Abstract
The airway microbiome has been implicated in the pathogenesis of laryngotracheal stenosis (LTS), yet prior studies using 16 S rRNA sequencing have limited sub-genus level resolution. Metagenomic whole genome shotgun sequencing (mWGS) allows for strain-level taxonomic and functional genomic analysis, providing detailed insights into specific organisms and pathways. A pilot study was conducted to explore the advantages and challenges of mWGS in investigating the airway metagenome in LTS. mWGS was conducted on 12 intraoperative swab samples from 8 LTS patients, divided into tracheostomy-dependent (n = 3) and non-tracheostomy (n = 5) groups, and 4 controls. Patient comorbidities, antibiotic use, and medications were documented. Biobakery workflows were used for taxonomic and functional profiling. Species-specific reference databases were constructed for 6 abundant species for strain-level analyses. LTS samples had decreased taxonomic diversity and were dominated by species with previously described roles in other chronic inflammatory processes such as Staphylococcus aureus, Streptococcus parasanguinis, Streptococcus mitis, and Corynebacterium pseudogenitalium. LTS samples were enriched for pathways involved in fatty acid biosynthesis and formaldehyde metabolism. Our results identified tracheostomy as an important potential confounder in airway metagenomics but show mWGS techniques are promising in uncovering microbiota correlates in LTS that could reveal disease-specific biomarkers, comorbidity links, and therapeutic targets.
Similar content being viewed by others
Introduction
Laryngotracheal stenosis (LTS) is the progressive narrowing of the laryngotracheal tract at the level of the glottis, subglottic space, or the trachea. The predominant etiology of LTS is iatrogenic, secondary to epithelial damage such as endotracheal intubation or, less commonly due to cartilage injury following tracheostomy. Idiopathic subglottic stenosis (iSGS) is a rare, recurrent fibroinflammatory condition primarily affecting middle-aged Caucasian females1. Treatment modalities of LTS present as a challenge due to their invasive nature, risk of re-stenosis1, and complications associated with tracheostomy1. To address these limitations, research in laryngology has proposed distinct pathophysiological mechanisms in iSGS and iLTS. Recent studies on subglottic stenosis propose that epithelial dysfunction promotes displacement of native microbiome2,3 secondary to defects in the epithelial barrier4,5, adaptive immune system response3,4, and tissue fibrosis6. Recent evidence showed an upregulation in the IL-17 A/IL237 signaling pathway in iSGS airway scar mucosa. This upregulation is characterized by an increase in INFγ producing γδT cells, involved in the induction and regulation of mucosal immunity7. This association with immune infiltration and activation within the lower airway suggests that dysbiosis could contribute to disease development through increased inflammatory signals in response to pathogens. Gelbard et al. (2017) illustrated this interaction by outlining the role of Mycobacterium tuberculosis complex infection in IL-17 A/γδT upregulation and subsequent tissue remodeling in patients with iSGS2.
Conventional culture-based approaches framed our understanding of infections as a single pathogenic entity, however, advances in culture-independent high-throughput sequencing studies have expanded our understanding of the host microbiome’s role in disease pathogenesis. Dysbiosis of the resident mucosal microbiota has demonstrated, mechanistic roles in other chronic inflammatory conditions with compromised epithelial integrity including inflammatory bowel disease8 and atopic dermatitis9. Studies have begun to explore similar pathways involved in airway function. In a 16 S rRNA amplicon analyses in patients with subglottic stenosis, microbial associations at the genus level were identified including Moraxella and Acinetobacter from the scar mucosa, with a potential protective association of Prevotella10. Consequently, drug-eluting tubes, stents, and implants have been developed to direct antimicrobials release and modulate the local microbiota11,12.
To date, the role of airway microbiome in stenosis relied primarily on 16 S rRNA amplicon sequencing. This technique identifies bacterial taxa at the genus level and overlooks information on strain-level diversity and functional profiling of the communities present. Metagenomic whole genome shotgun sequencing (mWGS) overcomes these limitations by enabling the identification of novel taxa, performing strain-level taxonomic and functional genomic analysis. This approach provides a more detailed view of the specific organisms and their involvement in disease mechanisms. To build on previous research on the role of microbiome in LTS we aimed to present a general overview on the feasibility and potential insight of mWGS in understanding microbial communities in LTS. We anticipated finding decreased microbial diversity within the LTS condition, as well as species, strains, or functional differences associated with the chronic inflammation characteristic of LTS, which could reveal disease-specific biomarkers, comorbidity links, and potential therapeutic targets. To address these questions, we generated a cross-sectional metagenomic study on airway samples from patients with LTS and a control group without airway stenosis.
Methods
A cross-sectional study was conducted with approval from the ethics committee of University of Cincinnati, under the protocol number 2022 − 0986. The procedures used adhere to the tenets of the Declaration of Helsinki.
Sample collection
Fifteen patients undergoing suspension for microlaryngoscopy between November 2022 and November 2023 at the University of Cincinnati Medical Center were recruited into either the LTS or control condition. Of the 15 collected samples, 3 patients were excluded: 1 from the control group and 1 from the LTS group due to low read counts, and 1 sample from the LTS group with tracheostomy due to systemic antibiotic use. In the LTS condition, we included 4 patients with LTS and five with LTS requiring tracheostomy. The second condition, serving as the control, included 4 patients without any history or signs of LTS or other airway stenosis undergoing the procedure for benign vocal fold lesions such as cysts, polyps, or sulcus vocalis, which have not been shown to have specific microbiome signatures13. Intraoperative samples were collected from the stenotic areas of the airway in the LTS group using a rigid flocked swab attached to a Dedo laryngoscope following standardized, sterile techniques to minimize contamination from the oral microbiome. For the control condition, samples were collected from the airways where no stenosis was present. Following informed consent, demographic data, including comorbidities, antibiotic use, and current medications, were collected through chart review and questionnaires.
Metagenomic sample extraction, library preparation and sequencing
Samples were collected in the operating room and immediately placed on dry ice to preserve DNA integrity. They were then stored at − 80 °C, ensuring long-term stability until they could be processed in batches. Sample processing, library preparation and sequencing were performed by the University of Cincinnati Genomics, Epigenomics and Sequencing Core Facility according to standard protocols.
DNA was isolated from the samples using the QIAamp DNA Microbiome Kits (Qiagen., Hilden, Germany) according to manufacturer protocol, to enrich for microbial DNA while minimizing the presence of host DNA. Following DNA extraction, sequencing libraries were constructed using the NEBNext Ultra II FS DNA Library Prep Kit (New England Biolabs., Ipswich, MA, USA). Resulting libraries were sequenced on an Illumina NextSeq 2000 (Illumina Inc., San Diego, CA, USA) with 2 × 61-bp paired end reads to a minimum sequencing depth of ~ 25 million reads/sample. Demultiplexing yielded 122.2 million forward and 121.1 million reverse reads across all samples. Individual samples ranged from ~ 31,000 to 24.2 million reads, with a median of 2.4 million per direction. Median read length was 61 bp for both forward and reverse reads. This sequencing depth has shown robustness in providing detailed characterization of taxonomic composition at the species and strain levels14,15. Samples with less than 10,000 high quality reads or less than 1000 mapped taxonomic reads were excluded. To evaluate sequencing depth efficiency in capturing taxonomic diversity, we generated alpha-rarefaction curves in QIIME 2 (version 2024.10)15,16 Alpha diversity analysis showed that Shannon values plateaued beyond 500,000 reads. To verify that sequencing depth sufficiently represented taxonomic diversity, we repeated our taxonomic diversity analysis at subsampling depths at 50,000 reads and 500,000 reads. On average, Shannon diversity increased by only 0.085 from 50,000 to 500,000 reads (Wilcoxon, p < 0.0001), and by 0.052 from 500,000 to full depth (Wilcoxon, p < 0.0001).
Metagenomic quality control
The BioBakery17 metagenomic workflows (Huttenhower Lab., MA, USA) were carried out for quality control, taxonomic profiling, and functional profiling of samples. Initially, Kneaddata was used for quality control of raw pair-ended fastq files from sequencing libraries. This step included quality checks and trimming of low-quality reads, and sequencing adapters using Trimmomatic (v0.33). Samples were then decontaminated from human reads using Bowtie2 (v2.2.9, hgb37dec_v0.1 human genome database) with ‘very sensitive’ parameter.
Species-level taxonomic profiling
The profiling of the microbial community composition of the metagenomic data was executed using MetaPhlan (v.4.1)18. MetaPhlan utilizes a comprehensive set of clade-specific marker genes in the mpa_vJun23 database using Bowtie2 (v 2.5, --very-sensitive parameter) alongside the ChocoPhlan database (version 202307). Resulting count tables of bacterial species were normalized and grouped into a joint table.
Strain heterogeneity
Bacteria exhibit profound functional diversity at the subspecies level. Colonization by different strains can have vastly different implications for the host19. The species E. coli encompasses both well-validated probiotic strains such as Nissile and epidemic pathogens such as OH157:H720,21. Microbe-microbe interactions conveying colonization resistance against pathogens such as S. aureus are often strain-specific22. We constructed species-specific reference databases by extracting genomes for 6 most abundant and widely prevalent species in our dataset including: Staphylococcus aureus, Corynebacterium accolens, Streptococcus mitis, Streptococcus parasanguinis, Corynebacterium striatum, Veillonella atypica, and Veillonella dispar. We mapped reads to custom databases of all currently available Refseq genomes (https://ftp.ncbi.nlm.nih.gov/genomes/) (as of March 2024) using Bowtie2 (v2.2.9) with “very sensitive mode”, allowing up to ten alignments per read (k = 10) to ensure thorough mapping. The resulting alignment data, stored in SAM files, were processed through Pathoscope (v2.0.7)23 to match reads to the closest strain.
Bacterial community diversity was assessed using alpha-diversity and beta-diversity indices. Alpha-diversity represented by the Shannon Diversity Index was compared across groups. To investigate the heterogeneity in microbiome composition between the different groups, we calculated Bray-Curtis distances of species level taxonomic composition between each sample and performed Principal Coordinate Analysis (PCoA) using the Vegan (v2.6-4) and Ecodist (v2.1.3) packages in R. Significant species-level composition differences between groups were identified using differential abundance analysis with the DESeq2 (v1.38.3) package in R (alpha = 0.05).
Functional analysis
Taxonomic profiles obtained from MetaPhlan (v.4.1) were merged and integrated into HUMAnN (v3.9)17to characterize the abundance of microbial metabolic pathways and molecular functions of our dataset. Gene families, pathway abundance, and coverage files were generated referencing the ChocoPhlan (v. 202307) nucleotide database and the UniRef90_201901b database of proteome genome identifiers using Diamond24. The pathway abundance and gene families’ outputs, initially represented as RPK (reads per kilobase), were normalized to counts per million (CPM) to address potential biases resulting from differences in reference gene length. Subsequently, the generated pathway abundance tables were grouped and represented based on MetaCyc metabolic pathways annotation25. For differential abundance of genetic pathways, the DESeq2 package was used to calculate log2 fold difference in pathway abundnace between groups, with significance determined using an adjusted alpha of 0.0001.
Strain heterogeneity analysis
To estimate strain diversity, we calculated the Shannon Diversity Index for strain-level taxonomic composition for each species investigated to compare the number and evenness of different strains represented. Differences in by-species strain diversity between samples was assessed using bi-directional Wilcoxon test with Bonferroni correction for multiple comparisons (alpha = 0.05).
Results
After processing in Kneaddata, a total of 1.39E9 (1.39 billion) raw reads yielded 2.22E8 (222 million) high-quality paired end reads. Airway and control samples had a mean of 11.88E6 (11.88 million) and 10.18E6 (10.18 million) paired end reads/sample respectively. Reads per sample at each stage of quality control and mapping are included in Supplementary Table S1.
Patients’ demographics
After excluding three samples, our analysis included 12 subjects divided into non-tracheostomy LTS (n = 5), LTS requiring tracheostomy (n = 3), and control (n = 4) groups. The LTS requiring tracheostomy group was predominantly female (66.67%). Non-tracheostomy LTS patients were younger on average (M = 43.2, SD = 14.82) with higher BMI (M = 37.01, SD = 10.92) compared to controls (47.75 years, SD = 12.5) and BMI (M = 29.49, SD = 4.54). Diabetes and GERD were notably more prevalent in the LTS requiring tracheostomy group (75%). Recent inhaler use was exclusive to LTS groups, with smoking history present in all controls (Table 1).
Species-level taxonomy
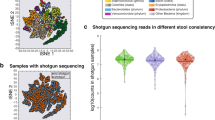
An initial analysis revealed a significant decrease in airway species-level diversity among LTS patients compared to controls, as measured by the Shannon diversity index (Fig. 1a, p = 0.028). Further stratification of LTS by tracheostomy presence revealed a stepwise decline in α-diversity from controls to the LTS group, and further to the LTS with tracheostomy group, however these differences did not rise to statistical significance (Supplementary Figure S1). We next performed Bray-Curtis Principal Coordinates Analysis (PCoA) to examine between samples differences in species composition. The first PCo1 and PCo2 accounted for 33% and 16.1% of the variance respectively (Fig. 1b). The control cohort was clustered together and overlapped with LTS without tracheostomy group, however the latter subgroup showed modest spread toward LTS with tracheostomy group. Because this exploratory analysis identified tracheostomy as a major co-variate of taxonomic composition within the LTS cohort, we subdivided the cohort for subsequent analysis into LTS with and without tracheostomy.
Species-level diversity of the airway microbiome in healthy airways vs. airways with LTS. (a) Shannon Index of Diversity represents the number and evenness of different species. Wilcoxon, alpha = 0.05. (b) Principal Coordinate Analysis of Bray-Curtis differences in species-level composition.
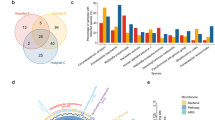
Visualizing the relative abundance of the most common taxa in our dataset (Fig. 2), we observed a loss of taxonomic diversity in LTS patients (Fig. 1a) with decreased proportion of lower-abundance microbes and a commensurate increase in organisms such as Staphylococcus aureus, Streptococcus mitis, Corynebacterium striatum, and Streptococcus parasanguinis. These changes are heterogenous, with different opportunistic pathogens dominating different samples. We also noted a trend towards less representation of Veillonella atypica, Veillonella dispar, and Corynebaceterium accolens.
Relative abundance of top 20 most abundant species. Each bar represents 100% of species assignments for one patient. All other species grouped under their respective kingdom (e.g. Bacteria).
Differential abundance profiles at the species level revealed that S. aureus, Corynebacterium pseudogenitalium, Anaerococcus vaginalis, and Actinomyces bouchesdhurhonesis were significantly enriched in LTS compared to control condition (Fig. 3). Among these, A. vaginalis and C.pseudogenitalium showed the largest effect sizes (log2 fold changes of 3.6 and 3.7, respectively), and were significantly more abundant in LTS patients with tracheostomy compared to those without (Fig. 3c). In contrast, S. mitis was less abundant in the tracheostomized LTS subgroup (log2 fold change − 2.5). Further comparisons between non-tracheostomized LTS patients and controls showed enrichment of S. aureus in the LTS group (log2 fold change 4.7), while Actinomyces sp. ICM47 and Schaliaa SGB17168 were more prevalent in the control cohort.
Differential abundance of species between groups. (a) All patients with LTS vs. patients with healthy airways, (b) LTS vs. healthy airways excluding LTS patients with tracheostomy, and (c) LTS patients with tracheostomy vs. LTS patients without tracheostomy. Differential abundances calculated using DESeq2, alpha = 0.05.
To further understand this diversity, samples from non-tracheostomized LTS were compared to controls. Staphylococcus aureus (log2FoldChange 4.7) was found to be significantly enriched in LTS, whereas Actinomyces sp. ICM47, and Schaliaa SGB17168 were more abundant in the control group (Fig. 3).
Functional analysis
An additional advantage of mWGS over 16 S amplicon sequencing is the ability to detect directly what bacterial genetic pathways are present in a sample, from which differences in the functional role of the commensal microbiome in health and disease states can be inferred. To account for the effect of tracheostomy as a potential confounding factor, we first compared non-tracheostomized LTS patients to healthy controls (Fig. 4a), then evaluated differences between LTS patients with and without tracheostomy (Fig. 4b). We identified significant enriched pathway in LTS mainly involved in fatty acid biosynthesis (PWY − 5972: stearate biosynthesis I, log2FoldChange = 6.9; PWY − 7388: octanoyl-[acyl-carrier protein] biosynthesis, log2FoldChange = 5.7), protein ubiquitination (PWY − 7511, log2FoldChange = 6.7), and formaldehyde metabolism (PWY − 1861: formaldehyde assimilation II, RuMP Cycle, log2FoldChange = 5.7; RUMP − PWY: formaldehyde oxidation I, log2FoldChange = 5.6) (Fig. 4a). In contrast, when comparing LTS patients with and without tracheostomy (Fig. 4b), depleted pathways were identified in tracheostomized patients including PWY − 5972: stearate biosynthesis I (log2FoldChange = − 7.4), PWY − 7832: ABH and Lewis epitope biosynthesis (log2FoldChange = − 8.6), and PWY66-201: nicotine degradation IV (log2FoldChange = − 7.9). After stratification, no pathways remained significantly different between LTS patients without tracheostomy and healthy controls.
Differential abundance of functional pathways between groups. (a) Log2fold change of significantly enriched pathways for all patients with LTS vs. controls. (b) Log2fold change of significantly enriched pathways for LTS patients with tracheostomy vs. LTS patients without tracheostomy. Differential abundances calculated using DESeq2, alpha = 0.0001.
Strain heterogeneity
Although we observed a trend for decreased strain-level diversity in S. mitis, S. parasanguinis, V. atypica, and V. dispar in LTS patients with tracheostomy compared to healthy controls (Fig. 5), none rose to statistical significance when corrected for multiple comparisons.
Strain-level diversity for species of interest in health airways vs. airways with LTS without and with tracheostomy. Shannon Index of Diversity represents the number and evenness of different strains. No differences are statistically significant for alpha = 0.05 with Bonferroni correction.
Discussion
We present a pilot study employing WGS to study the airway microbiome in iatrogenic LTS, where the study population was unmatched for demographics, comorbidities, disease severity, and smoking history. Our results reinforce some previous findings from 16 S studies and identify new potential microbial players and challenges for future investigations. Isolating specific microbiome co-variants of LTS is challenging both due to inter-individual heterogeneity of commensal communities and the high prevalence of comorbidities with LTS that have known associations in the mucosal microbiota. The inclusion of tracheostomized patients allowed us to capture a greater breadth of LTS severity. However, our analysis quickly identified tracheostomy as a major co-variate. Without a control cohort of tracheostomized patients without LTS, this introduced a potential confounding variable, and our study was not powered nor had redundant sample size to treat LTS patients with and without tracheostomy as separate cohorts. Indeed, the greatest variance in the taxonomic diversity of our dataset was observed in LTS patients with tracheostomy (Fig. 1b). Because greater LTS disease severity results in tracheostomy requirement, we cannot conclude whether trends most pronounced in these patients are directly associated with LTS or indirectly associated as a sequela of tracheostomy. Prior 16 S amplicon studies have demonstrated that tracheostomy is an independent variable in the airway microbiome, as it provides a more direct pathway for skin and environmental microbiome to enter the trachea26. However, to our knowledge this has not been studied with WGS, so the sub-genus taxonomic and functional implications for our dataset are unknown.
We note several dataset-specific limitations to this work. Similarly to previous studies, in our study both GERD and diabetes were more prevalent in LTS patients. The relationship between diabetes and the respiratory microbiota has not been characterized, but there may be implications that parallel the known interplay between the systemic immune dysregulation and decreased epithelial integrity in diabetes and the skin microbiota27. Another consideration for this dataset is that controls were undergoing microlaryngoscopy for benign vocal lesions, and all control patients had a smoking history. At 16 S resolution, smoking has been associated with decreased laryngeal microbiome diversity and increased streptococcal abundance which when overrepresented in our control group may therefore be decreasing the signal of these pro-inflammatory features associated with LTS in our dataset. Another limitation of note in this dataset is that despite passing our inclusion criteria and quality control metrics, several retained samples had relatively lower sequencing depth (1–5 × 103 mapped reads) with low mapped ratios (approximately 2–5%) and retained reads ratios as low as 0.1–0.2% (Supplementary Table S1). As such their taxonomic profiles should be interpreted with caution, particularly the possibility that some low-abundance organisms in the specimens may not have been captured. We believe the conclusions we draw are appropriate considering the robustness of our alpha diversity analysis to subsampling, that these samples are distributed among the cohorts, and the congruence of composition of higher abundance taxa within groups.
Despite these weaknesses, this study identified several metagenomic correlates of LTS worthy of discussion. The microbiota of LTS airways were heterogenous, but consistently exhibited reduced species level taxonomic diversity (Fig. 1a) with a reduced proportion of low-abundance bacteria and a reciprocal increase in species such as S. aureus, S. parasanguinis, S. mitis, C. pseudogenitalium, or A. vaginalis (Fig. 2). A 16 S study by Hillel et al. also noted decreased alpha diversity and increased Streptococcus in LTS in scarred regions in SGS10. Decreased microbiota diversity is observed broadly in other disease states, the causes of which are often multi-factorial including increased mucosal immune activation and microbe-microbe antagonism28, but loss of diversity can also precede onset of inflammatory disease states through the loss of immune-regulating microbial interactions29. Another consideration in patients with chronic inflammatory processes requiring healthcare exposure is increased frequency of antibiotic exposure, which diminishes commensal bacteria and promote the proliferation of pathogenic species such as Enterobacteriaceae and S. aureus30. A previous study investigating the tracheal microbiota in diabetic patients revealed an increased abundance of Prevotella and Porphyromonas, along with a decrease in Streptococcus, Neisseria, and Fusobacterium31.
Although the strongest species-level association of LTS in our dataset was S. aureus (Fig. 3), this is driven by outliers, two LTS samples (one with and one without tracheostomy) with over 80% representation of S. aureus each (Fig. 2). One possible explanation for these findings is recent antibiotic use in these two participants. Although LTS has been observed subsequent to MRSA tracheitis in neonates32, this study may be the first to suggest an association between S. aureus and LTS in adults. We do not know the temporal relationship between S. aureus colonization and stenosis development in these patients, but we note that when present it dominates the airway microbiome. Although present in a minority of our subjects, the known role of S. aureus in sustaining other chronic inflammatory processes such as eczema9 makes it a potential risk factor or therapeutic target in colonized patients.
Although our study was not designed for a systematic investigation of strain-level interactions, we identified trends that could be the product of strain-level interactions for future inquiry. Although not statistically significant, we observed C. accolens was prevalent in our study but had decreased relative abundance in LTS subjects (Fig. 2). This organism is known to inhibit mucosal barrier disruption by S. aureus in the nasal airway in a strain-specific fashion22. It is possible this organism may also be having a protective effect in LTS. In addition to providing colonization-resistance, Intra-individual strain diversity has been shown to suppress populational level virulence factor expression33. Our observed trend towards decreased strain diversity in Streptococcus and Veillonella species in tracheostomy-dependent LTS patients (Fig. 5) also raises suspicion for a competitive microbe-microbe interaction within or between species.
The airway microbiota of LTS patients were enriched for several functional pathways of biologic interest (Fig. 4). Two representatives of the RuMP pathway for formaldehyde metabolism were significantly increased. Formaldehyde detoxification has emerged as an important survival mechanism for microbes at the host-pathogen interface, both due to aldehyde products created by bacterial metabolism and from the inflammatory host respiratory burst34. The RuMP pathway has been detected in microbes from multiple phyla, including S. aureus. It makes sense that in a chronically inflamed airway this function would provide a selective advantage. Two fatty acid metabolism pathways were elevated in LTS, which Fan et al. also noted in their metabolomic LTS study35. While bacterial fatty acid metabolism is a major mechanism for gut microbiome interactions with the host metabolism and immunomodulation, its role in mucosal scarring has not been explored. Finally, a protein ubiquitination was enriched, which in bacteria likely represents a virulence factor that exploits host protein processing pathways36.
The findings of this pilot study add to the small but growing body of literature examining the role of the airway microbiome in the pathogenesis of LTS. Our results highlight tracheostomy as an important confounder when conducting microbiome research on patients with severe airway disease. However, these results also build on previous work and help illustrate that certain microbes functionally adapted to chronic inflammatory processes such as S. aureus and streptococci which may at least exploit the disrupted microbial niche for colonization advantage and raise suspicion that these colonized opportunistic pathogens could lead to inflammatory flares exacerbating the disease. Our future applications of WGS will be to recruit a recruit non-LTS tracheostomy cohort and expanded LTS tracheostomy cohorts with and without tracheostomy to better deparse sub-genus taxonomic and functional metagenomic implications of tracheostomy, as well as identify specific strains and virulence factor pathways associated with LTS.
Conclusion
In conclusion, our findings revealed decreased microbial diversity and specific bacterial species association with LTS, as well as distinct abundant pathways despite the added complexity from unmatched confounders. This study demonstrates the feasibility and application of mWGS in LTS and aims to inform future clinical study design by identifying potential therapeutic targets to interrupt cycle of re-stenosis and reduce burden of disease.
Data availability
The datasets generated and/or analyzed during the current study are available on publication in the NCBI repository, BioSample accession number: SAMN47550761.
References
Gelbard, A. et al. Causes and consequences of adult laryngotracheal stenosis. Laryngoscope 125 (5), 1137–1143 (2015).
Gelbard, A. et al. Molecular analysis of idiopathic subglottic stenosis for Mycobacterium species. Laryngoscope 127 (1), 179–185 (2017).
Lina, I. A. et al. Phenotypic epithelial changes in laryngotracheal stenosis. Laryngoscope 132 (11), 2194–2201 (2022).
Gelbard, A. et al. Idiopathic Subglottic Stenosis Arises at the Epithelial Interface of Host and Pathogen (Res Sq, 2023).
Hillel, A. T. et al. T-Helper 2 lymphocyte immunophenotype is associated with iatrogenic laryngotracheal stenosis. Laryngoscope 129 (1), 177–186 (2019).
Morrison, R. J. et al. Pathologic fibroblasts in idiopathic subglottic stenosis amplify local inflammatory signals. Otolaryngol. Head Neck Surg. 160 (1), 107–115 (2019).
Gelbard, A. et al. Idiopathic subglottic stenosis is associated with activation of the inflammatory IL-17A/IL-23 axis. Laryngoscope 126 (11), E356–E361 (2016).
Jakubczyk, D., Leszczyńska, K. & Górska, S. The effectiveness of probiotics in the treatment of inflammatory bowel disease (IBD)-A critical review. Nutrients, 12(7). (2020).
Boguniewicz, M. & Leung, D. Y. Recent insights into atopic dermatitis and implications for management of infectious complications. J. Allergy Clin. Immunol. 125 (1), 4–13 (2010). quiz 14 – 5.
Hillel, A. T. et al. Laryngotracheal microbiota in adult laryngotracheal stenosis. mSphere, 4(3). (2019).
Aronson, M. R. et al. Drug-Eluting endotracheal tubes for preventing bacterial inflammation in subglottic stenosis. Laryngoscope 132 (7), 1356–1363 (2022).
Cruz, D. R. D. et al. Drug delivery systems for wound healing treatment of upper airway injury. Expert Opinion on Drug Delivery: p. null-null.
Hanshew, A. S., Jetté, M. E. & Thibeault, S. L. Characterization and comparison of bacterial communities in benign vocal fold lesions. Microbiome 2 (1), 43 (2014).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded human Microbiome project. Nature 550 (7674), 61–66 (2017).
Callahan, B. J. et al. DADA2: High-resolution sample inference from illumina amplicon data. Nat. Methods. 13 (7), 581–583 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible Microbiome data science using QIIME 2. Nat. Biotechnol. 37 (8), 852–857 (2019).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with biobakery 3. eLife 10, e65088 (2021).
Blanco-Miguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species with MetaPhlAn 4. bioRxiv, : p. 2022.08.22.504593. (2022).
Zhu, A. et al. Inter-individual differences in the gene content of human gut bacterial species. Genome Biol. 16 (1), 82 (2015).
Karmali, M. A. Infection by Shiga toxin-producing Escherichia coli: an overview. Mol. Biotechnol. 26 (2), 117–122 (2004).
Sonnenborn, U. & Schulze, J. The non-pathogenic Escherichia coli strain Nissle 1917 – features of a versatile probiotic. Microb. Ecol. Health Disease. 21, 122–158 (2009).
Menberu, M. A. et al. In vitro and in vivo evaluation of probiotic properties of Corynebacterium accolens isolated from the human nasal cavity. Microbiol. Res. 255, 126927 (2022).
Francis, O. E. et al. Pathoscope: species identification and strain attribution with unassembled sequencing data. Genome Res. 23 (10), 1721–1729 (2013).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods. 18 (4), 366–368 (2021).
Caspi, R. et al. The metacyc database of metabolic pathways and enzymes - a 2019 update. Nucleic Acids Res. 48 (D1), D445–D453 (2019).
Pérez-Losada, M. et al. The Temporal dynamics of the tracheal Microbiome in tracheostomised patients with and without lower respiratory infections. PLoS One. 12 (8), e0182520 (2017).
Schwartz, D. J., Langdon, A. E. & Dantas, G. Understanding the impact of antibiotic perturbation on the human Microbiome. Genome Med. 12 (1), 82 (2020).
Hou, K. et al. Microbiota in health and diseases. Signal. Transduct. Target. Ther. 7 (1), 135 (2022).
Minerbi, A. & Shen, S. Gut Microbiome in anesthesiology and pain medicine. Anesthesiology 137 (1), 93–108 (2022).
Luchen, C. C. et al. Impact of antibiotics on gut Microbiome composition and resistome in the first years of life in low- to middle-income countries: A systematic review. PLoS Med. 20 (6), e1004235 (2023).
Cho, S. Y. et al. Metataxonomic investigation of the microbial community in the trachea and oropharynx of healthy controls and diabetic patients using endotracheal tubes. PLOS ONE. 16 (11), e0259596 (2021).
Yamada, Y. et al. Acquired subglottic stenosis caused by methicillin resistant Staphylococcus aureus that produce epidermal cell differentiation inhibitor. Arch. Dis. Child. Fetal Neonatal Ed. 84 (1), F38–F39 (2001).
Zhou, W. et al. Host-Specific evolutionary and transmission dynamics shape the functional diversification of Staphylococcus epidermidis in human skin. Cell 180 (3), 454–470e18 (2020).
Chen, N. H. et al. Formaldehyde stress responses in bacterial pathogens. Front. Microbiol. 7, 257 (2016).
Fan, Z. et al. Tracheal Microbiome and metabolome profiling in iatrogenic subglottic tracheal stenosis. BMC Pulm Med. 23 (1), 361 (2023).
Kim, M. et al. Bacterial effectors and their functions in the ubiquitin-proteasome system: insight from the modes of substrate recognition. Cells 3 (3), 848–864 (2014).
Funding
This research was funded by the University of Cincinnati Center for Clinical & Translational Science and Training Just in Time Funding.
Author information
Authors and Affiliations
Contributions
NA contributed to data collection, analysis, and writing; PJL contributed to data analysis, writing, and revision; CAK contributed to study development, data collection, and analysis; LLB contributed to revision; GRD contributed to data collection, revision, and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics community of the University of Cincinnati, under the protocol number 2022 − 0986.
Consent to participate and publish
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Awad, N., Larson, P.J., Sissoko, C.A. et al. Metagenomic whole genome shotgun analysis of the airway microbiome in laryngotracheal stenosis: a pilot study. Sci Rep 15, 26570 (2025). https://doi.org/10.1038/s41598-025-11468-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11468-y