Abstract
Recent developments in the registration of histology and micro-computed tomography (µCT) have broadened the perspective of pathological applications such as virtual histology based on µCT. This topic remains challenging because of the low image quality of soft tissue CT. Additionally, soft tissue samples usually deform during the histology slide preparation, making it difficult to correlate the structures between the histology slide and µCT. In this work, we propose a novel 2D-3D multi-modal deformable image registration method. The method utilizes an initial global 2D-3D registration using an ML-based differentiable similarity measure. The registration is then finalized by an analytical out-of-plane deformation refinement. The method is evaluated on datasets acquired from tonsil and tumor tissues. µCTs of both phase-contrast and conventional absorption modalities are investigated. The registration results from the proposed method are compared with those from intensity- and keypoint-based methods. The comparison is conducted using both visual and fiducial-based evaluations. The proposed method demonstrates superior performance compared to the other two methods.
Similar content being viewed by others
Introduction
Histology and CT are two imaging modalities commonly integrated in clinical applications. Histological imaging is widely used to examine tissue samples to study their morphology or composition on a microscopic scale, which is a key procedure in clinical pathology and histopathology. Various staining methods are used to enhance the contrast for different tissue components, such as lymphatic tissue, connective tissue, and muscle. In Germany, around 40 million samples are analyzed by 1800 pathologists each year for cancer diagnosis1. However, given the relatively long turnaround time (exceeding 3 days for the conventional, established standard procedure in pathology laboratories23) and the large number of tissue examinations per pathologist each year, there is an increasing need from the clinics to improve the efficiency of the current cancer diagnosis pipeline.
Focusing on improving the time cycle of the diagnosis pipeline, various studies on virtual histology using CT have been done45. Recent developments of the CT imaging setup6 enable the resolution under sub-micrometer scale using absorption and phase-contrast modalities. The novel CT imaging methods reveal the possibility of resolving micro-structures that are usually visible in histology images but invisible in conventional clinical CTs. This study was carried out in the scope of the research project named HORUS (High-resolution X-ray computed tomography and intelligent evaluation of histological tissue samples for diagnostics), supported by the German Ministry for Education and Research (BMBF), grant number 13GW0571A. In this project, histology images were taken from the conventional pathology workflow. In the meanwhile, a wide range of advanced µCT imaging methods has been explored to replace or support histological imaging in the conventional pathology workflow. Due to the intrinsic difficulty in high-resolution imaging of soft tissue with CT, only a limited number of CTs have reached sufficient resolution and contrast at the current stage. However, it remains interesting for the researchers and pathologists to register the histology and µCT images of the same patient for further pathological analysis or exploring multi-modal image synthesis, even though the datasets are limited in the beginning phase of the research. Moreover, with the histology image and the matching µCT slice, one can transfer annotations by pathologists from histology to µCT.
The process of searching for the matching µCT slice is non-trivial. The histology slides are acquired following the conventional sample preparation and sectioning procedure7. The histological sectioning is not always parallel to the surface of the sample block; a small angle (± 5 degrees) is manually selected by the pathologist while performing the cut8. During the sectioning process, the 2D slices undergo in- and out-of-plane deformation due to the blade motion9. To successfully enhance this classical, established laboratory workflow with CT-generated 3D images, this deformation must be unambiguously and computationally reversible, to create a clear link between coordinates in the µCT volume and actual locations in the physical tissue sample. The core challenge in matching the images involves implementing a 2D-3D multi-modal deformable image registration algorithm. Studies focusing on the 2D histology to 3D µCT registration have been conducted in recent years11. To overcome the limit when evaluating the similarity between images of different modalities, an approach using ML-based modality translation with cascaded plane selection and 2D-3D deformable registration has been investigated12. Moreover, a method using speeded-up robust features (SURF) and scale-invariant feature transform (SIFT), which were originally invented for optical image matching, has also been developed1314. This method was developed with bone CTs. However, µCTs of soft tissues have a lower contrast and a higher noise level in comparison to that of bone CTs. Moreover, a greater deformation exists between the µCT and histology slide of the soft tissues. SURF and SIFT are sensitive to non-rigid transformation and to the noise of the image15. The performance of SIFT, therefore, drops drastically with soft tissue images. A method targeting clinical soft tissue is not yet well established. The reproducibility of a fully ML-integrated registration framework is not guaranteed due to the lack of ground truth training data, particularly given the dramatically varying morphology between different tissues. Driven by the need to address this unresolved registration problem between soft tissue histology and µCT images, this study is carried out to explore new paradigms when immediate engineering solutions are not applicable due to the novel image modalities and limitations on the training dataset.
Previously, registration methods using intensity- and keypoint-based initialization were investigated13141617. The first experiment exploits a relatively small capture range of these methods. Recently, an ML-based differentiable similarity measure approximation (DISA)18 was introduced. Using the DISA image similarity metric enables a fast differentiable global registration in the initialization, which is beneficial for enlarging the capture range of the registration. DISA has a lightweight structure and a simple training objective. Moreover, we use a patch-based approach to train the DISA CNN. Therefore, the training of DISA CNN does not require an excessive amount of data. Targeting this research-oriented scenario where the number of initial datasets is limited, we propose a 2D-3D deformable registration algorithm that combines a DISA-based plane initialization with an analytic plane refinement optimization. In comparison to other histology to µCT registration methods using intensity- and keypoint-based initialization, our method exhibits improvement in both visual- and fiducial-based evaluation.
Methods
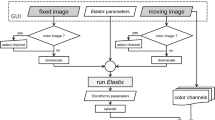
The workflow of the proposed registration algorithm and the DISA 2D CNN structure. (A) Image preprocessing with grayscale conversion and percentile normalization. (B) Initialization of the plane pose using 2D-3D registration on the feature maps of CT and histology. The original feature maps contain 16 channels. Here, only the first three channels are extracted for visualization as an RGB image. (C) Sampling plane refinement by optimizing over the plane pose parameters and the out-of-plane deformation. Control point pairs are displayed as yellow markers above and under the CT volume. (D) The structure of the DISA 2D CNN that generates a 16-channel feature map from an input image.
The 2D-3D histology to µCT registration algorithm is equivalent to searching an optimal sampling plane that extracts a slice from the CT volume that matches the 2D histology slide. The algorithm consists of the following parts: plane initialization, optimization of plane pose parameters, and optimization of the out-of-plane deformation. The workflow of the registration algorithm is demonstrated in Figure 1.
In the pre-processing phase (Figure 1A), the histology image is converted to grayscale by computing the mean of all three channels. Additionally, percentile normalization using a range of 1% to 99% is applied to both the grayscale histology image and the µCT.
Initialization
We previously implemented an intensity-based initialization approach16. This approach iteratively extracts 2D CT slices along one axis and runs a 2D-2D image registration with the histology slide. The optimal slice is then defined as the one that returns the highest similarity with the histology slide. Here, the local normalized cross-correlation (LNCC) is used as the similarity measure. Besides, we have also implemented a keypoint-based initialization method17. Following the approach originally invented by Chicherova et al.14, 2D slices are iteratively extracted from the CT volume. Keypoints from the CT slice and the histology slide are then detected using the SIFT feature detector. The detected features are later matched and used for the plane fitting to search for the initial plane. The results from these approaches are compared later with those of the DISA-based method.
In this work, a novel initialization approach is introduced. The initialization is replaced with a high-performance global image registration using DISA-\(\text {LC}^2\) similarity measure. The \(\text {LC}^2\) similarity measure19 quantifies whether the patches of the target image can be approximated by a linear combination of the intensities and the gradient magnitude of the corresponding patches of the source image. The patch-wise similarities are then aggregated by weighted averaging. While the \(\text {LC}^2\) similarity has been successfully used for multi-modal registration with a large capture range20, its inner use of pixel-wise least-squares fitting makes it non-differentiable and computationally expensive.
As a solution to this, Ronchetti et al.18 proposed to approximate the \(\text {LC}^2\) similarity with a dot-product between features produced by a convolutional neural network (CNN). Concretely, given a dataset of patches S, they train a CNN \(\phi\) to minimize the following approximation error
The DISA approach, by replacing the \(\text {LC}^2\) metric with dot-products, produces a much faster and differentiable similarity metric, which allows for global registration as required by our initialization problem.
In its original formulation, the DISA approach is not well-suited for 2D–3D registration, as it does not incorporate the orientation of the 2D slice into the feature vectors. To address this limitation, we leverage the observation that the histology slide is approximately parallel to the µCT slices. This allows us to apply a 2D DISA model to each individual CT slice and subsequently stack the resulting feature maps. This procedure effectively embeds an orientation prior into the DISA features.
Following the architecture design proposed by Ronchetti et al.18, our model consists of 2D convolutional layers with LeakyReLU activation function22 and residual blocks23, and employs BlurPool24 for downsampling, leading to a combined striding factor of 4. The detailed architecture of DISA 2D CNN is visualized in Figure 1D.
For each forward pass of the model, a 2D image is used as input, and a 2D 16-channel feature map is generated as output. Extracting the CT slices from the volume as the input images, the output 2D CT feature maps are later stacked into a volume, whereas the 2D histology feature map is obtained with the 2D histology image as input. The thickness of the histology feature map is set to the estimated thickness (1.5 µm) of the histology slide. Global 2D-3D image registration with DISA-\(\text {LC}^2\) is performed on the two feature maps, whereby the histology feature map is set as the moving image, as shown in Figure 1B. The resulting 3D transformation matrix of the histology feature map is then used as the initial guess for the later plane refinement optimization.
Plane refinement
The optimization problems in the plane fine-tuning phase can be defined as follows:
Assuming extracting the CT slices along the z-axis, \(\textbf{T} \in \mathbb {R}^3\) denotes the pose parameters that transform the sampling plane in 3D space (translation on z-axis, rotations on x-axis and y-axis). \(\textbf{T}_{opt}\) refers to the optimal pose parameters returned from Eq. (12). We use a free-form deformation model based on B-spline21. Two control points are assigned above and below the sampling plane as a pair. These pairs of control points are distributed in a grid pattern across the entire plane, as shown in Figure 1C. By introducing displacements to the control point pairs, out-of-plane deformation of the plane is achieved. \(\textbf{D} \in \mathbb {R}^N\) denotes the z-displacements of the control point pairs and N is the number of the grid points. \(I_{histo}\) is the histology slide, \(I_{ct}(\textbf{T})\) and \(I_{ct}(\textbf{T}_{opt}, \textbf{D})\) refer to the CT slices extracted from the sampling plane defined by the corresponding plane pose parameters and the out-of-plane deformation. The similarity between the 2D images after performing a 2D-2D in-plane deformable image registration is denoted as \(s(\cdot ,\cdot )\).
Experiments
The samples used in this work were human tonsil and tumor tissues prepared by the University Hospital of Düsseldorf. All methods to acquire such tissue samples were carried out in accordance with relevant guidelines and regulations. All experimental protocols were approved by the Ethics Committee Heinrich-Heine-University Düsseldorf, Medical Faculty, positively voted application number 2020-943 from 3rd June 2020. Due to the retrospective sampling from diagnostically closed cases and anonymised samples, the Ethics Committee Heinrich-Heine-University Düsseldorf, Medical Faculty, waived the need to obtain informed consent.
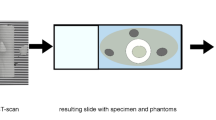
The samples were embedded in paraffin blocks, sent for CT scans, and then slices were extracted from the blocks to prepare for the histology slides. Both absorption and phase-contrast µCT were acquired and tested for the performance of the proposed algorithm. At the time of writing this paper, a limited amount (4 phase-contrast µCTs and 2 absorption µCTs) of µCT-histology image pairs were available for training and analysis. The sample orientation is fixed for all CTs, with the block surface being roughly perpendicular to the z-axis. Slices were extracted via the established standard slicing procedure in a pathology laboratory. Despite the actual cutting direction not being perfectly parallel to the surface, it intrinsically provides initial information to decide which axis to extract the CT slices for later processing.
Datasets and training
Two phase-contrast µCTs and the two corresponding histology slides are used for training the DISA 2D CNN. We use a total of 640 phase-contrast µCT slices with a spacing of 2.6 µm acquired at a commercial phase-contrast µCT setup, Exciscope25. Two histology slides prepared by the University Hospital of Düsseldorf are used. The training approach is unsupervised, and it does not require a ground truth matching between the CT and histology images. 640 CT-histology 2D image pairs are generated by reusing the same two histology slides. We resample the CT volumes and the histology images to a spacing of 10.4 µm and normalize the pixel values with a mean of 0 and standard deviation of 1. Following the original sampling and training strategy for 3D images18, we adapt the approaches for 2D images. The source-target patch sampling procedure is repeated 10000 times on each CT-histology image pair, which results in a total of 40000 pairs of patches. Note that the source-target patch pairs are not spatially registered, but are merely randomly sampled from both images. For each patch pair, we analytically compute the \(\text {LC}^2\) similarity value as the ground truth value to approximate. An \(\text {L}_2\) loss is used in the training, which is computed from the dot product of the feature maps from the DISA 2D CNN and the pre-computed ground truth \(\text {LC}^2\) value. Training converges to an average \(\text {L}_2\) error of 0.0090 and validation converges to 0.0095. The output of the CNN is a 16-channel feature map.
Registration and optimization
For the 2D-3D registration of the feature maps, as shown in Figure 1B, we use the BFGS26 optimizer. We equidistantly select 10 depths below the surface of the tissue block and repeat rigid and affine registration starting from the corresponding depth. The range of each registration trial is set to \(\pm 10^\circ\) and \(\pm 400\)µm. The best result among the 10 trials is then selected to continue with the plane refinement later.
The plane refinement optimizations are executed with the pre-processed CT volumes and histology slides. To improve the efficiency, we crop out the sample-free area in the CT volumes and the histology slides and resample the images to the same spacing as mentioned in section 3.1, which is approximately a downsampling factor of 4 for the CT volumes. We use derivative-free optimizer BOBYQA27 from the NLopt library28 for the plane refinement optimizations (Figure 1C). Each optimization step of the plane refinement optimization wraps a 2D-2D deformable registration. The 2D-2D deformable registration uses a B-spline free-form deformation with a \(4 \times 4\) control point grid, with LNCC as the similarity measure and BOBYQA as the optimizer. For the plane pose optimization, the iteration is aborted if the change of parameters falls below \(1\times 10^{-4}\). For the out-of-plane deformation optimization, a grid of \(4 \times 4\) for the control point pairs is used. To increase the search space, we repeat the optimization 5 times with a random initial guess for the control point displacements and terminate each after 80 iterations. The best result among the 5 optimizations is then selected.
Results and discussion
On a CT volume with a dimension of \(650 \times 580 \times 490\) and a resampled histology slide with a dimension of \(780 \times 620\), the runtime of the initialization is 2 seconds for the DISA approach, which is significantly more efficient than the intensity-based initialization (30 seconds) and keypoint-based initialization (22 seconds).
Registration cases
Preprocessed histology slides and the registered CT slices of different tissues and imaging modalities. (A) and (B) Tonsil tissues and phase-contrast CTs. (C) Tumor tissue and phase-contrast CT. (D) Tonsil tissue and absorption CT.
Comparison of registration results from different initialization approaches. (A) Preprocessed histology image. (B) Registration using intensity-based initialization. (C) Registration using keypoint-based initialization. (D) Registration using DISA initialization. (E) Registration using manual initialization.
The proposed algorithm is applied to the rest of the datasets that are excluded from the model training, including 3 phase-contrast µCT-histology pairs and 1 absorption µCT-histology pair. These datasets exhibit different anatomical structures and image qualities from the training data. An evaluation of this approach is conducted based on the comparison with the previously developed intensity- and keypoint-based methods.
The registered images from 4 datasets are shown in Figure 2. The CTs in Figures 2A and B are the histology slides and phase-contrast µCTs of tonsil tissues acquired at Exciscope. The initial spacing of the CT volumes is 2.6 µm. Figure 2C shows the histology slide and the phase-contrast CT of tumor tissue acquired with the TOMCAT beamline at the synchrotron setup at Paul Scherrer Institut29, with a spacing of 1.6 µm. An absorption µCT acquired at MITOS GmbH30 and the corresponding histology slide are shown in Figure 2D. It shows visually good registration on various tissue types (tonsil, Figure 2A, B, D, and tumor tissues, Figure 2C) and imaging modalities (absorption, Figure 2D, and phase-contrast CT, Figure 2A, B, C), which indicates the proposed method has the potential to be further applied to different tissues and imaging modalities without retraining the model. All the registration results demonstrate visually good registration of the proposed approach, especially on the macro structures.
Detailed comparison of the registration results from different approaches is shown in Figure 3. Note that these approaches only use different initialization methods, but they use the same method for plane refinement optimization, as mentioned in section 2.2. Figures 3B and C are the results from the previously developed approaches using intensity-based and keypoint-based initialization. In these results, the registration of specific micro-structures within the marked dashed square is not as robust as that of the larger structures outside of the square. Figure 3D is the registered CT slice acquired from the proposed approach, which exhibits a better registration of the micro-structures than the previous results. As a reference, we also manually locate an initial plane that visually matches the histology slide the best and start the plane refinement optimization from this manual initial guess. The corresponding result is presented in Figure 3E. To quantitatively compare the different approaches, we present the similarity values using LNCC and \(\text {LC}^2\) metrics between the histology slide and the registered CT slice from the sample displayed in Figure 2A. As shown in Table 1, the registration using the DISA initialization approach shows a higher value in both LNCC and \(\text {LC}^2\) in comparison with the intensity- and keypoint-based approaches, which align well with the visual comparison on the images.
DISA-\(\text {LC}^2\) similarity analysis
We conduct a post-registration DISA-\(\text {LC}^2\) similarity analysis to verify the optimal plane and the optimal similarity value. After obtaining the final optimal plane with registration using DISA-based initialization, we equidistantly sample CT slices apart from (translation on the z-axis) the optimal plane and compute the corresponding similarity metrics with the histology image. As shown in Figure 4A, B, and C, an interval of 200 µm, 100 µm, and 50 µm is taken, respectively, covering different ranges within the µCT volume. DISA-\(\text {LC}^2\), NCC, LNCC, and \(\text {LC}^2\) similarity metrics are computed. For a more intuitive comparison, the similarity values are normalized to the min and max values in the current range. In all three ranges, the similarity values of all metrics achieved the maximum at the optimal plane, with a trend to decrease while moving away from the optimal plane. It shows that DISA-\(\text {LC}^2\) is capable of evaluating feature correspondences under multiple scales. Despite the local maxima observed at certain positions, the registration converges to the global maximum owing to the global search in the DISA-based initialization.
The similarity values between the histology image and the CT slices extracted from various distances away from the optimal plane. DISA-\(\text {LC}^2\), NCC, LNCC, and \(\text {LC}^2\) are computed, normalized to the corresponding min and max values in the current range. (A) Similarity values of the histology image and the CT slices sampled from the range [−400 µm, 1000 µm] with an interval of 20 µm. (B) Similarity values from the range [−500 µm, 500 µm] with an interval of 200 µm. (C) Similarity values from the range [−250 µm, 250 µm] with an interval of 50 µm.
FRE analyses
An example of the 20 fiducial pairs selected on both modalities and the plot of the computed FRE values. (A) 20 manually annotated fiducials on the 2D histology. (B) Corresponding 20 fiducials on the CT volume. (C) FRE values of the registration results from four different initialization approaches and the final registration using the corresponding initialization approaches.
Since there’s no ground truth that we could use to benchmark the registration results, we utilize fiducial registration error (FRE) analysis to quantitatively evaluate the initialization and final registration results. Fiducial pairs corresponding to the same tissue structure are manually selected from the CT and histology. After performing the registration, the FRE is computed as the mean \(\text {L}_2\) distances between these fiducial marker pairs:
where n refers to the number of the fiducial marker pairs, \(\textbf{f}_{i, histo}\) and \(\textbf{f}_{i, ct}\) stand for the 3D world position of the corresponding fiducial markers i on the histology image and CT.
A total of 20 (\(n=20\)) fiducial pairs on the histology and CT are manually annotated for each patient dataset, as shown in Figure 5A and 5B. The FRE values after different initialization approaches are computed to compare the performance of these initialization approaches. The corresponding FRE values after the final registration are also calculated to quantify the registration accuracy, which are shown in the plot in Figure 5C. The FRE from the proposed DISA initialization approach is lower than the other initialization methods, which exhibits its superiority in the initial global search compared to the keypoint- and intensity-based initialization methods. Consequently, the final registration using DISA initialization yields a lower FRE than the other two approaches. The FRE of the DISA-based approach is comparable to that of the manual approach.
Conclusions and outlook
This work develops a novel algorithm for 2D-3D multi-modal image registration for an application scenario of limited datasets and no ground truth. With the proposed initialization method, the results demonstrate that the convergence of the optimization outperforms the intensity- and keypoint-based methods, with a significantly improved runtime of initialization. Registered ground truth image pairs are unnecessary; the training approach is unsupervised, using unregistered image patch pairs. With a relatively lightweight structure and a simple training objective, the DISA 2D CNN model can be trained with a modest amount of data, which makes it suitable for applications and research where datasets are limited in the first place. According to our test, the model also has the potential to be directly applied to images of different modalities, as well as images acquired from different tissue types, without retraining the model. The accuracy of the registration results is comparable to that of the manual results. It shows that the proposed method can potentially be applied to generate ground-truth registered image pairs, which are essential to training networks for multi-modal translation and synthesis.
Despite the orientation of the sample surface being utilized as initial information for the algorithm, the workflow can be further automated by integrating an intensity-based surface detection algorithm. The out-of-plane deformation is realized by applying the deformation model to the CT volume. The CT slices are then extracted from the deformed volume via a flat sampling plane, which is mathematically equivalent to directly deforming the sampling plane. The runtime of the plane refinement optimization could be drastically improved in the future by deforming the sampling plane instead of the CT volume. Moreover, the integration of DISA in the plane refinement phase will be accomplished. In histology-CT image pairs, one can observe more complicated deformation and discontinuity, such as folding and fracturing. Some local micro-structures are still not perfectly registered. The deformation at the micro-structures can be modelled better with a denser control point grid; however, the optimization will become more expensive with an increased degree of freedom. A future study is needed to improve the complexity of the deformation model while maintaining a decent computational efficiency. Moreover, the discontinuity in the actual deformation should also be considered. Further studies comparing the proposed approach and the other established ML-based 2D-3D deformable registration will be carried out, once we acquire a sufficient amount of datasets in the future to train larger models.
Data availability
Data is provided within the supplementary files.
Code availability
The source code for the proposed registration algorithm is included as a supplementary file.
References
Bruno, M., Laszló, F., Ralf, H., Svenja, B. & Tina, S. Number of pathologists in germany: Comparison with european countries, usa, and canada. Virchows Arch.478, 335–341 (2021).
Dias, M. M. & Barrett, A. W. Comparison of histopathological turnaround times for mandibulectomies, glossectomies, and incisional biopsies of the tongue. Br. J. Oral Maxillofac. Surg.61(2), 131–135 (2023).
Sharma, A., Nishadham, V., Gupta, P., Gupta, G., Sharma, D., Goel, S., Pasricha, S., Kamboj, M. & Mehta, A.: Evaluation of turnaround times of diagnostic biopsies: A metric of quality in surgical pathology. International Journal of Surgical Pathology 0(0).
Hieber, S. E. et al. Tomographic brain imaging with nucleolar detail and automatic cell counting. Sci. Rep.6(1), 32156 (2016).
Lee, J. Y. et al. Microvascular imaging of the unstained human superior colliculus using synchrotron-radiation phase-contrast microtomography. Sci. Rep.12(1), 9238 (2022).
Pacilè, S. & Tromba, G. Introduction to x-ray micro-tomography In (eds Giuliani, A. & Cedola, A.) 19–39 (Springer, 2018).
Funkhouser, W.K.: Chapter 11 - pathology: The clinical description of human disease. In: Coleman, W.B., Tsongalis, G.J. (eds.) Molecular Pathology, Second edition edn., pp. 217–229. Academic Press, Rockville, MD, USA (2018).
Dey, P. Tissue microtomy: Principle and procedure 41–50 (Springer, 2018).
Schulz, G. et al. Three-dimensional strain fields in human brain resulting from formalin fixation. J. Neurosci. Methods202(1), 17–27 (2011).
Sotiras, A., Davatzikos, C. & Paragios, N. Deformable medical image registration: A survey. IEEE Trans. Med. Imaging32(7), 1153–1190 (2013).
Nolte, P. et al. Current approaches for image fusion of histological data with computed tomography and magnetic resonance imaging. Radiol. Res. Pract.https://doi.org/10.1155/2022/6765895 (2022).
Leroy, A. et al. Structuregnet: Structure-guided multimodal 2d–3d registration. In Medical Image Computing and Computer Assisted Intervention - MICCAI 2023 (eds Greenspan, H. et al.) 771–780 (Springer, Cham, 2023).
Chicherova, N., Fundana, K., Müller, B., Cattin, P.C. Histology to \(\mu\)ct data matching using landmarks and a density biased ransac. In Medical Image Computing and Computer-Assisted Intervention - MICCAI 2014 (eds Golland, P. et al.) 243–250 (Springer, Cham, 2014).
Chicherova, N. et al. Automatic deformable registration of histological slides to \(\mu\)ct volume data. Journal of Microscopy 271(1), 49–61 (2018).
Lowe, D. G. Distinctive image features from scale-invariant keypoints. Int. J. Comput. Vis.60(2), 91–110 (2004).
Chen, e.a.: Deformable Registration of \(\mu\)CT and Histology Slide. In: Proceedings of the 6th International Symposium on Medical Applications of X-ray Phase-Contrast & Photon-Counting, Technical University of Munich, July 2023 (2023).
Chen, e.a.: 2D-3D Deformable Image Registration of Histology Slide and Phase-contrast Micro-CT. In: Proceedings of The 6th International Conference on X-ray and Neutron Phase Imaging with Gratings, Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences, April 2024 (2024).
Ronchetti, M., Wein, W., Navab, N., Zettinig, O. & Prevost, R. Disa: Differentiable similarity approximation for universal multimodal registration. In Medical Image Computing and Computer Assisted Intervention - MICCAI 2023 (eds Greenspan, H. et al.) 761–770 (Springer, Cham, 2023).
Wein, W., Brunke, S., Khamene, A., Callstrom, M. R. & Navab, N. Automatic CT-ultrasound registration for diagnostic imaging and image-guided intervention. Med. Image Anal.12(5), 577–585 (2008).
Wein, W., Ladikos, A., Fuerst, B., Shah, A., Sharma, K. spsampsps Navab, N.: Global registration of ultrasound to mri using the lc2 metric for enabling neurosurgical guidance 16(Pt 1), 34–41 (2013).
Rueckert, D. et al. Nonrigid registration using free-form deformations: Application to breast mr images. IEEE Trans. Med. Imaging18(8), 712–721 (1999).
Maas, A.L.: Rectifier nonlinearities improve neural network acoustic models. In: Proceedings of the International Conference on Machine Learning (ICML) (2013).
He, K., Zhang, X., Ren, S. & Sun, J.: Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 770–778 (2016).
Zhang, R.: Making convolutional networks shift-invariant again. In: Chaudhuri, K., Salakhutdinov, R. (eds.) Proceedings of the 36th International Conference on Machine Learning. Proceedings of Machine Learning Research, vol. 97, pp. 7324–7334 (2019).
Exciscope. https://exciscope.com/
Fletcher, R.: Newton-like methods. In: Alex, A. (ed.) Practical Methods of Optimization, pp. 44–79. John Wiley & Sons, Ltd, The Atrium, Southern Gate Chichester, England PO19 8SQ (2000). Chap. 3.
Powell, M.J.D.: The BOBYQA algorithm for bound constrained optimization without derivatives. Technical Report NA2009/06, Department of Applied Mathematics and Theoretical Physics, Cambridge University, Cambridge, UK (2009).
Johnson, S.G.: The NLopt nonlinear-optimization package. https://github.com/stevengj/nlopt (2007).
TOMCAT, Swiss Light Source, Paul Scherrer Institute. https://www.psi.ch/en/sls/tomcat.
MITOS GmbH. https://www.mitos.de/en/services.html.
Acknowledgements
This study was partially supported by the German Ministry for Education and Research (BMBF), grant number 13GW0571A, “HORUS”. Collaborating institutions: ImFusion GmbH, Munich; DORNER Health IT Solutions, Muellheim; Fraunhofer EMI, Efringen-Kirchen; Institute for Surgical Pathology, Medical Center-University of Freiburg; Institute of Pathology, Heinrich Heine University and University Hospital of Duesseldorf. The authors wish to thank Prof. Julia Herzen for her suggestions on the phase-contrast micro-CT acquisition and manuscript publication.
Author information
Authors and Affiliations
Contributions
J.C. and M.W. conceived the study and designed the methodology. M.R. and V.N. reviewed the methodology and provided suggestions for refinement. J.C., V.S., M.K., C.L. and M.G. conducted the imaging experiments and collected the data. J.C. and V.N. performed the data analysis. S.M. and M.S. provided critical insights in the data analysis and contributed to the interpretation of the results. J.C. wrote the manuscript and prepared all the figures, with input from all authors. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, J., Ronchetti, M., Stehl, V. et al. 2D-3D deformable image registration of histology slide and micro-CT with DISA-based initialization. Sci Rep 15, 25972 (2025). https://doi.org/10.1038/s41598-025-11583-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11583-w
Keywords
This article is cited by
-
2D-3D deformable image registration of histology slide and micro-CT with DISA-based initialization
Scientific Reports (2025)