Abstract
Epithelial ovarian cancer (EOC) metastasizes predominantly through multicellular aggregates known as spheroids, which disseminate within the peritoneal cavity and initiate secondary disease upon reattachment at distant sites. EOC spheroids resist detachment-induced cell death by upregulating stress responses including AMP-activated protein kinase (AMPK) signaling and AMPK-dependent macroautophagy (autophagy), highlighting these pathways as potential therapeutic targets. Previously, we used a pharmacological approach to putatively identify Ca2+/calmodulin-dependent protein kinase kinase 2 (CAMKKβ, encoded by CAMKK2) as the primary activator of AMPK in EOC spheroids. Herein we have generated CAMKK2 knockout EOC cell lines via CRISPR–Cas9 genome editing to confirm this function of CAMKKβ and explore the impacts of its loss using in vitro and in vivo models of metastatic EOC. CAMKK2 knockout spheroids exhibited decreased AMPK activation, autophagic flux, cell viability, and metastatic potential relative to parental spheroids, and intraperitoneal xenograft tumours lacking CAMKKβ grew slower than their CAMKKβ-intact counterparts. Effect magnitudes varied between cell line models, suggesting context-dependent roles for CAMKKβ in EOC and rationalizing further studies to characterize the underlying mechanisms. Altogether, our findings highlight CAMKKβ as an important contributor to metabolic reprogramming in EOC spheroids and as a potential therapeutic target in the setting of advanced disease.
Similar content being viewed by others
Introduction
Among gynecological cancers, epithelial ovarian cancer (EOC) is the most lethal1. This is largely attributable to late detection, as approximately 75% of EOC diagnoses occur after metastasis1,2 and 70% of these patients die within 5 years1,2,3. Standard of care for advanced EOC comprises surgical debulking and platinum/taxane chemotherapy, as it has remained for nearly three decades1,3,4,5,6. Most patients initially respond well to standard treatment1,7, but chemoresistant relapse is exceedingly common and inevitably fatal due to a lack of effective second-line treatments1,3,7. Ultimately, the bleak landscape for treatment of advanced EOC highlights an urgent need to identify targetable vulnerabilities in this disease.
Metastasis of EOC within the peritoneal cavity is understood to occur primarily via multicellular aggregates called spheroids, which form through the passive aggregation of cells that have exfoliated from the primary tumour4,8,9,10. Supporting this model for EOC metastasis, spheroids containing viable cancer cells are often present in the ascites of advanced-stage EOC patients11,12,13,14. Moreover, in vitro studies have demonstrated that EOC spheroids attach readily to components of the extracellular matrix found in the peritoneum and displace mesothelial cells efficiently11,15,16. Other studies characterizing EOC spheroid biology have determined that aggregation drives the activation of various stress response pathways, enabling constituent cells to resist detachment-induced programmed cell death (anoikis) and survive metastatic transit8,17. Our group has previously characterized the regulation of several stress responses in EOC spheroids, including reversible entry into a quiescent state18,19activation of AMP-activated protein kinase (AMPK)20,21,22induction of macroautophagy (herein referred to as autophagy)22,23,24and reversion of the Warburg effect25.
Activation of AMPK occurs via phosphorylation of AMPKα at Thr172—a process canonically mediated by liver kinase B1 (LKB1) in response to bioenergetic stress (i.e., ATP depletion)26,27,28,29. Once activated, AMPK acts to restore bioenergetic homeostasis by phosphorylating an array of downstream targets, inhibiting anabolic pathways (e.g., lipid and protein biosynthesis) and upregulating catabolic processes (e.g., autophagy)26,27,28,29. Although spheroids possess physical properties that are strongly associated with bioenergetic stress (i.e., hypoxia, nutrient deficiency)30,31 we have shown that AMPK activation in EOC spheroids does not occur through the canonical LKB1-mediated pathway. Indeed, we have putatively identified Ca2+/calmodulin-dependent protein kinase kinase 2 (CAMKKβ) as the kinase primarily responsible for AMPK activation in EOC spheroids, rather than LKB121,22.
Previously, we have shown that CAMKKβ-dependent activation of AMPK drives autophagy induction in EOC spheroids and that pharmacological inhibition of CAMKKβ reduces cell viability in EOC spheroids22. However, the overall importance of CAMKKβ-AMPK signaling in facilitating EOC metastasis has yet to be investigated. In this study, we have genetically ablated CAMKKβ in EOC cell lines and characterized the associated impacts on growth and stress response signaling in vitro and in a mouse xenograft model of metastatic EOC. In doing so, we have identified that CAMKKβ is an important mediator of EOC growth in both contexts, suggesting CAMKKβ is a viable therapeutic target in the setting of advanced EOC.
Results
CAMKK2 ablation attenuates AMPK activation and downstream signaling in EOC spheroids
It has been demonstrated that CAMKKβ-mediated AMPK phosphorylation is induced by matrix detachment in various non-EOC cell lines32 and we have shown that inhibiting CAMKKβ with STO-609 blocks AMPK phosphorylation in EOC spheroids21,22. STO-609 is known to inhibit kinases other than CAMKKβ33,34 however, so we sought to confirm our prior observations using a more specific targeting approach. Thus, we generated CAMKK2 knockout (CAMKK2 KO) OVCAR8 and HeyA8 cell lines using CRISPR–Cas9-mediated genome editing to evaluate the impacts of CAMKKβ ablation with improved specificity. After confirming CAMKKβ loss (Fig. 1A), we assessed AMPK activation in spheroids by quantifying changes in phosphorylation of AMPKT172. Detachment-induced AMPK activation was diminished significantly in CAMKK2 KO spheroids relative to parental spheroids (Fig. 1B), confirming CAMKKβ is the primary mediator of AMPK activation in EOC spheroids.
Loss of CAMKKβ prevents activation of AMPK and ULK1 in EOC spheroids. a Confirmation of CRISPR/Cas9-mediated knockout of CAMKK2 via western blot in OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO adherent cells (Adh) and spheroids (Sph). # indicates a non-specific band. Vinculin was used as a loading control. b, c Western blot analysis and densitometric quantification of b AMPKT172 and c ULK1S555 phosphorylation in OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO cells under adherent and spheroid culture conditions.
Quantified western blot data represent mean normalized phospho-protein abundance ± SEM for n = 5 (p-AMPKT172) or n = 4 (p-ULK1S555) independent experiments. Signal for each phospho-target was normalized against the respective total protein signal. Data are expressed relative to the parental adherent sample, for which the mean was set to 1. Data were analyzed via two-way ANOVA and groups with shared lineage (e.g., parental Adh vs. parental Sph) or shared culture condition (e.g., parental Adh vs. CAMKK2 KO Adh) were compared using the Holm-Šídák method (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
After confirming that AMPK activation in EOC spheroids was significantly reduced due to genetic ablation of CAMKKβ, we sought to evaluate the associated effects on signaling targets downstream of AMPK. However, over 100 unique targets of AMPK have been identified in diverse systems26,27,28,29 necessitating a focused approach. Since CAMKKβ-AMPK signaling drives autophagy induction in EOC spheroids22 we chose to assess the effect of CAMKKβ loss on the activation of Unc-51-like autophagy activating kinase (ULK1) – a target of AMPK with a central role in autophagy induction27. Indeed, AMPK is known to activate ULK1 via phosphorylation of ULK1S555 and other key residues35. Detachment-induced phosphorylation of ULK1S555 was significantly weaker in CAMKK2 KO spheroids than in parental spheroids, much like that of AMPKT172 (Fig. 1C). The effects of CAMKKβ loss on the regulation of other canonical AMPK targets were less consistent; relative to parental spheroids, acetyl-CoA carboxylase (ACC)S79 phosphorylation was reduced in OVCAR8 CAMKK2 KO spheroids but not in HeyA8 CAMKK2 KO spheroids, while RaptorS792 phosphorylation was maintained in both CAMKK2 KO spheroid models (Supplementary Fig. S1). Nonetheless, the loading control-normalized abundance of each phospho-species we assessed (p-AMPKT172, p-ULK1S555, p-ACCS79, and p-RaptorS792) was lower in CAMKK2 KO spheroids relative to parental spheroids (Supplementary Fig. S2). Taken together, these findings demonstrate that loss of CAMKKβ is sufficient to block detachment-induced activation of AMPK in EOC spheroids and attenuate signal transduction to its downstream targets.
CAMKKβ loss attenuates autophagy activation in EOC spheroids
Since detachment-induced phosphorylation of ULK1S555 was blunted in CAMKK2 KO spheroids, we next sought to determine if autophagy induction was interrupted as a result. To do so, we assessed the regulation of two commonly-used markers of autophagy activity – sequestosome 1 (p62) and microtubule-associated protein 1 light chain 3 beta (LC3B). When autophagy is upregulated, p62 levels typically decrease and the ratio of lipidated to unlipidated LC3B (i.e., LC3BII: I) increases36. However, upon probing for these markers, we identified no differences in p62 abundance between parental and CAMKK2 KO spheroids and found that LC3BII: I ratio was higher in CAMKK2 KO spheroids than in parental spheroids (Fig. 2A). Notably, while increased LC3BII: I ratio is typically indicative of autophagy induction, accumulation of LC3BII can also indicate inhibition of autophagic flux at a later step (i.e., failure of autolysosome formation/fusion)36. We also considered that sustained loss of CAMKKβ may have elicited long-term signaling adaptations in the CAMKK2 KO cell lines, leading to altered regulation of p62 and LC3B. Thus, the effects of CAMKKβ loss on autophagy could not be discerned clearly based on these experiments. In turn, we addressed these limitations by evaluating the effects of acute CAMKK2 disruption on autophagy induction in EOC spheroids using a functional, reporter-based assay.
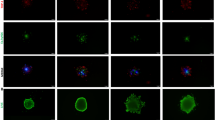
Loss of CAMKKβ attenuates autophagy activity in EOC spheroids. a Western blot analysis and densitometric quantification of p62 abundance and LC3BII: I ratio in OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO adherent cells (Adh) and spheroids (Sph). b Representative images and SPoRTS analysis of mCherry, EGFP, and mCherry/EGFP ratio from mCherry-EGFP-LC3B (autoR)-expressing OVCAR8 (OVCAR8-autoR) and HeyA8 (HeyA8-autoR) spheroids under control (siNT) and CAMKKβ knockdown (siCAMKKβ) conditions. c Western blot analysis and densitometric quantification of p62 abundance and LC3BII: I ratio in OVCAR8-autoR and HeyA8-autoR adherent cells and spheroids under control and CAMKKβ knockdown conditions. # indicates a non-specific band. Quantified western blot data represent mean normalized p62 abundance or mean LC3BII: I ratio ± SEM for n = 3 independent experiments. α-tubulin was used for normalization. For p62, data are expressed relative to the parental adherent sample, for which the mean was set to 1. Data were analyzed via two-way ANOVA and groups with shared lineage (e.g., parental Adh vs. parental Sph) or shared culture condition (e.g., parental Adh vs. CAMKK2 KO Adh) were compared using the Holm-Šídák method (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). For b, SPoRTS data represent mean ratio of integrated mCherry signal to integrated EGFP signal ± SEM for n = 3 independent experiments, each comprising 5 individual spheroids (15 spheroids in total). Data were analyzed via two-way repeated measures ANOVA (++p < 0.01, ++++p < 0.0001). Scale bars: 500 μm.
We have previously utilized the tandem fluorescent autophagy biosensor mCherry-EGFP-LC3B (autoR) to investigate autophagy in EOC spheroids22,23 and used the same approach here to assess the impact of transient CAMKKβ loss. Mechanistically, delivery of autoR to the acidic lysosome results in a preferential decrease in EGFP fluorescence due to the difference in acid sensitivity between EGFP and mCherry37. Accordingly, induction of autophagy in cells expressing autoR drives an increase in the ratio of mCherry signal/EGFP signal, which is readily detectable via fluorescence imaging22,23. Thus, we used siRNA-mediated gene silencing to target CAMKK2 in autoR-expressing OVCAR8 (OVCAR8-autoR) and HeyA8 (HeyA8-autoR) cells, seeded the resulting cells to form spheroids, and utilized a modified version of Spatial Profiling of Ratiometric Trends in Spheroids (SPoRTS) to monitor changes in spheroids’ mCherry/EGFP ratio over time38. Relative to the control condition (siNT), knockdown of CAMKK2 (siCAMKKβ) caused a striking reduction in mCherry/EGFP ratio in both OVCAR8-autoR and HeyA8-autoR spheroids (Fig. 2B), confirming that acute CAMKKβ loss was sufficient to attenuate autophagy in EOC spheroids. Of note, siCAMKKβ also elicited a marked change in the spatial regulation of autophagy within HeyA8-autoR spheroids, suggesting CAMKKβ is particularly relevant for autophagy induction in the spheroid core. (Supplementary Fig. S3). Despite this functional confirmation that CAMKKβ loss diminished autophagic flux in spheroids, analysis of p62 abundance and LC3B processing revealed no differences between siNT- and siCAMKKβ-treated spheroids (Fig. 2C), mirroring our findings from CAMKK2 KO spheroids (Fig. 2A). Taken together, these results demonstrate that CAMKKβ is required for autophagy in EOC spheroids and imply a regulatory role for this kinase or its effectors downstream of p62 and LC3B in the autophagy pathway.
Loss of CAMKKβ reduces EOC spheroid viability
In a previous report, we showed that treatment with STO-609 blocks autophagy and reduces cell viability in EOC spheroids, suggesting that CAMKKβ could be a feasible target for therapeutic targeting in advanced EOC22. However, as noted earlier, STO-609 has the potential to cause off-target effects33,34. Since genetic targeting of CAMKKβ recapitulated our previous findings with respect to autophagy, we sought to determine if the same was true for spheroid viability. To assess spheroid viability, trypan blue exclusion cell counting was used to compare the number of viable cells in parental and CAMKK2 KO spheroids at time points of 3, 5, and 7 days after seeding. CAMKKβ loss was associated with a significant reduction in viability of OVCAR8 spheroids at the 7-day time point, and in HeyA8 spheroids at each time point (Fig. 3A), corroborating our previous findings. Notably, we also found that the proliferative rate of CAMKK2 KO cells was slower than that of parental cells under adherent conditions (Supplementary Fig. S4). Taken together, these findings highlight a crucial role for CAMKKβ in the growth and viability of EOC cells, regardless of culture condition.
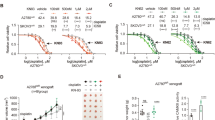
Loss of CAMKKβ reduces EOC spheroid viability and has cell line-dependent effects on spheroid invasion and cell migration. a Representative images and viable cell number of OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO spheroids at time points of 3, 5, and 7 days after seeding to spheroid culture conditions, as determined via trypan blue exclusion cell counting. b Representative images of OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO spheroids prior to reattachment (starting size), ZT-GFP mesothelial cell monolayers 48 h after spheroid reattachment was initiated (cleared area), and quantification of normalized cleared area. c Representative images and quantification of scratch closure over time in OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO monolayers. For a, data represent mean cell count after 3, 5, or 7 days in spheroid culture ± SEM for n = 4 independent experiments. Data were analyzed via unpaired two-tailed student’s t-tests and the Holm-Šídák method was used to control for multiple comparisons (*p < 0.05, ***p < 0.001, ****p < 0.0001). For b, data represent mean normalized cleared area ± SEM for 3 independent experiments, each comprising 8 individual spheroids (24 spheroids in total). Cleared area was normalized on a per-spheroid basis to the cross-sectional area of the spheroid prior to initiating reattachment. Data were analyzed via two-tailed student’s t-tests. For c, data represent mean scratch closure as a percentage of the initial scratch area ± SEM for 3 independent experiments, each comprising 3 fields of view from each of 2 wells (18 fields of view in total). Data were analyzed via two-way repeated measures ANOVA (++p < 0.01). Scale bars: a, c 1 mm, b 500 μm (equal scale for starting size and cleared area).
We next sought to determine if the impacts of CAMKKβ loss on autophagy and spheroid viability were associated with changes in the metastatic properties of spheroids. Spheroid invasion following reattachment is thought to be a key element of EOC metastasis11,15,16 so we assessed the effects of CAMKKβ loss on the invasive capacity of spheroids using an established mesothelial clearance assay16,39. In the present study, spheroids were allowed to invade monolayers of GFP labeled ZT human mesothelial cells and the cleared area was measured 48 h later. After normalizing cleared area to spheroid starting size to account for differences in spheroid viability due to loss of CAMKKβ (as recommended39), we identified a strong trend toward decreased invasive potential in OVCAR8 CAMKK2 KO spheroids but observed no effect in HeyA8 CAMKK2 KO spheroids (Fig. 3B). We reasoned these outcomes could reflect general differences in cell motility due to loss of CAMKKβ. Scratch wound closure analysis supported this hypothesis, as cell motility was significantly compromised in OVCAR8 CAMKK2 KO cells but not in HeyA8 CAMKK2 KO cells (Fig. 3C). Taken together, these findings suggest that loss of CAMKKβ is associated with moderately diminished metastatic potential in OVCAR8, but not in HeyA8.
Loss of CAMKK2 slows EOC tumour growth in vivo
The results of our in vitro work highlight CAMKKβ as an important factor in the induction of autophagy and maintenance of viability in EOC spheroids, and indicate a potential role for CAMKKβ in mediating motility-based metastatic processes in this context. Thus, we next assessed the impacts of CAMKKβ ablation on tumour growth in vivo. STO-609 has been used to inhibit CAMKKβ in previous xenograft studies, wherein STO-609 treatment has slowed or arrested tumour growth40,41,42,43. While promising, off-target activity of STO-609 (noted earlier33,34) could have contributed to these results, requiring confirmation using more target-specific approaches. This uncertainty has largely been addressed through xenograft studies of non-ovarian cancers (e.g., breast, liver, prostate) in which CAMKK2 was ablated genetically42,43,44 but this has not been applied in a xenograft model of EOC.
To assess the impacts of CAMKKβ loss on disease progression in vivo, we utilized a mouse intraperitoneal (IP) xenograft model, which mimics the exfoliation-reattachment mechanism of EOC metastasis and produces tumours with histology and distribution similar to that observed in patients with advanced EOC8,45. Xenografts were established via IP injection of parental or CAMKK2 KO cells stably expressing firefly luciferase and tdTomato, enabling non-invasive disease monitoring using bioluminescence imaging (BLI). BLI analysis revealed that OVCAR8 xenografts grew at similar rates irrespective of CAMKK2 status, whereas HeyA8-derived tumours lacking CAMKKβ grew significantly slower than CAMKKβ-intact tumours (Fig. 4A). Notably, these outcomes paralleled the results of our in vitro analyses in which growth deficits due to loss of CAMKKβ were more pronounced in HeyA8 than in OVCAR8 (Fig. 3A, Supplementary Fig. S4).
Loss of CAMKKβ has context-dependent effects on disease progression in xenograft models. a Representative images depicting radiance and quantification of total photon flux obtained via in vivo bioluminescence imaging (BLI) of NOD SCID mice intraperitoneally engrafted with firefly luciferase-expressing OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, or HeyA8 CAMKK2 KO cells. b Survival analysis of hosts indicated in (a). For a, data represent mean total photon flux ± SEM for n = 8 (OVCAR8 parental, HeyA8 parental, and HeyA8 CAMKK2 KO) or n = 7 (OVCAR8 CAMKK2 KO) mice. BLI data were analyzed via two-way repeated measures ANOVA (+++p < 0.001) and the Holm-Šídák method was used to compare individual time points (*p < 0.05, **p < 0.01). For b, survival curves were compared using the log-rank test.
Based on BLI analysis, we did not expect survival to differ between hosts bearing OVCAR8 parental and CAMKK2 KO xenografts but anticipated a survival benefit for hosts bearing HeyA8-CAMKK2 KO xenografts in comparison to those bearing parental xenografts. However, we were surprised to find that loss of CAMKKβ was not associated with a difference in survival among either group (Fig. 4B). It is possible that differences in tumour localization underly the absence of a survival benefit in hosts bearing HeyA8-CAMKK2 KO-derived tumours, as miliary disease and tumours on the reproductive tract were more common in this cohort than in hosts bearing CAMKKβ-intact tumours (Supplementary Fig. S5). Regardless of the underlying mechanism, these findings illustrate a context-dependent role for CAMKKβ in the progression of advanced EOC that warrants further investigation in future studies.
Discussion
There have been few treatment advances for patients with EOC since the current standard of care was established in the 1990s1,3,4,5,6 constituting a marked disparity between EOC and many other cancers for which the treatment landscape has changed vastly in the same period (e.g., implementation of immunotherapy46,47,48 which is notoriously ineffective in EOC49,50,51). Thus, the factors that distinguish EOC from other cancers, like its spheroid-mediated metastatic mechanism4,8,9,10 must be taken into consideration when pursuing novel treatment options.
EOC spheroids maintain metastatic capacity by upregulating various stress responses8,17 many of which influence cell metabolism18,20,21,22,23,24,25,52,53,54,55,56,57. Thus, metabolic reprogramming is a key area for potential intervention in the context of metastatic EOC. AMPK activation is among the metabolic stress responses induced in EOC spheroids20,21,22 and we previously used a pharmacological approach to gather evidence implicating CAMKKβ as the primary mediator of AMPK activation in this setting21,22. Here, we used CRISPR–Cas9-mediated genome editing to improve target specificity and show conclusively that CAMKKβ is the primary mediator of AMPK activation in EOC spheroids. Of note, it has been shown that LKB1—the canonical activator of AMPK—can also contribute to AMPK activation upon matrix detachment32 presenting an opportunity to utilize experimental models with double knockout of LKB1 and CAMKKβ in future work. Indeed, models lacking both LKB1 and CAMKKβ will enable an even deeper assessment of the importance of AMPK signaling in EOC spheroids.
AMPK regulates cellular metabolism through a vast array of downstream effectors26,27,28,29 and CAMKK2 KO EOC spheroids exhibited reduced signaling from AMPK to its targets in the present study. These targets included ULK1 – a key regulator of autophagy, a stress response we and others have previously shown is required to maintain EOC spheroid viability22,23,24,58. Consistent with this, we have shown that reduced activation of ULK1 in CAMKK2 KO spheroids was associated with a loss of autophagy activity and a reduction in spheroid viability in the current study. Notably, neither short- nor long-term CAMKKβ loss elicited changes in the regulation of p62 or LC3B in spheroids, implying CAMKKβ loss may have reduced autophagic flux by inhibiting autophagy at a later step. Consistent with this hypothesis, CAMKKβ has been linked to lysosomal acidification59 and AMPK has been linked to lysosomal biogenesis60. A deficiency in either of these processes represents a plausible mechanism for downstream inhibition of autophagy in CAMKK2 KO spheroids, but further studies will be required to discern the mechanism of action at play in this setting. Altogether, these findings support the assertion that autophagy is a critical requirement for EOC spheroid survival while highlighting CAMKKβ and AMPK as potential therapeutic targets in the setting of advanced EOC.
Fatty acid synthesis and protein translation are also regulated by AMPK via ACC and Raptor, respectively26,27,28,29 and AMPK-mediated phosphorylation of both species was blunted in CAMKK2 KO spheroids. Though we did not investigate the functional impacts of CAMKKβ loss on AMPK-regulated biosynthetic pathways in this study, our findings provide an actionable foundation for future studies designed to characterize and exploit metabolic vulnerabilities that may arise in response to CAMKKβ inhibition. For example, AMPK-mediated phosphorylation of ACCS79 promotes fatty acid β-oxidation61 which has been identified as a metabolic dependency in EOC spheroids62. Hypothetically, precluding fatty acid utilization in EOC spheroids via inhibition of CAMKKβ represents a means of forcing reliance on alternative pathways, establishing a base for combination strategies targeting collateral lethality.
Throughout this study, the impacts of CAMKKβ loss on growth potential and viability of cells, spheroids, and tumours were more severe for HeyA8 than for OVCAR8, consistent with findings from our previous study in which we inhibited CAMKKβ using STO-60922. However, molecular analyses showed that the effects of CAMKKβ loss on AMPK and its downstream targets were consistent between OVCAR8 and HeyA8, yielding no clear explanation for the noted differences in growth potential and viability. Importantly, we have focused on CAMKKβ solely in relation to AMPK signaling, owing to activation of AMPK in EOC spheroids and its known role in the regulation of autophagy. However, CAMKKβ regulates targets other than AMPK as well. Indeed, CAMKKβ mediates the activation of protein kinase B (AKT), calcium/calmodulin-dependent protein kinase I (CAMKI), and CAMKIV63,64,65—each of which can promote cell proliferation43,66,67,68. In the context of our findings, this suggests HeyA8 may be more reliant on CAMKKβ effectors for proliferation and survival than OVCAR8, representing a potential mechanism for our results and rationale for future studies.
The differences we observed in tumour site distribution between parental and CAMKK2 KO tumours suggest CAMKKβ facilitates EOC survival and growth within certain metastatic niches in a cell line-specific manner. This property may explain why there was no survival benefit for hosts engrafted with HeyA8 CAMKK2 KO cells (versus HeyA8 parental cells) despite a drastic reduction in tumour burden as measured by BLI. Namely, mesenteric tumours, which were more prevalent in HeyA8 CAMKK2 KO hosts, were associated with symptoms of bowel obstruction (enlarged stomach, loss of body weight) and the occurrence of humane endpoints at relatively low tumour burden. Corroborating these findings, CAMKKβ and its effectors have been shown to influence tumour distribution in previous studies. For instance, CAMKKβ loss altered tumour site distribution in an orthotopic xenograft model of triple-negative breast cancer42 and phosphorylation of ACC by AMPK and salt-inducible kinase 2 has been shown to promote omental metastasis in xenograft models of EOC69. Altogether, it is clear that the roles of CAMKKβ and AMPK in cancer growth and metastasis are complex and context-dependent.
While CAMKKβ ablation impacted growth and viability to a greater extent in HeyA8 than in OVCAR8, the opposite was true for migration and invasion. Notably, roles have been identified for CAMKKβ and its effectors in these processes70 and recent studies linking CAMKKβ with regulation of the actin cytoskeleton have elucidated a potential mechanism underlying our observations42,71. Therefore, we reason that differential effects of CAMKKβ loss on migration and invasion in OVCAR8 and HeyA8 likely reflect cell-line specific differences in the regulation of these processes. This hypothesis is supported by findings from our recent study, in which STRADA knockdown attenuated spheroid invasion (mesothelial clearance) more drastically for HeyA8 spheroids than for OVCAR8 spheroids72. Taken together, our findings suggest that in vitro growth potential may be a better predictor of disease trajectory for the IP xenograft model than in vitro migratory and invasive capacity, providing helpful context for planning similar xenograft studies in future.
Finally, there is evidence to suggest whole-body inhibition of CAMKKβ (i.e., untargeted delivery of a pharmacological inhibitor) may have advantages over disease-selective targeting. Indeed, studies employing immune-intact genetically engineered and syngeneic mouse models have reported reduced rates of disease progression in Camkk2−/− versus wildtype cohorts, linking these outcomes to potentiation of the antitumour immune response41,73. However, despite these promising findings, certain immune cells increase CAMKKβ expression to maintain their tumour infiltrating capacity, implying CAMKKβ inhibition could negatively impact antitumour immunity as well74. Nonetheless, this approach could be especially impactful if capable of sensitizing EOC to immunotherapeutic treatment options, rationalizing further investigation.
It is worth noting that this work drew comparisons between CAMKK2 KO cell lines and their respective parental cell lines, the latter of which were not exposed to Cas9 (e.g., using non-targeting guides). Thus, the potential for off-target Cas9 activity is a limitation that should be considered when interpreting the findings of this study. However, potential off-target effects of Cas9 were mitigated by ensuring its expression was transient and without selective pressure, and that multiple knockout clones were pooled to generate each CAMKK2 KO cell line, aligning with established best practices75,76. Thus, while our approach cannot guarantee the absence of clone-specific off-target effects, we expect the potential impact of such effects to be minimized.
In summary, we have confirmed that CAMKKβ is the primary mediator of AMPK activation in EOC spheroids and shown that loss of CAMKKβ diminishes EOC spheroid growth potential and viability. Deleterious effects of CAMKKβ ablation were present both in vitro and in vivo, and effect size disparities between cell lines likely reflect diminished signaling not only through AMPK, but through other effectors of CAMKKβ as well. Altogether, our findings demonstrate the importance of CAMKKβ signaling in advanced EOC and provide the basis for studies further interrogating the effects of CAMKKβ loss on the specific signaling pathways regulated by its various effectors.
Materials and methods
Cell lines and culture conditions
The OVCAR8 and HeyA8 cell lines were obtained from the MD Anderson Cancer Center. Both cell lines and their derivatives were cultured in RPMI-1640 medium (Gibco, 22400089) supplemented with 10% fetal bovine serum (FBS) (Wisent, 098–150). GFP-labeled ZT human mesothelial cells were a gift from M. Iwanicki (Stevens Institute of Technology, NJ, USA) and were cultured in DMEM/F12 medium (Gibco, 11330032) supplemented with 10% FBS. Cells were maintained in a humidified atmosphere of 5% CO2 at 37 °C. All cell lines were authenticated by short tandem repeat analysis performed by The Centre for Applied Genomics (The Hospital for Sick Children, Toronto, Ontario, Canada) and routinely tested for mycoplasma in-house using the Universal Mycoplasma Detection Kit (ATCC, 30–1012 K). Spheroid formation was achieved passively by seeding single-cell suspensions into ultra-low attachment (ULA)-treated culture plates (Corning, various; see Supplementary Table S1).
Plasmids
pCDH-EF1-Luc2-P2A-tdTomato was a gift from Kazuhiro Oka (Addgene plasmid #72486; RRID: Addgene_72486). pBABE-puro.mCherry-EGFP-LC3B was a gift from Jayanta Debnath (Addgene plasmid #22418; RRID: Addgene_22418)77. To facilitate antibiotic selection and place cDNA expression under the control of a CMV promoter, the mCherry-EGFP-LC3B cDNA was cloned into pcDNA3 by using the Q5 Site-Directed Mutagenesis Kit (New England Biolabs, E0554) to perform the following mutagenesis reactions (mutagenic primer sequences provided below): (1) convert the unique NgoMIV restriction site in pBABE-puro.mCherry-EGFP-LC3B to a HindIII site using primer pair A/B, (2) convert the unique SalI restriction site in pBABE-puro.mCherry-EGFP-LC3B to a BamHI site using primer pair C/D. Once conversion of the restriction sites had been confirmed via diagnostic digest, the mCherry-EGFP-LC3B cDNA was excised from the mutated version of pBABE-puro.mCherry-EGFP-LC3B using HindIII and BamHI, then ligated into pcDNA3 that had been digested with the same enzymes to yield pcDNA3.mCherry-EGFP-LC3B. Agarose gel extractions were performed using the QIAEX II Gel Extraction Kit (Qiagen, 20021) and ligations were done using T4 DNA Ligase (New England Biolabs, M0202) according to the manufacturers’ protocols.
Mutagenic primer pairs (5’➝3’)
Generation of CAMKK2 KO cell lines
Parental OVCAR8 and HeyA8 cells were seeded to 6-well tissue culture-treated plates as indicated in Supplementary Table S1 and medium was changed the following day. Two days after seeding, cells were transfected with CAMKKβ CRISPR/Cas9 KO plasmid (h) (Santa Cruz Biotechnology, sc-400928) using Lipofectamine LTX with PLUS (ThermoFisher, A12621) according to the manufacturer’s suggested protocol for transfection in a 6-well plate (9 µL/well LTX reagent, 2.5 µL/well PLUS reagent). Two days after transfection, cells were subjected to fluorescence-activated cell sorting at the London Regional Flow Cytometry Facility (LRFCF; Robarts Research Institute, London, Ontario, Canada) and sorted to 96-well tissue culture-treated plates at a density of 1 cell/well. When generating the CAMKK2 KO cell lines, multiple widely-accepted strategies were utilized to minimize the likelihood that off-target effects of Cas9 expression could influence the overall phenotype75,76. Namely, clones with stable GFP expression were excluded from consideration to ensure the final CAMKK2 KO cell lines only contained cells that been exposed to Cas9 transiently and without selective pressure, and after verifying CAMKKβ ablation in CAMKK2 KO clones by western blot, equal proportions of at least 4 independent clones were pooled to generate each cell line.
Generation of firefly luciferase/tdTomato-expressing cell lines
OVCAR8 parental, OVCAR8 CAMKK2 KO, HeyA8 parental, and HeyA8 CAMKK2 KO cells were transfected with pCDH-EF1-Luc2-P2A-tdTomato using the transfection protocol described under Generation of CAMKK2 KO cell lines. Populations stably expressing firefly luciferase and tdTomato (-luc2/tdT) were established through two sequential rounds of fluorescence-activated cell sorting at the LRFCF. All xenograft experiments were conducted using these cell lines, but -luc2/tdT designations were omitted from the associated text for readability.
Generation of -autoR cell lines
OVCAR8 parental and HeyA8 parental cells were transfected with pcDNA3.mCherry-EGFP-LC3B using the transfection protocol described under Generation of CAMKK2 KO cell lines. Stable populations were established via selection with 500 µg/mL G418 sulfate (Wisent, 450 − 130) for approximately one week, beginning two days after transfection. Since many cells from each population failed to maintain transgene expression after G418 selection, a single round of fluorescence-activated cell sorting was performed at the London Regional Flow Cytometry Facility (LRFCF; Robarts Research Institute, London, Ontario, Canada) to enrich each population for transgene-positive cells.
SPoRTS analysis
Time course fluorescence images of OVCAR8-autoR and HeyA8-autoR spheroids (mCherry and EGFP channels) were captured using an IncuCyte S3 live cell analysis instrument with exposure times of 400 ms (mCherry) and 300 ms (EGFP). Analyses of whole-spheroid mCherry/EGFP fluorescence ratio (Fig. 2B) were conducted using a modified version of SPoRTS, the automated ratiometric image analysis platform we published recently38. Briefly, this modified version of SPoRTS maintains identical frameworks for image processing, thresholding, and data aggregation as the original, but the spatial component has been removed. Modified SPoRTS was used to calculate whole-spheroid mCherry/EGFP ratios based on total spheroid fluorescence from the respective channels (i.e., total mCherry fluorescence divided by total EGFP fluorescence). Spatial analyses of mCherry-EGFP ratio (Supplementary Fig. S3) were conducted using the comprehensive method described in the original version of SPoRTS. Default settings for SPoRTS were used for all analyses in this study. Both versions of SPoRTS are available at https://github.com/mjborrelli/SPoRTS.
siRNA-mediated gene silencing
Cells were seeded to 6-well tissue culture-treated plates as indicated in Supplementary Table S1. The next day, DharmaFECT1 Transfection Reagent (Horizon Discovery, T-2001) was used at a final dilution of 1:500 to deliver ON-TARGETplus siRNAs at a final concentration of 20 nM in a volume of 2 mL. The day after transfection, 3 mL of medium was added to each well to bring the total volume to 5 mL. The following day, cells were lifted, counted, and seeded for experiments. The following ON-TARGETplus siRNA preparations were purchased from Horizon Discovery: Non-targeting Pool (siNT, D-001810-10) and CAMKK2 SMARTpool (siCAMKKβ, L-004842-00).
Preparation of protein lysates
Sample lysis was performed using modified RIPA buffer: 50 mM HEPES (pH 7.4), 150 mM NaCl, 10% (v/v) glycerol, 1.5 mM MgCl2, 0.1 mM EGTA, 1% (w/v) deoxycholic acid, 1% (v/v) Triton X-100, 0.1% (w/v) SDS, 1 mM Na3VO4, 10 mM NaF, 1 mM PMSF, 10 mM β-glycerophosphate, 10 mm Na4P2O7, 1× SIGMAFAST protease inhibitor cocktail (Sigma, S8820). For adherent samples, cells were seeded to 100 mm tissue culture-treated plates as indicated in Supplementary Table S1 and cultured under standard conditions for 72 h. Plates were then transferred to ice and cells were washed once with ice-cold 1× Dulbecco’s phosphate-buffered saline (DPBS), then lysed by scraping in modified RIPA buffer. For spheroid samples, cells were seeded to 6-well ULA-treated plates as indicated in Supplementary Table S1 and cultured under standard conditions for 72 h. Spheroids were then collected in conical tubes on ice, washed once with ice-cold 1× DPBS, and lysed in modified RIPA buffer. Crude lysates were kept on ice and vortexed periodically for 30 min, then sonicated for a single 10 s pulse using a Misonix XL-2000 probe sonicator set to minimum power (~ 1 W). Lysates were then centrifuged at 21,000 × g for 20 min at 4 ºC and protein concentrations in the resulting supernatants were determined using the Bio-Rad Protein Assay (Bio-Rad, #5000006) according to the manufacturer’s instructions.
Immunoblotting
Immunoblotting was performed using the Bio-Rad Mini-PROTEAN II Electrophoresis System, according to the manufacturer’s instructions, using gels cast in-house. For each sample, 20 µg of protein was resolved via SDS-PAGE (6% acrylamide for p-ACCS79 and ACC, 8% acrylamide for all other targets) and transferred to PVDF membranes (MilliporeSigma, IPVH00010). Membranes were blocked for 1 h with either 5% skim milk or 5% bovine serum albumin (BSA; BioShop, ALB001) in TBS-T (137 mM NaCl, 20 mM Tris, 0.1% Tween-20, pH 7.6), washed extensively with TBS-T, then incubated overnight with primary antibodies diluted in the supplier’s recommended diluent at 4 ºC with gentle rocking (primary antibody conditions are provided in Supplementary Table S2). The following day, each membrane was incubated with the species-appropriate HRP-conjugated secondary antibody (Cytiva, NA931 or NA934) at a dilution of 1:10,000 in the same diluent as the primary antibody for 1 h under ambient conditions with gentle rocking. Immunoreactive bands were visualized with a ChemiDoc Imaging System (Bio-Rad, 12003153) using the ECL method with Immobilon Forte Western HRP substrate (MilliporeSigma, WBLUF0500). When necessary, the “Mild stripping” protocol from Abcam was used to strip membranes for sequential probing. Densitometry analysis was done using Image Lab software (Bio-Rad, ver. 6.1.0).
Trypan blue cell counting
Cells were seeded to 24-well ULA-coated plates as described in Supplementary Table S1. At the indicated time points, spheroids were transferred 1:1 to microcentrifuge tubes and were washed once with 1× DPBS. After removing as much residual DPBS as possible, 50 µL of trypsin/EDTA (0.25%/2.21 mM; Wisent, 325 − 043) was added to each well, spheroids were resuspended by pipetting up and down, and tubes were placed in a 37 ºC water bath to dissociate. After incubating for 20 min, spheroids were pipetted up and down vigorously to ensure complete dissociation to a single-cell suspension. Trypsin was then neutralized by adding 50 µL of FBS to each tube and pipetting to mix. Finally, 100 µL of trypan blue solution (0.4%; Gibco, 15250061) was added to each tube with gentle mixing, and cell counts were determined using a TC20 Automated Cell Counter (Bio-Rad, 1450102).
Doubling time analysis
Cells were seeded into 96-well tissue culture-treated plates as described in Supplementary Table S1 and phase contrast images were acquired using an IncuCyte ZOOM live cell analysis instrument at 3-hour intervals for a total of 72 h (10×/0.30 objective, 4 fields of view per well). Confluence was determined using IncuCyte software (ver. 2018 A) and doubling times were determined using GraphPad Prism (ver. 10.2.3; RRID: SCR_002798) via least squares regression.
Scratch wound closure
Cells were seeded into 24-well tissue culture-treated plates as indicated in Supplementary Table S1. Two days later, a P200 tip was used to create a uniform scratch in the cell monolayer within each well. Wells were then washed once with 1 × DPBS to remove debris and fresh medium was added. Beginning immediately thereafter, phase contrast images of each well were captured using an IncuCyte S3 System at 2-hour intervals for a total of 72 h (4× objective, 3 fields of view per well). Scratch area was identified using the Trainable Weka Segmentation plugin (RRID: SCR_001214)78 for ImageJ (ver. 1.54k; RRID: SCR_002285) and confirmed by visual inspection. The percentage of the initial scratch area that had been covered by cells at each time point for each field of view was then calculated to determine scratch closure.
Mesothelial clearance
OVCAR8 and HeyA8 cells were seeded to 96-well round bottom ULA-treated plates as described in Supplementary Table S1. The next day, 96-well tissue culture-treated plates (Falcon, 353072) were coated with 1.5 µg/well rat tail collagen (30 µg/mL in 70% ethanol, 50 µL/well) (Gibco, A1048301). After leaving the plates to coat for 3 h under standard tissue culture conditions, they were rinsed once with 1× DPBS and stored in 1× DPBS at 4 ºC overnight. The following day, GFP-labeled ZT cells were seeded into the coated plates as indicated in Supplementary Table S1. The day after (approx. 24 h after seeding ZT cells and 72 h after seeding spheroids), OVCAR8 and HeyA8 spheroids were imaged using an IncuCyte S3 System then transferred 1:1 to wells containing ZT monolayers. Fluorescent signal from the GFP-labeled ZT monolayer was imaged using the IncuCyte S3 48 h after spheroids were transferred, and the area of the monolayer than had been cleared by the spheroid was quantified using the fuzzy select tool in the GNU Image Manipulation Program (GIMP) (ver. 2.10.38; RRID: SCR_003182). The same method was used to determine the pre-transfer cross sectional area of spheroids. The ZT monolayer area cleared by the spheroid was then divided by the pre-transfer cross sectional area of the respective spheroid to calculate the normalized cleared area.
Xenograft studies
All animal experiments were conducted according to the approved Animal Use Protocol (2023 − 138) by the Animal Care Committee of Western University (London, Ontario, Canada). As such, all experiments involving animals were performed in accordance with ARRIVE guidelines. Female NOD SCID mice (n = 7–8 per cohort, 8–10 weeks old; Charles River Laboratories, strain code 304, RRID: IMSR_CRL:394) were inoculated with 150 µL of DPBS containing 2.0 × 106 cells (OVCAR8-derived luc2/tdT cell lines) or 1.0 × 106 (HeyA8-derived luc2/tdT cell lines) cells via intraperitoneal injection. Disease progression was monitored via BLI using an IVIS Lumina S5 Imaging System. BLI measurements were acquired once weekly, beginning one week after xenograft injection. For each round of BLI, mice were injected with 75 mg/kg of D-luciferin (Revvity, 122799) in sterile filtered 1× DPBS (50 mg/mL) 5 min prior to initiating measurements. Living Image software (ver. 4.7.3; RRID: SCR_014247) was used to quantify bioluminescence. For survival analysis, mice were monitored daily following xenograft injection and euthanized by CO2 asphyxiation using standard criteria for humane endpoints (i.e., impaired breathing, hunched posture, lethargy, excessive ascites, extreme weight loss). Chow (Envigo, 2919) and water were provided ad libitum.
Light microscopy
Brightfield images of spheroids were captured using a Leica DMI 4000 B inverted microscope equipped with a Leica DFC450 camera (2.5×/0.07 objective, 2560 × 1920 pixels, 2.2 μm/pixel).
Approval statement
Ethical approval for animal experiments (protocol # 2023 − 138) was obtained from the Animal Care Committee of Western University (London, ON, Canada). The study was conducted in strict accordance with ARRIVE 2.0 guidelines to ensure the ethical and accurate reporting of animal research. All methods were performed in accordance with the relevant guidelines and regulations.
Statistical analysis
In all instances, data represent mean ± SEM. Statistical analyses were performed using GraphPad Prism (ver. 10.2.3; RRID: SCR_002798). The statistical tests used for individual analyses have been identified in the respective figure legends, and all statistical testing was two-tailed with the threshold for significance set at p < 0.05.
Data availability
All data from this study are available from the corresponding author upon reasonable request.
References
Lheureux, S., Gourley, C., Vergote, I. & Oza, A. M. Epithelial ovarian cancer. Lancet 393, 1240–1253 (2019).
Torre, L. A. et al. Ovarian Cancer Stat. 2018 CA Cancer J. Clin 68, 284–296 (2018).
Cannistra, S. A. Cancer of the ovary. N Engl. J. Med. 351, 2519–2529 (2004).
Lengyel, E. Ovarian Cancer development and metastasis. Am. J. Pathol. 177, 1053–1064 (2010).
Piccart, M. J. et al. Randomized intergroup trial of Cisplatin–Paclitaxel versus Cisplatin–Cyclophosphamide in women with advanced epithelial ovarian cancer: Three-Year results. JNCI J. Natl. Cancer Inst. 92, 699–708 (2000).
McGuire, W. P. et al. Cyclophosphamide and cisplatin compared with Paclitaxel and cisplatin in patients with stage III and stage IV ovarian Cancer. N Engl. J. Med. 334, 1–6 (1996).
Vaughan, S. et al. Rethinking ovarian cancer: recommendations for improving outcomes. Nat. Rev. Cancer. 11, 719–725 (2011).
Lengyel, E. et al. Epithelial ovarian cancer experimental models. Oncogene 33, 3619–3633 (2014).
Mikuła-Pietrasik, J., Uruski, P., Tykarski, A. & Książek, K. The peritoneal ‘soil’ for a cancerous ‘seed’: a comprehensive review of the pathogenesis of intraperitoneal cancer metastases. Cell. Mol. Life Sci. CMLS. 75, 509–525 (2018).
Shield, K., Ackland, M. L., Ahmed, N. & Rice, G. E. Multicellular spheroids in ovarian cancer metastases: biology and pathology. Gynecol. Oncol. 113, 143–148 (2009).
Burleson, K. M. et al. Ovarian carcinoma Ascites spheroids adhere to extracellular matrix components and mesothelial cell monolayers. Gynecol. Oncol. 93, 170–181 (2004).
Davidson, B. et al. Expression of Membrane-Type 1, 2, and 3 matrix metalloproteinases messenger RNA in ovarian carcinoma cells in serous effusions. Am. J. Clin. Pathol. 115, 517–524 (2001).
Tarin, D. et al. Mechanisms of human tumor metastasis studied in patients with peritoneovenous shunts. Cancer Res. 44, 3584–3592 (1984).
Miceska, S., Škof, E., Gašljević, G. & Kloboves-Prevodnik, V. Morphological and immunocytochemical characterization of tumor spheroids in Ascites from High-Grade serous carcinoma. Cells 12, 2390 (2023).
Burleson, K. M., Hansen, L. K. & Skubitz, A. P. N. Ovarian carcinoma spheroids disaggregate on type I collagen and invade live human mesothelial cell monolayers. Clin. Exp. Metastasis. 21, 685–697 (2005).
Iwanicki, M. P. et al. Ovarian Cancer spheroids use Myosin-Generated force to clear the mesothelium. Cancer Discov. 1, 144–157 (2011).
Cai, Q., Yan, L. & Xu, Y. Anoikis resistance is a critical feature of highly aggressive ovarian cancer cells. Oncogene 34, 3315–3324 (2015).
Correa, R. J. M., Peart, T., Valdes, Y. R., DiMattia, G. E. & Shepherd, T. G. Modulation of AKT activity is associated with reversible dormancy in ascites-derived epithelial ovarian cancer spheroids. Carcinogenesis 33, 49–58 (2012).
Buensuceso, A., Borrelli, M. J., Ramos Valdés, Y. & Shepherd, T. G. Reversible downregulation of MYC in a spheroid model of metastatic epithelial ovarian cancer. Cancer Gene Ther. 1–12. https://doi.org/10.1038/s41417-024-00850-z (2024).
Peart, T. et al. Intact LKB1 activity is required for survival of dormant ovarian cancer spheroids. Oncotarget 6, 22424–22438 (2015).
Buensuceso, A., Ramos-Valdes, Y., DiMattia, G. E. & Shepherd, T. G. AMPK-Independent LKB1 activity is required for efficient epithelial ovarian Cancer metastasis. Mol. Cancer Res. MCR. 18, 488–500 (2020).
Laski, J. et al. Activated CAMKKβ-AMPK signaling promotes autophagy in a spheroid model of ovarian tumour metastasis. J. Ovarian Res. 13, 58 (2020).
Singha, B. et al. Inhibiting ULK1 kinase decreases autophagy and cell viability in high-grade serous ovarian cancer spheroids. Am. J. Cancer Res. 10, 1384–1399 (2020).
Correa, R. J. M. et al. Combination of AKT Inhibition with autophagy Blockade effectively reduces ascites-derived ovarian cancer cell viability. Carcinogenesis 35, 1951–1961 (2014).
Frederick, M. I. et al. Metabolic adaptation in epithelial ovarian cancer metastasis. Biochim. Biophys. Acta BBA - Mol. Basis Dis. 1870, 167312 (2024).
Herzig, S. & Shaw, R. J. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell. Biol. 19, 121–135 (2018).
Trefts, E. & Shaw, R. J. AMPK: restoring metabolic homeostasis over space and time. Mol. Cell. 81, 3677–3690 (2021).
Steinberg, G. R. & Carling, D. AMP-activated protein kinase: the current landscape for drug development. Nat. Rev. Drug Discov. 18, 527–551 (2019).
Steinberg, G. R. & Hardie, D. G. New insights into activation and function of the AMPK. Nat. Rev. Mol. Cell. Biol. 24, 255–272 (2023).
Hirschhaeuser, F. et al. Multicellular tumor spheroids: an underestimated tool is catching up again. J. Biotechnol. 148, 3–15 (2010).
Jamieson, L. E., Harrison, D. J. & Campbell, C. J. Chemical analysis of multicellular tumour spheroids. Analyst 140, 3910–3920 (2015).
Sundararaman, A., Amirtham, U. & Rangarajan, A. Calcium-Oxidant signaling network regulates AMP-activated protein kinase (AMPK) activation upon matrix deprivation. J. Biol. Chem. 291, 14410–14429 (2016).
O’Byrne, S. N. et al. In depth analysis of kinase cross screening data to identify CAMKK2 inhibitory scaffolds. Molecules 25, 325 (2020).
Tokumitsu, H. et al. STO-609, a specific inhibitor of the Ca(2+)/calmodulin-dependent protein kinase kinase. J. Biol. Chem. 277, 15813–15818 (2002).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
J Klionsky, D. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17, 1–382 (2021).
Kimura, S., Noda, T. & Yoshimori, T. Dissection of the autophagosome maturation process by a novel reporter protein, tandem Fluorescent-Tagged LC3. Autophagy 3, 452–460 (2007).
Borrelli, M. J., Kolendowski, B., DiMattia, G. E. & Shepherd, T. G. Spatiotemporal analysis of ratiometric biosensors in live multicellular spheroids using sports. Cell. Rep. Methods. 5, 100987 (2025).
Davidowitz, R. A., Iwanicki, M. P. & Brugge, J. S. In vitro mesothelial clearance assay that models the early steps of ovarian Cancer metastasis. J. Vis. Exp. JoVE. 3888 https://doi.org/10.3791/3888 (2012).
Massie, C. E. et al. The androgen receptor fuels prostate cancer by regulating central metabolism and biosynthesis. EMBO J. 30, 2719–2733 (2011).
Racioppi, L. et al. CaMKK2 in myeloid cells is a key regulator of the immune-suppressive microenvironment in breast cancer. Nat. Commun. 10, 2450 (2019).
Mukherjee, D. et al. Targeting CaMKK2 inhibits actin cytoskeletal assembly to suppress Cancer metastasis. Cancer Res. 83, 2889–2907 (2023).
Lin, F. et al. The camKK2/camKIV relay is an essential regulator of hepatic cancer. Hepatology 62, 505–520 (2015).
Lin, C. et al. Inhibition of CAMKK2 impairs autophagy and castration-resistant prostate cancer via suppression of AMPK-ULK1 signaling. Oncogene 40, 1690–1705 (2021).
Mitra, A. et al. In vivo tumor growth of high-grade serous ovarian cancer cell lines. Gynecol. Oncol. 138, 372–377 (2015).
Blank, C. U. et al. Neoadjuvant nivolumab and ipilimumab in resectable stage III melanoma. N Engl. J. Med 391, 1696–1708 (2024).
Paz-Ares, L. et al. Pembrolizumab plus chemotherapy for squamous Non-Small-Cell lung Cancer. N Engl. J. Med. 379, 2040–2051 (2018).
The CRI Anna-Maria Kellen Clinical Accelerator team. Regulatory approval timeline of active immunotherapies. Cancer Res. Institute (2024). https://www.cancerresearch.org/regulatory-approval-timeline-of-active-immunotherapies
Odunsi, K. Immunotherapy in ovarian cancer. Ann. Oncol. 28, viii1–viii7 (2017).
Kandalaft, L. E., Odunsi, K. & Coukos, G. Immunotherapy in ovarian cancer: are we there yet?? J. Clin. Oncol. 37, 2460–2471 (2019).
Siminiak, N., Czepczyński, R., Zaborowski, M. P. & Iżycki, D. Immunotherapy in ovarian Cancer. Arch. Immunol. Ther. Exp. (Warsz). 70, 19 (2022).
Sato, M. et al. Spheroid cancer stem cells display reprogrammed metabolism and obtain energy by actively running the Tricarboxylic acid (TCA) cycle. Oncotarget 7, 33297–33305 (2016).
Frederick, M. I. et al. Proteomic and phosphoproteomic reprogramming in epithelial ovarian Cancer metastasis. Mol. Cell. Proteom. 22, 100660 (2023).
Kim, Y. S. et al. Context-dependent activation of SIRT3 is necessary for anchorage-independent survival and metastasis of ovarian cancer cells. Oncogene 39, 1619–1633 (2020).
Ding, Y. et al. Molecular characteristics and tumorigenicity of ascites-derived tumor cells: mitochondrial oxidative phosphorylation as a novel therapy target in ovarian cancer. Mol. Oncol. 15, 3578–3595 (2021).
Ward, A. S., Hall, C. N., Tree, M. O. & Kohtz, D. S. Spheroid architecture strongly enhances miR-221/222 expression and promotes oxidative phosphorylation in an ovarian cancer cell line through a mechanism that includes restriction of miR-9 expression. Mol. Biol. Rep. 51, 275 (2024).
Feng, X. et al. ASS1 enhances Anoikis resistance via AMPK/CPT1A-mediated fatty acid metabolism in ovarian Cancer. Cancer Lett. 217082 https://doi.org/10.1016/j.canlet.2024.217082 (2024).
Wang, Q. et al. Autophagy Is Indispensable for the Self-Renewal and Quiescence of Ovarian Cancer Spheroid Cells with Stem Cell-Like Properties. Oxid. Med. Cell. Longev. 7010472 (2018). (2018).
Stewart, L. M. et al. CaMKK2 facilitates Golgi-associated vesicle trafficking to sustain cancer cell proliferation. Cell. Death Dis. 12, 1–12 (2021).
Malik, N. et al. Induction of lysosomal and mitochondrial biogenesis by AMPK phosphorylation of FNIP1. Science 380, eabj5559 (2023).
Tomas, E. et al. Enhanced muscle fat oxidation and glucose transport by ACRP30 globular domain: Acetyl–CoA carboxylase Inhibition and AMP-activated protein kinase activation. Proc. Natl. Acad. Sci. 99, 16309–16313 (2002).
Sawyer, B. T. et al. Targeting fatty acid oxidation to promote Anoikis and inhibit ovarian Cancer progression. Mol. Cancer Res. 18, 1088–1098 (2020).
Marcelo, K. L., Means, A. R. & York, B. The Ca2+/Calmodulin/CaMKK2 axis: nature’s metabolic camshaft. Trends Endocrinol. Metab. 27, 706–718 (2016).
Gocher, A. M. et al. Akt activation by Ca2+/calmodulin-dependent protein kinase kinase 2 (CaMKK2) in ovarian cancer cells. J. Biol. Chem. 292, 14188–14204 (2017).
Dai, S. et al. Calcium/calmodulin-dependent protein kinase kinase 2 mediates pleiotropic effects of epidermal growth factor in cancer cells. Biochim. Biophys. Acta Mol. Cell. Res. 1869, 119252 (2022).
Schmitt, J. M., Wayman, G. A., Nozaki, N. & Soderling, T. R. Calcium activation of ERK mediated by calmodulin kinase I. J. Biol. Chem. 279, 24064–24072 (2004).
Kahl, C. R. & Means, A. R. Regulation of Cyclin D1/Cdk4 complexes by Calcium/Calmodulin-dependent protein kinase I. J. Biol. Chem. 279, 15411–15419 (2004).
Hoxhaj, G. & Manning, B. D. The PI3K–AKT network at the interface of oncogenic signalling and cancer metabolism. Nat. Rev. Cancer. 20, 74–88 (2020).
Miranda, F. et al. Salt-Inducible kinase 2 couples ovarian Cancer cell metabolism with survival at the Adipocyte-Rich metastatic niche. Cancer Cell. 30, 273–289 (2016).
Villalobo, A. & Berchtold, M. W. The role of calmodulin in tumor cell migration, invasiveness, and metastasis. Int. J. Mol. Sci. 21, 765 (2020).
Tojkander, S., Ciuba, K. & Lappalainen, P. CaMKK2 regulates mechanosensitive assembly of contractile actin stress fibers. Cell. Rep. 24, 11–19 (2018).
Trelford, C. B. et al. LKB1 and STRADα promote epithelial ovarian Cancer spheroid cell invasion. Cancers 16, 3726 (2024).
Huang, W. et al. Calcium/Calmodulin dependent protein kinase kinase 2 regulates the expansion of Tumor-Induced Myeloid-Derived suppressor cells. Front. Immunol. 12, 754083 (2021).
Juras, P. K. et al. Increased CaMKK2 expression is an adaptive response that maintains the fitness of tumor-infiltrating natural killer cells. Cancer Immunol. Res. 11, 109–122 (2023).
Tycko, J., Myer, V. E. & Hsu, P. D. Methods for optimizing CRISPR-Cas9 genome editing specificity. Mol. Cell. 63, 355–370 (2016).
Kimberland, M. L. et al. Strategies for controlling CRISPR/Cas9 off-target effects and biological variations in mammalian genome editing experiments. J. Biotechnol. 284, 91–101 (2018).
N’Diaye, E. N. et al. PLIC proteins or ubiquilins regulate autophagy-dependent cell survival during nutrient starvation. EMBO Rep. 10, 173–179 (2009).
Arganda-Carreras, I. et al. Trainable Weka segmentation: a machine learning tool for microscopy pixel classification. Bioinformatics 33, 2424–2426 (2017).
Acknowledgements
We thank Dr. Kristin Chadwick for her assistance with cell sorting at the LRFCF. pBABE-puro.mCherry-EGFP-LC3B was a gift from Jayanta Debnath and pCDH-EF1-Luc2-P2A-tdTomato was a gift from Kazuhiro Oka. This research was supported by donations to the Mary and John Knight Translational Ovarian Cancer Research Unit and to the London Run for Ovarian Cancer, by the Natural Sciences and Engineering Research Council of Canada (graduate scholarship, M.J.B.), and by the Canadian Institutes of Health Research (grant #451405, T.G.S).
Author information
Authors and Affiliations
Contributions
Conceptualization, M.J.B. and T.G.S.; Software, M.J.B.; Formal Analysis, M.J.B., A.B.; Investigation, M.J.B., A.B., Y.R.V., T.P.A.J., J.H.; Writing – Original Draft: M.J.B.; Writing – Review & Editing – M.J.B., A.B., Y.R.V., T.P.A.J., J.H., T.G.S.; Visualization: M.J.B., A.B.; Supervision, T.G.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Borrelli, M.J., Buensuceso, A., Ramos Valdes, Y. et al. CAMKKβ supports growth and viability of epithelial ovarian cancer in vitro and in vivo. Sci Rep 15, 25913 (2025). https://doi.org/10.1038/s41598-025-11584-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11584-9