Abstract
Brain tumors are caused by the abnormal growth of cells in the brain, which are often irregular in shape. Growing at such a rate of about 1.4% per day, the abnormal rate of growth accounts for an invisible illness and depressive behavioral changes; hence, brain tumors have become a major cause of increased death rates among adults worldwide. The prognosis of a brain tumor can be improved significantly if the tumor is diagnosed early using different imaging modalities. Out of these various imaging systems, Magnetic Resonance Imaging (MRI) is the most commonly used diagnostic modality because it is non-invasive and provides clear visualization of brain tissues. This study proposes a novel brain MRI image classification framework involving image enhancement, adaptive decomposition, statistical feature extraction, and a supervised classification method. First, the contrast of magnetic resonance is improved through the Tuned Single-Scale Retinex (TSSR) technique. Afterward, these enhanced images are decomposed through Empirical Wavelet Transform (EWT) so that informative subbands can be extracted. From each subband, energy and entropy features (Shannon and Tsallis) are computed and concatenated into a feature vector. This feature set is used to train Support Vector Machine (SVM) and LPBoost classifiers. The model was evaluated on a binary-class brain MRI dataset comprising 280 images sourced from the Harvard Medical School and Kaggle repositories. Experimental results demonstrate that the proposed framework achieves a classification accuracy of 96.43%, with a true positive rate (TPR) of 100% and a true negative rate (TNR) of 77.78%, outperforming several state-of-the-art methods. The primary contributions include the introduction of EWT-based statistical features for brain MRI classification and the use of TSSR for enhanced image quality, offering a robust and generalizable solution for medical image analysis.
Similar content being viewed by others
Introduction
Brain tumors are abnormal masses of tissue that arise due to the uncontrolled growth of cells in the brain or its surrounding structures. These tumors can disrupt normal brain functions, leading to a range of symptoms, including persistent headaches, seizures, cognitive impairments, and motor dysfunctions1. Brain tumors are broadly categorized into two main types: benign and malignant. Benign tumors are generally slow-growing and non-cancerous, whereas malignant tumors are aggressive and capable of invading surrounding tissues2.
The classification of brain tumors is critical for understanding their biological behavior and guiding appropriate treatment strategies. Tumors are often classified based on their origin, histological type, and molecular characteristics. Primary brain tumors, such as gliomas, meningiomas, and pituitary adenomas, originate within the brain, while secondary (metastatic) tumors arise from cancers in other parts of the body that have spread to the brain3. The World Health Organization (WHO) grading system further refines this classification, ranging from Grade I (least aggressive) to Grade IV (most aggressive), based on tumor aggressiveness and prognosis3.
Magnetic Resonance Imaging (MRI) is a pivotal tool in the diagnosis and management of brain tumors. This advanced imaging technology utilizes powerful magnetic fields and radio waves to produce highly detailed images of the brain, enabling the precise visualization of tumor characteristics4. Unlike CT scans or X-rays, MRI does not use ionizing radiation, making it a safer option for repeated imaging. MRI is invaluable for identifying the size, location, and extent of brain tumors. It helps distinguish between tumor types, evaluate their impact on surrounding structures, and guide treatment decisions5. Therefore, recently, researchers have utilized MRI images to detect brain tumors through computer-aided detection (CAD) methods6.
The structure of this study is organized to ensure clarity and coherence in presenting the research. “Related works” provides a detailed exploration of existing methods for brain tumor detection, emphasizing their limitations. “Proposed methodology” offers the innovative aspects of the approach and describes its detailed implementation. “Experimental results” presents a comparison of the simulation outcomes of the implemented model with those of existing techniques. It also includes an analysis of factors that contribute to the success and efficiency of the suggested strategy. Finally, “Discussions” summarizes the main findings and highlights the effectiveness and potential impact of the proposed methodology in advancing brain tumor detection.
Related works
The incidence of brain tumor cases is rapidly increasing worldwide. If these tumors are not identified in their early stages, patients may experience severe medical complications. To address this challenge, numerous computer-aided detection (CAD) techniques have been developed in recent years7,8. This discussion highlights several recent methodologies while summarizing the challenges associated with these approaches.
Zhang et al.9 introduced an innovative approach for classifying brain MRI images by utilizing wavelet energy (WE) feature extraction in combination with a K-nearest neighbor (KNN) learning model. Similarly, Zhou et al.10 proposed an automated method for brain MRI classification, leveraging wavelet entropy (WN) features in conjunction with the Naive Bayes (NB) classifier. Yang et al.11 employed WE features alongside a kernel-based support vector machine (SVM) to effectively distinguish brain abnormalities in MRI scans. Additionally, Bahadure et al.12 developed a computer-aided diagnostic (CAD) system designed for brain tumor detection using Berkeley wavelet transform (BWT), discrete cosine transform (DCT), and genetic algorithms (GA).
In another study, Abadi et al.13 presented a diagnostic framework incorporating the gray-level co-occurrence matrix (GLCM) and a singular value decomposition-based radial basis function neural network (SVD-RBFNN). Gokulalakshmi et al.14 proposed an advanced classification model that integrates discrete wavelet transform (DWT) and SVM techniques to analyze brain MRI images. Mudda et al.15 introduced a more refined brain tumor classification method by combining gray-level run-length matrix (GLRLM), center-symmetric local binary patterns (CS-LBP), and artificial neural networks (ANN). For more accurate diagnostics, Ansari et al.16 created an enhanced statistical machine learning model that integrates GLCM, DWT, and SVM.
Togaçar et al.17 and Kaur et al.18 proposed deep learning-based automated techniques for brain tumor detection, demonstrating the efficacy of neural networks in medical imaging. Furthermore, Kumar et al.19 developed an automated detection and classification methodology using DWT, principal component analysis (PCA), and kernel-based SVM (K-SVM). Amin Abdollahi et al.20 advanced brain tumor classification by employing an enhanced convolutional neural network (CNN) optimized with a nonlinear Levy chaotic moth flame optimizer (NLCMFO). Janardhana Rao et al.21 applied a pre-trained ResNet-18 model for analyzing brain MRI images, while Mohsen Ahmadi et al.22 proposed a customized CNN model paired with PCA for brain tumor detection in MRI scans.
Sowjanya et al.23 developed a novel methodology for the classification of brain tumors using histogram-based local shape and texture features followed by supervised learning models. Abdullah et al.24 presented a machine learning (ML)-based framework for the grading of brain tumors. Rahman et al.25 implemented an EfficientNet-B5 architecture to discriminate normal and pathological brain MRI images. Yanning et al.26 proposed a new framework with scarce labels, namely Double-SimCLR, to improve the robustness of the brain tumor detection system.
By analyzing the issues outlined above, it is evident that while these methods have significantly advanced brain tumor detection, they still face challenges such as a heavy reliance on feature engineering, high computational demands, sensitivity to dataset size, and the risk of overfitting. These limitations underscore the need for further refinement to improve the robustness and scalability of such techniques. To address these challenges, we propose a novel framework for classifying brain MRI images. This framework incorporates Tuned Single-Scale Retinex (TSSR), Empirical Wavelet Transform (EWT), and statistical measures such as energy and entropy (Shannon and Tsallis) to enhance diagnostic accuracy.
Key contributions
The key contributions of the proposed approach are as follows:
-
1.
A Retinex-based enhancement model is employed to enhance the quality of brain MRI images, thereby improving the classification accuracy of the model.
-
2.
A novel feature set is proposed, combining the Empirical Wavelet Transform (EWT) with energy and entropy measures. The performance of this feature set is compared against traditional wavelet-based energy and entropy features to demonstrate its efficiency. This combination is a unique aspect of the proposed framework, introduced for the first time in the classification of brain tumors from MRI images.
-
3.
The effectiveness of the proposed model is validated using various wavelet basis functions and benchmarked against widely recognized approaches.
-
4.
To highlight the significance of the proposed Retinex model, its performance is assessed under different noise conditions and geometric transformations. This evaluation constitutes a key contribution of the proposed approach.
Proposed methodology.
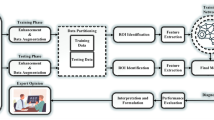
The proposed approach consists of four key stages. First, contrast enhancement is achieved through the TSSR technique to enhance image quality. Second, the enhanced images are decomposed using EWT to capture essential frequency components. Third, energy and entropy features are extracted from the decomposed images to provide meaningful representations. Finally, the extracted features are classified using supervised machine learning approaches, as illustrated in Fig. 1.
Working flow of the proposed framework.
Contrast enhancement
MRI sequences are vital diagnostic tools for analyzing medical images and visualizing the internal structure of the human body. However, during the image acquisition process, MRI images are often affected by various types of noise27, primarily due to environmental factors such as illumination variability. This noise can result in the loss of critical information and degrade image quality, making it challenging for medical practitioners to accurately diagnose diseases. Therefore, obtaining high-quality MRI images is essential for enhancing detection accuracy. To address this issue, a TSSR approach has been implemented28. This method significantly enhances the contrast and brightness of MRI brain images without compromising fine details, offering superior performance compared to existing techniques29. The mathematical formulation of the TSSR is provided below:
where \(\:I(m,n)\) is the original image; L is the dimensions of the image; G and \(\:\widehat{G}\) denotes the gradient along \(\:m\) and \(\:n\) direction; N illustrates the normalization factor; \(\:h(m,n)\)is the modified Gaussian function; ‘\( * \)’ indicates the convolution operator; \(\:\xi\:\)is an arbitrary constant which controls the brightness and contrast of an image. In the TSSR method, the parameter \(\:\xi\:\) controls image brightness and contrast. In this work, \(\:\xi\:\) = 1.5 is selected after evaluating several brain MRI images to enhance tumor visibility without introducing noise or edge artifacts. This value preserved anatomical detail while improving contrast between healthy and abnormal regions. Our choice is consistent with prior studies28, which recommend values between 1.2 and 2 for effective MR image enhancement. The effectiveness of β = 1.5 is visually supported in Fig. 2. It was observed that the TSSR method enhances the visual details of the tumor-affected region when compared to the original images. Consequently, the proposed model demonstrates improved effectiveness differentiating tumors in enhanced MR brain images. After that, decompose the enhanced image into low and high-frequency components using the EWT. Algorithm 1 represents the pseudo-code of the suggested enhanced model.
Contrast enhancement: (a,b) Original images; (c,d) enhanced images.
Image decomposition
Recently, wavelets and their geometric extensions, such as Ridgelet30, Curvelet31, and Contourlet32 transforms, have been widely applied in the analysis of MR images. These methods rely on fixed basis functions, which may not be optimal for characterizing the image. This limitation can be overcome by employing adaptive decomposition techniques. In line with this, we introduce the Empirical Wavelet Transform (EWT)33, which combines the strengths of both Empirical Mode Decomposition (EMD)34 and Wavelet Transform (WT)35. The primary advantage of EWT lies in its ability to accurately capture the local characteristics (or features) of an image. The EWT approach can be summarized as follows:
1) Divide the Fourier spectrum.
2) Building an empirical wavelet filter-bank.
3) EWT representation.
Divide the Fourier spectrum
Here, initially, we employed a Fast-Fourier Transform (FFT) to obtain spectrum of a signal over a frequency range of\(\left[ {0,\pi } \right]\). For boundaries, the corresponding Fourier spectrum is segmented into N frequency bands as shown in Fig. 3 and they are represented by\({\lambda _n}=\left[ {{\omega _{n - 1}},{\omega _n}} \right]\), where\(n=1,2,3, \ldots ,N\). From this, the complete Fourier spectrum is can be obtained by \( \cup _{{n=1}}^{N}{\lambda _n}=\left[ {0,\pi } \right]\). Now, we develop the filter-banks to implement EWT.
Building an empirical wavelet filter bank
To construct EWT, we define a bank of N wavelet filters including one low-pass (LP) and \(N - 1\)band-pass (BP) filters as shown in Fig. 3 using the idea utilized in building Meyer wavelets37. Then, the empirical scaling and wavelet functions of one-dimensional EWT (1D-EWT) using various basis functions are given by.
Principle of Fourier spectrum decomposition (adopted from work in36).
-
a)
Ridgelet and tensor:
-
b)
Littlewood-Paley:
and, when \(n{\text{ }} \ne {\text{ }}N - 1:\)
and, when \(n{{ = }}N - 1:\)
Here, the arbitrary function\(\beta \left( z \right)\), the parameter \(\alpha \)and the transition phase \({T_n}\) are defined as follows
EWT representation
The EWT \(E_{g}^{\varepsilon }\) is defined similar to that of the conventional wavelet transform. The detailed \(E_{g}^{\varepsilon }\left( {n,t} \right)\)and approximated \(E_{g}^{\varepsilon }\left( {0,t} \right)\)coefficients of empirical wavelets are obtained by performing inner product between original signal\(g\left( t \right)\), empirical wavelet\({\Psi _n}\), and empirical scaling function\({\Phi _n}\), is given below
where \({\mathcal{F}^{ - 1}}\left( . \right)\)is the inverse Fourier transform. The original signal can be reconstructed by applying the inverse EWT as
Building upon the 1D-EWT, the Littlewood-Paley, Ridgelet, and Tensor-based 2D-EWTs38 have been developed.
Parameters selection
To ensure consistency in feature extraction across brain MRI images, the number of EWT subbands was fixed at four. This was achieved using a local maximum detection scheme that identifies dominant spectral components within the Fourier domain. Four subbands offered a trade-off between detailed frequency analysis and manageable feature dimensionality.
In addition, three basis functions were employed in the EWT framework: Littlewood-Paley, Ridgelet, and Tensor-based designs. These were selected based on their unique capabilities:
-
a.
Littlewood-Paley: Effective in capturing smooth variations, gradual transitions, and rounded boundary details. This basis function is particularly suitable for highlighting tumor contours, which often present with smooth but irregular shapes.
-
b.
Ridgelet: Specialized in detecting linear and directional structures, making it ideal for characterizing elongated tumor features or streak-like patterns observed in some brain abnormalities.
-
c.
Tensor-based basis: Capable of extracting multidirectional and edge-dense regions by combining several orientations. It is well-suited for detecting sharp transitions, intersecting features, and complex tumor textures, which are typical in advanced pathological cases.
These theoretical advantages are supported by the empirical results presented in Tables 3, 4 and 5, where each EWT basis function exhibits distinct strengths under various test conditions. Additionally, Figs. 4, 5 and 6 illustrate the effectiveness of each basis in capturing specific image features. These observations confirm that each EWT approach contributes significantly to accurately characterizing brain MR images. Algorithm 2 represents the pseudo code of EWT decomposition approach.
Components of 2D Littlewood-Paley EWT for N = 4.
Components of 2D Ridgelet EWT for N = 4.
Components of 2D Tensor EWT for\({N_R},{N_C}=4\).
Feature extraction
Features play a key role in the classification of brain MR images. Among all the available features, texture features39 are extensively utilized since they can significantly capture the variations within the image. In this work we consider energy and entropy measures to classify the brain MR images as healthy and pathological.
Energy
Energy is an estimation of the similarity/closeness of the image, which is defined as follows
Entropy
Entropy quantifies the variations contained in the image and thus illustrating the texture details. In this work, we employed Shannon entropy40 and Tsallis entropy41 due to their complementary characteristics. Shannon entropy effectively captures the uncertainty and information content in the image, making it suitable for detecting texture variations. On the other hand, Tsallis entropy generalizes Shannon entropy and provides better sensitivity to long-range interactions and nonlinearities, which are often present in pathological brain tissues. The combination of these two measures ensures a more comprehensive quantification of textural heterogeneity in MRI subbands. The mathematical formulations of each measure are presented below:
-
a.
Shannon entropy:
It evaluates the uncertainty or randomness about the information content of an image40 and is formulated as
where L gives the number of gray values and \({p_j}\)indicates the probability of gray value j.
-
b.
Tsallis entropy:
It is a generalization of traditional Boltzmann-Gibbs entropy41 and is expressed as
where \(\gamma \) represents a entropy index which is always a real value. Note that, if\(\gamma \to 1\), then \({H_T}\)approached to \({H_S}\).
Wavelet-based energy and entropy features, along with Curvelet-entropy features, have been widely used in the classification of MR brain images9,10,1142,43,44,45,]-46. However, to the best of our knowledge, empirical wavelet-energy and entropy features have not been explored in the context of brain MR image classification. In this study, we first estimate the energy and entropy from each empirical wavelet subband. These features are then organized into a row feature vector by linearly concatenating them. This extracted feature vector is subsequently fed into a classifier to distinguish between pathological and healthy brain MR images.
where \(EWT - E\)represents the empirical wavelet-energy feature, \(EWT - {H_S}\)denotes the empirical wavelet-Shannon entropy feature, \(EWT - {H_T}\)gives the empirical wavelet-Tsallis entropy feature.
From each brain MRI image, we extracted a total of 36 features, resulting in a feature vector of size 1 × 36. A Kruskal-Wallis (KW) test was conducted to find the feature relevance. This is a nonparametric test that ranks the data to test for statistical differences. The test results are presented in Table 1. All features had p-values less than 0.05 (e.g., p = 0.032), demonstrating the statistical significance of the features. This signifies that the extracted local texture features can be used for further analysis. Then, the feature vectors were fed into conventional classifiers for discriminating between normal and abnormal brain MRI scans. Algorithm illustrates the pseudo code of the presented feature extraction method.
Classification
In this study, supervised machine learning techniques such as Support Vector Machine (SVM)47 and LPBoost48 are utilized to identify the pathology in brain MR images. Further, assess the performance of these models using several established metrics, including the true positive rate (TPR), true negative rate (TNR), positive predictive value (PPV), F-score, area under the curve (AUC), and accuracy. Algorithm 4 shows the pseudo code of the presenting classification framework, whereas Algorithm gives the main work-flow of the implemented model.
Experimental results
This study used brain MRI images obtained from the Harvard Medical School (HMS) dataset49 and the Kaggle brain tumor repository50. The combined dataset includes 280 images, comprising 235 pathological and 45 healthy samples. To maintain class distribution integrity during evaluation, both 70/30 and 80/20 train-test splits were performed using stratified sampling, which is illustrated in Table 2.
All MRI images were resized to a fixed resolution of 256 × 256 pixels to ensure uniformity across the processing pipeline. Before processing of these images, we performed intensity normalization to map pixel values into the [0, 1] range, and TSSR-based contrast enhancement to highlight critical tumor features without amplifying noise. The dataset includes a diverse mix of T1- and T2-weighted scans, covering a variety of tumor types, anatomical locations, and intensity profiles. This diversity enhances the clinical relevance and generalizability of the proposed method to real-world diagnostic applications.
Simulation settings
In this study, to identify an optimal feature set, the Empirical Wavelet Transform (EWT) is constructed using various basis functions, such as Littlewood-Paley, Ridgelet, and Tensor. Due to the adaptive nature of the EWT, the number of subbands may vary from one image to another, which is not ideal for classification tasks. To address this, we generate four subbands using the ‘localmax’ detection approach. Furthermore, to evaluate the effectiveness of the image enhancement, the proposed methodology is tested under different noise and geometric transformation conditions. The parameters for these conditions are summarized in Table 3. All simulations are performed on an Intel® Core™ i3-5005U CPU @ 2 GHz using MATLAB 2020a.
Simulation results
In this subsection, we present and discuss the findings from the proposed and existing methods. To assess the performance of our model, a series of simulations were conducted using the dataset outlined in Table 2. The classification outcomes presented in Tables 4, 5 and 6 reveal consistent trends and critical observations across various EWT basis functions, tested under different image distortions and classifier types. Across each of the EWT variants and perturbed under various transformations such as Gaussian noise and impulse noise, rotation, shear, and translation, the SVM classifier has maintained an extremely high TPR, sometimes achieving a clean 100%. This indicates that SVM is highly effective in detecting abnormal brain MR images, making it a dependable option for clinical diagnostics. On the other hand, whereas LPBoost seemed sometimes to lag behind SVM in TPR, it behaved more consistently with respect to TNR, PPV, and Accuracy. LPBoost benefited more in reducing false positives, particularly under geometric distortions with impulse noise (greater TNR than SVM), which might come into play where a good compromise is sought between sensitivity and specificity.
A comparative evaluation of the three EWT basis functions reveals the following:
-
From Table 4, we observed that the Littlewood-Paley EWT was proficient at extracting boundary and shape-related information, producing reasonable results. However, it exhibited some inconsistency in TNR, especially under intense geometric transformations.
-
From Table 5, we conclude that the Ridgelet EWT outperformed Littlewood-Paley in texture-rich conditions, demonstrating better accuracy and TNR, particularly in noisy environments, and showed notable synergy with the LPBoost classifier.
-
From Table 6, we discovered that Tensor-based EWT stood out as the most resilient and high-performing approach, maintaining excellent TPR and balanced TNR values across all distortions and classifiers. It consistently delivered top-tier results in both pristine and degraded image conditions.
A significant enhancement across all EWT configurations was observed when the TSSR technique was applied. For example, the combination of TSSR and Tensor-EWT with LPBoost achieved the highest recorded accuracy of 96.43%, alongside a TPR of 100% and a TNR of 77.78%. This highlights the effectiveness of TSSR in amplifying visual features crucial for accurate tumor detection. In summary, the integrated framework of TSSR + Tensor-EWT + LPBoost emerges as the most effective setup for brain MRI classification. It offers a well-balanced performance in terms of sensitivity and specificity, and demonstrates consistent reliability under varying image conditions—surpassing many existing methodologies.
To further validate the theoretical advantages of each EWT basis function, we conducted a series of experiments under various noise conditions and geometric transformations. The performance of the proposed framework using Littlewood-Paley, Ridgelet, and Tensor basis functions was systematically evaluated and is summarized in Tables 4, 5 and 6. These results not only reinforce the distinct strengths of each basis function described earlier but also confirm that their structural characteristics directly influence classification effectiveness. Specifically, the Littlewood-Paley basis, with its emphasis on shape and boundary features, performs well in delineating tumor outlines, particularly when enhanced by TSSR. Similarly, the Ridgelet basis demonstrates strong performance in scenarios requiring texture sensitivity, aligning with its ability to capture corner and directional information. Notably, the Tensor basis, which offers multi-directional edge and line feature extraction, consistently achieves the highest accuracy across all test conditions. This empirical evidence validates our design choice and highlights the practical advantage of combining TSSR enhancement with Tensor-based EWT for robust and accurate brain tumor classification.
Ablation study
In this study, we compared the performance of the complete framework (TSSR + EWT + statistical features + classifier) with a baseline variant that excluded the TSSR module (i.e., using raw MRI images directly with EWT and the same classification pipeline). As detailed in Tables 4, 5 and 6 of the manuscript, the inclusion of TSSR consistently improved performance across all metrics. For instance, under the 80/20 train-test split with Tensor-based EWT and LPBoost, the complete model achieved an accuracy of 96.43%, TPR of 100%, and TNR of 77.78%. In contrast, without TSSR, the same configuration yielded noticeably lower accuracy and TNR values. This demonstrates that TSSR significantly enhances the visual contrast and edge definition of brain MR images, leading to more discriminative features during EWT decomposition and ultimately improving classification. Thus, the ablation confirms that TSSR is a critical component in the pipeline, substantially contributing to the improved robustness and sensitivity of the proposed framework. We have clarified this in the revised manuscript to better highlight the impact of each individual module.
Comparative analysis
The proposed framework is more complete with rigorous statistical validation to support its performance claims. The KW statistical test was applied to the features extracted-energy, Shannon entropy, and Tsallis entropy-and returned a p-value of 0.032, confirming their statistical significance for classification. Statistical comparisons of classification measures TPR, TNR, and accuracy were also made across competing methods through the same test, and the results again showed significant differences (p < 0.05), confirming that the enhancements offered by the proposed method are not merely due to chance but are statistically relevant. Tables 4, 5 and 6 cover the studies of the proposed system, where the framework showed consistent performance with all the noise types and geometric distortions tested, supporting evidence of its robustness and reliability.
The recent advances in statistical approaches on which the method is based, which are necessary to benchmark the method’s performance in comparison to other techniques used in analogous applications presented in the literature, are shown in Table 7 using the 80/20 train-test splitting ratio. Results show that the fabulous framework does the best in classification efficiency of any method, with a perfect TPR of 100%, 77.78% TNR, and 96.43% accuracy. This surpasses those recorded in former works, thus indicating that the solution stands as an effective and general one. The framework is useful and reliable for brain MR image analysis and classification, that is, to achieve the identification of both normal and pathological states quickly and reliably.
To ensure statistical validity of the observed performance improvements over existing techniques, we conducted KW- test for classifier comparison. The results confirmed that the performance gains of the proposed framework are statistically significant (p < 0.05), supporting our claim of superior classification performance.
Discussions
To evaluate the significance of the presented model, a sequence of experiments were conducted on a two-class brain MR image dataset. The proposed framework has major implications for developing clinical diagnostics and research in medical image processing. The approach uses TSSR to enhance image contrast; thereafter, it employs EWT followed by energy and entropy-driven feature extraction to develop a robust yet efficient method for classifying normal brain tissue from the abnormal. The results demonstrated that the framework performed well with both classifiers, particularly with the Little-Wood Paley and tensor-based empirical wavelets, as shown in Tables 4 and 6. Among the two classifiers, LPBoost achieved superior performance, consistently exceeding 90% accuracy across all features. In this way, the system can act as a computer-aided diagnosis (CAD) tool in locales that may have limited specialized medical competence.
Advantages
The key pros of the proposed framework are as follows:
-
1.
The use of the TSSR technique preserved the meaningful information (edge) of brain MR images, improving classification accuracy, as reflected in Tables 4, 5 and 6.
-
2.
The EWT effectively extracted relevant local texture details from the images.
-
3.
Entropy and energy features were derived from each empirical wavelet subband to accurately classify the images by capturing critical information.
-
4.
The combination of EWT, entropy, and energy features efficiently highlighted the intrinsic details of pathological brain MR images, enabling the proposed framework to outperform existing methods, as represented in Table 7.
-
5.
Another advantage lies in the model’s resilience to noise and geometric transformation and its consistent performance levels.
These advantages underline the robustness and efficiency of the suggested model in brain MR image classification.
Limitations
Despite several advantages of the proposed model, there are still a few limitations present, including:
-
1.
Currently, the model has been set to evaluate the binary classification of cases between healthy and pathological. There is scope to extend this framework into multi-class classification tasks in the future, such as differentiation among glioma, meningioma, and pituitary tumors.
-
2.
Currently, the dataset used in this study was gathered from two sources, i.e., Harvard and Kaggle; yet, the dataset can still be considered relatively limited in size. Hence, this, in turn, could limit the ability of the model to generalize into broader clinical scenarios. Therefore, bigger and more diverse datasets will be trained with, in the forthcoming phases, to increase generalizability.
-
3.
Furthermore, strategies will be put in place in the forthcoming phases of this study, such as cross-validation and validation on external datasets, which are very pivotal in judging the model’s stability and reliability.
Conclusion
This study introduces a novel framework for classifying brain MR images by integrating TSSR, EWT, and entropy and energy features (Shannon and Tsallis). The TSSR technique enhances the brightness and contrast of brain MR images, improving their visual quality. Following this, EWT is employed to decompose the enhanced images into meaningful subbands using various basis functions. From these subbands, entropy and energy features are extracted to construct a feature vector capable of accurately identifying pathological brain MR images by revealing hidden details. The extracted features are then classified using SVM and LPBoost to differentiate healthy and pathological brain MR images. Comprehensive experiments were conducted on a brain MR image dataset to evaluate the proposed approach. The experimental outcomes reveal that the framework outperforms existing methods, demonstrating its robustness and efficiency. Additionally, the methodology is versatile and can be extended to detect other medical conditions, such as glaucoma and Parkinson’s disease, highlighting its potential for broader applications in medical image analysis.
Data availability
The datasets used/or analysed during the current study are publicly available at http://www.med.harvard.edu/AANLIB/ and https://www.kaggle.com/navoneel/brain-mri-images-for-brain-tumor-detection/metadata.
References
Pichaivel, M. et al. An overview of brain tumor. Brain Tumors. 1, 1–10 (2022).
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: A summary. Neuro-oncology 23.8, 1231–1251 (2021).
Radbruch, A. & Bendszus, M. Advanced MR imaging in neuro-oncology. Clin. Neuroradiol. 25, 143–149 (2015).
Schaller, B. J., Modo, M. & Buchfelder, M. Molecular imaging of brain tumors: a Bridge between clinical and molecular medicine? Mol. Imaging Biology. 9, 60–71 (2007).
Abd-Ellah, M. et al. Design and implementation of a computer-aided diagnosis system for brain tumor classification. In 2016 28th International Conference on Microelectronics (ICM). (IEEE, 2016).
Saritha, S. & Prabha, A. A comprehensive review: segmentation of MRI images—brain tumor. Int. J. Imaging Syst. Technol. 26 (4), 295–304 (2016).
Saman, S. & Swathi Jamjala Narayanan. Survey on brain tumor segmentation and feature extraction of MR images. Int. J. Multimedia Inform. Retr. 8 (2), 79–99 (2019).
Zhang, G. et al. Automated classification of brain MR images by wavelet-energy and k-nearest neighbors algorithm. In 2015 Seventh International Symposium on Parallel Architectures, Algorithms and Programming (PAAP). (IEEE, 2015).
Zhou, X. et al. Detection of pathological brain in MRI scanning based on wavelet-entropy and naive Bayes classifier. In International Conference on Bioinformatics and Biomedical Engineering. (Springer, 2015).
Yang, G. et al. Automated classification of brain images using wavelet-energy and biogeography-based optimization. Multimedia Tools Appl. 75 (23), 15601–15617 (2016).
Bahadure, N., Bhaskarrao, A. K., Ray & Pal, H. Comparative approach of MRI-based brain tumor segmentation and classification using genetic algorithm. J. Digit. Imaging. 31 (4), 477–489 (2018).
Abadi, A. M., Wustqa, D. U. & Nurhayadi, N. Diagnosis of brain cancer using radial basis function neural network with singular value decomposition method. Int. J. Mach. Learn. Comput. 9 (4), 527–532 (2019).
Gokulalakshmi, A. et al. ICM-BTD: improved classification model for brain tumor diagnosis using discrete wavelet transform-based feature extraction and SVM classifier. Soft. Comput. 24 (24), 18599–18609 (2020).
Mudda, Mallikarjun, R., Manjunath & Krishnamurthy, N. Brain tumor classification using enhanced statistical texture features. IETE J. Res. 68, 3695–3706 (2020).
Ansari, M. A., Mehrotra, R. & Agrawal, R. Detection and classification of brain tumor in MRI images using wavelet transform and support vector machine. J. Interdiscip. Math. 1–12 (2020).
Toğaçar, M., Ergen, B. & Zafer Cömert. BrainMRNet: brain tumor detection using magnetic resonance images with a novel convolutional neural network model. Med. Hypotheses. 134, 109531 (2020).
Kaur, T. & Tapan Kumar, G. Deep convolutional neural networks with transfer learning for automated brain image classification. Mach. Vis. Appl. 31, 1–16 (2020).
Mandle, A., Kumar, S. P., Sahu & Gupta, G. Brain tumor segmentation and classification in MRI using clustering and kernel-based SVM. Biomedical Pharmacol. J. 15 (2), 699–716 (2022).
Dehkordi, A. et al. Brain tumor detection and classification using a new evolutionary convolutional neural network. arXiv preprint: arXiv:2204.12297 (2022).
Sunkara, J. et al. An evaluation of medical image analysis using image segmentation and deep learning techniques. J. Artif. Intell. Cloud Comput. 388(2), 2–8 . 10.47363/JAICC/2023 (2023).
Ahmadi, M. et al. Detection of brain lesion location in MRI images using convolutional neural network and robust PCA. Int. J. Neurosci. 133 (1), 55–66 (2023).
Sowjanya, Kotte, K. R., Reddy & Raveena, M. A new distinctive methodology for the classification of brain MR images using histogram based local feature descriptors. Int. J. Comput. Digit. Syst. 13 (1), 1–1 (2023).
Asiri, A. A. et al. Machine learning-based models for magnetic resonance imaging (mri)-based brain tumor classification. Intell. Autom. Soft Comput. 36, 299–312 (2023).
Rahman, M. et al. A comparative analysis and visualizable of MRI image type-based brain tumor classification using transfer learning models. In 6th International Conference on Electrical Engineering and Information & Communication Technology (ICEEICT). (IEEE, 2024).
Ge, Y. et al. A novel framework for multimodal brain tumor detection with scarce labels. IEEE J. Biomed. Health Inform. (2024).
Boyat, A., Kumar & Brijendra Kumar Joshi A review paper: Noise models in digital image processing. arXiv preprint arXiv:1505.03489 (2015).
Al-Ameen, Z. Ameliorating the dynamic range of magnetic resonance images using a tuned Single-Scale retinex algorithm. Int. J. Signal. Process. Image Process. Pattern Recognit. 9 (7), 285–292 (2016).
Garg, G. & Juneja, M. A survey of denoising techniques for multi-parametric prostate MRI. Multimedia Tools Appl. 78 (10), 12689–12722 (2019).
Do, M. N. & Vetterli, M. The finite ridgelet transform for image representation. IEEE Trans. Image Process. 12 (1), 16–28 (2003).
Candes, E. et al. Fast discrete curvelet transforms. Multiscale Model. Simul. 5(3), 861–899 (2006) .
Do, M. N. & Vetterli, M. Contourlets. Studies in Computational Mathematics. Vol. 1083–105 (Elsevier, 2003).
Gilles, J. Empirical wavelet transform. IEEE Trans. Signal Process. 61, 3999–4010 (2013).
Rilling, G., Flandrin, P. & Goncalves, P. On empirical mode decomposition and its algorithms. In IEEE-EURASIP Workshop on Nonlinear Signal and Image Processing. Vol. 3. NSIP-03, Grado (I) (2003).
Shensa, M. J. The discrete wavelet transform: wedding the a trous and Mallat algorithms. IEEE Trans. Signal Process. 40 (10), 2464–2482 (1992).
Polinati, S. & Dhuli, R. Multimodal medical image fusion using empirical wavelet decomposition and local energy maxima. Optik 205, 163947 (2020).
Daubechies, I. Ten Lectures on Wavelets (Society for Industrial and Applied Mathematics, 1992).
Gilles, J., Tran, G. & Osher, S. 2D empirical transforms. Wavelets, ridgelets, and curvelets revisited. SIAM J. Imaging Sci. 7 (1), 157–186 (2014).
Khalid, S., Khalil, T. & Nasreen, S. A survey of feature selection and feature extraction techniques in machine learning. 2014 Science and Information Conference. IEEE, (2014).
Bromiley, P. A., Thacker, N. A. & Bouhova-Thacker, E. Shannon entropy, Renyi entropy, and information. Stat. Inf. Ser. (2004).
Tsallis, C. Nonadditive entropy: the concept and its use. Eur. Phys. J. A. 40 (3), 257 (2009).
Wang, S. et al. Application of stationary wavelet entropy in pathological brain detection. Multimedia Tools Appl. 77 (3), 3701–3714 (2018).
Nayak, D. et al. Application of fast curvelet Tsallis entropy and kernel random vector functional link network for automated detection of multiclass brain abnormalities. Comput. Med. Imaging Graph. 77, 101656 (2019).
Bansal, S. Nature-inspired-based multi-objective hybrid algorithms to find near-OGRs for optical WDM systems and their comparison. In Handbook of Research on Biomimicry in Information Retrieval and Knowledge Management. (IGI Global, 2018).
Bansal, S., Kumar, S. & Bhalla, P. A novel approach to WDM channel allocation: Big bang–big crunch optimization. In The Proceeding of Zonal Seminar on Emerging Trends in Embedded System Technologies (ETECH) Organized by The Institution of Electronics and Telecommunication Engineers (IETE), Chandigarh Centre, Chandigarh, India. 80–81. August 23–24. (2013).
Bansal, S. Nature-inspired hybrid multi-objective optimization algorithms in search of near-OGRs to eliminate FWM noise signals in optical WDM systems and their performance comparison. J. Institution Eng. (India): Ser. B. 102 (4), 743–769 (2021).
Vapnik, V. The Nature of Statistical Learning Theory (Springer, 2013).
Warmuth, M. K., Liao, J. & Rätsch, G. Totally corrective boosting algorithms that maximize the margin. In Proceedings of the 23rd International Conference on Machine Learning. (2006).
https://www.kaggle.com/navoneel/brain-mri-images-for-brain-tumor-detection/metadata
Acknowledgements
The authors acknowledge that the research Universiti Grant, Universiti Kebangsaan Malaysia, Geran Translasi: UKM-TR2024-10 conducting the research work. Moreover, this research acknowledges Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2025R10), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
P R Kumar, K Prakash, B R S Teja, K A Manoj, made substantial contributions to design, analysis and characterization. A Bhuvaneswar, D D S Kumar, S Bansal, and S Kumar, participated in the conception, application, experimental validation and critical revision of the article for important intellectual content. M R I Faruque and K S A Mugren provided necessary instructions for analytical expression, data analysis for practical use and critical revision of the article purposes.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kumar, P.R., Prakash, K., Teja, B.R.S. et al. A novel brain MRI classification framework integrating tuned single scale retinex and empirical wavelet entropy features. Sci Rep 15, 29765 (2025). https://doi.org/10.1038/s41598-025-13281-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-13281-z