Abstract
Tumor immune microenvironment plays a crucial role in determining the prognosis of lung adenocarcinoma (LUAD), with the interaction of immune cells within this microenvironment contributing to a poorer prognosis. We sought to investigate the causal relationship and underlying biological mechanisms between immune cell characteristics and LUAD to offer new insights for enhancing treatment strategies. We evaluated the association between immune cell characteristics and LUAD using Mendelian randomization (MR) analysis based on genome-wide association studies summary statistics. Sensitivity analysis was performed to verify the robustness of MR results. Immune cell infiltration analysis and machine learning on bulk RNA-sequencing data were conducted to further identify immune cells associated with LUAD. Prognostic genes of LUAD were identified using single-cell RNA-sequencing data and high dimensional weighted gene co-expression network analysis. MR analysis identified three immune cell characteristics associated with LUAD, including CCR2 on granulocyte, CD25 on CD45RA + CD4 not Treg, and plasmacytoid dendritic cell, and sensitivity analysis confirmed the robustness of the associations. Machine learning identified neutrophils, hematopoietic stem cells, B cells, and myeloid progenitor cells as key immune cell characteristics related to LUAD. Neutrophil was identified as the target cell of LUAD based on the MR analysis and machine learning. Subcequently, single-cell RNA-sequencing mapped the immune microenvironment of LUAD and identified down-regulated neutrophil. In addition, robust neutrophil-macrophage communication in LUAD was revealed using the CellChat package. Finally, nine neutrophil-related prognostic genes of LUAD were identified, three of which potentially regulated neutrophil-macrophage communication. This study showed a significant correlation between neutrophil and LUAD, particularly highlighting the neutrophil-macrophage communication. This finding may enhance our comprehension of LUAD immune microenvironment and hopefully promote the discovery of new immunotherapeutic targets for LUAD and the development of related drugs.
Similar content being viewed by others
Introduction
The GLOBOCAN 2022 study reveals that lung cancer ranks as the most prevalent form of cancer worldwide, accounting for 2.48 million new cases and resulting in 1.8 million fatalities1. Lung adenocarcinoma (LUAD) is the most common subtype of lung cancer. Regardless of tumor histology and driver mutation status, immunotherapy represented by immune checkpoint inhibitors (ICIs) plays a impressive anti-tumor effect, and has become an important means of LUAD treatment. However, immunotherapy has limitations, including low patient response rates to ICIs, limited benefits for only a few patients, and high rates of acquired resistance resulting in poor long-term outcomes2. The primary impediment to the effecacy of ICIs is the heterogeneity of tumor immune microenvironment. Research has demonstrated that the presence of immunosuppressive cells, a lack of tumor antigens, deficiencies in antigen-presenting cells, impaired T cell infiltration, and activation of immunosuppressive signaling pathways collectively contribute to the heterogeneity of the tumor immune microenvironment, which diminished the immune system’s capacity to identify and target tumor cells, thereby constraining the efficacy of ICIs and potentially leading to immune evasion3.
However, due to the current limited understanding of the LUAD immune microenvironment, strategies to improve immunotherapy efficacy remain unclear. Researches are focused on finding new immune checkpoints, or reliable ICIs efficacy prediction tools at molecular level. Programmed death ligand 1 (PD-L1) was recognized as a significant component of the immune microenvironment in LUAD. While initially considered a biomarker for predicting the efficacy of ICIs in patients with LUAD, the sensitivity of PD-L1 expression for this purpose was found to be modest4. Furthermore, prediction tools for assessing the efficacy of immunotherapy in LUAD, based on the expression of multiple genes in tumor tissue or serum, have not yet been validated through real-world studies5,6,7. While research focused on the molecular level has not led to breakthroughs, perspectives have focused on the interactions and connections within the tumor microenvironment to understand the complexity of tumors from macroscopic perspective8. A recent study revealed an abundance of mesenchymal macrophages, derived from blood monocytes, within the immune microenvironment of LUAD. These macrophages interact with tumor-associated fibroblasts and tumor cells, correlating with adverse prognosis in LUAD9. Meanwhile, prior research has revealed abnormal infiltration of various immunocytes in LUAD tissues and blood, encompassing T cells, B cells, macrophages, neutrophils, dendritic cells, natural killer cells, and monocytes10,11,12,13,14,15,16. These abnormally infiltrated immunocytes interact with LUAD cells, playing diverse roles in the initiation and progression of LUAD, which raised great concerns about the immune cells characteristics. Enhanced understanding of the role of immune cell characteristics in the pathogenesis would deepen etiology comprehension, advancing LUAD diagnosis and treatment strategies. However, the relationship between immune cell characteristics and LUAD is still poorly understood, including their causal roles and mechanisms in tumor development. A recent study found that neutrophil chemotactic function decreased in patients with lung cancer. Meanwhile, the degree of neutrophil infiltration in the microenvironment of LUAD was shown to be negatively correlated with survival of patients17. In addition, neutrophils may reshape the tumor microenvironment and promote immune escape by releasing neutrophil extracellular traps (NETs)18. This sparked interest in the role of neutrophils in LUAD’s immune microenvironment.
Herein, we utilized Mendelian Randomization (MR) analysis, an etiological causal inference technique using genetic variations as instrumental variables, to exclude confounders and achieve causal inference19. A recent MR study identified 15, 31, and 11 immune cell characteristics that causally associate with lung cancer, non-small cell lung cancer, and small cell lung cancer, yet their causal links to LUAD remain unexplained20. Another study revealed 36 immune cell characteristics associated with LUAD, but this research was based on the analysis of only one dataset, thus leaving the findings vulnerable to chance21. Therefore, in this study, we soundly used multiple datasets for MR analysis, focusing on the causal association between immune cell characteristics and LUAD. Through further application of machine learning to bulk RNA-sequencing, we identified the most pertinent immune cell characteristics related to LUAD. Additionally, at single-cell level, we captured the intricate communication patterns and signature gene modules of immune cells within the LUAD immune microenvironment, evaluating their predictive capabilities for LUAD prognosis. Ultimately, this study aims to provide an understanding of the relationship between immune cell characteristics and LUAD, thereby offering novel insights and strategies for investigating the biological mechanisms of LUAD, and hopefully enable more patients with LUAD to benefit from immunotherapy.
Materials and methods
Study design
This study was conducted in four key steps. Firstly, we employed MR analysis to explore the causal associations between immune cell characteristics and LUAD. Secondly, utilizing machine learning, we analyzed bulk RNA-sequencing data of LUAD to identify its characteristic immune cell profiles. Thirdly, we mapped the immune cell landscapes of LUAD using single-cell RNA-sequencing data. Finally, through high dimensional weighted gene co-expression network analysis (hdWGCNA) analysis of the neutrophil expression matrix in LUAD, we identified prognostic genes for LUAD. The MR study followed the STROBE-MR guidelines and adhered to the key principles of the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines22. The MR study adheres to three core assumptions: (1) Relevance: The instrumental variables (IVs) must demonstrate a strong correlation with the exposure factor, namely immune cell signatures. (2) Independence: The IVs should not be correlated with any known or unknown confounding factors that are associated with both immune cell signatures and LUAD. (3) Exclusivity: The IVs should influence LUAD solely through the mediation of circulating immune cells and not through any other pathways23. This study utilized publicly available GWAS summary statistics and RNA-sequencing data, without the need for collecting new data, thus rendering additional ethical approval unnecessary. The flowchart outlining the research process is presented in Fig. 1.
The flowchart of this study.
Data sources
To avoid publication bias from racial heterogeneity, we used genome-wide association studies (GWAS) summary statistics and RNA-sequencing data from European (white people) populations. The GWAS summary statistics of immune cell characteristics utilized in this study are publicly accessible from the GWAS catalog, with accession numbers ranging from GCST0001391 to GCST0002121. This dataset comprises 731 immune cell signatures, encompassing absolute cell counts (AC, n = 118), median fluorescence intensity reflecting surface antigen levels (MFI, n = 389), morphological parameters (MP, n = 32), and relative cell counts (RC, n = 192). The GWAS summary statistics for LUAD were sourced from IEU OpenGWAS database (https://gwas.mrcieu.ac.uk/). The ieu-a-984 dataset encompassed 11,245 LUAD cases and 54,619 controls with 10,345,176 SNPs, while the ieu-a-965 dataset comprised 3442 LUAD cases and 14,894 controls with 8,881,354 SNPs. Bulk RNA-seq data of 386 Caucasian LUAD patients and 61 healthy controls were obtained from The Cancer Genome Atlas (TCGA) database (https://portal.gdc.com/). Additionally, bulk RNA-seq data (GSE118370, GSE140797, GSE32863, and GSE31210) of LUAD cancer tissues and paracancer tissues were obtained from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). Single-cell RNA-seq data (GSE117570) of LUAD patients, comprising 3 cases of LUAD and 3 healthy controls, were retrieved from the GEO database.
MR statistical analysis
We rigorously screened IVs for the MR study: (1) Set P < 1.0 × 10–5 as the significance threshold for immune cell characteristics. (2) Calculated linkage disequilibrium among SNPs using 1000 Genomes European data, retaining r2 = 0.001 within 10,000 kb. (3) Excluded SNPs with minor allele frequency < 0.01. (4) Removed SNPs related to confounders like smoking. (5) Discarded SNPs with F-statistic < 10. (6) Eliminated palindromic SNPs. (7) Excluded immune cell traits with < 3 SNPs. Finally, 573 and 709 immune cell characteristics from ieu-a-965 and ieu-a-984 datasets were selected for MR analysis.
Subsequently, we conducted MR analyses to assess the causal effects of immune cell characteristics on LUAD. The inverse-variance weighted (IVW) method was adopted as the primary approach, while additional analyses such as MR-Egger, Simple mode, Weighted median, and Weighted mode were performed to evaluate the robustness of the IVW results. The Wald ratio method was employed to estimate the effect of each SNP. Since the outcome (occurrence of LUAD) is a binary categorical variable, the Odds Ratio (OR) was used to demonstrate the potential causal relationship. Cochrane’s Q test was utilized to assess the heterogeneity among SNPs. The MR-Egger-Intercept test was applied to evaluate horizontal pleiotropy. MR-PRESSO was employed to detect outliers. The leave-one-out analysis was conducted as a sensitivity analysis to assess whether significant results were driven by a single SNP.
All statistical analyses were conducted in the R environment (v4.3.1). The TwoSampleMR package (v0.5.8) served as the primary tool for conducting the MR study. The significance threshold was set at P < 0.05. To mitigate the risk of false positives, we applied FDR correction to the P values, and an Adjusted P < 0.06 was considered statistically significant.
Bulk RNA-sequencing data analysis
LUAD’s STAR-counts data and corresponding clinical information were downloaded from the TCGA database. Then, we extracted the data in TPM format and performed log2(TPM + 1) normalization processing. After preserving the samples with both bulk RNA-seq data and clinical information, a total of 447 samples (386 LUAD and 61 Normal) were subsequently analyzed. The normalize.quantiles function in the preprocessCore package was used for data normalization, and the removeBatchEffect function in the limma package was used to remove batch effects. Utilizing the Limma package (v3.40.2), we examined the differential expression of mRNAs. P values adjusted by Benjamini–Hochberg method were analyzed to correct for false positives, and mRNAs satisfying the criteria of “Adjusted P < 0.05 and log2(FC) > 1 or log2(FC) < -1” were defined as differentially expressed genes (DEGs). To analyze the biological functions of these DEGs, we employed the ClusterProfiler package. Furthermore, we leveraged the immunedeconv package (v2.1.3) and utilized EPIC, TIMER, CIBERSORT, QUANTISEQ, MCPCOUNTER, and XCELL to assess the differences in immune cell distributions between LUAD and normal samples. Four machine learning algorithms, including Random Forest (RF), Support Vector Machine (SVM), Extreme Gradient Boosting (XGboost), and LightGBM, were employed to evaluate and rank the distinguishing power of immune cell characteristics between LUAD patients and healthy controls. RF and SVM were completed using the Wekemo Bioincloud (https://www.bioincloud.tech)24, and the XGboost and LightGBM were conducted using the SPSSPRO platform (https://www.spsspro.com/). The sample proportion of training set and test set was 0.7 and 0.3 respectively. In addition, tenfold cross-checks were performed to improve the confidence of the results. Subsequently, we cross-validated the immune cell characteristics that exhibited high distinguishing ability across the four algorithms, thereby identifying target cells highly associated with the occurrence of LUAD.
Single‑cell RNA-sequencing data analysis
For the analysis of single-cell RNA-seq data, R software (v4.3.1) was employed. Specifically, the process entailed: (1) normalization of data using the “NormalizeData” and “ScaleData” functions from the Seurat package (v5.0), followed by dimensionality reduction via the “FindVariableFeatures” and “RunPCA” functions; (2) removal of batch effects through the application of the “RunHarmony” function; (3) clustering and visualization of cells using the “FindClusters” and “UMAP” functions; (4) identification of cell subpopulations with the aid of the CellMarker 2.0 (http://bio-bigdata.hrbmu.edu.cn/CellMarker/) and PanglaoDB (https://panglaodb.se/index.html) databases; (5) identifying the biological functions of marker genes in each cell type using the “ClusterGVis” and "org.Hs.eg.db" packages; (6) analysis of intercellular communication using the CellChat software package (v1.5.0); and (7) molecular docking, leveraging the CB-Dock2 platform (https://cadd.labshare.cn/cb-dock2/php/index.php), to assess the efficacy of ligand-receptor interactions between cells, with the structures of all proteins downloaded from the Protein Data Bank (https://www1.rcsb.org/).
Identifying neutrophil-associated genes in LUAD single‑cell RNA-seq
We utilized hdWGCNA package (v0.3.01) to identify neutrophil-related module genes associated with LUAD. Specifically, the process involved the following steps: (1) Constructing an expression matrix from Seurat object using the “SetupForWGCNA” function; (2) Establishing metacells for the neutrophil expression matrix using the “MetacellsByGroups” function, with the nearest neighbor cell count set at k = 25 and a maximum of 10 cells shared between two metacells; (3) Normalizing the metacell expression matrix using the “NormalizeMetacells” function; (4) Setting the neutrophil expression matrix as the expression matrix required for network analysis with the “SetDatExpr” function; (5) Simulating the similarity between the co-expression network and the scale-free topology under different soft-thresholding powers using the “TestSoftPowers” function and visualizing the results with the “PlotSoftPowers” function; (6) Constructing the co-expression network using the “ConstructNetwork” function; (7) Calculating the module eigengenes with the “ModuleEigengenes” function; (8) Analyzing the biological functions of the module genes through the Metascape platform (https://www.metascape.org); (9) Building and validating a neutrophil gene-related LUAD prognosis model to predict 1-, 3-, and 5-year overall survival rates through multi-factor Cox iterative regression analysis using data in the TCGA cohort and GSE31210 cohort. The prognosis model was conducted using the survival package in R software, and the p-value and HR with 95% CI used in the Kaplan–Meier curve were generated by log-rank tests and univariate cox proportional hazards regression.
Results
Mendelian randomization revealed that granulocyte, CD4 T cell, and plasmacytoid dendritic cell were causally associated with LUAD
In the ieu-a-984 dataset, the number of IVs for each immune cell characteristic ranged from 3 to 24, with F-statistic values spanning from 19.58 to 442.37, indicating no weak instrument variable bias (Supplementary file 1: Table s1). Similarly, for the ieu-a-965 dataset, the IV counts for each immune cell feature varied between 3 and 430, accompanied by F-statistic values ranging from 19.56 to 486.21, suggesting the absence of weak instrument variable bias (Supplementary file 1: Table s2).
In the ieu-a-984 dataset, utilizing the results of the IWV analysis, we initially identified 33 immune cell characteristics that have a potential impact on the occurrence of LUAD. After applying FDR correction, we confirmed that 29 of these immune cell characteristics significantly influence the development of LUAD (Fig. 2A and Supplementary file 1: Table s3). In the ieu-a-965 dataset, utilizing the results of the IWV analysis, we initially identified 54 immune cell characteristics that have a potential impact on the occurrence of LUAD. After applying FDR correction, we confirmed that 8 of these immune cell characteristics significantly influence the development of LUAD (Fig. 2B and Supplementary file 1: Table s4).
Results of MR analysis. (A) Result in ieu-a-984 dataset; (B) Result in ieu-a-965 dataset.
We identified overlapping immune cell characteristics in ieu-a-984 and ieu-a-965: CCR2 on granulocyte, CD25 on CD45RA + CD4 not Treg, and Plasmacytoid DC. Furthermore, we conducted an inverse MR analysis to explore the causal relationship between LUAD and these cells, and the results revealed no significant relationship between them (Supplementary file 1: Figure s1). Consequently, we designate these three immune cell characteristics as potential causal factors associated with LUAD. To further establish the reliability of our MR results, we conducted a series of sensitivity analyses (Supplementary file 1: Table s5-7). The Cochrane’s Q test revealed no heterogeneity among the SNPs associated with these immune cell characteristics (P > 0.05). The MR-Egger-Intercept test indicated no horizontal pleiotropy among the SNPs (P > 0.05). Additionally, the MR-PRESSO analysis showed no outliers among the SNPs (P > 0.05). Finally, the leave-one-out test suggested that the observed causal effects were not driven by a single SNP (P > 0.05).
Bulk RNA-sequencing data analysis identified neutrophil as target cell of LUAD
The baseline characteristics of included data were shown in Supplementary file 1: Table s8. Utilizing bulk RNA-seq, we identified 909 upregulated DEGs and 1621 downregulated DEGs in LUAD (Fig. 3A). These DEGs were primarily enriched in pathways such as Cell Adhesion Molecules and Hematopoietic Cell Lineage, biological processes including Extracellular Matrix Organization and Extracellular Structure Organization, cellular components like Collagen-containing Extracellular Matrix and Collagen Trimer, and molecular functions such as Extracellular Matrix Structural Constituent and Glycosaminoglycan Binding (Fig. 3B). Based on the MR analysis, we focused on the distribution differences of granulocytes, dendritic cells, and CD4+ cells between LUAD cases and healthy controls. Overall, neutrophils and dendritic cells exhibited significantly decreased infiltration in LUAD, while CD4+ cells were significantly upregulated (Fig. 3C and Supplementary file 1: Table s9). Subsequently, we employed four machine learning algorithms to identify the 20 most discriminative immune cell characteristics distinguishing LUAD cases from healthy controls (Fig. 3D and Supplementary file 1: Table s10). Through cross-validation, neutrophil was identified as the target cell (Fig. 3E). Moreover, data from the GEO database alse showed that neutrophil was down-regulated in LUAD (Fig. 3F).
Results of bulk RNA-seq analysis. (A) DEGs of LUAD; (B) Functional enrichment analysis of DEGs; (C) Difference of immune cell infiltration in LUAD microenvironment; (D) Machine learning identifies key immune cells of LUAD; (E) Neutrophil was identified as the target cell of LUAD through cross-validation; (F) Neutrophil was validated to decrease in LUAD tissue based on data from GEO database.
Single‑cell RNA-sequencing data analysis revealed robust neutrophil-macrophage communication in the immune microenvironment of LUAD
Following stringent quality control metrics, the data underwent preprocessing steps such as normalization and batch correction. Utilizing UMAP technology based on the top 30 principal components, we clustered the high-dimensional single-cell RNA-seq, resulting in 11 distinct cell clusters that exhibited varying gene expression patterns at a resolution of 0.5 (Supplementary file 1: Table s11). These clusters were annotated using CellMarker 2.0 and PanglaoDB, identifying them as Epithelial cell (Markers: KRT7, CAV1, SFTPA1), Immune cell (Markers: IL7R, CD96, LTB, S100A2, PLA2G7, CCL2, S100A8, SPP1, GZMA, PRF1, KLRG1, FABP4, MME, GPD1, CLEC10A, FCER1A), Secretory cell (Markers: SCGB3A2, PIGR), SLC16A7 + cell (Marker: APOBEC3A), Fibroblast-like cell (Marker: PTGDS), Lonocyte cell (Marker: THBS1), and Ciliated cell (Markers: RSPH1, TPPP3, PIFO, MORN2, CAPS) (Fig. 4A). Notably, we observed significant differences in the distribution of Immune cells between LUAD and healthy controls (Fig. 4A). Subsequently, the immune cells were classified into seven cell types (B cell, dendritic cell, macrophage, neutrophil, NK cell, T cell, and Unknown), and the respective marker genes of each cell were detailed in Fig. 4B. Strikingly, the neutrophil population was significantly depleted in LUAD compared to healthy controls (Fig. 4C).
Results of single-cell RNA-seq analysis. (A) Differences in the distribution of different cell types between LUAD and normal; (B) Heatmap showing the top 5 DEGs in each immune cell type; (C) The difference of immune cells distribution between LUAD and normal people; (D) The number of immune cell communication in LUAD and normal; (E) The strength of immune cell communication in LUAD and normal; (F) Communication between neutrophils and other immune cells in LUAD and normal; (G) Communication between neutrophils and macrophages was significantly enhanced in LUAD; (H) Molecular docking revealed good binding energy between ligands on neutrophils and receptors on macrophages.
To investigate the intercellular interactions among immune cells in LUAD microenvironment, we employed the CellChat package to perform a cell-to-cell communication analysis. Macrophages, neutrophils, dendritic cells, and Unknown cells exhibited higher communication frequencies with other cell types. In contrast, in healthy controls, macrophages, neutrophils, dendritic cells, and NK cells demonstrated higher communication frequencies (Fig. 4D). We observed a greater communication intensity between neutrophils and other cells in LUAD (Fig. 4E). Subsequently, we delved into the communication between neutrophils and the remaining immune cells. Compared to healthy controls, neutrophils in the LUAD group exhibited stronger communication with macrophages (Fig. 4F). This difference primarily stemmed from the varied binding of neutrophil ligands (HLA-DPA1, HLA-DPB1, HLA-DQA1, HLA-DQB1, HLA-DRA, HLA-DRB1) to macrophage receptors (CD4) (Fig. 4G, Supplementary file 1: Table s12, and Supplementary file 2: Figure s2). Molecular docking analysis revealed favorable ligand-receptor binding energies (Fig. 4H and Supplementary file 1: Table s13). In summary, while there is a reduced neutrophil population in the LUAD microenvironment, these neutrophils maintain a robust communication relationship with macrophages.
Identifying neutrophil-associated genes in LUAD single‑cell RNA-sequencing data
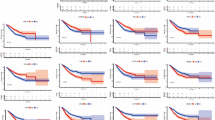
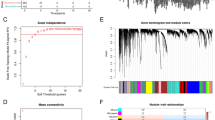
A comprehensive co-expression network was constructed using the hdWGCNA package to identify genes that exhibit a strong association with neutrophils in LUAD. With a soft threshold β set at 20, chosen for its biological relevance and adherence to the scale-free network criteria (Fig. 5A and Supplementary file 1: Table s14), we identified five distinct gene modules related to neutrophils (Fig. 5B). A UMAP projection offered further insights into the co-expression patterns of genes within these modules (Fig. 5C). Among them, the brown module (Neutrophil-M3) stood out as having the most profound correlation with neutrophils (Fig. 5D,E). Enrichment analysis of this module revealed that the core genes were predominantly involved in functions such as regulated exocytosis, secretory granule formation, secretory vesicle trafficking, and neutrophil degranulation (Fig. 5F). Through rigorous multivariate Cox regression analysis, we pinpointed nine crucial prognostic genes: S100A11, CFL1, FNDC3B, GAPDH, TPM4, VCAN, S100A8, CTSL, and SH3BGRL3 (Supplementary file 1: Table s15). Utilizing these prognostic genes, we formulated a neutrophil-gene-based prognostic model for LUAD: Riskscore = (0.4635)*S100A11 + (0.6175)*CFL1 + (− 0.2691)*FNDC3B + (0.3799)*GAPDH + (− 0.2783)*TPM4 + (0.2584)*VCAN + (− 0.1068)*S100A8 + (0.2435)*CTSL + (− 0.3201)*SH3BGRL3. Patients were divided into high and low groups based on the median riskscore. Notably, patients with a high risk score had a median survival time of 2.7 years, whereas those with a low risk score had a median survival time of 4.9 years. The model demonstrated robust performance, achieving AUCs of 0.721, 0.703, and 0.71 for predicting 1-, 3-, and 5-year overall survival rates, respectively (Fig. 5G). We validated the prognostic riskscore in an external cohort and found that there was a statistically significant difference in the overall survival rate between the high-risk group and the low-risk group in the GSE31210 cohort (HR 2.95, P = 5.9e-3). Moreover, this riskscore can predict the 5-year overall survival rate of relatively accurately (AUC = 0.69) (Fig. 5H).
Identifying neutrophil-associated genes using hdWGCNA. (A) Set the soft threshold β to 20 based on biological relevance and scale-free network criteria; (B) The neutrophil-related genes were divided into five modules; (C) Co-expression network of neutrophil-related genes; (D) Neutrophil-M3 was the gene module most strongly associated with neutrophil; (E) Core genes of the Neutrophil-M3 module; (F) Functional enrichment of the core genes of Neutrophil-M3 module; (G) Multivariate Cox regression analysis identified neutrophil-related prognostic genes of LUAD in the TCGA cohort; (H) External validation of the neutrophil-related prognostic genes in the GSE 31210 cohort.
Discussion
This study used MR analysis to examine the causal links between 731 immune cell traits and LUAD in two cohorts, finding that CCR2 on granulocytes, CD25 on CD45RA + CD4 not Treg, and Plasmacytoid DC were linked to higher LUAD risk. Machine learning analysis of LUAD bulk RNA-seq identified neutrophils, hematopoietic stem cells, B cells, and myeloid progenitor cells as crucial for LUAD detection. Single-cell RNA-seq analysis revealed a decrease in neutrophils within the LUAD immune microenvironment and strong interactions with macrophages. Nine neutrophil-related genes (S100A11, CFL1, FNDC3B, GAPDH, TPM4, VCAN, S100A8, CTSL, SH3BGRL3) were significantly linked to LUAD prognosis.
Neutrophils play a crucial role in LUAD development. Neutrophils are among the most critical immune cells within the human body. Upon the invasion of foreign pathogens or the onset of endogenous inflammation, neutrophils are the initial immune cells to migrate from peripheral circulation to the site of pathology. They combat pathogens through mechanisms such as direct phagocytosis, the release of reactive oxygen species (ROS), and a process known as “suicidal” NETosis. During NETosis, neutrophils expel their nuclear DNA, which combines with histones and antimicrobial enzymes to form a fibrous network structure known as neutrophil extracellular traps (NETs). These NETs serve to capture and neutralize pathogens25. In addition to killing pathogens, NETs also promote tumor metastasis by regulating the cell cycle and shielding tumor cells from immune attacks26,27,28,29. Clinical studies indicated that NETs-related gene expression can predict survival and distant metastasis in patients with LUAD but not squamous cell carcinoma30,31. NETs also serve as independent predictors of LUAD immunotherapy efficacy, with high expression linked to shorter progression-free survival and higher chances of disease progression and immune-related side effects32,33. Animal studies revealed that NETs suppress CD4 and CD8 T cells, while promoting CD4 T cells to become Tregs, aiding immune escape34,35. NETs inhibitors could boost the tumor-reducing effect of ICIs on CD8 T cells, increase CD8 T cells in tumors, restore local immunity, and prevent metastasis36,37,38,39. In addition, chronic lung inflammation could cause NETs infiltration, reactivating dormant lung metastatic cancer cells post-ICI therapy, leading to invasive metastasis40. Cell experiments revealed that NETs can induce M2 macrophage polarization, enhancing LUAD A549 cell proliferation and invasion, while inhibiting the immune response of monocytes, CD8 T cells, and NK cells34,41. NETs also express PD-L1, leading to T cell dysfunction and tumor immune escape40. Thus, neutrophils, through NETs, are crucial in LUAD development.
At the single-cell level, we observed strong communication between neutrophils and macrophages in LUAD. Research indicated that these innate immune cells inhibit the cytotoxic effects of other immune cells and interact with tumor cells to support their growth, metastasis, and drug resistance42. The interaction between neutrophils and macrophages has been shown in inflammatory environments. In the early stages of inflammation, macrophages release cytokines to recruit neutrophils, which can then activate macrophages. In the later stage, macrophages engulf apoptotic neutrophils to lessen inflammation, with neutrophil apoptosis promoting anti-inflammatory macrophage phenotypes43,44. Additionally, neutrophils influence macrophage polarization and proliferation via colony-stimulating factor 145. Recent investigations have demonstrated that neutrophil-macrophage interactions also occur within tumor microenvironment. Macrophages may induce neutrophils to release NETs through the secretion of cytokines such as IL-1β, IL-6, IL-18, and TNF-α, thereby facilitating the migration and invasion of LUAD cells41. Conversely, NETs can modulate macrophage polarization via TREM1 induction, thereby contributing to the progression of gastric cancer46. Additionally, NETs enhance macrophage recruitment in vivo, which subsequently inhibits the growth of bladder cancer tumors47. In the immune microenvironment of LUAD, NETs enhance the invasion and migration of A549 cells through an inflammation maintained by macrophages41. Neutrophil elastase from NETs promotes M2 macrophage polarization by down-regulating PTEN, aiding LUAD tumor growth and A549 cell proliferation, migration, and invasion48. The above evidence suggests neutrophils and macrophages interact via NETs to advance LUAD.
At the genetic level, we identified nine neutrophil-associated LUAD prognostic genes (S100A11, CFL1, FNDC3B, GAPDH, TPM4, VCAN, S100A8, CTSL, and SH3BGRL3). S100A8, S100A11 and GAPDH have been shown to regulate the function of neutrophils and macrophages. S100A8, a pro-inflammatory protein from neutrophils, activates fibroblasts in tumor cells by accumulating oxidized lipids, leading to LUAD recurrence49. It also modifies PD-L1 histones in macrophages, reducing CTL anti-tumor activity and enabling tumor immune escape50. Additionally, macrophages increase S100A8 expression in LUAD LLC cells, promoting liver metastasis51. S100A11 has been shown to facilitate the secretion of inflammatory mediators, including IL-6 and TNF, by neutrophils, thereby amplifying the inflammatory response52. Furthermore, S100A11 present in extracellular vesicles derived from osteosarcoma cells has the capacity to activate macrophages, leading to the secretion of CXCL2 and subsequent induction of lung metastasis53. Notably, the expression of S100A11 is modulated by NETs52. Recent studies indicate that inflammatory stimulation reduces GAPDH activity in neutrophils, promoting NET formation54. In tumors, GAPDH expression inversely correlates with neutrophil infiltration55. Reduced GAPDH activity in macrophages may drive anti-inflammatory aerobic glycolysis56. GAPDH oxidation might activate inflammasomes and IL-1β release, leading to pro-inflammatory macrophage metabolic reprogramming57. Unfortunately, no study has found the effect of these genes on neutrophil-macrophage communication, and further research is needed in the future.
This study presents several notable strengths. Initially, a thorough multi-dimensional analysis was performed utilizing a vast dataset to investigate the causal relationship between immune cell characteristics and LUAD. This analysis also involved mapping the overall immune cell landscape of LUAD and identifying crucial connections in the development of LUAD at single cell level. Significant, emphasis was placed on the contribution of neutrophil and associated genes to the onset of LUAD, offering a fresh viewpoint for deciphering the immune landscape of LUAD. However, there are several limitations. The GWAS summary statistics utilized were derived exclusively from European populations, thereby limiting the generalizability of our findings to other ethnic groups, and this study relied solely on LUAD bulk RNA-sequencing data from TCGA database. Integrating data from additional databases and clinical sample sequencing would enhance the robustness of our results. Lastly, the findings of this study require validation through further experimental investigations. In future, we will investigate neutrophil-macrophage communication in the immune microenvironment of LUAD, and develop therapeutic targets and drugs through in-vivo and in-vitro experiments.
Conclusion
In conclusion, our comprehensive data analysis has revealed the immune cell characteristics linked to the progression of LUAD, emphasizing the crucial involvement of neutrophils and associated genes in the immune microenvironment of LUAD. These results serve as a valuable foundation for further exploration of the tumor immune microenvironment of LUAD, presenting new opportunities for researchers to study the etiology of LUAD and develop potential therapeutic interventions. Hopefully, this study will promote the discovery of new immunotherapeutic targets for LUAD and the development of related drugs.
Data availability
The datasets analysed during the current study are available in the IEU OpenGWAS database (https://gwas.mrcieu.ac.uk/), the TCGA database (https://portal.gdc.com/), and the GEO database (https://www.ncbi.nlm.nih.gov/geo/).
Abbreviations
- GWAS:
-
Genome-wide association studies
- hdWGCNA:
-
High dimensional weighted gene co-expression network analysis
- ICI:
-
Immune checkpoint inhibitor
- IV:
-
Instrumental variable
- IVW:
-
Inverse-variance weighted
- LUAD:
-
Lung adenocarcinoma
- MR:
-
Mendelian randomization
- NETs:
-
Neutrophil extracellular traps
- OR:
-
Odds ratio
- PD-L1:
-
Programmed death ligand 1
- TCGA:
-
The Cancer Genome Atlas
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74(3), 229–263. https://doi.org/10.3322/caac.21834 (2024).
Lahiri, A. et al. Lung cancer immunotherapy: progress, pitfalls, and promises. Mol. Cancer 22(1), 40. https://doi.org/10.1186/s12943-023-01740-y (2023).
Tang, T. et al. Advantages of targeting the tumor immune microenvironment over blocking immune checkpoint in cancer immunotherapy. Signal Transduct. Target Ther. 6(1), 72. https://doi.org/10.1038/s41392-020-00449-4 (2021).
Schoenfeld, A. J. et al. Clinical and molecular correlates of PD-L1 expression in patients with lung adenocarcinomas. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 31(5), 599–608. https://doi.org/10.1016/j.annonc.2020.01.065 (2020).
He, J., Luan, T., Zhao, G. & Yang, Y. Fusing WGCNA and machine learning for immune-related gene prognostic index in lung adenocarcinoma: Precision prognosis, tumor microenvironment profiling, and biomarker discovery. J. Inflamm. Res. 16, 5309–5326. https://doi.org/10.2147/JIR.S436431 (2023).
Xiang, Y. et al. Macrophage-related gene signatures for predicting prognosis and immunotherapy of lung adenocarcinoma by machine learning and bioinformatics. J. Inflamm. Res. 17, 737–754. https://doi.org/10.2147/JIR.S443240 (2024).
Luo, J., An, J., Jia, R., Liu, C. & Zhang, Y. Identification and verification of metabolism-related immunotherapy features and prognosis in lung adenocarcinoma. Curr. Med. Chem. 32(7), 1423–1441. https://doi.org/10.2174/0109298673293414240314043529 (2025).
Swanton, C. et al. Embracing cancer complexity: Hallmarks of systemic disease. Cell 187(7), 1589–1616. https://doi.org/10.1016/j.cell.2024.02.009 (2024).
Hu, Z. et al. Dissecting the single-cell transcriptome network of macrophage and identifies a signature to predict prognosis in lung adenocarcinoma. Cell Oncol. Dordr Neth. 46(5), 1351–1368. https://doi.org/10.1007/s13402-023-00816-7 (2023).
Cui, C. et al. Neoantigen-driven B cell and CD4 T follicular helper cell collaboration promotes anti-tumor CD8 T cell responses. Cell 184(25), 6101-6118.e13. https://doi.org/10.1016/j.cell.2021.11.007 (2021).
Bahabayi, A. et al. Expression of GPR56 reflects a hypoactivated state of circulating B cells and is downregulated in B cell subsets in patients with early-stage lung adenocarcinoma. Immunology 173(1), 172–184. https://doi.org/10.1111/imm.13819 (2024).
Wang, Z. et al. Bone-metastatic lung adenocarcinoma cells bearing CD74-ROS1 fusion interact with macrophages to promote their dissemination. Oncogene 43(28), 2215–2227. https://doi.org/10.1038/s41388-024-03072-7 (2024).
Ancey, P. B. et al. GLUT1 expression in tumor-associated neutrophils promotes lung cancer growth and resistance to radiotherapy. Cancer Res. 81(9), 2345–2357. https://doi.org/10.1158/0008-5472.CAN-20-2870 (2021).
Schenkel, J. M. et al. Conventional type I dendritic cells maintain a reservoir of proliferative tumor-antigen specific TCF-1+ CD8+ T cells in tumor-draining lymph nodes. Immunity 54(10), 2338-2353.e6. https://doi.org/10.1016/j.immuni.2021.08.026 (2021).
Park, M. D. et al. TREM2 macrophages drive NK cell paucity and dysfunction in lung cancer. Nat. Immunol. 24(5), 792–801. https://doi.org/10.1038/s41590-023-01475-4 (2023).
Hai, Y. et al. High postoperative monocyte indicates inferior Clinicopathological characteristics and worse prognosis in lung adenocarcinoma or squamous cell carcinoma after lobectomy. BMC Cancer 18(1), 1011. https://doi.org/10.1186/s12885-018-4909-1 (2018).
Yang, Y. et al. Neutrophil chemotaxis score and chemotaxis-related genes have the potential for clinical application to prognosticate the survival of patients with tumours. BMC Cancer 24(1), 1244. https://doi.org/10.1186/s12885-024-12993-1 (2024).
Fan, L. L. et al. GNGT1 remodels the tumor microenvironment and promotes immune escape through enhancing tumor stemness and modulating the fibrinogen beta chain-neutrophil extracellular trap signaling axis in lung adenocarcinoma. Transl. Lung Cancer Res. 14(1), 239–259. https://doi.org/10.21037/tlcr-2024-1200 (2025).
Li, W. et al. Complex causal association between genetically predicted 731 immunocyte phenotype and osteonecrosis: a bidirectional two-sample Mendelian randomization analysis. Int. J. Surg. Lond. Engl. 110(6), 3285–3293. https://doi.org/10.1097/JS9.0000000000001327 (2024).
Zhu, H., Chen, C., Guo, H., Zhang, B. & Hu, Q. The causal role of immune cells on lung cancer: A bi-directional Mendelian randomization (MR) study. Aging 16(11), 10063–10073. https://doi.org/10.18632/aging.205917 (2024).
Xu, S. et al. Causal association between immune cells and lung cancer risk: A two-sample bidirectional Mendelian randomization analysis. Front Immunol. 15, 1433299. https://doi.org/10.3389/fimmu.2024.1433299 (2024).
Skrivankova, V. W. et al. Strengthening the reporting of observational studies in epidemiology using mendelian randomization: The STROBE-MR statement. JAMA 326(16), 1614–1621. https://doi.org/10.1001/jama.2021.18236 (2021).
Qian, J., Xu, H., Liu, J. & Zheng, Y. Associations of cholecystectomy with the risk of gastroesophageal reflux disease: a Mendelian randomization study. Int. J. Surg. Lond. Engl. 110(10), 6836–6840. https://doi.org/10.1097/JS9.0000000000001806 (2024).
Gao, Y., Zhang, G., Jiang, S. & Liu, Y. X. Wekemo Bioincloud: A user-friendly platform for meta-omics data analyses. iMeta 3(1), 175. https://doi.org/10.1002/imt2.175 (2024).
Brinkmann, V. et al. Neutrophil extracellular traps kill bacteria. Science 303(5663), 1532–1535. https://doi.org/10.1126/science.1092385 (2004).
Wang, J. et al. Neutrophil extracellular traps-inhibiting and fouling-resistant polysulfoxides potently prevent postoperative adhesion, tumor recurrence, and metastasis. Adv. Mater. Deerfield Beach Fla. 36(31), e2400894. https://doi.org/10.1002/adma.202400894 (2024).
Szczerba, B. M. et al. Neutrophils escort circulating tumour cells to enable cell cycle progression. Nature 566(7745), 553–557. https://doi.org/10.1038/s41586-019-0915-y (2019).
Zhang, Y. et al. Interleukin-17-induced neutrophil extracellular traps mediate resistance to checkpoint blockade in pancreatic cancer. J. Exp. Med. 217(12), e20190354. https://doi.org/10.1084/jem.20190354 (2020).
Jiang, Z. Z. et al. Neutrophil extracellular traps induce tumor metastasis through dual effects on cancer and endothelial cells. Oncoimmunology 11(1), 2052418. https://doi.org/10.1080/2162402X.2022.2052418 (2022).
Zhang, L., Zhang, X., Guan, M., Yu, F. & Lai, F. In-depth single-cell and bulk-RNA sequencing developed a NETosis-related gene signature affects non-small-cell lung cancer prognosis and tumor microenvironment: results from over 3,000 patients. Front Oncol. 13, 1282335. https://doi.org/10.3389/fonc.2023.1282335 (2023).
Zhang, Y. et al. A signature for pan-cancer prognosis based on neutrophil extracellular traps. J. Immunother. Cancer 10(6), e004210. https://doi.org/10.1136/jitc-2021-004210 (2022).
Petrova, M. et al. Tumor neutrophil extracellular traps and pretreatment neutrophils in association with progression-free survival in patients with metastatic non-small cell lung cancer receiving pembrolizumab alone or with chemotherapy. J. Clin. Oncol. 40(16), e21099. https://doi.org/10.1200/JCO.2022.40.16_suppl.e21099 (2022).
Guo, J. et al. A combined model of serum neutrophil extracellular traps, CD8+ T cells, and tumor proportion score provides better prediction of PD-1 inhibitor efficacy in patients with NSCLC. FEBS J. 291(15), 3403–3416. https://doi.org/10.1111/febs.17144 (2024).
Kaltenmeier, C. et al. Neutrophil extracellular traps promote T cell exhaustion in the tumor microenvironment. Front Immunol. 12, 785222. https://doi.org/10.3389/fimmu.2021.785222 (2021).
Wang, H. et al. Regulatory T-cell and neutrophil extracellular trap interaction contributes to carcinogenesis in non-alcoholic steatohepatitis. J. Hepatol. 75(6), 1271–1283. https://doi.org/10.1016/j.jhep.2021.07.032 (2021).
Teijeira, Á. et al. CXCR1 and CXCR2 chemokine receptor agonists produced by tumors induce neutrophil extracellular traps that interfere with immune cytotoxicity. Immunity 52(5), 856-871.e8. https://doi.org/10.1016/j.immuni.2020.03.001 (2020).
Xia, Y. et al. AAV-mediated gene transfer of DNase I in the liver of mice with colorectal cancer reduces liver metastasis and restores local innate and adaptive immune response. Mol. Oncol. 14(11), 2920–2935. https://doi.org/10.1002/1878-0261.12787 (2020).
Chen, D., Liang, H., Huang, L., Zhou, H. & Wang, Z. Liraglutide enhances the effect of checkpoint blockade in lung and liver cancers through the inhibition of neutrophil extracellular traps. FEBS Open Bio 14(8), 1365–1377. https://doi.org/10.1002/2211-5463.13499 (2024).
Chen, D. et al. Exenatide enhanced the antitumor efficacy on PD-1 blockade by the attenuation of neutrophil extracellular traps. Biochem. Biophys. Res. Commun. 619, 97–103. https://doi.org/10.1016/j.bbrc.2022.06.052 (2022).
Albrengues, J. et al. Neutrophil extracellular traps produced during inflammation awaken dormant cancer cells in mice. Science 361(6409), eaa04227. https://doi.org/10.1126/science.aao4227 (2018).
Zhang, L. et al. Neutrophil extracellular traps facilitate A549 cell invasion and migration in a macrophage-maintained inflammatory microenvironment. BioMed Res. Int. 2022, 8316525. https://doi.org/10.1155/2022/8316525 (2022).
Güç, E. & Pollard, J. W. Redefining macrophage and neutrophil biology in the metastatic cascade. Immunity 54(5), 885–902. https://doi.org/10.1016/j.immuni.2021.03.022 (2021).
Liu, X. et al. In situ neutrophil apoptosis and macrophage efferocytosis mediated by Glycyrrhiza protein nanoparticles for acute inflammation therapy. J. Control Release Off. J. Control Release Soc. 369, 215–230. https://doi.org/10.1016/j.jconrel.2024.03.029 (2024).
Horckmans, M. et al. Neutrophils orchestrate post-myocardial infarction healing by polarizing macrophages towards a reparative phenotype. Eur. Heart J. 38(3), 187–197. https://doi.org/10.1093/eurheartj/ehw002 (2017).
Braza, M. S. et al. Neutrophil derived CSF1 induces macrophage polarization and promotes transplantation tolerance. Am. J. Transplant Off. J. Am. Soc. Transplant. Am. Soc. Transpl. Surg. 18(5), 1247–1255. https://doi.org/10.1111/ajt.14645 (2018).
Yu, C., Zhou, G., Shi, Z., Yu, L. & Zhou, X. TREM1 facilitates the development of gastric cancer through regulating neutrophil extracellular traps-mediated macrophage polarization. Dig. Liver Dis. Off. J. Ital. Soc. Gastroenterol. Ital. Assoc. Study Liver 56(7), 1237–1247. https://doi.org/10.1016/j.dld.2023.12.002 (2024).
Liu, K. et al. BCG-induced formation of neutrophil extracellular traps play an important role in bladder cancer treatment. Clin. Immunol. Orlando Fla. 201, 4–14. https://doi.org/10.1016/j.clim.2019.02.005 (2019).
Song, S. et al. ELANE promotes M2 macrophage polarization by down-regulating PTEN and participates in the lung cancer progression. Immunol. Invest. 52(1), 20–34. https://doi.org/10.1080/08820139.2022.2115379 (2023).
Perego, M. et al. Reactivation of dormant tumor cells by modified lipids derived from stress-activated neutrophils. Sci Transl Med. 12(572), eabb5817. https://doi.org/10.1126/scitranslmed.abb5817 (2020).
Li, Z. et al. Proinflammatory S100A8 induces PD-L1 expression in macrophages, mediating tumor immune escape. J. Immunol. Baltim Md 1950 204(9), 2589–2599. https://doi.org/10.4049/jimmunol.1900753 (2020).
Lim, S. Y., Yuzhalin, A. E., Gordon-Weeks, A. N. & Muschel, R. J. Tumor-infiltrating monocytes/macrophages promote tumor invasion and migration by upregulating S100A8 and S100A9 expression in cancer cells. Oncogene 35(44), 5735–5745. https://doi.org/10.1038/onc.2016.107 (2016).
Navrátilová, A. et al. S100A11 (calgizzarin) is released via NETosis in rheumatoid arthritis (RA) and stimulates IL-6 and TNF secretion by neutrophils. Sci. Rep. 11(1), 6063. https://doi.org/10.1038/s41598-021-85561-3 (2021).
Deng, C. et al. Extracellular-vesicle-packaged S100A11 from osteosarcoma cells mediates lung premetastatic niche formation by recruiting gMDSCs. Cell Rep. 43(2), 113751. https://doi.org/10.1016/j.celrep.2024.113751 (2024).
Li, Y. et al. Neutrophil metabolomics in severe COVID-19 reveal GAPDH as a suppressor of neutrophil extracellular trap formation. Nat. Commun. 14(1), 2610. https://doi.org/10.1038/s41467-023-37567-w (2023).
Shen, C., Li, W. & Wang, Y. Research on the oncogenic role of the house-keeping gene GAPDH in human tumors. Transl. Cancer Res. 12(3), 525–535. https://doi.org/10.21037/tcr-22-1972 (2023).
Liao, S. T. et al. 4-Octyl itaconate inhibits aerobic glycolysis by targeting GAPDH to exert anti-inflammatory effects. Nat. Commun. 10(1), 5091. https://doi.org/10.1038/s41467-019-13078-5 (2019).
Yoo, H. J. et al. MsrB1-regulated GAPDH oxidation plays programmatic roles in shaping metabolic and inflammatory signatures during macrophage activation. Cell Rep. 41(6), 111598. https://doi.org/10.1016/j.celrep.2022.111598 (2022).
Acknowledgements
The authors acknowledge the following datasets: IEU OpenGWAS (https://gwas.mrcieu.ac.uk/), TCGA (https://portal.gdc.com/), GEO (https://www.ncbi.nlm.nih.gov/geo/), Wekemo Bioincloud (https:// www.bioincloud.tech), SPSSPRO platform (https://www.spsspro.com/), CellMarker 2.0 (http://bio-bigdata.hrbmu.edu.cn/CellMarker/), PanglaoDB (https://panglaodb.se/index.html), CB-Dock2 (https://cadd.labshare.cn/cb-dock2/php/index.php), Protein Data Bank (https://www1.rcsb.org/), and Metascape (https://www.metascape.org) for data sharing.
Funding
This work was supported by the National Natural Science Foundation of China (No. 82305188); the China Postdoctoral Science Foundation (No. 2022MD723715); and the Natural Science Foundation of Sichuan Science and Technology Department (No. 23NSFSC6246).
Author information
Authors and Affiliations
Contributions
Conceptualization: Jing Guo, Xiang Xiao, Xuan-Yu Wu. Data curation: Xiang Xiao, Xuan-Yu Wu, Jing-Qi Zhang. Formal analysis: Xiang Xiao, Xuan-Yu Wu, Jing-Qi Zhang. Funding acquisition: Jing Guo. Investigation: Xiang Xiao, Xuan-Yu Wu, Feng-Ming You. Methodology: Xuan-Yu Wu, Wen-Yuan Li, Jing Guo. Project administration: Jing Guo. Supervision: Feng-Ming You, Jing Guo. Validation: Jing Guo, Xiang Xiao. Writing—original draft: Xiang Xiao, Xuan-Yu Wu. Writing—review & editing: Xiang Xiao, Xuan-Yu Wu, Jing-Qi Zhang, Jing Guo.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study utilized publicly available GWAS summary statistics and RNA-sequencing, without the need for collecting new data, thus rendering additional ethical approval unnecessary.
Consent for publication
All authors give their consent for this work to be published.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xiao, X., Wu, XY., Zhang, JQ. et al. Exploring the impact of neutrophils on lung adenocarcinoma using Mendelian randomization and transcriptomic study. Sci Rep 15, 23835 (2025). https://doi.org/10.1038/s41598-025-08490-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-08490-5
Keywords
This article is cited by
-
Prognostic role of CDCA3 in lung adenocarcinoma with immune infiltration
Scientific Reports (2026)