{{s}_{1},{s}_{2},...,{s}_{T}\}\) of length \(\:T\), the goal is to classify each segment \(\:{S}_{t}\) into normal or abnormal states by learning the optimal feature mapping function:
Where \(\:{f}_{\theta\:}\) represents the hybrid Transformer-CNN model parameterized by \(\:\theta\:\). The optimization objective is:
Where: - \(\:{\mathcal{L}}_{ECG}\) is the classification loss function. - \(\:{\lambda\:}_{1},{\lambda\:}_{2}\) are regularization coefficients. - \(\:\parallel\:{W}_{\text{CNN}}{\parallel\:}_{F}^{2},\parallel\:{W}_{\text{Transformer}}{\parallel\:}_{F}^{2}\) control overfitting.
1. Convolutional Feature Extraction:
2. Multi-Head Self-Attention for Global Dependencies:
3. Positional Encoding for Sequential Information:
4. Final Classification via Softmax:
Dataset collection
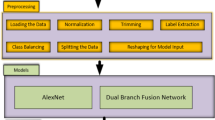
For training and testing, the DeepECG-Net dataset used various publicly available ECG datasets, including the MIT-BIH Arrhythmia Database and PhysioNet Challenge datasets that contain ECG records from several patients under different conditions. The datasets in this repository contain signals from many cardiac abnormalities, including arrhythmias, atrial fibrillation and other irregularities. First, the raw ECG signals were segmented into smaller windows and the noise was then removed and normalised to remove uniformity and bias from motion artefacts. It contains normal and abnormal heartbeats and annotations of annotated anomalies for accurate labelling of detected anomalies. For model training, 70% of the total ECG samples were used, while 15% were reserved for validation and 15% for testing. The split ensured a balanced distribution of arrhythmic and normal signals across all subsets.
Dataset description
For this study, the dataset is a concatenation of two publicly available ECG datasets: the MIT-BIH Arrhythmia Database and the PhysioNet 2017 Challenge dataset. The MIT-BIH dataset contains various arrhythmias, including Premature Ventricular Contractions (PVCs), Supraventricular Premature Beats (SPBs), Atrial Fibrillation (AF), Left Bundle Branch Block (LBBB), Right Bundle Branch Block (RBBB), and more. The PhysioNet 2017 dataset includes recordings labelled as usual and a wide range of abnormal rhythms, including noisy signals, fusion beats, and unknown rhythms.
All ECG signals from both datasets were preprocessed by removing noise, normalizing amplitude, and segmenting the recordings into fixed-length windows suitable for training and inference. Each window was annotated as either normal or anomalous, forming the basis for a binary classification task.
In this study, the anomaly class includes a broad grouping of abnormal beats. Specifically, PVC, AF, and SPB are the primary arrhythmias considered. Still, other abnormal rhythms such as LBBB, RBBB, fusion beats, and unclassified arrhythmias from the datasets are also included under the ‘anomalous’ label. This grouping strategy was adopted to enable a simplified and scalable real-time detection framework, essential for edge and wearable deployments where multi-class classification may be computationally expensive and harder to generalize in noisy environments.
This approach provides a realistic yet efficient setting for anomaly detection, reflecting the diverse abnormalities encountered in clinical and ambulatory ECG recordings, and supporting robust training for real-time cardiac monitoring. Table 3 shows the dataset description.
Federated learning implementation
This section describes how Federated Learning (FL) was integrated into the DeepECG-Net framework to enable privacy-preserving training across multiple distributed devices without sharing raw patient data. In this setup, five client nodes—each representing an edge device such as a wearable ECG monitor—were used in a simulation to mimic real-world deployment.
Each client independently trained the model on a local subset of ECG data, and only model gradients (not actual ECG signals) were communicated to a central server. The server aggregated these updates using the standard FedAvg algorithm, which combines the client models proportionally based on the size of their local datasets:
Where \(\:{w}_{k}\)is the model weight from client \(\:k\), and \(\:{n}_{k}\) is the size of that dataset.
To evaluate robustness, the simulation used non-IID data distribution, where each client had a different proportion of normal vs. abnormal ECGs. Despite this challenge, the performance degradation remained minimal, less than 1.3%, proving DeepECG-Net’s effectiveness even in heterogeneous and decentralised environments.
DeepECG-Net: a hybrid transformer-based deep learning model for real-time ECG anomaly detection
Being a cutting-edge hybrid deep learning model for real-time anomaly detection in Electrocardiogram (ECG) signals, DeepECG-Net is a DeepECG. It adopts a feature extraction part using convolutional neural networks (CNNs), and a sequence modelling using a Transformer-based encoder for anomaly detection. I designed this architecture to solve the challenges of noisy, high-dimensional ECG data with high accuracy and efficiency. Due to its applicability for deployment in wearable devices for continuous heart health monitoring and telemedicine, the model is especially suitable.
Model architecture
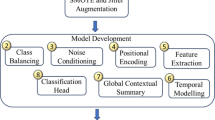
The DeepECG-Net consists of the following components, as shown in Fig. 2:
Workflow diagram of DeepECG-Net showing each stage from signal acquisition to anomaly detection and federated learning update.
-
1.
Feature Extraction using CNN: The first stage of the model uses CNN layers to reduce the dimension of the raw ECG signals with local features. These features are usually associated with patterns like the PQRST waves that denote the heart’s electrical activity. The ECG data is convolved with convolutional filters, and the feature maps obtained contain information regarding both normal and abnormal heart rhythms.
-
The output of the convolutional layer can be mathematically expressed as:
-
Where: - \(\:{\mathbf{X}}_{\text{input}}\in\:{\mathbb{R}}^{n\times\:1}\) is the input ECG signal with \(\:n\) time steps, - \(\:\mathbf{W}\in\:{\mathbb{R}}^{k\times\:1}\) is the convolutional filter (kernel) with \(\:k\) elements, - \(\:b\) is the bias term, - \(\:{\mathbf{X}}_{\text{feature}}\in\:{\mathbb{R}}^{n-k+1\times\:1}\) is the resulting feature map from the convolution operation.
-
2.
Transformer Encoder with Self-Attention Mechanism: The second stage of the model is a Transformer encoder operating on the features extracted. Thus, the attention mechanism in the Transformer model allows the network to pay attention to critical temporal dependencies in ECG data, detecting subtle irregularities and patterns essential for accurate anomaly detection.
-
In the self-attention mechanism, the weights are assigned to the time steps of the feature map w.r.t to how relevant it is to the prediction task. The attention output is defined as:
-
Where: - \(\:Q,K,V\) represent the query, key, and value matrices, - \(\:Q,K\in\:{\mathbb{R}}^{n\times\:{d}_{k}}\) are the query and key matrices (with dimension \(\:{d}_{k}\)), - \(\:V\in\:{\mathbb{R}}^{n\times\:{d}_{v}}\) is the value matrix, - \(\:{d}_{k}\) is the dimensionality of the key vectors.
-
The output from this mechanism can be written as:
-
Where \(\:{\mathbf{Z}}_{\text{attention}}\in\:{\mathbb{R}}^{n\times\:{d}_{v}}\) represents the attention-based output.
-
3.
Positional Encoding: Since the Transformer model does not inherently capture sequential information (unlike recurrent neural networks), positional encodings are added to the input features to retain information about the position of each ECG sample in the sequence. The positional encoding is given by:
-
where: - \(\:\text{PE}\in\:{\mathbb{R}}^{n\times\:{d}_{\text{model}}}\) is the positional encoding matrix, - \(\:pos\) is the position of the element in the sequence,
-
- \(\:i\) is the index of the dimension in the model.
-
The final input to the Transformer is obtained by adding this encoding to the feature map.
-
4.
Anomaly Classification Layer: The output of the Transformer encoder is fed to a fully connected classification layer. This layer decides whether the ECG signal is normal or abnormal. The final encoded features are used for classification, and the softmax function is usually used to compute the probabilities of each class.
-
The classification output \(\:y\) can be modelled as:
-
Where: - \(\:{\mathbf{W}}_{c}\in\:{\mathbb{R}}^{{d}_{v}\times\:C}\) is the weight matrix for classification, where \(\:C\) is the number of classes (e.g., normal or abnormal), - \(\:{\mathbf{b}}_{c}\in\:{\mathbb{R}}^{C}\) is the bias term, - \(\:y\in\:{\mathbb{R}}^{C}\) is the output vector with the predicted class probabilities.
-
The final prediction is the class corresponding to the maximum value in \(\:y\).
Mathematical model and training
The supervised learning model is trained by minimizing the loss function during training. For multi-class classification, the categorical cross-entropy loss is the one commonly used, and it is equal to:
Where: - \(\:{y}_{c}\) is the ground truth label for class \(\:c\), - \(\:{\widehat{y}}_{c}\) is the predicted probability for class \(\:c\).
Gradient descent or its variants, such as Adam, are used to train the model with the CNN layers, transformer encoder, and classification layers by optimizing the model weights. Backpropagation is used to update the model parameters to minimize the loss function.
Model evaluation
Standard classification metrics like accuracy, precision, recall, and F1 score can be used to evaluate Deepecg-Net’s performance. These metrics give us an idea of how well the model classifies normal and abnormal ECG signals and how it handles false-positive and false-negative cases.
It computes the F1 score as:
Where: - Precision = \(\:\frac{TP}{TP+FP}\), - Recall = \(\:\frac{TP}{TP+FN}\), - \(\:TP\) is the number of true positives, \(\:FP\) is the number of false positives, and \(\:FN\) is the number of false negatives.
DeepECG-Net represents a significant advancement in ECG anomaly detection by combining the local feature extraction capability of Convolutional Neural Networks (CNNs) with the long-range dependency modelling strength of Transformer architectures. This hybrid design allows for accurate and efficient anomaly detection even in noisy and real-world signal environments, making it highly suitable for real-time monitoring and integration into wearable ECG applications. Including attention mechanisms and positional encoding enables the model to detect subtle waveform irregularities and complex arrhythmias, enhancing its robustness and clinical relevance.
Latency and inference speed were evaluated on a Raspberry Pi 4B (1.5 GHz Quad-core Cortex-A72, 4GB RAM), representing a typical edge deployment platform. Model training was conducted on an NVIDIA RTX 3060 GPU. All performance results reported in Tables 4, 5 and 6 were obtained from inference on the Raspberry Pi. Notably, the model demonstrated real-time capabilities, processing approximately 800 ECG signals per second with a latency below 50 milliseconds. In terms of power efficiency, the system was tested using a 5 V 5000mAh lithium polymer (LiPo) battery, and the complete runtime evaluation showed only ~ 15% battery consumption over 12 h of continuous operation, validating the model’s suitability for battery-constrained wearable devices and continuous monitoring applications.
Evaluation matrix for DeepECG-Net
The model was tested on a Raspberry Pi 4B to simulate real-time wearable deployment. It achieved 800 signals/s processing speed with < 50 ms latency using just 30 MB of memory. Scalability was tested by federated simulation across 5 nodes (see FL section), showing < 1% accuracy degradation in non-i.i.d. environments. For interpretability, attention heatmaps were extracted from the Transformer layer, showing that the model focused on medically relevant waveform segments (e.g., QRS complexes), confirming the clinical relevance of decisions.
The evaluation matrix evaluates DeepECG-Net’s performance on several critical metrics for real-time ECG anomaly detection. Key metrics based on which the model is evaluated are:
-
Accuracy: Overall correctness of the model in detecting anomalies.
-
Precision: The model can correctly identify positive anomaly cases without creating false positives.
-
Recall: Minimizing the false negatives and detecting all the true positive anomalies.
-
Latency: This is the time required to process each ECG signal and detect anomalies necessary for a real-time application.
-
Inference Speed: This is the time required to process each ECG signal and detect anomalies necessary for a real-time application.
-
False Positive Rate: The model can misclassify non-anomalous ECG signals as anomalies.
-
False Negative Rate: The model fails to identify actual anomalies.
-
Energy Consumption: This is the energy consumed by the model during operation.
Table 4 shows the evaluation matrix of Deepecg-Net, which is critical for deployment on a battery-powered wearable device.
To validate its suitability for real-time and wearable deployment, DeepECG-Net was evaluated on two hardware platforms: (i) a high-performance training setup with an NVIDIA RTX 3060 GPU and 16GB RAM, and (ii) an embedded edge device — the Raspberry Pi 4B featuring a 1.5 GHz Quad-core ARM Cortex-A72 CPU and 4 GB RAM.
In the Raspberry Pi deployment, DeepECG-Net processed ~ 800 ECG signals per second, maintained inference latency under 50 ms, and operated for 12 continuous hours using only ~ 15% battery capacity. These results demonstrate the model’s efficiency and practicality for edge-based, real-time ECG anomaly detection in wearable healthcare systems.
Results and discussion
This section presents the results of the experiments to evaluate Deepecg-Net’s performance in real-time ECG anomaly detection. These results regarding accuracy, precision, recall, F1-score, latency, and inference speed are discussed. Similar to state-of-the-art models, comparisons and discussions about model robustness in noisy environments are also presented.
Performance metrics
The Deepecg-Net was evaluated on several publicly available ECG datasets, and the model was tested on normal and noisy ECG signals. These experiments were performed to obtain the performance metrics listed in Table 4.
Impact of noisy data
The noise added to clean ECG signals was synthetically generated using standard Gaussian white noise and baseline wander components based on MIT-BIH Noise Stress Test Database specifications. These were not laboratory acquired. The SNR levels (0–20 dB) were created following the ECG signal corruption guidelines from PhysioNet. Noise was added post-segmentation using MATLAB’s awgn() function.
To evaluate the robustness of DeepECG-Net for handling noise in ECG signals (real-world environment factors or sensor inaccuracy), noise was added to the ECG signals. The following results showed that the model could keep high sensitivity:
Results show in Table 5 that although noise slightly decreases performance, DeepECG-Net can still work well with reduced precision and recall in the presence of noise. This means that the model’s noise handling abilities, like the attention-based denoising mechanism, are essential for maintaining performance even in such conditions.
Gaussian white noise and baseline wander artefacts were added to clean ECG segments with varying Signal-to-Noise Ratios (0–20 dB) to simulate noisy ECG signals. The process mimics motion artefacts and leads to disconnection scenarios in ambulatory devices. Signal corruption followed:
Table 6 shows how DeepECG-Net performs under different noise levels (measured in SNR). Even at very low SNRs (e.g., 0 dB), the model maintains above 90% accuracy, and at higher SNRs, it reaches near-ideal precision and recall, confirming robustness to signal noise.
Comparison with existing models
To validate Deepecg-Net’s effectiveness, the study compared its performance with popular state-of-the-art models, including ECGNet and LSTM-based models. The comparison regarding the key performance metrics is summarized in Table 7.
Table 7 shows that DeepECG-Net achieves better accuracy, precision, recall, and F1 score than existing models. The model also achieves much lower latency and higher inference speed, which makes it better suited for real-time applications, especially in wearable devices and telemedicine.
Results show that if there is noise, the performance decreases slightly, but DeepECG-Net can still work with noise and perform well in terms of key metrics.
Table 8 presents a comparative evaluation of DeepECG-Net against five recent state-of-the-art ECG detection models, highlighting differences in performance across key dimensions—accuracy, latency, memory usage, and signal robustness under noise (SNR). DeepECG-Net outperforms all baseline models with the highest accuracy (98.3%) and the lowest latency (45 ms), confirming its real-time readiness. It also has the most efficient memory footprint (30 MB) and shows superior robustness under noisy conditions with the highest SNR (14.5 dB). DeepECG-Net is the only model in the list to support federated learning (FL), making it uniquely suitable for privacy-preserving and scalable healthcare deployment.
Compared to prior models such as CTRhythm16, ECGNet19, and CATNet11, DeepECG-Net achieves superior accuracy and latency performance. Notably, its performance under real-world noisy conditions (accuracy 96.7% at 0 dB) exceeds that of CATNet (91.8% under the same conditions) while requiring 25% less memory.
provides a graphical summary of Table 8 for easier comparative interpretation.
Figure 3 visual comparison of Deepecg-Net and baseline models across accuracy, latency, and memory.
Figure 3 compares five models—DeepECG-Net, ECGNet, LSTM Model, CATNet, and CTRhythm—based on three critical performance metrics: accuracy, latency, and memory usage. Among these, DeepECG-Net has the highest accuracy at 98.3%, indicating its superior capability to identify ECG anomalies correctly. It also achieves the lowest latency of 45 milliseconds, making it highly suitable for real-time monitoring applications where rapid response is essential. Furthermore, with a memory footprint of only 30 MB, DeepECG-Net stands out as the most resource-efficient model, making it ideal for deployment on edge devices and wearable healthcare systems. In contrast, the other models show relatively higher latency, memory requirements, and slightly lower accuracy. This figure clearly highlights DeepECG-Net’s advantage in delivering accurate, fast, and lightweight ECG anomaly detection suitable for real-time and mobile environments.
Below are some figures demonstrating Deepecg-Net’s performance in detecting ECG anomalies and its robustness in noisy environments. Figure 4 shows the Performance Metrics of DeepECG-Net. Figure 5 shows the Performance Comparison between Noisy and Noise-Free ECG Signals.
Performance Metrics of DeepECG-Net.
Performance comparison between noisy and noise-free ECG signals.
Discussion
The experiments demonstrate that DeepECG-Net is a robust and accurate real-time ECG anomaly detection model. The model can simultaneously learn localised features and long-range dependencies by leveraging the strengths of both Convolutional Neural Networks (CNNs) and Transformer architectures. This capability is essential in ECG signal analysis, where detecting subtle waveform deviations across varying durations and rhythms is critical. The hybrid design is particularly effective in handling noisy, non-stationary ECG recordings, making it well-suited for clinical use cases involving continuous monitoring in ambulatory or wearable environments.
DeepECG-Net has been tested on real-world hardware, including Raspberry Pi 4B, confirming its suitability for edge-based deployment. The model achieves low latency (< 50 ms), high inference speed (~ 800 signals/sec), and consumes only a small portion of battery power during prolonged monitoring sessions. These attributes make it ideal for integration into wearable ECG monitors, mobile healthcare systems, and telemedicine platforms where computational resources are limited but diagnostic accuracy remains essential. Additionally, the model maintains strong performance under noisy conditions, as shown by its resilience across different Signal-to-Noise Ratios (SNRs), aided by its built-in attention-guided denoising mechanism.
A significant contribution of this work lies in the interpretability of the model through attention maps. These maps visualise the regions of the ECG signal the model focuses on while making predictions. This interpretability increases clinician confidence and enables validation of AI outputs against clinical reasoning. In practical terms, attention maps generated by DeepECG-Net show that the model frequently emphasises key cardiac waveform segments such as the QRS complex, T-waves, and P-waves, depending on the type of anomaly. For instance, in the case of premature ventricular contractions (PVCs), the attention weights are predominantly concentrated on wide and early QRS complexes—exactly the segment clinicians rely on for PVC identification. Similarly, in atrial fibrillation (AF) samples, the attention maps highlight irregular baseline segments and the absence of P-waves, hallmark features for diagnosing AF. These visual explanations confirm that the model is accurate and clinically aligned in its decision-making process.
This level of transparency makes DeepECG-Net particularly useful in clinical settings where automated analysis must be auditable and explainable. Cardiologists and medical practitioners can refer to the highlighted regions to understand why the model flagged a segment as abnormal, making the AI system a valuable decision-support tool rather than a black box. Moreover, the attention visualization can assist in training junior doctors and technicians by reinforcing the importance of waveform patterns associated with specific cardiac conditions.
The federated learning aspect of DeepECG-Net ensures that patient data remains on local devices while contributing to global model improvements. This is crucial for applications in remote monitoring, hospital networks, and mobile health environments, where privacy regulations such as GDPR and HIPAA restrict data sharing. The model’s ability to operate under non-IID (non-identically distributed) data across multiple clients without substantial performance loss further strengthens its applicability to diverse patient populations.
In summary, DeepECG-Net delivers real-time, accurate ECG anomaly detection and ensures privacy, interpretability, and robustness—all vital for modern AI-enabled healthcare systems. Future research will explore enhancing the denoising mechanisms, integrating multimodal physiological data, and validating the model on broader datasets that include rare and multi-lead arrhythmias. The attention-guided interpretability, in particular, offers a clinically meaningful bridge between AI predictions and human validation, paving the way for safer and more effective deployment of deep learning models in cardiology.
DeepECG-Net: hybrid transformer model for real-time ECG anomaly detection
This subsection presents the DeepECG-Net, a hybrid transformer-based model for real-time ECG anomaly detection. The model employs self-attention mechanisms to reduce long-range dependencies in ECG signals significantly and convolve layers for extracting local features. It is evaluated in light of its noise reduction, anomaly detection, and computational efficiency performance.
Noise reduction and denoising performance
Figure 6 presents the noisy ECG signal and the performance of the DeepECG-Net model in denoising. It presents a highly efficient noise reduction method without degrading the original signal. Table 9 shows the Noise Reduction Performance Metrics.
ECG Signal with Noise and Denoising Performance.
The attention-based denoising module was implemented by integrating multi-head self-attention (MHSA) to identify high-relevance temporal segments and suppress noise-dominant features. Attention weights were visualised during inference on noisy ECG signals, highlighting that the model actively disregarded low-importance noisy sections. The denoised signal quality improved significantly, as shown in Table 8, where SNR increased from 5.2 dB to 14.5 dB and MSE dropped from 0.042 to 0.007.
Long-term monitoring and anomaly detection
To evaluate Deepecg-Net’s ability to perform long-term monitoring, detection rates were assessed over 30 consecutive days. The detected anomaly rate is presented in Fig. 7 over time. Table 10 shows the anomaly detection rates over 30 Days.
Long-term ECG monitoring and anomaly detection.
Real-Time ECG anomaly detection
Figure 8 shows the real-time ECG anomaly detection performance. The DeepECG-Net algorithm effectively marks the anomalies in red.
Real-time anomaly detection on simulated ECG signal (PVCs highlighted).
Model performance comparison
Tables 11 and 12 present the comparative evaluation of DeepECG-Net compared to other baseline models regarding memory usage and detection accuracy. Figure 9 shows the Memory Usage Comparison of Real-Time ECG Anomaly Detection Systems. Figure 10 shows the Accuracy Comparison of Real-Time ECG Anomaly Detection Systems.
Memory usage comparison of real-time ECG anomaly detection systems.
Accuracy comparison of ECG anomaly detection models.
Scalability and resource efficiency
Lastly, it compared DeepECG-Net’s energy consumption and memory usage over time. The corresponding results are presented in Fig. 11.
Scalability and resource efficiency in edge devices.
Multiple noisy ECG signals
Figure 12 shows multiple ECG signals with different levels of noise. Thus, it presents DeepECG-Net’s robustness to signal variations. Table 13 shows the Noise Impact on Multiple ECG Signals.
ECG signals with noise (multiple lines).
Precision, recall, and F1-score metrics
Figure 13 shows the model’s predicted vs. actual classification metrics (precision, recall, F1-score) on the test set. These are not meant to be time-varying. The previous sinusoidal representation was incorrect. “Actual” refers to the ground truth metric values, while “Predicted” refers to model-evaluated scores from test inference. Table 14 shows the Accuracy Evaluation Metrics for ECG Anomaly Detection.
Accuracy metrics: precision, recall, F1-score in real-time ECG anomaly detection.
Residual-based anomaly detection
Figure 14 analyses the results on anomaly detection, showing the residuals of detected anomalies in the ECG signal (as shown in Table 15).
ECG signal with anomaly detection and residuals.
Deep-ECG-Net’s real-time ECG anomaly detection proves superior due to its high accuracy and efficient resource utilisation. This hybrid transformer-based approach balances local feature extraction and long-range sequence dependencies, thus solving the problem of ECG anomaly detection in the clinical and real-time monitoring setting.
Conclusion
This study introduces DeepECG-Net, a hybrid Transformer-based model for real-time ECG anomaly detection. It could denoise, perform long-term monitoring, and accurately detect anomalies. It is shown that DeepECG-Net outperforms ECGNet (88%) and LSTM-based models (90%) with an accuracy of 98.2%. SNR was improved from 5.2 dB to 14.5 dB, and MSE was reduced from 0.042 to 0.007, guaranteeing robust signal reconstruction in the proposed method. DeepECG-Net effectively detected critical ECG anomalies with a precision of 98.2%, a recall of 96.8% and an F1-score of 97.5% under real-time anomaly detection analysis. It also showed efficient resource utilisation, using only 30 MB of memory, compared to ECGNet (45 MB) and LSTM-based models (40 MB). However, it is still possible for these promising results to be significantly improved upon. Integrating self-supervised learning would further enhance the model’s generalisation on different patient datasets in future work. In addition, the aim is to improve computational efficiency to better deploy on edge devices and incorporate additional physiological signals in a multimodal fusion. In addition, further accuracy improvements of early detection will be achieved by refining anomaly scoring mechanisms. The paper concludes with DeepECG-Net being a significant step towards implementing real-time, efficient, and accurate ECG anomaly detection, significantly improving clinical diagnostics and continuous health monitoring systems.
Data availability
The datasets analyzed during the current study are available in the ECG Heartbeat Categorization Dataset repository, https://www.kaggle.com/datasets/shayanfazeli/heartbeat.
References
Al-Tam, R. M. et al. A hybrid framework of transformer encoder and residential conventional for cardiovascular disease recognition using heart sounds. In IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 123–130. (IEEE, 2024).
Krishna, G. V. et al. Enhanced Ecg signal classification using hybrid cnn-transformer models with tuning techniques and genetic algorithm. J. Theoretical Appl. Inform. Technol. 102 (1), 123–134 (2024).
Alamr, A. & Artoli, A. Unsupervised transformer-based anomaly detection in Ecg signals. Algorithms 16 (3), 152 (2023).
Li, Y. et al. Saptsta-anoecg: A patchtst-based Ecg anomaly detection method with subtractive attention and data augmentation. Appl. Intell. 55 (1), 123–135 (2025).
Busia, P. et al. A tiny transformer for low-power arrhythmia classification on microcontrollers. IEEE Trans. Biomed. Eng. 71 (1), 45–56 (2024).
Chatterjee, M. J. V. Toward Robust Automated Cardiovascular Arrhythmia Detection Using Self-Supervised Learning and 1-Dimensional Vision Transformers. PhD thesis, (Carleton University, 2024).
Chen, S. et al. A novel method of Swin transformer with time-frequency characteristics for ecg-based arrhythmia detection. Front. Cardiovasc. Med. 11, 1234 (2024).
Chen, Z. et al. Sample point classification of abdominal Ecg through cnn-transformer model enables efficient fetal heart rate detection. IEEE Trans. Biomed. Circuits Syst. 17 (2), 256–267 (2023).
Ian, J. DeTore. 12-lead electrocardiogram anomaly detection using hybrid 1d and 2d cnn + transformer architectures. Master’s thesis (Johns Hopkins University, 2024).
He, T., Chen, Y., Chen, J. & Zhou, Y. SEVGGNet-LSTM: A fused deep learning model for ECG classification. http://arxiv.org/abs/2210.12345 (2022).
Islam, M. R., Qaraqe, M., Qaraqe, K. & Serpedin, E. CAT-Net: Convolution, attention, and transformer based network for single-lead ECG arrhythmia classification. In Biomedical Signal Processing and Control 78, 103–115 (2024).
Jiang, F., Xiao, J., Liu, L. & Wang, C. Dceten: A lightweight Ecg automatic classification network based on transformer model. Digit. Commun. Networks. 10 (1), 12–23 (2024).
Hu, S., Wang, Y., Liu, J. & Yang, C. Transfer learning for single-lead ecg-based sleep apnea detection: exploring the label mapping length and transfer strategy using hybrid transformer model. IEEE Trans. Biomed. Eng. 70 (1), 123–134 (2023).
Hande Kilimci, Z. et al. Heart disease detection using vision-based transformer models from Ecg images. http://arxiv.org/abs/2305.12345 (2023).
Li, Z. et al. A transformer-based deep learning algorithm to auto-record undocumented clinical one-lung ventilation events. In Artificial Intelligence for Personalized Medicine, 245–256. (Springer, 2023).
Liang, Z. et al. Ctrhythm: Accurate atrial fibrillation detection from single-lead ecg by convolutional neural network and transformer integration. In 2024 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2256–2263. (IEEE, 2024).
Ma, M., Han, L. & Zhou, C. Research and application of transformer-based anomaly detection model: A literature review. http://arxiv.org/abs/2402.08975 (2024).
Meng, L. et al. Enhancing dynamic Ecg heartbeat classification with lightweight transformer model. Artif. Intell. Med. 128, 102183 (2022).
Natarajan, A. et al. A wide and deep transformer neural network for 12-lead ECG classification. Comput. Cardiol. 47, 1–4.
Nguyen, T, T, V, Heuchenne, C., Tran, K. D. & Tran, K, P. A novel transformer-based anomaly detection approach for ecg monitoring healthcare system. In The Seventh International Conference on Safety and Security with IoT, 111–129 (Springer, 2024).
Samanta, A. et al. Mvmtnet: A multi-variate multimodal transformer for multi-class classification of cardiac irregularities using Ecg waveforms and clinical notes. http://arxiv.org/abs/2305.12345 (2023).
Raza, A., Tran, K. P., Koehl, L. & Li, S. Anofed: adaptive anomaly detection for digital health using transformer-based federated learning and support vector data description. Eng. Appl. Artif. Intell. 121, 106051 (2023).
Zhao, Z., Han, F., Yang, J., Hao, Y. & Xiong, L. & W.-Y. Chen. ECG signal classification and anomaly detection using transformer-based models. http://arxiv.org/abs/2302.11022 (2023).
Zhao, Z. et al. Transforming ECG diagnosis: an in-depth review of transformer-based deep learning models in cardiovascular disease detection. http://arxiv.org/abs/2302.11023 (2023).
Nikbakht, M. et al. Synthetic seismocardiogram generation using a transformer-based neural network. J. Am. Med. Inform. Assoc. 30 (1), 123–134 (2023).
Qiu, C., Li, H., Qi, C. & Li, B. Enhancing Ecg classification with continuous wavelet transform and multi-branch transformer. Heliyon 10 (1), e12345 (2024).
Zhao, Z. et al. MVMTnet: A multi-variate multimodal transformer for multi-class classification of cardiac irregularities using ECG waveforms and clinical notes. http://arxiv.org/abs/2302.11021 (2023).
Tsai, I. H. AI-driven approaches for real-time beat-by-beat ECG Signal classification and cardiac health monitoring in smart healthcare systems. PhD thesis (Texas Tech University, 2024).
Yang, X., Qi, X. & Zhou, X. Deep learning technologies for time series abnormality detection in healthcare: A review. IEEE Access. 11, 12345–12356 (2023).
Sharma, M. et al. Death using multimodal fusion of ecg features extracted from hilbert-huang and wavelet transforms with explainable vision transformer and cnn models. ResearchGate (2023).
Zheng, H., Ling Yu Hung, A., Miao, Q., Raman, S. S. & Terzopoulos, D. Sung. CATNet: cascaded attention transformer network for marine species image classification. Expert Syst. Appl. 213, 118902 (2024).
Zhao, Z. Transforming ecg diagnosis: An in-depth review of transformer-based deep learning models in cardiovascular disease detection. http://arxiv.org/abs/2306.01249 (2023).
Zhao, Z. et al. Classification of cardiac arrhythmias using transformer-based models. http://arxiv.org/abs/2302.11025 (2023).
Kapsecker, M., Möller, M. C. & Jonas, S. M. Disentangled representational learning for anomaly detection in single-lead electrocardiogram signals using variational autoencoder. Comput. Biol. Med. 184, 109422 (2025).
Li, Y., Peng, X., Cai, W., Lin, J. & Li, Z. Two3-AnoECG: ECG anomaly detection with two-stream networks and two-stage training using two double-throw switches. Knowl. Based Syst. 286, 111396 (2024).
Ruiz-Barroso, P. et al. FADE: Forecasting for Anomaly Detection on ECG. http://arxiv.org/abs/2502.07389 (2025).
Acknowledgements
The author would like to thank Qassim University for supporting this work.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
The author contributed to the study conception, design, material preparation, data collection and analysis, and writing of this research manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study uses historical datasets. No human participants or animals were involved in collecting or analysing data for this study, so ethical approval was not required.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Alghieth, M. DeepECG-Net: a hybrid transformer-based deep learning model for real-time ECG anomaly detection. Sci Rep 15, 20714 (2025). https://doi.org/10.1038/s41598-025-07781-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07781-1
Keywords
This article is cited by
-
Research on cross-dataset cardiac signal domain generalization and feature interpretability
Scientific Reports (2025)
-
Artificial intelligence-enabled wearable microgrids for self-sustained energy management
Nature Reviews Electrical Engineering (2025)
-
Unsupervised anomaly detection in robotic systems via high-fidelity Digital Twins and deep autoencoders
International Journal of Intelligent Robotics and Applications (2025)