Abstract
The Arabidopsis Toxicos en Levadura (ATL) family is a widely distributed and evolutionarily conserved group of RING-type ubiquitin ligase in plants, with crucial roles in plant growth, development, and stress response. Although ATL gene families have been identified in numerous plants, a genome-wide identification and comprehensive analysis of their characteristics and functions are still needed in tomato (Solanum lycopersicum), a key model for studying fruit development, ripening, and stress responses. By employing a bioinformatics approach, here, we identified 86 candidate members of the SlATL gene family, constructed their phylogenetic tree, and classified these into seven subfamilies based on gene structure similarity and conserved motifs. Chromosome locations and collinearity analysis further demonstrated the structural correlations and evolutionary conservation of the SlATL family. Cis-acting elements analysis indicated that the SlATL family may be closely associated with functions such as hormone responses, environmental stress, and plant growth and development. Gene ontology (GO) functional annotation of the SlATL family, along with some individual members and their interacting proteins, revealed insights into their biological roles. The findings lay a foundation for further exploration of the SlATL family in tomato growth and stress resilience.
Similar content being viewed by others
Introduction
Ubiquitination, an important post-translational modification in eukaryotes, regulates a wide range of essential cellular activities involving three key enzymes: E1 ubiquitin-activating, E2 ubiquitin-conjugating, and E3 ubiquitin-ligase enzymes1,2. The E3 ubiquitin ligases play a pivotal role in the ubiquitination pathway by specifically targeting substrates for ubiquitination. The Arabidopsis Toxicos en Levadura (ATL) family is a specialized group of E3 ubiquitin ligases in plants, named for its first member, AtATL2, which induces a lethal phenotype in yeast when overexpressed3. The ATL family proteins are characterized by three distinct motifs: an N-terminal hydrophobic region, a GLD motif, and a RING-H2 type domain. The RING-H2 domain, a typical zinc finger protein domain, contains cysteine (Cys) and histidine (His) residues that form a specific sequence. In tomatoes, this motif is represented as Cys-X2-Cys-X15-Cys-X1-His-X2-His-X2-Cys-X10-Cys-X2-Cys, where X denotes any amino acid4. The RING-H2 domain participates in the ubiquitination process, influences protein stability and function, and is essential for various biological processes. Currently, 100 members of the ATL family have been identified in Arabidopsis5, while rice contains genes encoding 121 proteins classified within the ATL family6.
Recent advances and refinements in innovative technologies, such as high-throughput sequencing, contemporary gene editing methods, heterologous expression systems, and bioinformatics, have propelled research on ATL gene functions significantly. Current literature evidence suggests that ATL genes are involved in regulating plant responses to hormone signaling and various stress conditions, such as high salinity, drought, and low temperature. For instance, AtATL61 positively regulates the abscisic acid (ABA)-mediated drought stress response7, whereas AtATL31 and AtATL6 regulate salt tolerance in Arabidopsis in an ABA pathway-independent manner8. Overexpression of MdATL16 in apples increases the salt tolerance of apple calli, and heterologous expression of MdATL16 in tomatoes enhances resistance to salt stress9. Similarly, the heterologous expression of the sweet potato IbATL38 in Arabidopsis improves salt stress tolerance10. ATL genes are also involved in plant tolerance to low-temperature stress. Knockout of OsATL38 in rice enhances resistance to low-temperature stress, indicating a negative regulatory role for OsATL38 in this pathway11. In Arabidopsis, overexpression of AtATL80 significantly increases sensitivity to low-temperature stress and acts as a negative regulator of phosphate mobilization12. Finally, ATL genes are crucial for plant defense responses to pathogens. For example, AtATL2, a plasma membrane-integrated protein, actively participates in the defense responses of Arabidopsis against fungal pathogens13.
Tomato (Solanum lycopersicum) is a widely popular vegetable crop known for its flavorful fruits and rich nutritional contents. It also serves as a model plant for studying fruit development, ripening, and stress responses. Given the crucial role of ATL proteins in plant growth and development, a comprehensive exploration of the ATL gene family in tomatoes could provide a molecular foundation for improving tomato quality and resilience to a diverse range of stressful stimuli. The insights of this work could serve as a valuable reference for expanding ATL gene families research on other fruits and vegetables.
Results
Genome-wide identification and protein physicochemical properties of SlATLs
Employing Hidden Markov Model (HMM) and transmembrane hydrophobic region screening, we identified 86 members of the SlATL family, with 78 possessing a single transmembrane hydrophobic region, seven containing two, and only Solyc12g005020 harboring three.
As shown in Fig. 1A and Supplementary Table S1, the physical and chemical properties for 86 of the SlATL family members and their protein lengths ranged from 130 amino acids (aa) (Solyc03g115465) to 479 aa (Solyc07g006360). More specifically, 34 proteins were between 100 and 200 aa long, 23 ranged between 200 and 300 aa, 26 were between 300 and 400 aa, and only three proteins exceeded 400 aa in length. The molecular weights of these proteins varied between 14.37 and 53.32 kDa. In addition, the isoelectric point (pI) values ranged from 4.71 (Solyc06g064440) to 9.28 (Solyc07g006400), with 49 SlATLs exhibiting a pI < 7 and 37 with pI > 7.
Physicochemical property analysis of SlATL family proteins. (A) Distribution of amino acid numbers in SlATL proteins; (B) Number of hydrophilic/hydrophobic proteins, stable/unstable proteins, and acidic/basic proteins.
The instability index serves as an experimental estimate of protein stability, with proteins having an instability index below 40 considered as stable. Among all the SlATL family members identified, the instability index values ranged from 21.23 to 79.9. Only 12 SlATLs were predicted to be stable proteins, accounting for 13.95% of the family, indicating that most SlATLs were likely unstable. Additionally, the aliphatic index ranged from 62.89 to 113.6, with an average of 88.10. Furthermore, the grand average of hydropathicity values indicated that 51 SlATLs were hydrophilic, and 35 were hydrophobic (Fig. 1B).
Phylogenetic tree, conserved motif and gene structure analyses
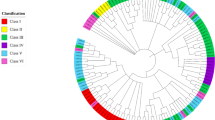
Based on the types and characteristics of the conserved motifs in SlATL proteins, the 86 SlATL members were classified into 7 groups (Group I to VII) (Fig. 3A), with each group containing 6, 19, 6, 33, 12, 4, and 6 family members, respectively (Fig. 2). Group III uniquely possessed the GUB_WAK_bind domain, which is associated with cell wall-associated kinases, potentially involved in defense and immune signaling pathways. Group VI contained the cell wall-related Gly-zipper_YMGG domain, whereas no major unique domains were found in the other groups (Supplementary Fig. S3), suggesting a considerable degree of functional conservation within the SlATL family members.
Phylogenetic tree of the SlATL family. The SlATL family is classified into seven distinct groups (I, II, III, IV, V, VI, VII) based on conserved motifs and gene structure features, with each category represented by a unique color.
We detected 10 distinct conserved motifs (motifs 1–10) among SlATL ligases (Fig. 3A). Notably, all members contained motif 1, characteristic of the RING domain, and 80 members also contained motif 2. The conserved order and positional features of these motifs suggest the conservation of cellular functions during evolutionary process of SlATL proteins. Motif 3 was mainly found in Group V, motifs 4 and 8 were primarily encountered within Group III, while motifs 6 and 7 were present in Group VI. Finally, motifs 9 and 10 were only present within Group VII members. These unique conserved motifs could indicate potentially distinct biological functions for proteins harboring.
Conserved block diagram and presence of the GLD motif in the tomato SlATL family using MEME online server. (A, B) Each colour represents different motifs, letters indicate the amino acid sequences of the motifs, gray lines represent non-conserved amino acid sequences, and the bottom scale represents the number of amino acids. I to VII represent groups. (C) Number of conserved varieties of the GLD motif in the SlATL family.
Eukaryotic genes are generally classified into intron-less (no introns), intron-poor (three or fewer introns per gene) or intron-rich14. Gene structural analysis of SlATLs (Fig. 4) revealed that they could be divided into three groups. Among these, 63 SlATL members were intron-less, and 19 were introns-poor, including Solyc01g091770, Solyc05g055390, Solyc08g082680, Solyc11g007530, Solyc08g082310, and Solyc06g064440, each containing two introns. These two groups accounted for approximately 95.3% of SlATL family members, primarily located in Group I, III, IV, V, and VII, which suggests functional conservation within SlATL gene family numbers. The remaining four members belonged to the intron-enrich group. Of these, Solyc12g005020, Solyc02g083400, Solyc02g062040, and Solyc03g033330 harbored four introns; all four members of Group VI belonged to this group. The observed structural differences likely contribute to the functional diversity of the gene family members and reflect their evolutionary expansion.
Gene structure analysis of SlATLs. Yellow denotes coding regions (CDS), green denotes untranslated regions (UTR), lines represent introns, and the bottom scale shows the direction and length of the gene sequence.
Chromosomal localization, gene density, and collinearity analysis of the SlATL family members
As illustrated in Fig. 5, except for Solyc00g217960, which occupied a locus on an unplaced scaffold not mapped to a specific chromosome, we found the remaining 85 SlATL family members to be distributed across 12 tomato chromosomes, predominantly near the telomeres and in regions of high gene density, with only a few located in lower gene density areas.
Chromosome localization and gene density of the SlATL family. The scale on the left shows chromosome length. SlATL members are marked with short black lines and labeled in red. The color gradient on the chromosomes, ranging from blue to red, represents gene density, with red indicating high gene density and blue indicating low gene density.
Synteny analysis revealed duplication relationships among the genes, as shown in Fig. 6A. Approximately 55.8% of the SlATL family members were involved in duplication events within the tomato genome. A total of 36 syntenic gene pairs were identified among SlATL members, with Solyc02g082420, Solyc02g087040, Solyc03g123680, Solyc04g074790, and Solyc04g009780 demonstrating two syntenic relationships, while Solyc01g095810 showed three. The non-synonymous/synonymous substitution ratio (Ka/Ks), a common metric for assessing the nature of gene duplication and evolutionary patterns, was calculated for duplicated SlATL gene pairs(Fig. 6B). All Ka/Ks values were less than 1, indicating that purifying selection predominated in the evolution of this gene family. This result suggests that the expansion of the gene family may be a result of linkage-specific genomic duplication, with purifying selection acting to eliminates harmful mutations while retaining core functional domains.
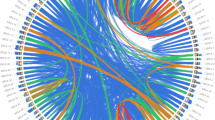
Collinearity analysis. (A) Collinearity analysis of ATL gene in tomato species. The outer scale estimates chromosome length. Synteny relationships among SlATL family members are indicated by red lines, while other gene synteny relationships are shown with gray lines. The gene duplication analysis with Ka and Ks relation of SlATLs. (B) Chromosomes are labeled with black numbers 1–12. (C) ATL gene collinearity between tomato and Arabidopsis genomes.
Excluding Chr5 and ChrUn, gene collinearity analysis between tomato and Arabidopsis shows that 41 SlATL genes in tomato were homologous to 85 genes in Arabidopsis (Fig. 6C). Among them, Chr6 contained the majority of homologous genes, with seven SlATL members and 12 homologs; in contrast, Chr11 contained the fewest, with only two SlATL genes and two homologs. This interspecies collinearity analysis of ATL genes provided insights into the genomic evolutionary mechanisms and functional conservation of ATL genes between tomato and Arabidopsis.
Cis-regulatory elements analysis
As shown in Fig. 7, 33 types of cis-regulatory elements were identified and classified into five types: 1441 promoter-related (CAAT-box), 230 environmental stress-related, 74 binding site-related, 479 hormone response, and 94 development-related, totaling 2318 cis-regulatory elements. Among these, we focused on hormone response and environmental stress-related elements within the promoter regions of SlATL family members. The hormone response elements consisted mainly of auxin- (AuxRE, AuxRR-core, TGA-box and TGA-element), gibberellin- (GARE-motif, P-box and TATC-box), salicylic acid- (TCA-element and SARE), MeJA- (CGTCA-motif and TGACG-motif), and ABA-responsive elements (ABRE). All family members, except Solyc01g091770 and Solyc09g075320, contained hormone response elements. Environmental stress-related elements include anaerobic-related (ARE, GC-motif), salt or drought stress (DRE, HD-Zip 1), low temperature (LTR), defense (TC-rich repeats), and wound response-related (WUN-motif) elements. These results suggest that SlATL family members may regulate tomato environmental stress tolerance through various hormones signaling pathways (detailed information on the cis-regulatory elements in the SlATL genes promoter regions was provided in Supplementary Table S2).
Cis-regulatory elements analysis of the SlATL family. (A) Classification and statistics of cis-acting elements. Thirty-three kinds of cis-acting elements were divided into five categories (①-⑤). The number in the grid represents the number of elements, and the color from blue to red represents the number of elements from fewer to more. (B) The different colored barplot represented the sum of the cis-acting elements in each category, and number represents the quantity of cis-regulatory elements.
Analysis of SlATL genes expression in different tissues and fruit developmental stages of tomato
As illustrated in Fig. 8, the expression patterns of the SlATL family members were tissue specific. For example, Solyc03g033330, Solyc04g081890, Solyc11g007530, Solyc08g082310, and Solyc11g010 330 exhibited higher expression levels in the roots, whereas Solyc01g066430 and Solyc03g112340 were more abundant in the leaves. Additionally, Solyc04g082690, Solyc02g082420, Solyc12g005020, Solyc10g011880, and Solyc07g006400 were enriched in shoots. Similarly, Solyc02g082420, Solyc03g112340, Solyc06g064440, Solyc10g011880, Solyc12g087840, Solyc11g005290, Solyc11g010330, and Solyc03g033330 exhibited the highest expression levels in flowers. Moreover, certain genes demonstrated expression patterns closely related to fruit ripening. For example, Solyc11g010330 expression was elevated during fruit development and ripening, while Solyc02g082420 showed higher expression during fruit growth. The expression levels of Solyc01g066430 and Solyc03g112340 considerable rose from the mature green stage to the breaker +10 d stage, with Solyc00g217960 peaking during the breaker stage. Notably, compared with the breaker stage, the expression of Solyc04g081890 (SlATL23) increased 160-fold after 10 days. Therefore, these genes might play important roles in tomato fruit development and ripening.
Gene expression analysis of the SlATL family in different tomato tissues using RNA-seq data from the Tomato Functional Genomics database. MG represents the mature green stage, B represents the breaker stage, and B10 represents 10 days after breaker stage. Relative expression levels are normalized based on the Z-score, with the heatmap colors indicating expression levels: blue for lower expression and red for higher expression.
Gene ontology functional annotation of the SlATL family proteins
To explore the functional classification of the SlATL family, we conducted a gene ontology (GO) functional annotation analysis of the family proteins (Fig. 9). Our results showed that the SlATL family could be divided in three groups based on the enriched GO terms returned: biological processes (BP), cellular components (CC), and molecular functions (MF). Among these, BP was the most abundant with 35 GO terms, followed by MF. Within BP, organic substance metabolism, various protein catabolic processes, organonitrogen compound metabolic and catabolic and protein ubiquitination, protein metabolism, hydrolysis, and protein modifications were widely observed; In MF, the regulation of catalytic activity, ubiquitin-protein transferase activity, ubiquitin-protein ligase activity, acyltransferase activity, and aminoacyltransferase activity was most prevalent; In the CC classification, SlATL proteins were predominantly annotated to the plasma membrane and cell periphery.
GO functional annotation of the SlATL family proteins.
Prediction of SlATL-interacting proteins and their GO functional annotation
As E3 ubiquitin ligases, the SlATL family members are involved in the regulation of biological processes by ubiquitinating substrate proteins. Therefore, to gain a deeper understanding of the functions of the SlATL family, we predicted the interacting proteins of 12 highly expressed SlATLs in the tomato fruits (Fig. 10) and subsequently performed GO functional annotation of the identified interacting proteins (Fig. 11). As shown in Fig. 11, 132 interacting proteins were predicted between 12 SlATLs. Among these, Solyc00g217960, Solyc02g082420, Solyc03g112340, Solyc06g064440, and Solyc11g010330 displayed unique interacting partners, with 1, 6, 16, 6, and 10 interacting proteins, respectively. Notably, Solyc02g083400 and Solyc03g033330 shared the same of interactors, which were distinct from those of the other family members. Additionally, the interacting proteins of Solyc04g081890 and Solyc12g094385, were highly similar, with the only difference being that Solyc04g081890 could interact with Solyc07g065810.
PPI of 12 SlATL proteins and their interacting proteins. Circle size represents the number of proteins that can interact with the protein; Circle colors represent different SlATL members; Proteins interacting with the same SlATL members are shown in the same color; Red circles indicate proteins that can interact with multiple SlATL members.
GO functional annotation of interacting proteins of 10 SlATL family proteins.
To gain deeper insights into the functions of proteins encoded by highly expressed genes in tomato fruit, we performed GO functional annotation for the interactors of 12 SlATL family proteins (Fig. 11). Among these, ten family proteins were annotated. In biological processes, the interacting proteins of Solyc01g066430 had the most extensive functional annotations, including cellular and organic substance metabolism and catabolic, protein ubiquitination, ABA hormone responses, and osmotic stress. Our findings indicated Solyc04g081890 and Solyc12g094385 to be involved in cellular ion homeostasis, lithium ions responses, and seed germination. Solyc11g010330 played a role in maintaining floral, inflorescence, apical meristem identity, as well as regulating gene expression, while Solyc11g066510 was likely involved in phenylpropanoid metabolism and as a negative regulator of cytokinin-activated signaling pathways and other cellular processes. In terms of molecular functions and cellular components, the interaction proteins of Solyc01g066430 may associate with ATP- and metal ions- binding, or have ubiquitin transferase, and ubiquitin ligase activities. Additionally, the interaction proteins of other SlATL family members also exhibited MAP kinase kinase activity and zinc ion transmembrane transporter activity.
Discussion
E3 ubiquitin ligas are the largest family in the ubiquitin–proteasome degradation mechanism and can regulate the ubiquitination of substrates15. The ATL ubiquitin ligase family is important for plant growth, hormone regulation, and responses to biotic and abiotic stresses. Research suggests that ATL genes are involved in hormone signaling, maintaining ion balance within cells, and managing various stress responses16. In this study, we identified 86 members of the SlATL family using bioinformatics methods. Notably, the RING domain, a key protein–protein interaction domain, was present in ATL E3 ubiquitin ligases. In addition, all 86 SlATLs contained at least one transmembrane hydrophobic region. The presence of varying numbers of transmembrane hydrophobic regions suggested that different SlATL proteins might target distinct cellular compartments17. For instance, AtATL55, which includes a single transmembrane hydrophobic region, is likely an unstable protein within the lipid raft domain of the Arabidopsis plasma membrane18. Conversely, AtATL9, which is characterized by two transmembrane hydrophobic regions, is involved in pathogen defense and is localized in the endoplasmic reticulum19. Similarly, AtATL25 has multiple transmembrane regions and interacts with the thylakoid membrane of chloroplasts20. Research has shown that the RING domain is widely distributed among plants such as Arabidopsis21, Brassica rapa22, rice23, soybean (Glycine max L.)17, and Ostreococus tauri24. However, the GLD motif cannot serve as a universal marker for identifying ATL family members, as it is not present in all ATL proteins. As shown in Fig. 3C, 32 members of the SlATL family possessed the conserved GLD motif, while 35 possessed either a GLx or GxD motif, and the remaining 19 lacked a GLD-type motif entirely. In the grape ATL family, Chen et al. have reported 17 ATL proteins to contain the full GLD motif, 23 having two conserved residues, and 12 possessing conserved glycine residues only at specific positions25. Similarly, among the 2132 identified ATL family members across 24 plant species, approximately 82% contained a conserved GLD motif17. Furthermore, the hydrophobic transmembrane domains and RING domains of ATL proteins are arranged sequentially, suggesting evolutionary conservation that may help maintain similar protein functions26,27,28.
Gene duplication events are common in plants, with whole-genome and segmental duplications occurring once or multiple times during the evolution of plant species29,30,31. These events are considered the main factors in the expansion of gene families over time and are influenced by both the internal environment and biological factors of the host organism32. The SlATL family has relatively low complexity in gene structure, with approximately 95.3% of SlATLs being intron-less or having few introns. Most members shared comparable exon–intron structures, with intron enrichment or loss distributed similarly across various groups. Notably, the majority of intron-enriched SlATLs were found in Group VI. This suggests that the exon–intron structures of these genes may have originated from tandem or segmental duplication events in the SlATL genes family, which typically share similar gene structures with minor variations. Conserved motifs analysis showed that several conserved motifs are present in most SlATL family members, which can preserve the core function of the gene during evolution, while the addition of new conserved motifs may contribute to functional diversification among family members, particularly in specific signal transduction pathways. Collinearity analysis identified numerous SlATL genes clustered at specific regions within the tomato genome, with similar mapping patterns to the Arabidopsis genome The gene family influenced by sub-functionalization and neofunctionalization of duplicate genes, has undergone expansion in structural diversity and gene expression33. Over time, conserved sequences may undergo functional divergence, enhancing the plant’s adaptability to its environment.
Promoter analysis predicted the relationship between the SlATL gene family and responses to environmental stress, hormone regulation, and developmental processes. Analysis of cis-regulatory elements upstream of the promoter regions revealed that the SlATL family contains numerous hormone-responsive, environmental stress-related, and development-related elements. For example, 12 SlATL family members were predicted to have MYBHv1 binding sites, which are commonly involved in the regulation of environmental stress responses, such as drought. MYBHv1 functions as a mediator of ABA activity, helping protect plants from drought stress34. In addition to MYBHv1 binding sites, seven MYB-binding domains found among SlATL family members, are considered to help regulate expression of flavonoid biosynthesis genes, while 31 MYB-binding sites were associated with the drought-related stress response. Studies have shown that MYB transcription factors are essential for plant stress tolerance and flavonoid biosynthesis35. Elements related to environmental stress are widely distributed among SlATL family members. Of these, Solyc02g082420 contained a cis-regulatory element involved in responses to dehydration, low temperature, and salt stress, and is highly expressed in tomato buds, flowers, and immature fruits, suggesting a role in tolerance development in these tissues. Additionally, 42 members possess cis-regulatory elements related to defense and stress responses, and 28 members have cis-regulatory elements associated with low-temperature responses. This indicates that the SlATL family is likely involved mainly in plant defense and stress responses. Song et al. discovered that StATL2 negatively regulated cold tolerance in potatoes36. Under drought conditions, overexpression of PbrATL18 significantly improved drought resistance in pears37, Similarly, AtATL78 played a dual role in Arabidopsis environmental stress responses by upregulating drought- and downregulating the low temperature stress-induced response38. The involvement of the ATL family in plant stress tolerance is closely linked to regulation of plant hormone signaling. Our analysis revealed that the most abundant hormone-response elements were those for MeJA and ABA. Previous studies have reported the regulatory effects of these two hormones on ATL functions in plants, for example, MeJA treatment increasing the expression of the ATL protein MaRING1 in bananas, which enhances fruit cold resilience39. Similarly, ABA treatment strongly induced the expression of AtATL12, enhancing Arabidopsis’ resistance to salt stress40, while ABA-dependent signaling significantly suppressed the expression of AtATL7841. In addition to their roles in abiotic stress responses, some ATL family members regulate plant growth and development. For instance, AtATL5 modulates Arabidopsis seed longevity42, while OsATL24 inhibits cell deaths caused by cytokinin and superoxide accumulation in meristematic tissue, thereby promoting root development43. Analysis of cis-acting elements strongly supports the notion that the SlATL genes family is commonly associated with stress responses, hormone regulation, and fruit ripening pathways in plants.
To gain a comprehensive understanding of the ATL gene family’s expression patterns across different tomato tissues, we utilized the latest Tomato Functional Genomics Database 44. Several genes showed higher expression levels during tomato fruit development and ripening, suggesting a potential role for SlATL family members in fruit maturation. Gene expression analysis further corroborated promoter analysis of the ATL gene family, with Solyc04g081890 (ATL23) showing increased expression during fruit ripening. Additionally, key elements involved in hormone regulation and stress response were identified, suggesting that this gene may regulate both fruit ripening and stress tolerance. Our results indicate that the ATL gene family constitutes an important group of E3 ubiquitin ligases, predicted to participate in tomato tissue development, stress responses, and hormone signaling pathways, which is consistent with the findings described by Shu et al45. To refine the above results, we performed functional annotation analysis for SlATL family members and their potential target proteins. The results indicate that through their ubiquitin conjugation functions, SlATL E3 ligases are extensively involved in the transfer and linkage of ubiquitin molecules, the intracellular degradation and metabolism of macromolecules, such as organic nitrogen compounds and proteases, as well as the post-translational modifications of proteins. Additionally, their interacting proteins are broadly involved in cellular metabolism, hormone responses, stress responses, ion binding, and developmental processes, potentially contributing to cellular composition. This further suggests that the ATL family may play a crucial role in regulating plant growth and development, hormone signaling, and stress responses46.
Although the genomic characteristics of the ATL family have been investigated in several plant species, the functions for many of the SlATL family members remain to be elucidated. This study employed bioinformatics methods to analyze the genes and encoded proteins of the SlATL family members, investigating their evolutionary relationships, expression patterns, and predicted the protein functions to enhance our understanding of their roles in tomato growth and development. Our work offers a comprehensive overview of the ATL gene family in tomatoes, providing foundational insights for future research on ATL ubiquitin ligases in tomatoes and other plants in greater detail.
Methods
Sequence retrieval and analyses
To identify members of the tomato SlATL family, we obtained the SL4.0 tomato genome and ITAG4.0 annotation files from the Sol Genomics Network database (https://solgenomics.net/)47. Next, we retrieved the Pfam database number for the RING domain (PF13639)37, and used it to obtain the HMM file for this domain from the website (http://pfam.xfam.org/)48. The TBtools49 HMM search function was used to screen for proteins containing the RING domain in tomatoes, with an E-value < 10–5, resulting in 283 proteins that met these criteria. We then screened the 283-protein list for hydrophobic regions via TMHMM-2.0 (https://www.healthtech.dtu.dk/), which returned 83 hits with both RING and hydrophobic domains. To validate the identified proteins, we visualized their structures with SMART (http://smart.embl.de/)50, and multiple sequence alignments were conducted by CLUSTALW website (https://www.genome.jp/tools-bin/clustalw)51 (Supplementary Fig. S1). Additionally, by consulting other studies, we identified three additional ATL family genes and obtained their corresponding gene IDs using the BLAST function on the Sol Genomics Network website52. In total, 86 SlATL family genes were identified.
The protein sequences for all SlATL family members were retrieved from the Sol Genomics Network website, and their physicochemical properties, such as amino acid numbers, molecular weight, pI, grand average of hydropathicity, instability index, and lipid aliphatic index, were analyzed on the Expasy (https://web.expasy.org/protparam/)53.
Phylogenetic relationships, gene structure, and conserved motif analysis
Protein sequences of the SlATL family were aligned and analyzed using the MEGA 11 software. A phylogenetic tree was constructed using the Maximum Likelihood (ML) method with a bootstrap value of 1000, and the tree was further refined by the ITOL website (https://itol.embl.de/)54. The gene structure of SlATLs was explored by GSDS 2.0 website (https://gsds.gao-lab.org/)55 and visualized with TBtools. The conserved motif analysis was conducted using the MEME Suite v5.5.5 (https://meme-suite.org/meme/)56, with the motif value set to 10, and draw the sequence logo diagram of motifs 1–10 (Supplementary Fig.S2).
Chromosomal localization, gene density, and collinearity analysis
The lengths of the 12 tomato and 5 Arabidopsis chromosomes were extracted from the downloaded ITAG4.0 and TAIR 10 files (https://www.arabidopsis.org/)57. We utilized TBtools for analysis and visualization of the chromosomal location, gene density, and collinearity, as well as to calculate Ka and Ks values for ATL family genes.
Cis-regulatory element analysis
The 2000 bp sequences upstream of the SlATLs were extracted from the tomato genome sequence and ITAG4.0 file, and then uploaded to the PlantCare website (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/)58 to predict the regulatory elements in the promoter regions. The identified cis-regulatory elements were organized according to their reported functions and classified into five categories: promoter-, environmental stress-, binding site-, hormone response-, and development-related elements. Results were visualized via TBtools.
Analysis of SlATL genes expression in different tissues and fruit developmental stages of tomato
The fragment per million exons mapping (FPKM) data for the SlATL family members in roots, leaves, buds, flowers, young fruits (1 cm, 2 cm, and 3 cm), as well as mature green, breaker, and breaker + 10 d fruits of the Heinz 1706 tomato variety, were obtained from the Tomato Functional Genomics Database (TFGD) (http://ted.bti.cornell.edu/)44, with accession number D00459 for analysis. The expression patterns of the SlATL family were visualized using TBtools to create heatmaps (Supplementary Table S3).
Gene ontology (GO) functional annotation analysis
GO analysis was used to obtain information regarding the SlATL family and its interacting proteins, including the BP, CC, and MF GO terms, using the eggNOG-mapper web server (http://eggnog-mapper.embl.de/)60.
Analysis of protein interaction network for SlATLs
We used the online database Search Tool for the Retrieval of Interacting Genes/Proteins (STRING; version 11.5; https://version-11-5.string-db.org/)61, with default settings and the medium required score (0.400), to investigate candidate interacting proteins of SlATL members with high expression levels in tomato fruits. All resulting protein–protein interaction (PPI) networks were created using Cytoscape v3.10.2, and a GO functional annotation bubble chart was generated via ChiPlot (https://www.chiplot.online/).
Data availability
The data presented in this study are openly available in the Tomato Functional Genomics Database (TFGD), reference number D004.
References
Song, L. & Luo, Z. Q. Post-translational regulation of ubiquitin signaling. J. Cell Biol. 218, 1776–1786 (2019).
Yu, F. F., Wu, Y. R. & Xie, Q. Ubiquitin–proteasome system in ABA signaling: From perception to action. Mol. Plant 9, 21–33 (2016).
Aguilar-Henonin, L., Bravo, J. & Guzmán, P. Genetic interactions of a putative Arabidopsis thaliana ubiquitin-ligase with components of the Saccharomyces cerevisiae ubiquitination machinery. Curr. Genet. 50, 257–268 (2006).
Kud, J. et al. Functional characterization of RING-type E3 ubiquitin ligases in vitro and in planta. J. Vis. Exp. 154, e60533 (2019).
Jiménez-López, D., Muñóz-Belman, F., González-Prieto, J. M., Aguilar-Hernández, V. & Guzmán, P. Repertoire of plant RING E3 ubiquitin ligases revisited: New groups counting gene families and single genes. PLoS ONE 13, e0203442 (2018).
Serrano, M., Parra, S., Alcaraz, L. D. & Guzmán, P. The ATL gene family from Arabidopsis thaliana and Oryza sativa comprises a large number of putative ubiquitin ligases of the RING-H2 type. J. Mol. Evol. 62, 434–445 (2006).
Yang, R. R. et al. E3 ubiquitin ligase ATL61 acts as a positive regulator in abscisic acid mediated drought response in Arabidopsis. Biochem. Biophys. Res. Commun. 528, 292–298 (2020).
Du, M. S., Lu, D. P. & Liu, X. T. The Arabidopsis ubiquitin ligases ATL31 and ATL6 regulate plant response to salt stress in an ABA-independent manner. Biochem. Biophys. Res. Commun. 685, 149156 (2023).
Yuan, P. H. et al. The RING-H2 type E3 ubiquitin ligase gene MdATL16 positively regulates salt tolerance in transgenic tomato and apple. Environ. Exp. Bot. 220, 105689 (2024).
Du, B. et al. A novel sweetpotato RING-H2 type E3 ubiquitin ligase gene IbATL38 enhances salt tolerance in transgenic Arabidopsis. Plant Sci. 304, 110802 (2021).
Cui, L. H. et al. OsATL38 mediates mono-ubiquitination of the 14-3-3 protein OsGF14d and negatively regulates the cold stress response in rice. J. Exp. Bot. 73, 307–323 (2022).
Suh, J. Y. & Kim, W. T. Arabidopsis RING E3 ubiquitin ligase AtATL80 is negatively involved in phosphate mobilization and cold stress response in sufficient phosphate growth conditions. Biochem. Biophys. Res. Commun. 463, 793–799 (2015).
Kim, D. et al. The auto-regulation of ATL2 E3 ubiquitin ligase plays an important role in the immune response against Alternaria brassicicola in Arabidopsis thaliana. Int. J. Mol. Sci. 25, 2388 (2024).
Liu, H., Lyu, H. M., Zhu, K. K., Van de Peer, Y. V. & Cheng, Z. M. The emergence and evolution of intron-poor and intronless genes in intron-rich plant gene families. Plant J. 105(4), 1072–1082 (2021).
Buetow, L. & Huang, D. T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 17(10), 626–642 (2016).
Vierstra, R. D. The ubiquitin–26S proteasome system at the nexus of plant biology. Nat. Rev. Mol. Cell Biol. 10, 385–397 (2009).
Aguilar-Hernández, V., Aguilar-Henonin, L. & Guzmán, P. Diversity in the architecture of ATLs, a family of plant ubiquitin-ligases, leads to recognition and targeting of substrates in different cellular environments. PLoS ONE 6, e23934 (2011).
Lin, S. S. et al. RING1 E3 ligase localizes to plasma membrane lipid rafts to trigger FB1-induced programmed cell death in Arabidopsis. Plant J. 56, 550–561 (2008).
Berrocal-Lobo, M. et al. ATL9, a RING zinc finger protein with E3 ubiquitin ligase activity implicated in chitin-and NADPH oxidase-mediated defense responses. PLoS ONE 5, e14426 (2010).
Azevedo, J. et al. Intraplastidial trafficking of a phage-type RNA polymerase is mediated by a thylakoid RING-H2 protein. Proc. Natl. Acad. Sci. 105, 9123–9128 (2008).
Stone, S. L. et al. Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol. 137, 13–30 (2005).
Alam, I. et al. Genome-wide identification, evolution and expression analysis of RING finger protein genes in Brassica rapa. Sci. Rep. 7, 1–11 (2017).
Lim, S. D. et al. A gene family encoding RING finger proteins in rice: their expansion, expression diversity, and co-expressed genes. Plant Mol. Biol. 72, 369–380 (2010).
Gao, Y. et al. Genome-wide analysis of RING finger proteins in the smallest free-living photosynthetic eukaryote Ostreococus tauri. Mar. Genomics 26, 51–61 (2016).
Ariani, P. et al. Genome-wide characterisation and expression profile of the grapevine ATL ubiquitin ligase family reveal biotic and abiotic stress-responsive and development-related members. Sci. Rep. 6, 38260 (2016).
Bashton, M. & Chothia, C. The geometry of domain combination in proteins. J. Mol. Biol. 2002(315), 927–939 (2002).
Apic, G., Gough, J. & Teichmann, S. A. Domain combinations in archaeal, eubacterial and eukaryotic proteomes. J. Mol. Biol. 310, 311–325 (2001).
Hegyi, H. & Gerstein, M. Annotation transfer for genomics: Measuring functional divergence in multi-domain proteins. Genome Res. 11, 1632–1640 (2001).
Soltis, D. E. et al. Polyploidy and angiosperm diversification. Am. J. Bot. 96, 336–348 (2009).
French. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature, 449, 463–467 (2007) .
Vision, T. J., Brown, D. G. & Tanksley, S. D. The origins of genomic duplications in Arabidopsis. Science 290, 2114–2117 (2000).
Chang, D. & Duda, T. F. Jr. Extensive and continuous duplication facilitates rapid evolution and diversification of gene families. Mol. Biol. Evol. 29(8), 2019–2029 (2012).
Rastogi, S. & Liberles, D. A. Subfunctionalization of duplicated genes as a transition state to neofunctionalization. BMC Evol. Biol. 5, 1–7 (2005).
Alexander, R. D., Wendelboe-Nelson, C. & Morris, P. C. The barley transcription factor HvMYB1 is a positive regulator of drought tolerance. Plant Physiol. Biochem. 142, 246–253 (2019).
Abubakar, A. S. et al. Genome wide characterization of R2R3 MYB transcription factor from Apocynum venetum revealed potential stress tolerance and flavonoid biosynthesis genes. Genomics 114, 110275 (2022).
Song, Q. P. et al. StATL2-like could affect growth and cold tolerance of plant by interacting with StCBFs. Plant Cell Rep. 41, 1827–1841 (2022).
Lin, L.K., Chen, Q.M., Yuan, K.L., Xing, C.H., Qiao, Q.H., Huang, X.S., Zhang, S.L. PbrATL18, an E3 ubiquitin ligase identified by genome-wide identification, is a positive factor in pear resistance to drought and Colletotrichum fructicola infection. Horticultural Plant Journal, (2023).
Suh, J. Y. et al. Arabidopsis Tóxicos en Levadura 78 (AtATL78) mediates ABA-dependent ROS signaling in response to drought stress. Biochem. Biophys. Res. Commun. 469, 8–14 (2016).
Chen, J. et al. Molecular characterization of a cold-responsive RING-H2 finger gene from banana fruit and its interaction with MaMYC2a. Postharvest Biol. Technol. 98, 48–55 (2014).
Kong, F. & Ramonell, K. M. Arabidopsis toxicos en levadura 12 modulates salt stress and ABA responses in arabidopsis thaliana. Int. J. Mol. Sci. 23, 7290 (2022).
Kim, S. J. & Kim, W. T. Suppression of Arabidopsis RING E3 ubiquitin ligase AtATL78 increases tolerance to cold stress and decreases tolerance to drought stress. FEBS Lett. 587, 2584–2590 (2013).
He, W. P. et al. E3 ligase ATL5 positively regulates seed longevity by mediating the degradation of ABT1 in Arabidopsis. New Phytol. 239, 1754–1770 (2023).
Mochizuki, S. et al. Ubiquitin ligase EL5 maintains the viability of root meristems by influencing cytokinin-mediated nitrogen effects in rice. J. Exp. Bot. 65, 2307–2318 (2014).
Fei, Z. et al. Tomato functional genomics database: A comprehensive resource and analysis package for tomato functional genomics. Nucleic Acids Res 39(1), D1156–D1163 (2010).
Shu, K. & Yang, W. E3 ubiquitin ligases: Ubiquitous actors in plant development and abiotic stress responses. Plant Cell Physiol. 58(9), 1461–1476 (2017).
Wu, M. et al. ATL protein family: Novel regulators in plant response to environmental stresses. J Agric Food Chem 71(51), 20419–20440 (2023).
Fernandez-Pozo, N. et al. The sol genomics network (SGN): from genotype to phenotype to breeding. Nucleic Acids Res 43(D1), D1036–D1041 (2015).
Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res 49(D1), D412–D419 (2021).
Chen, C. et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol Plant 16(11), 1733–1742 (2023).
Letunic, I., Khedkar, S. & Bork, P. SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res. 49(D1), D458–D460 (2021).
Thompson, J. D., Gibson, T. J. & Higgins, D. G. Multiple sequence alignment using ClustalW and ClustalX. Curr. Protoc. Bioinform 1, 2–3 (2003).
Kosarev, P., Mayer, K. F. & Hardtke, C. S. Evaluation and classification of RING-finger domains encoded by the Arabidopsis genome. Genome Biol. 3, 1–12 (2002).
Gasteiger, E., Hoogland, C., Gattiker, A., Duvaud, S. E., Wilkins, M. R., Appel, R. D., Bairoch, A. Protein identification and analysis tools on the ExPASy server. The Proteomics Protocols Handbook, 571–607, (2005).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: Recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52(W1), W78–W82 (2024).
Hu, B. et al. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 31(8), 1296–1297 (2015).
Bailey, T. L. et al. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res 37(2), W202–W208 (2009).
Lamesch, P. et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic Acids Res 40(D1), D1202–D1210 (2012).
Lescot, M. et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1), 325–327 (2002).
Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635 (2012).
Huerta-Cepas, J. et al. eggNOG 50: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47(D1), D309–D314 (2019).
Szklarczyk, D. et al. The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 51(D1), D638–D646 (2023).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grants 32172278, 32272384), the Natural Science Foundation of Shandong Province (grant ZR2024MC063) and the United Arab Emirates University (UAEU) Program for Advanced Research (UPAR) (grant 12R059).
Author information
Authors and Affiliations
Contributions
X.K., J.S., Y.J. Y.L. and S.H. designed the research. X.K. carried out the experiments. X.K. wrote the manuscript. F.L., X.Z. and Z.F.R.A. polished the language and helped to draft the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kong, X., Li, F., Shang, J. et al. Genome-wide analysis of the RING-H2 E3 ubiquitin ligase SlATL family in tomato. Sci Rep 15, 27987 (2025). https://doi.org/10.1038/s41598-025-13292-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-13292-w