Abstract
Plant growth–promoting rhizobacteria (PGPR) are soil microorganisms through which phytohormones and other bioactive compounds are produced, thereby enhancing plant growth and stress tolerance. In this study, a novel PGPR strain was identified from the rhizosphere of Lycium chinense seedlings, which produce protein-rich fruit. Whole-genome sequencing and annotation revealed that the genome of this strain, designated Pseudomonas sp. A-2, consists of a 6.65-Mb circular chromosome with 5,980 predicted protein-coding sequences. Comparative genomic analysis classified the strain within the genus Pseudomonas. The A-2 strain genome encodes proteins involved in indole-3-acetic acid (IAA) biosynthesis and signaling pathways, which was validated through IAA detection assays and quantitative analyses. Plant growth rates were significantly enhanced by the A-2 strain treatment, with increases of 3-fold in Arabidopsis, 1.5-fold in tobacco, and 1.35-fold in peanut. In Arabidopsis thaliana, expression of key genes associated with lateral and adventitious root formation was induced by the A-2 strain treatment, including ARFs, AMI1, TAA1, YUCs, IBRs, TOB1, and ECH2. Moreover, enhanced tolerance to salt stress was conferred by the A-2 strain treatment, as evidenced by improved biomass accumulation, chlorophyll content, antioxidant enzyme activity, and reduced lipid peroxidation. Levels of total soluble sugars, including trehalose, were elevated in the A-2 strain treated plants, suggesting a role in osmotic adjustment under stress. The plant growth–promoting and stress-alleviating properties of Pseudomonas sp. A-2 highlight its potential application as an effective biological agent for sustainable agriculture.
Similar content being viewed by others
Introduction
Fertilizers have long been indispensable in modern agriculture for enhancing crop productivity and meeting global food demands. However, their excessive or imbalanced use has raised concerns over environmental sustainability, soil degradation, and reduced crop quality. As a promising eco-friendly alternative, plant growth–promoting rhizobacteria (PGPR) have been increasingly studied due to their ability to improve plant performance by interacting with root systems without establishing obligate symbiotic relationships1,2.
Various types of PGPR have been studied for their abilities to produce indole-3-acetic acid (IAA), synthesize siderophores, produce 1-aminocyclopropane-1-carboxylate deaminase (ACC) deaminase (EC 3.5.99.7), solubilize phosphate, fix nitrogen, and suppress fungal pathogens. Reported PGPR genera include Azospirillum3Bacillus4Brevundimonas5Enterobacter, Klebsiella, Serratia6,7Pseudomonas8,9Rhizobium9,10and Streptomyces11. In addition, PGPR are known to contribute to plant protection12improve soil structure13utilize soil nutrients to stimulate plant growth14produce plant growth regulators15and control or inhibit plant pathogens16,17. The plant growth–promoting effects of PGPR have been documented in numerous studies involving various plant species, including Arabidopsis18,19pea20rice21,22soybean23tobacco24tomato25and wheat26.
PGPR have also been shown to participate in the biosynthesis of auxins27,28. Auxins are key regulators of plant development, modulating root and shoot architecture, lateral root (LR) formation, and tropic responses29,30,31,32,33,34. Tryptophan is the principal precursor of IAA35,36,37,38. IAA is synthesized from tryptophan through four distinct pathways: (1) the indole-3-acetamide (IAM) pathway, (2) the indole-3-pyruvic acid (IPA) pathway, (3) the tryptamine (TAM) pathway, and (4) the indole-3-acetaldoxime (IAOx) pathway39,40,41,42,43,44. The involvement of various soil microorganisms in IAA synthesis has been demonstrated both in pure cultures and soil environments45,46,47. It has also been reported that some PGPR harbor the iaaM gene, enabling high-level IAA production under specific environmental conditions48,49. Auxin-producing PGPR (e.g., Bacillus and Azotobacter) have been shown to significantly enhance crop yields and growth traits such as root and shoot development in various species, including maize, cucumber, and Arabidopsis19,50,51,52.
In addition to promoting growth under optimal conditions, PGPR have been recognized for their capacity to alleviate abiotic stress in plants. Salt stress is a major agricultural constraint, leading to osmotic imbalance, ion toxicity, and oxidative damage53. Certain salt-tolerant PGPR (ST-PGPR) have been shown to mitigate the effects of salinity by enhancing antioxidant capacity, improving osmotic adjustment, and modulating stress-responsive gene expression54,55. Moreover, PGPR have been reported to confer tolerance to other abiotic stresses, such as osmotic and drought stresses, which similarly impair plant growth by inducing water deficit, oxidative stress, and metabolic imbalance56,57. Through mechanisms such as the accumulation of compatible solutes, activation of antioxidant systems, and modulation of stress-related signaling pathways, PGPR are known to contribute to enhanced plant resilience under these challenging environmental conditions58.
While many studies have focused on the phenotypic effects of PGPR, relatively few have addressed the genetic basis of their functional traits. Whole-genome sequencing (WGS) has enabled comprehensive characterization of microbial genomes and has revealed key genes associated with PGPR functions, including nitrogen metabolism, hormone production, and stress resistance59,60,61,62,63. The integration of WGS with physiological and molecular analyses is considered essential to understand how PGPR influence plant development at the systems level.
In this study, a novel PGPR strain was isolated from the rhizosphere of Lycium chinense, and its ability to promote plant growth and enhance salt stress tolerance was investigated. Auxin-related activity was assessed, and effects on physiological parameters and gene expression were evaluated in multiple plant species. Through a multidisciplinary approach combining genomics, transcriptomics, and plant physiology, the functional role of this strain was elucidated, and its potential as a multifunctional biofertilizer for sustainable agriculture was explored.
Materials and methods
Bacterial strain isolation and DNA Preparation
Soil samples were collected from the rhizosphere of L. chinense located in Dongan-ri, Okcheon-eup, Okcheon Country, Chungcheongbuk-do, Republic of Korea (36.31°N, 127.59°E). For each sample, 1 g soil, including roots, was mixed with 9 mL distilled water and serially diluted 3 times at a 1:10 ratio, resulting in 1:10, 1:100, and 1:1000 dilutions. These dilutions were then spread onto various media: Bennet agar (1 g/L yeast extract, 1 g/L peptone B, 2 g/L tryptone, 10 g/L dextrose, 30 g/L bacto agar, pH, 7.3 ± 0.1), potato dextrose agar (PDA; 24 g/L potato dextrose broth, 30 g/L bacto agar, pH 5.6 ± 0.1), glucose yeast peptone (2 g/L yeast extract, 5 g/L peptone, 0.5 g/L MgSO4·7H2O, 1 g/L KH2PO4, 20 g/L glucose, 10 g/L bacto agar, pH 5.6 ± 0.1), or yeast glucose (5 g/L yeast extract, 10 g/L glucose, 10 g/L bacto agar, pH 5.6 ± 0.1). Plates were incubated at 28, 30, or 37 °C for 2 days. Microorganisms were isolated based on the morphological characteristics of colonies and grown in pure cultures. A single colony of a novel strain was cultured in liquid PDA medium at 30 °C for 24 h with shaking at 180 rpm. The cultured strain was stored as a 25% glycerol stock at − 80 °C. To select promising microbial candidates, colonies that inhibited the growth of neighboring microbes were prioritized, as these were presumed to exhibit competitive or antagonistic traits. Several isolates were shortlisted based on this criterion and subsequently subjected to preliminary plant growth promotion assays. A single strain that consistently promoted plant growth was selected for further experiments.
Total genomic DNA was extracted using a Quant-iT™ PicoGreen® dsDNA Kit (Cat. No. P7589, Thermo Fisher Scientific, Waltham, MA, USA), following the manufacturer’s instructions. Genomic DNA integrity was assessed via 0.7% agarose gel electrophoresis, and DNA purity was evaluated using a NanoDrop UV–Vis Spectrophotometer (Cat. No. ND-2000, Thermo Fisher Scientific). DNA concentrations were quantified using a Qubit dsDNA HS Quantification Assay Kit (Cat. No. Q32854, Thermo Fisher Scientific) and a Qubit 4 Fluorometer (Cat. No. Q33238, Thermo Fisher Scientific). For sequencing, 4 µg DNA was used, and sequencing was conducted using both Illumina (San Diego, CA, USA) and Oxford Nanopore Technologies (ONT; Oxfordshire, UK) platforms.
Bacterial strain sequencing and assembly
Libraries for long-read sequencing were prepared with end repair, dA-tailing, barcode ligation, and adapter ligation using the NEBNext® Ultra™ End Repair/dA-Tailing Module [Cat. No. E7546, New England Biolabs Co. (NEB), Ipswich, MA, USA], FFPE Repair Mix NEBNext Quick Ligation Module (Cat. No. E6056, NEB), and Native Barcoding Kit (Cat. No. SQK-NBD114.24, ONT). Ligated DNA was purified, and long-read sequencing was conducted using a MinION Mk1C device R10.4.1 (Cat. No. MIN-101 C, ONT) with a SpotON Flow Cell (Cat. No. FLO-MIN114, ONT), according to the manufacturer’s instructions and managed via MinKNOW software v4.2.5 (ONT). Adapter sequences were removed using Guppy basecalling and Porechop v0.2.464 (https://github.com/rrwick/Porechop). For long contig assembly, the Flye v2.9.2-b178665 program was used, and taxonomic classification was performed with Kraken2 v2.1.266 (https://github.com/DerrickWood/kraken2). Homopolish v0.3.467 was employed to correct sequencing errors. Genome assembly completeness was evaluated using Benchmarking Universal Single-Copy Ortholog (BUSCO) v5.4.468. Circlator v1.5.569 was used to position the dnaA gene at the start of the contig. Gene prediction and annotation were completed with PROKKA v1.14.670.
Short-read sequencing was performed to refine bacterial strain genomic sequences. Bacterial strain genomic DNA paired-end libraries with 350-bp inserts were generated using the TruSeq Nano DNA High Throughput Preparation Kit (Cat. No. 20015965, Illumina) and sequenced at Macrogens Co. (Seoul, Korea) using Illumina Sequencing by Synthesis technology. SPAdes v3.15.071 (http://cab.spbu.ru/software/spades/) was used for genome assembly.
Comparative genomic analysis
Average Nucleotide Identity (ANI) and Genome-to-Genome Distance Calculator (GGDC) values were calculated to evaluate the genomic relatedness between the A-2 strain and closely related strains. ANI and GGDC analyses were performed using the standalone version of the Orthologous Average Nucleotide Identity tool (OAT) downloaded from72 (https://www.ezbiocloud.net/tools/orthoani). In addition, Average Amino Identity (AAI) was calculated using ezAAI v1.2.3 (https://github.com/endixk/ezaai), and digital DNA-DNA hybridization (dDDH) values were estimated using Genome-to-Genome Distance Calculator v3.0 (https://ggdc.dsmz.de/distcalc2.php). Comprehensive measures of overall genomic similarity were provided by these analyses to support species delineation of the A-2 strain.
Scanning electron microscopy (SEM)
Samples were fixed in 2.5% paraformaldehyde–glutaraldehyde (4 °C, phosphate buffer, pH 7.2) for 4 h, followed by three washes with 0.1 M phosphate buffer (pH 7.2) for 10 min each. Subsequently, samples were post-fixed in 1% OsO4 (25 °C, 0.1 M phosphate buffer, pH 7.2) for 1 h, followed by several washes with 0.1 M phosphate buffer (pH 7.2) and dehydration in increasing ethanol concentrations. Substitution with isoamyl acetate and critical point drying were performed, and a 20-nm-thick coating was applied using an SC502 sputter coater. The specimens were observed at 5 kV using the FEI Quanta 250 FEG scanning electron microscope (FEI, Hillsboro, Oregon, USA).
Measurement of IAA production by bacterial strains
Bacterial strains were grown at 30 °C for 1 day with shaking at 180 rpm. Cultures were grown in liquid PDA with or without 100 mg/L L-tryptophan (L-Trp). A 5 mL sample of cultured bacteria was centrifuged at 2,700 × g for 15 min at 4 °C. The supernatant (1 ml) was mixed with 2–3 drops of ortho-phosphoric acid and 4 mL of Salkowski’s reagent (1 mL 0.5 M FeCl3 dissolved in 50 mL 35% HClO4)73. Salkowski’s reagent allows for the detection of substances belonging to the indole class. Samples were incubated at 30 °C for 30 min in the dark, and absorbance was measured at 535 nm. The IAA concentration in each bacterial strain was determined using a calibration curve generated using synthetic IAA.
Quantitative analysis of IAA production
Ultra-performance liquid chromatography (UPLC) analysis was conducted to confirm the identity of IAA produced by bacterial strain. Cells were isolated from a 5 mL bacterial culture by centrifugation for 15 min at 2,700 × g at 4 °C. The pH of the supernatant was adjusted to 2.8 with 1 M HCl, followed by three successive extractions with ethyl acetate. The dried extract was dissolved in 1 mL methanol. Samples were analysed using ACQUITY™ UPLC I-Class (Waters, Milford, MA, USA) and an ACQUITY UPLC® BEH C18 column 1.7 μm (2.1 mm × 100 mm, Waters). IAA was separated using a mobile phase of acetonitrile with 0.1% formic acid (v/v) and water. The gradient was initiated at 5% acetonitrile at a flow rate of 0.4 mL/minute, increasing to 40% over the first 8 min and then to 100% over 0.3 min. The injection volume for all samples was 1 µL, and the column temperature was maintained at 35 °C. The UPLC system was coupled with a Waters XEVO-QTOF-mass spectrometer (Micromass, Manchester, UK) equipped with an electrospray ionization scan mode. The mass conditions for multiple reaction monitoring mode were as follows: capillary voltage, 2.3 kV; source temperature, 110 °C; desolvation temperature, 350 °C; desolvation gas flow, 800 L/hour; and cone voltage, 40 V.
Bacterial culture and plant growth experiments
Bacterial strains were cultured on PDA medium at 30 °C for 24 h. The optical density at 600 nm (OD600) was measured, and the culture was adjusted to an OD600 of 0.5, equivalent to 3–4 × 106 colony-forming units mL− 1. In vitro experiments were conducted on Arabidopsis thaliana (ecotype Columbia-0). Seeds were sterilized by treatment with 70% ethanol for 1 min, followed by a solution of 1% sodium hypochlorite mixed with 0.05% Triton X-100 for 10 min, and rinsing eight times with sterile triple-distilled water (3DW). After sterilization, seeds were cold-treated at 4 °C for 2 days, then sown on square dishes (120 × 120 mm) containing half-strength Murashige and Skoog (MS) medium supplemented with 20 g/L sucrose and 10 g/L phyto agar. A total of 20 seeds were sown on each plate, and plates were incubated in a growth chamber at 25 ± 0.2 °C with a 16-hour light/8-hour dark photoperiod. The germination rate was assessed 4 days after sowing, and 12 uniformly growing seedlings were selected for subsequent experiments. Bacterial strains were cultured for 24 h. To ensure even exposure of the root zone, 60 µL of bacterial suspension was divided into small droplets and spot-applied along the agar surface, with one droplet placed per grid cell beneath each Arabidopsis seedling. PDA liquid medium was used as the control. The fresh weights of leaves and roots were measured at 1, 3, 5, and 7 days after treatment (DAT). Samples were collected in clear, 1.5 mL tubes and stored at − 80 °C until further analysis. In hypocotyl excision experiments, 5-day-old seedlings grown on germination medium were excised at the hypocotyl–root junction to induce adventitious root (AR) formation and bacterial strains were applied as described above. The fresh weights and data on AR formation were collected at 10 DAT.
Nicotiana tabacum cv. Xanti was also assessed. Seeds were sterilized and sown on square plates containing medium, as described for A. thaliana. A total of 12 seeds were sown per plate, and plates were incubated under the same growth conditions as described for A. thaliana. The germination rate was assessed after 10 days, and 10 uniformly germinated seedlings were selected for experiments. For N. tabacum, 100 µL of bacterial suspension was applied to the center of the agar surface and allowed to absorb before placing the seedlings. Due to the larger seedling size and longer treatment period, full-surface application led to occasional overgrowth and tissue damage. Central application was therefore used to minimize contamination while maintaining effective exposure. Data on fresh weights and root development were collected at 14 DAT.
Peanut (Arachis hypogaea) seeds were soaked in water for 1 day, placed on wet tissue for 2 additional days, and subsequently planted in 32-plug trays. The seedlings were grown for 4 days additional days (7 days total after sowing) before bacterial treatments began. A microbial solution (bacterial culture diluted 1:5 with 3DW) was applied by adding 25 mL microbial solution to each tray compartment at 3-day intervals over 14 total days. The control group was treated with PDA medium diluted 1:5 with 3DW.
Salinity stress treatment and physiological measurements
Plant growth condition
For the salinity stress treatment, 4-day-old A. thaliana seedling were transferred to germination medium supplemented with varying concentrations of NaCl (0, 100, and 150 mM). The plates were incubated for 14 days under the same growth conditions as in the previous experiment, following bacterial treatment. In the A-2 strain treated group, bacterial suspension (OD600 = 0.1) was evenly applied to the surface of the medium and allowed to absorb. Each treatment group contained 12 seedlings per plate, with three seedlings considered as one biological replicate.
Chlorophyll quantification
Chlorophyll content was measured to assess the physiological status of leaf tissue under stress. Approximately 10–40 mg of fresh leaf tissue was weighed and incubated in 1 mL of absolute ethanol for 24 h in the dark at room temperature. Following extraction, the supernatant was separated by centrifugation, and absorbance was measured at 645 nm and 663 nm using a UV–Vis spectrophotometer. Total chlorophyll content was calculated using the following equation : Chlorophyll (µg mg⁻¹ FW) = (20.2 × A₆₄₅ + 8.02 × A₆₆₃) / fresh weight (mg). Each treatment included three biological replicates, and each replicate consisted of three seedlings.
Superoxide dismutase (SOD) activity
Fresh aerial tissues were immediately frozen in liquid nitrogen, finely ground with a mortar and pestle, and homogenized in cold lysis buffer (50 mM potassium phosphate, 0.1 mM EDTA, 0.5% Triton X-100, pH 7.4). The homogenate was centrifuged at 12,000 × g for 5 min at 4 °C, and the resulting supernatant was used for protein quantification and antioxidant enzyme assays. SOD activity was measured using a commercial SOD Assay Kit (Sigma-Aldrich, Cat. No. MAK528, St. Louis, MO, USA) according to the manufacturer’s instructions. The assay was performed in a 96-well microplate format. Briefly, 20 µl of the crude enzyme extract was added to each well, followed by 160 µl of Working Solution and 20 µl of Xanthine Oxidase Solution. After adding all reagents, the absorbance at 440 nm was immediately measured using a microplate reader (Tecan Group Ltd., Männedorf, Switzerland), without additional incubation, following the kit protocol. One unit of SOD activity was defined as the amount of enzyme causing a 50% inhibition of the reduction of WST-1 reagent, as described in the kit protocol. All experiments were conducted with three biological replicates, each containing three seedlings.
Free proline content determination
Proline content was quantified according to the acid-ninhydrin method. Fresh shoots (~ 40 mg) were homogenized in 3% sulfosalicylic acid and centrifuged at 12,000 × g for 6 min. The resulting supernatant was reacted with acid ninhydrin reagent (1.25% ninhydrin in glacial acetic acid and 6 M phosphoric acid) and incubated at 100 °C for 1 h. After cooling, the chromophore was extracted with toluene, and absorbance was read at 520 nm. Proline concentrations were calculated from a standard curve using L-proline and expressed as µmol g⁻¹ fresh weight (FW).
Lipid peroxidation assay
Malondialdehyde (MDA) levels were measured as an indicator of lipid peroxidation using the Lipid Peroxidation (MDA) Assay Kit (Abcam, ab118970, Cambridge, UK), following the manufacturer’s protocol. Briefly, ground shoot tissue was homogenized in lysis solution centrifuged, and the supernatant was used for the assay. Absorbance was measured at 532 nm. MDA levels were calculated based on a standard curve generated using the provided MDA standard.
Quantification of total soluble sugar (TSS) and Trehalose
Total soluble sugars were quantified using the phenol–sulfuric acid method described by Dubois et al.74. Briefly, 200 µL of the sample extract was mixed with 200 µL of 5% phenol solution and 1 mL of concentrated sulfuric acid. The mixture was incubated at room temperature for 30 min, and absorbance was measured at 490 nm using a UV–Vis spectrophotometer.
Trehalose content was determined using the anthrone–sulfuric acid method as described by Magazin and Popovic75with trehalose as a standard. Absorbance was measured at 620 nm after color development, and results were expressed as mg trehalose per gram fresh weight (mg/g FW).
RNA isolation and quantitative reverse transcription polymerase gain reaction (RT-qPCR) analysis
Total RNA was extracted from A. thaliana root samples collected from three independent experiments: at 1, 3, 5, and 7 DAT; at 10 DAT; and at 14 DAT under salt stress conditions (0, 100, and 150 Mm NaCl), using the HiYield™ Total RNA Mini Kit (Cat. No. YRP100, RBC, Banqiao, Taiwan). RNA concentration and purity were assessed with a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). cDNA was synthesized by reverse transcription of 1 µg total RNA with the EcoDry™ Premix Oligo dT Kit (Cat. No. 639543, Takara Bio Inc., Kyoto, Japan). In a total reaction volume of 25 µL, 1 µL cDNA template was combined with 2×TB Green Fast qPCR Mix (Takara Bio Inc.), and auxin-related gene expression levels were quantified by RT-qPCR, according to the manufacturer’s instructions. PCR samples were amplified using a Thermal Cycler Dice Real-Time PCR system (Takara Bio Inc.). The RT-qPCR cycling conditions were as follows: initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 10 s. A melting curve analysis was performed with dissociation steps at 95 °C for 15 s, 60 °C for 30 s, and 95 °C for 15 s. The PCR efficiency for each primer pair was calculated based on the equation E = [10(−1/slope) − 1] × 100%, using the slope of the standard curve (maximum CT value of < 40). The value obtained from this equation should be in the range of 90–110%76. Relative gene expression levels were calculated using the comparative threshold cycle (2−ΔΔCt) method and normalized to the expression of actin2 mRNA as an internal reference. Primers (Supplementary Table 6) were designed using GenScript Real-time PCR (TaqMan) Primer and Probes Design Tool (https://www.genscript.com/tools/real-time-pcr-taqman-primer-design-tool), with lengths of 20–26 nucleotides, a target product size of 50–400 bp, and annealing temperatures ranging from 52 °C to 60 °C. RT-qPCR experiments were conducted with three biological replicates.
Statistical analysis
For all experiments, data were analysed by Student’s t-test with equality of variance, and differences between samples treated under control conditions or with bacterial strains were considered significant at P values equal to or less than 0.05. All experiments were performed with at least three biological replicates.
Results and discussion
Microorganism Preparation and whole-genome sequencing
Microorganisms were isolated from the rhizosphere surrounding L. chinense seedlings based on colony morphology. One bacterial strain was designated as ‘A-2’, which formed round, mucilaginous, pale-yellow colonies on PDA agar plates (Fig. 1a). SEM analysis revealed cylindrical cells with flagella, indicating motility (Fig. 1b).
Characteristics of Pseudomonas sp. A-2. (a) Circos plot showing the completed assembly and annotation data generated using Proksee (CGview). From the outer to the inner circle : coding domain sequence (CDS), tRNA genes, tmRNA genes, rRNA genes, GC content, GC skew, base pairs (bp). (b) Morphology of the A-2 strain colonies on an agar plant after 24 h of growth at 30 °C. (c) Scanning electron microscopy image of the A-2 strain grown for 24 h at 30 °C, shown at 10,000x magnification. (d) Phylogenetic tree of Pseudomonas species, based on whole-genome sequencing of the 16S rRNA region, showing the relationships among various strains within the genus. The species highlighted with a pale orange and labeled as ‘Pseudomonas sp. A-2’ indicates the specific strain of interest. The tree was generated in CLC Main Workbench 22 using the neighbor-joining method with 1,000 bootstrap replicates. The scale bar represents 0.005 substitutions per nucleotide position.
Whole-genome sequencing (WGS) was performed using Nanopore (1.26 Gb, 188.6× coverage) and Illumina (2.48 Gb, 376.4× coverage) platforms. The assembled genome was found to comprise a single circular chromosome of 6.6 Mb (Fig. 1c), containing 5,896 protein-coding genes, 16 rRNA genes, 67 tRNA genes, and 1 tmRNA gene. BUSCO analysis confirmed 98.7% completeness (772 genes) (Table 1). Genome annotation revealed the presence of the tryptophan-2-monooxygenase (iaaM)77essential for IAM-mediated IAA biosynthesis, along with other IAA-related genes, including trpA–F (encoding tryptophan biosynthesis enzymes) and enoyl-CoA hydratase 1 (ech1) (Table 2). The EggNOG-based functional categorization is shown in Supplementary Fig. 4.
A phylogenetic tree based on the 16S rRNA gene sequence placed the A-2 strain within the genus Pseudomonas (Fig. 1d). Comparative genomic analyses with ten phylogenetically related Pseudomonas strains revealed ANI values ranging from 75.0 to 85.03%, below the species threshold of 95%78 (Supplementary Fig. S3a). Consistently, dDDH values calculated via GGDC ranged from 0.14 to 0.20 (Supplementary Fig. S3b), well below the 70% species delineation cutoff. Hierarchical clustering of ANI and dDDH values (Supplementary Fig. S3c and d) further supported the classification of the A-2 strain as a novel genome species within Pseudomonas.
Characterization of IAA production by Pseudomonas sp. A-2
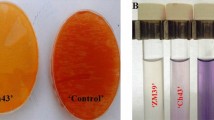
As Pseudomonas species are known to produce IAA, the newly isolated the A-2 strain was evaluated for its IAA production potential following the identification of IAM-related genes through genome annotation. Growth curve analysis revealed that maximum IAA production was achieved during the stationary phase, which was reached after approximately 20 h of culture (Fig. 2a). In the presence of 100 mg/L L-Trp, the strain produced 16.67 mg/L IAA, as measured by Salkowski’s reagent-based colorimetric assay (Fig. 2b, c, and Supplementary Table 2). The retention time for bacterially derived IAA at 280 nm was determined to be 5.68 min, which corresponded to that of the synthetic IAA standard (Fig. 2d and e).
Analysis of indole-3-acetic acid (IAA) extraction from Pseudomonas sp. A-2. (a) Growth curve of the A-2 strain grown in liquid potato dextrose agar (PDA) at 30 °C and 200 rpm over 24 h, with optical density values measured every 2 h. (b) Standard curve of synthetic IAA concentrations with Salkowski’s reagent. (c) IAA concentration of the A-2 strain isolates grown in the presence and absence of tryptophan, measured by spectrophotometry. Blank: liquid PDA medium mixed with Salkowski’s reagent. (d) Chromatogram of synthetic IAA (12.5 ppm). (e) Chromatogram of IAA extracted from the A-2 strain grown in the presence of 100 mg/mL tryptophan, showing a retention time at 280 nm. (f) Ultra-performance liquid chromatography was used to measure IAA concentrations following the A-2 strain isolate growth in the presence and absence of tryptophan. *P < 0.05; ****P < 0.0001. The results are presented as the mean ± standard deviation (n = 3).
For more accurate quantification, UPLC analysis was performed using individual standard curves, and the IAA concentration was determined to be 10.98 mg/L (Fig. 2f). Although the values obtained by Salkowski’s method and UPLC did not completely coincide, the difference between the two measurements was not substantial, supporting the validity of the overall IAA production estimation. Given that the Salkowski’s assay is a colorimetric screening tool with limited specificity, the UPLC-derived value is considered to be more accurate and reliable. Notably, although some microbes require L-Trp as a precursor for auxin biosynthesis79 the A-2 strain was capable of producing IAA even in the absence of L-Trp (Fig. 2c, f, and Supplementary Fig. 6), indicating the presence of an L-Trp independent biosynthetic pathway.
Effect of pseudomonas sp. A-2 on plant growth and lateral root formation
The effects of the A-2 strain treatment on A. thaliana were assessed by measuring fresh weight, LR number, and LR thickness between DAT1 and DAT7 (Fig. 3). Compared to untreated plants, the A-2 strain treatment led to a 1.3-fold increase in fresh weight, a 3.2-fold increase in LR number, and a 1.4-fold increase in LR thickness at DAT7 (Fig. 3a–d and Supplementary Table 3). LRs were first observed on day 3, and data collection was initiated on day 5. The A-2 strain, known to produce IAA, enhanced LR development near the treatment site while reducing primary root (PR) length, consistent with previous findings that auxin levels above 10⁻⁸ M suppress PR growth and stimulate LR formation80. However, when the A-2 strain was applied to the upper portion of the plant, PR length remained unaffected and LR formation resembled that of untreated controls (Fig. 3e). This suggests that the active compound is not a volatile organic compound (VOC), as VOCs are known to influence plant hormone levels81and high IAA levels can elevate ethylene, inhibiting root growth82. The absence of inhibitory effects implies that the IAA concentration produced by the A-2 strain is below the critical threshold.
Plant growth response to treatment with Pseudomonas sp. A-2. (a) Plant morphology of A. thaliana was observed 1, 3, 5, and 7 days after treatment (DAT) with a suspension of the A-2 strain applied at the middle of the vertical plate. (b) Fresh weight of A. thaliana was measured DAT 1, 3, 5, and 7 with the A-2 strain. (c) Lateral root number and (d) root thickness were measured DAT5 and DAT7 with the A-2 strain. (e) Growth differentiation based on the treatment position of the A-2 strain: growth was assessed DAT7 with a suspension solution applied at the top of the vertical plate. (f) Morphology of N. tabacum was observed DAT14 with a suspension of the A-2 strain applied at the middle of the vertical plate. (g) Fresh weight of N. tabacum was measured DAT14 with the A-2 strain. (h) Plant morphology of peanut plants DAT14 with a suspension of the A-2 strain applied to the soil. (i) Fresh weights of peanut plants were measured DAT14. Significant differences between control (C.) and Pseudomonas sp. A-2 treated plants (P. A-2) were evaluated using Student’s t-test : *P < 0.05; ** P < 0.01; *** P < 0.001; **** P < 0.0001. The results are presented as the mean ± standard deviation (n = 3).
Growth-promoting effects were also observed in N. tabacum, a model crop species frequently used in plant-microbe interaction studies. Application of the A-2 strain at the middle of the growth plate resulted in a 2-fold increase in total growth by DAT14, including elongation of both LR and PR (Fig. 3f, g, and Supplementary Table 4), whereas only PR elongation was noted in the control group.
Pot-based experiments were conducted to evaluated the effects of the A-2 strain on peanuts (A. hypogaea), a leguminous crop in which LR development and nodule formation originate from distinct cell types but are functionally interconnected, contributing to growth and pod development. Peanut sprouts also contain resveratrol, an antioxidant known for scavenging reactive oxygen species (ROS)83,84. Resveratrol is associated with hormonal pathways, including the biosynthesis and regulation of IAA and other auxins, while also scavenging ROS, providing a rationale for the experiment. Given that the A-2 strain enhances both root and shoot development, peanut was selected as a biologically and agronomically relevant system to evaluate whether root-promoting PGPR activity translates into increased biomass in a geocarpic legume where subterranean development is crucial for productivity. Compared with the untreated controls, the A-2 strain treated group exhibited enhanced shoot and root development and a 1.35-fold increase in total biomass (Fig. 3h and i).
Quantitative measurement of gene expression
To evaluate whether auxin biosynthesis in A. thaliana roots was influenced by treatment with the A-2 strain, RT-qPCR was performed on root samples collected at various time points after treatment. The expression level of auxin response factor 19 (ARF19), a central regulator of the auxin signaling pathway, was not significantly altered until DAT5, but was markedly elevated by DAT7 in the A-2 strain treated plants (Fig. 4a). ARF7 and ARF19 are critical for85,86 and play precise roles in the initiation of LR formation85,87. Therefore, the observed increases in expression suggests that the A-2 strain modulated auxin signaling during later developmental stages, leading to enhanced transcription of relevent genes.
Expression analysis of auxin and cytokinin biosynthesis genes in A. thaliana using RT-qPCR. (a) Expression levels of genes in the tryptophan to IAA synthesis pathway during early plant growth. A. thaliana seedlings were cultured on germination medium, and roots were sampled 1, 3, 5, and 7 days after treatment (DAT) with a suspension of the A-2 strain applied at the middle of the vertical plate. (b) Expression levels of genes in the cytokinin biosynthesis pathway during early growth stages. A. thaliana seedlings were cultured on germination medium, and roots were sampled DAT1, 3, 5, and 7. Relative gene expression levels were normalized to the untreated control (without the A-2 strain) and calibrated for each respective day. Significant differences between control (C.) and Pseudomonas sp. A-2 treated plants (P. A-2) were evaluated using Student’s t-test : *P < 0.05, **P < 0.01, ***P < 0.001. The results are presented as the mean ± standard deviation (n = 3).
To elucidated the biosynthetic pathways contributing to IAA production, the expression of several IAA biosynthesis genes was assessed. These included AMI1 (encoding amidase 1; EC 3.5.1.4) involved in the IAM pathway, and TAA1 (encoding tryptophan aminotransferase of Arabidopsis 1; EC 2.6.1.27), YUCCA1 (YUC, encoding Flavin monooxygenase; EC 1.14.13.168), and YUC2 from the IPA pathway. The expression of these genes, representing both IAM and IPA pathways, showed 3–6-fold increases in the A-2 strain treated plants at DAT7 (Fig. 4a). AMI1 is essential for auxin homeostasis and supports growth and stress tolerance88while TAA1, YUC1, and YUC2 contribute to IAA synthesis via the IPA pathway, acting cooperatively36. Given that TAA1 and YUCCAs are associated with the formation of auxin gradients essential for LR primordia (LRP) development, it is likely that the A-2 strain supplies an exogenous source of auxin during early LRP formation and stimulatneously promotes endogenous auxin biosynthesis.
Conversely, no significant changes were observed in AAO1 (encoding Arabidopsis aldehyde oxidase 1; EC 1.2.3.1), a key enzyme in the TAM pathway, or in NIT1 (encoding nitrilase 1; EC 3.5.5.1), involved in the IAOx pathway89suggesting that these pathways do not contribute to the incresed IAA levels observed (Fig. 4a).
IAA perception is mediated by auxin-binding protein (ABP) receptors, while its cellular import and export are regulated by auxin influx carriers (AUX) and efflux carriers (PIN), respectively. In this study, the expression of AUX1, which facilitates auxin import from vascular tissues and supports LR formation90,91remained unchanged following the A-2 strain treatment, indicating that auxin uptake mechanisms were not affected (Fig. 4a). Similarly, ABP1, which functions in auxin signaling rather than polar transport, showed no significant expression change. In contrast, PIN1, encoding a key auxin efflux carrier required for root development92,93was progressively upregulated, suggesting that the A-2 strain treatment may influence auxin distribution within root tissues. Although gain-of-function mutants of ABP1 have been shown to disrupt PIN polarity94the mechanism underlying the observed increase in PIN1 expression remains unclear and may involve complex auxin regulatory networks.
To further examine the impact of the A-2 strain treatment on cytokinin pathways in A. thaliana roots, RT-qPCR was performed on root samples collected at multiple time points (Fig. 4b). Among the cytokinin signaling genes analysed, only Arabidopsis response regulator 1 (ARR1) was significantly upregulated (2.5-fold at DAT7). As cytokinins regulate root architecture, and ARR1 mediates cytokinin signaling to repress auxin activity in quiescent center cells, its upregulation suggests enhanced cytokinin-meditaed signaling that may promote cell division and influence differentiation95. Additionally, AHK2, encoding a cytokinin receptor known to suppress LR formation in wild-type plants, was also upregulated at DAT7. However, no significant changes in LR induction were observed, implying that the increse in AHK2 expression likely reflects elevated cytokinin levels or sensitivity induced by the A-2 strain treatment, rather than a direct effect on LR development.
Adventitious root formation effects induced by pseudomonas sp. A-2 treatment
IAA production by the A-2 strain was confirmed, and its treatment was shown to enhance LR formation and overall growth in A. thaliana. Auxins, particularly IAA, are recognized as essential for both LR and AR formation. In plants, indole-3-butyric acid (IBA), a precursor of IAA, is converted into IAA, which is known to regulate AR development by triggering cellular reorganization in the hypocotyl to establish AR meristems96. To evaluate the effect of the A-2 strain treatment on AR formation, hypocotyl–root junctions of 5-day-old seedlings were excised, and the basal regions were treated with the A-2 strain (Fig. 5a). As a result, a 1.2-fold increase in fresh weight and a 2-fold increase in AR number were observed compared to untreated controls (Fig. 5b-d).
Regeneration of adventitious roots in A. thaliana induced by treatment with Pseudomonas sp. A-2. (a) In the assay for adventitious root formation induced by excision, the roots of 5-day-old seedlings were excised above the root–shoot junction and the medium was treated with the A-2 strain suspension. Observations and data collection were performed 10 days after hypocotyl excision and treatment (DAT) with the A-2 strain suspension on a vertical plate with A. thaliana. (b) Plant morphology, (c) fresh weight, and (d) number of adventitious roots were assessed DAT10. (e) Expression levels of genes involved in the IBA-to-IAA conversion pathway in A. thaliana were assessed using RT-qPCR in roots sampled DAT10. Relative gene expression levels were normalized to the untreated control (without the A-2 strain) and calibrated. Significant differences between control (C.) and Pseudomonas sp. A-2 treated plants (P. A-2) were evaluated using Student’s t-test : *P < 0.05, **P < 0.01. The results are presented as the mean ± standard deviation (n = 3).
To determine whether the conversion of IBA to IAA was affected by the A-2 strain treatment, RT-qPCR analysis was performed. The IBA transporter TOB1, which is responsible for the vacuolar sequestration of IBA and restriction of AR formation97was found to be upregulated 2.5-fold in the A-2 strain treated plants. This expression pattern was consistent with the upregulation of cytokinin-responsive genes such as ARR1 and AHK2, both of which have been reported to induce TOB1 expression. Nevertheless, AR formation was significantly enhanced by the A-2 strain treatment, indicating that TOB1-mediated repression of AR formation was not sufficient to counteract the promotive effects of the treatment.
Genes encoding key enzymes involved in IBA-to-IAA conversion, including ECH2 (encoding enoyl-CoA hydratase 2; EC 4.2.1.17) and the IBA response (IBR) proteins (IBR1, IBR3, and IBR10)98,99,100,101were also examined. The expression of IBR1, IBR10, and ECH2 was found to be upregulated, whereas no significant change was observed in IBR3 expression. These results suggest that AR formation was promoted by the A-2 strain treatment through the enhancement of IBA-to-IAA conversion and increased auxin biosynthesis, despite concurrent activation of cytokinin signaling pathways.
Pseudomonas sp. A-2 enhances salt stress tolerance in A. thaliana
To evaluate this potential, A. thaliana seedlings were cultivated on medium containing 0, 100, or 150 mM NaCl, and physiological as well as molecular responses were assessed after 14 days of the A-2 strain treatment (Figs. 6 and 7). Under non-stress conditions, shoot and root growth were slightly promoted by the A-2 strain treatment (Fig. 6b and c). Under salt stress, growth inhibition was markedly alleviated, as evidenced by significantly higher shoot and root fresh weights in the A-2 strain treated plants compared to untreated controls, indicating improved osmotic stress tolerance. Chlorophyll content, a key indicator of photosynthetic capacity, was better maintained in the A-2 strain treated plants under salt stress (Fig. 6d), suggesting protection of the photosynthetic machinery. In parallel, lipid peroxidation, measured via MDA content, was significantly reduced, indicating mitigation of oxidative stress (Fig. 6g). SOD activity was found to be elevated, particularly at 150 mM NaCl, reflecting enhanced ROS scavenging capacity (Fig. 6e). In support of this observation, antioxidant-related genes such as katA (catalase) and trxC (thioredoxin) were identified in the A-2 genome (Supplementary Table 1). Catalase is a key enzyme that decomposes hydrogen peroxide (H2O2) into water and oxygen, thereby protecting cells from oxidative damage102. Thioredoxin also plays a crucial role in maintaining redox homeostasis under stress conditions103. The presence of these genes suggests that the A-2 strain may directly contribute to enhanced ROS detoxification capacity in the plant through microbial enzymatic activity or redox-modulating signals.
Effects of Pseudomonas sp. A-2 on growth performance and physiological responses of A. thaliana under salt stress. (a) Morphological comparison of A. thaliana seedlings grown under different NaCl concentrations (0, 100, and 150 mM) with or without the A-2 strain treatment for 14 days. (b–g) Quantitative measurements of physiological parameters in shoots: shoot fresh weight (b), root fresh weight (c), total chlorophyll content (d), superoxide dismutase (SOD) activity (e), proline content (f), and malondialdehyde (MDA) content (g) of A. thaliana seedlings. (h–i) Soluble sugar content (h, Total Soluble Sugar; mg g− 1 FW) and trehalose content (i, mg g− 1 FW) under different salt stress conditions. Each bar represents the mean ± SD of three biological replicates. Significant differences between control (C.) and Pseudomonas sp. A-2 treated plants (P. A-2) were evaluated using Student’s t-test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001).
Expression analysis of salt-responsive genes in A. thaliana under NaCl stress with or without Pseudomonas sp. A-2 treatment. RT-qPCR analysis was conducted on five salt stress–responsive genes: RD20 (a), RD29A (b), RD29B (c), and KIN1 (d) in 10-day-old A. thaliana seedlings co-cultivated with or without the A-2 strain (OD₆₀₀ = 0.1) under NaCl treatments (0, 100, and 150 mM) for 3, 6, and 12 h. Gene expression levels were calculated using the 2^–ΔCT method and normalized against actin2 as an endogenous control. Data represent the mean ± standard deviation of three biological replicates. Significant differences between control (C.) and Pseudomonas sp. A-2 treated plants (P. A-2) were evaluated using Student’s t-test : *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. The results are presented as the mean ± standard deviation (n = 3).
Several salt stress–related genes, including opuAA, opuAB, osmY, osmV, and betB, were identified in the genome of the A-2 strain (Table 2). These genes are known to be involved in the transport and biosynthesis of osmoprotectants. For example, opuAA and opuAB are predicted to encode components of the OpuA ABC transporter system, which facilitates the uptake of glycine betaine. This compound functions as an important osmoprotectant in Bacillus subtilis under hyperosmotic stress conditions104. In addition, the presence of betB suggests a possible capacity for the endogenous synthesis of glycine betaine105. Taken together, these genomic features suggest that the A-2 strain may contribute to salt stress alleviation by producing compatible solutes and antioxidative metabolites. Furthermore, several regulatory genes were annotated in the A-2 genome, which may be involved in microbial signal transduction or the transcriptional regulation of stress responses. Among them, genes predicted to encode helix-turn-helix (HTH) type transcriptional regulators such as zntR, benM, betI, and yddM were identified106 (Supplementary Table 1). A gene encoding a sensor histidine kinase, rcsC, which functions as part of a two-component regulatory system, was also detected. These regulatory components have been reported to mediate environmental sensing and gene expression control in bacteria106,107and may also contribute to the modulation of plant–microbe interactions. Although plant gene expression was not directly assessed in this study, the presence of these elements supports the possibility that the A-2 strain may influence host stress signaling pathways through microbial sensing systems or metabolite-mediated interactions.
While only SOD activity was assessed in this study, further analyses of antioxidant-related genes and enzymes would help elucidate the underlying molecular mechanisms. Additionally, proline levels, which contribute to osmotic adjustment, were significantly increased in the A-2 strain treated plants under salt stress (Fig. 6f). Furthermore, total soluble sugars, including trehalose, which contribute to osmotic regulation and help protect cells under salt stress, were significantly increased in the A-2 strain treated plants compared to controls (Fig. 6h and i). These results suggest that the A-2 strain enhances salt stress tolerance in A. thaliana by promoting the accumulation of key osmoprotectants that support cellular osmotic balance and stress mitigation.
At the transcriptional level, salt stress was found to strongly induce the expression of RD20, RD29A, RD29B, and KIN1 in untreated control plants, whereas the induction of these genes was attenuated in the A-2 strain treated plants (Fig. 7). In previous studies, it was demonstrated that plants deficient in RD29A and RD29B genes show a greater increase in biomass under salt stress108. Therefore, the results of this study suggest that the reduced expression of RD29A and RD29B in the JBR18-treated group may contribute to the enhanced biomass accumulation ability of Arabidopsis under salt stress. This indicates that stress signaling was modulated, and transcriptional stress responses were attenuated by the A-2 strain, consistent with previous reports suggesting that PGPR mitigate salt stress by enhancing antioxidant defenses, modulating stress-responsive gene expression, and promoting the accumulation of osmoprotectants109.
Collectively, these findings demonstrate that salt stress tolerance in A. thaliana was enhanced by the A-2 strain through the protection of cellular functions, reinforcement of antioxidant defenses, and modulation of stress-inducible gene expression. Although osmotic and drought stress conditions were also tested, no consistent or significant improvements were observed in the A-2 strain treated plants under these conditions (data not shown). These results suggest that the stress-alleviating effects of the A-2 strain may be selectively exerted under salt stress conditions, possibly reflecting strain-specific stress response mechanisms.
Conclusions
In this study, whole-genome sequencing and functional characterization of the novel A-2 strain was conducted, revealing its potential as an eco-friendly biofertilizer. The A-2 strain was identified as a new taxon within the Pseudomonas genus and was shown to activate key genetic pathways involved in root development by promoting IAA biosynthesis and inducing the expression of genes associated with both lateral and adventitious root formation. In addition to promoting growth under non-stress conditions, the A-2 strain enhanced A. thaliana tolerance to salt stress by improving biomass accumulation, chlorophyll content, antioxidant enzyme activity, proline levels, and the accumulation of total soluble sugars, including trehalose, which function as osmoprotectants. Lipid peroxidation was reduced, and the expression of salt-responsive genes was modulated, indicating improved physiological adaptation and a role in stress signal regulation. Collectively, these findings demonstrate the multifaceted plant growth–promoting and stress-alleviating properties of the A-2 strain, underscoring its potential as a sustainable microbial agent to enhance crop productivity and resilience under adverse environmental conditions.
Data availability
The genome assembly of the strain named Pseudomonas sp. A-2 has been deposited in the Korea Bioinformation Center (https://www.kobic.re.kr/kobic), Korean Read Archive (KRA), under accession ID KAE23766264 and KAE23766265. The strain was deposited as a patent microorganism at the Korean Collection for Type Cultures (KCTC), and the deposit number is KCTC19172P.
Change history
22 October 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-24636-x
Abbreviations
- 3DW:
-
Sterile triple-distilled water
- AAI:
-
Average Amino Identity
- AAO1:
-
Arabidopsis aldehyde oxidase 1
- ABP1:
-
Auxin-binding protein 1
- AMI1:
-
Amidase 1
- ANI:
-
Average nucleotide identity
- AR:
-
Adventitious root
- ARF:
-
Auxin response factor
- ARR1:
-
Arabidopsis response regulator 1
- DAT:
-
Days after treatment
- dDDH:
-
Digital DNA-DNA hybridization
- ECH2:
-
Enoyl-CoA hydratase 2
- GGDC:
-
Genome-to-genome distance calculator
- IAA:
-
Indole-3-acetic acid
- IAM:
-
Indole-3-acetamide
- IAOX:
-
Indole-3-acetaldoxime
- IBA:
-
Indole-3-butyric acid
- IBR:
-
IBA response
- IPA:
-
Indole-3-pyruvic acid
- L-Trp:
-
L-tryptophan
- LR:
-
Lateral root
- MDA:
-
Malondialdehyde
- MS:
-
Murashige and skoog medium
- PDA:
-
Potato dextrose agar
- PGPR:
-
Plant growth–promoting rhizobacteria
- PIN:
-
Pin-formed proteins
- PR:
-
Primary root
- SEM:
-
Scanning electron microscopy
- SOD:
-
Superoxide dismutase
- TAA:
-
Tryptophan aminotransferase of arabidopsis
- TAM:
-
Tryptamine
- TOB1:
-
Transporter of IBA 1
- UPLC:
-
Ultra-performance liquid chromatography
- VOC:
-
Volatile organic compound
- WGS:
-
Whole-genome sequencing
- YUC:
-
YUCCA
References
Kloepper, J. W., Leong, J., Teintze, M. & Schroth, M. N. Enhanced plant growth by siderophores produced by plant growth-promoting rhizobacteria. Nature 286, 885–886 (1980).
Kim, J. A., Song, J. S., Jeong, M. H., Park, S. Y. & Kim, Y. Biocontrol of rice diseases by microorganisms. Res. Plant. Dis. 27, 129–136 (2021).
Mariotti, L., Scartazza, A., Curadi, M., Picciarelli, P. & Toffanin, A. Azospirillum baldaniorum Sp245 induces physiological responses to alleviate the adverse effects of drought stress in purple Basil. Plants 10, 1141 (2021).
Oh, K. et al. Plant Growth-Promoting activity characteristics of Bacillus strains in the rhizosphere. Microbiol. Biotechnol. Lett. 49, 403–412 (2021).
Ratnaningsih, H. R. et al. IAA and ACC deaminase producing-bacteria isolated from the rhizosphere of pineapple plants grown under different abiotic and biotic stresses. Heliyon 9, e16306 (2023).
Brunetti, C. et al. Effects of plant growth-promoting rhizobacteria strains producing ACC deaminase on photosynthesis, isoprene emission, ethylene formation and growth of Mucuna pruriens (L.) DC. in response to water deficit. J. Biotechnol. 331, 53–62 (2021).
Carlos, M. H. J., Stefani, P. V. Y., Janette, A. M., Melani, M. S. S. & Gabriela, P. O. Assessing the effects of heavy metals in ACC deaminase and IAA production on plant growth-promoting bacteria. Microbiol. Res. 188–189, 53–61 (2016).
Ali, S. Z., Sandhya, V. & Rao, L. V. Isolation and characterization of drought-tolerant ACC deaminase and exopolysaccharide-producing fluorescent Pseudomonas Sp. Ann. Microbiol. 64, 493–502 (2014).
Chen, Z. et al. Plant growth-Promoting rhizobacteria with ACC deaminase activity enhance maternal lateral root and seedling growth in Switchgrass. Front. Plant. Sci. 12, 1–14 (2022).
Alinia, M. et al. Co-application of ACC deaminase-producing rhizobial bacteria and melatonin improves salt tolerance in common bean (Phaseolus vulgaris L.) through ion homeostasis. Sci. Rep. 12, 1–13 (2022).
Abbasi, S., Sadeghi, A. & Safaie, N. Streptomyces alleviate drought stress in tomato plants and modulate the expression of transcription factors ERF1 and WRKY70 genes. Sci. Hortic. 265, 109206 (2020).
Rijavec, T. & Lapanje, A. Hydrogen cyanide in the rhizosphere: not suppressing plant pathogens, but rather regulating availability of phosphate. Front. Microbiol. 7, 1–14 (2016).
Qurashi, A. W. & Sabri, A. N. Bacterial exopolysaccharide and biofilm formation stimulate Chickpea growth and soil aggregation under salt stress. Braz J. Microbiol. 43, 1183–1191 (2012).
Akhtar, S. S. et al. Bacillus licheniformis FMCH001 increases water use efficiency via growth stimulation in both normal and drought conditions. Front. Plant. Sci. 11, 1–12 (2020).
Vejan, P., Abdullah, R., Khadiran, T. & Ismail, S. Nasrulhaq boyce, A. Role of plant growth promoting rhizobacteria in agricultural sustainability: A review. Molecules 21, 1–17 (2016).
Glick, B. R. Plant growth-promoting bacteria: mechanisms and applications. Scientifica 963401 (2012). (2012).
Tilocca, B., Cao, A. & Migheli, Q. Scent of a killer: microbial volatilome and its role in the biological control of plant pathogens. Front. Microbiol. 11, 41 (2020).
Chu, T. N., Tran, B. T. H., Van Bui, L. & Hoang, M. T. T. Plant growth-promoting rhizobacterium Pseudomonas PS01 induces salt tolerance in Arabidopsis Thaliana. BMC Res. Notes. 12, 1–7 (2019).
Alexander, A., Singh, V. K., Mishra, A. & Halotolerant, P. G. P. R. Stenotrophomonas maltophilia BJ01 induces salt tolerance by modulating physiology and biochemical activities of arachis hypogaea. Front. Microbiol. 11, 1–12 (2020).
Gupta, A. et al. ACC deaminase produced by PGPR mitigates the adverse effect of osmotic and salinity stresses in Pisum sativum through modulating the antioxidants activities. Plants 11, 3419 (2022).
Xiao, A. W., Li, Z., Li, W. C. & Ye, Z. H. The effect of plant growth-promoting rhizobacteria (PGPR) on arsenic accumulation and the growth of rice plants (Oryza sativa L). Chemosphere 242, 125–136 (2020).
Kumar, A., Singh, S., Mukherjee, A., Rastogi, R. P. & Verma, J. P. Salt-tolerant plant growth-promoting Bacillus pumilus strain JPVS11 to enhance plant growth attributes of rice and improve soil health under salinity stress. Microbiol. Res. 242, 126616 (2021).
Kumari, S., Vaishnav, A., Jain, S., Varma, A. & Choudhary, D. K. Bacterial-Mediated induction of systemic tolerance to salinity with expression of stress alleviating enzymes in soybean (Glycine max L. Merrill). J. Plant. Growth Regul. 34, 558–573 (2015).
Kim, J. S. et al. Gene expression profile affected by volatiles of new plant growth promoting rhizobacteria, Bacillus subtilis strain JS, in tobacco. Genes Genomics. 37, 387–397 (2015).
Kang, S. M. et al. Indole-3-acetic-acid and ACC deaminase producing leclercia adecarboxylata MO1 improves Solanum lycopersicum L. growth and salinity stress tolerance by endogenous secondary metabolites regulation. BMC Microbiol. 19, 1–14 (2019).
Nawaz, A. et al. Potential of salt tolerant PGPR in growth and yield augmentation of wheat (Triticum aestivum L.) under saline conditions. Front. Microbiol. 11, 1–12 (2020).
Shah, G. et al. Indole pyruvate decarboxylase gene regulates the auxin synthesis pathway in rice by interacting with the indole-3-acetic acid–amido synthetase gene, promoting root hair development under cadmium stress. Front. Plant. Sci. 13, 1–13 (2022).
Zhang, C. et al. Serratia marcescens PLR enhances lateral root formation through supplying PLR-derived auxin and enhancing auxin biosynthesis in Arabidopsis. J. Exp. Bot. 73, 3711–3725 (2022).
Alvarez, J. P., Goldshmidt, A., Efroni, I., Bowman, J. L. & Esheda, Y. The NGATHA distal organ development genes are essential for style specification in Arabidopsis. Plant. Cell. 21, 1373–1393 (2009).
Berckmans, B. et al. Auxin-dependent cell cycle reactivation through transcriptional regulation of Arabidopsis E2Fa by lateral organ boundary proteins. Plant. Cell. 23, 3671–3683 (2011).
Gallavotti, A. The role of auxin in shaping shoot architecture. J. Exp. Bot. 64, 2593–2608 (2013).
Su, Y. H., Liu, Y. B., Bai, B. & Zhang, X. S. Establishment of embryonic shoot-root axis is involved in auxin and cytokinin response during Arabidopsis somatic embryogenesis. Front. Plant. Sci. 5, 1–9 (2015).
Yun, F., Liu, H., Deng, Y., Hou, X. & Liao, W. The role of light-regulated auxin signaling in root development. Int. J. Mol. Sci. 24, 5253 (2023).
Zhang, Y. et al. Molecular mechanisms of diverse auxin responses during plant growth and development. Int. J. Mol. Sci. 23, 12495 (2022).
Kumar, K. V., Singh, N., Behl, H. M. & Srivastava, S. Influence of plant growth promoting bacteria and its mutant on heavy metal toxicity in Brassica juncea grown in fly Ash amended soil. Chemosphere 72, 678–683 (2008).
Mashiguchi, K. et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl. Acad. Sci. U S A. 108, 18512–18517 (2011).
Spaepen, S. & Vanderleyden, J. Auxin and plant-microbe interactions. Cold Spring Harb Perspect. Biol. 3, 1–13 (2011).
Mano, Y. & Nemoto, K. The pathway of auxin biosynthesis in plants. J. Exp. Bot. 63, 2853–2872 (2012).
Chandler, J. W. Local auxin production: A small contribution to a big field. BioEssays 31, 60–70 (2009).
Woodward, A. W., Bartel, B. & Auxin regulation, action, and interaction. Ann. Bot. 95, 707–735 (2005).
Pollmann, S., Müller, A. & Weiler, E. W. Many roads lead to ‘auxin’: of nitrilases, synthases, and amidases. Plant. Biol. 8, 326–333 (2006).
Mano, Y. et al. The AMI1 gene family: Indole-3-acetamide hydrolase functions in auxin biosynthesis in plants. J. Exp. Bot. 61, 25–32 (2010).
Normanly, J. Approaching cellular and molecular resolution of auxin biosynthesis and metabolism. Cold Spring Harb Perspect. Biol. 2, 1–17 (2010).
Zhao, Y. Auxin biosynthesis and its role in plant development. Annu. Rev. Plant. Biol. 2, 49–64 (2010).
Arshad, M. & Frankenberger, W. T. Plant growth-regulating substances in the rhizosphere: microbial production and functions. Adv. Agron. 62, 45–151 (1997).
Barazani, O. & Friedman, J. Is IAA the major root growth factor secreted from plant-growth-mediating bacteria? J. Chem. Ecol. 25, 2397–2406 (1999).
Biswas, J. C., Ladha, J. K., Dazzo, F. B., Yanni, Y. G. & Rolfe, B. G. Rhizobial inoculation influences seedling Vigor and yield of rice. Agron. J. 92, 880–886 (2000).
Jameson, P. E. Cytokinins and auxins in plant-pathogen interactions - An overview. Plant. Growth Regul. 32, 369–380 (2000).
Persello-Cartieaux, F., Nussaume, L. & Robaglia, C. Tales from the underground: molecular plant-rhizobacteria interactions. Plant. Cell. Environ. 26, 189–199 (2003).
Saeed, K. S., Ahmed, S. A., Hassan, I. A. & Ahmed, P. H. Effect of bio-fertilizer and chemical fertilizer on growth and yield in cucumber (cucumis sativus) in green house condition. Pakistan J. Biol. Sci. 18, 129–134 (2015).
Wintermans, P. C. A., Bakker, P. A. H. M. & Pieterse, C. M. J. Natural genetic variation in Arabidopsis for responsiveness to plant growth-promoting rhizobacteria. Plant. Mol. Biol. 90, 623–634 (2016).
Zerrouk, I. Z. et al. Growth and aluminum tolerance of maize roots mediated by auxin- and cytokinin-producing Bacillus toyonensis requires Polar auxin transport. Environ. Exp. Bot. 176, 104064 (2020).
Acosta-Motos, J. R. et al. Plant responses to salt stress: adaptive mechanisms. Agronomy 7, 1–38 (2017).
Ali, B. et al. PGPR-Mediated salt tolerance in maize by modulating plant physiology, antioxidant defense, compatible solutes accumulation and Bio-Surfactant producing genes. Plants 11, 345 (2022).
Li, H. P., Ma, H., Bin & Zhang, J. L. Halo-tolerant plant growth-promoting bacteria-mediated plant salt resistance and microbiome-based solutions for sustainable agriculture in saline soils. FEMS Microbiol. Ecol. 101, fiaf037 (2025).
Sandhya, V., Grover, Z. A. S., Reddy, M., Venkateswarlu, B. & G. & Alleviation of drought stress effects in sunflower seedlings by the exopolysaccharides producing Pseudomonas Putida strain GAP-p45. Biol. Fertil. Soils. 46, 17–26 (2009).
Vurukonda, S. S. K. P., Vardharajula, S., Shrivastava, M. & SkZ, A. Enhancement of drought stress tolerance in crops by plant growth promoting rhizobacteria. Microbiol. Res. 184, 13–24 (2016).
Frazzetto, G. Dialogue between the disciplines is thriving. Nature 458, 702 (2009).
Jaryenneh, J., Schoeniger, J. S. & Mageeney, C. M. Genome sequence and characterization of a novel Pseudomonas Putida phage, MiCath. Sci. Rep. 13, 1–9 (2023).
Verhoeve, V. I. et al. Genome sequencing of Pseudomonas aeruginosa strain M2 illuminates traits of an opportunistic pathogen of burn wounds. G3 Genes, Genomes, Genet. 12, jkac073 (2022).
Zhang, Y. & Palmgren, M. Gene-editing in plants no longer requires tissue culture. Front. Agric. Sci. Eng. 7, 229–230 (2020).
Ayangbenro, A. S., Olanrewaju, O. S., Amoo, A. E. & Babalola, O. O. Pseudomonas Koreensis AB36 from a gold mine: genomic insights into adaptation to extreme conditions and potential plant growth promotion. Biologia 79, 3677–3694 (2024).
Rikame, T., Borde, M. W. & Genome Functional annotation and comparative genomics of plant Growth-Promoting bacteria Pseudomonas aeruginosa (NG61) with potential application in Agro-Industry. Curr. Microbiol. 79, 1–12 (2022).
Chen, Z., Erickson, D. L. & Meng, J. Benchmarking long-read assemblers for genomic analyses of bacterial pathogens using Oxford nanopore sequencing. Int. J. Mol. Sci. 21, 1–27 (2020).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Wood, D. E., Salzberg, S. L. & Kraken Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Huang, Y. T., Liu, P. Y. & Shih, P. W. Homopolish: a method for the removal of systematic errors in nanopore sequencing by homologous Polishing. Genome Biol. 22, 1–17 (2021).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Hunt, M. et al. Circlator: automated circularization of genome assemblies using long sequencing reads. Genome Biol. 16, 1–10 (2015).
Seemann, T. & Prokka Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Lee, I., Kim, Y. O., Park, S. C. & Chun, J. OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103 (2016).
Ehmann, A. The Van URK-Salkowski reagent - a sensitive and specific chromogenic reagent for silica gel thin-layer chromatographic detection and identification of Indole derivatives. J. Chromatogr. A. 132, 267–276 (1977).
Dubois, M., Gilles, K. A., Hamilton, J. K., Rebers, P. T. & Smith, F. Colorimetric method for determination of sugars and related substances. Anal. Chem. 28, 350–356 (1956).
Magazin, N. & Popovic, M. Determination of Trehalose in plant tissues by anthrone-sulfuric acid method. Archives Biol. Sci. 62, 423–427 (2010).
Radonić, A. et al. Reference gene selection for quantitative real-time PCR analysis in virus infected cells: SARS Corona virus, yellow fever virus, human Herpesvirus-6, Camelpox virus and cytomegalovirus infections. Virol. J. 2, 1–5 (2005).
Palm, C. J., Gaffney, T. & Kosuge, T. Cotranscription of genes encoding indoleacetic acid production in Pseudomonas syringae subsp. Savastanoi. J. Bacteriol. 171, 1002–1009 (1989).
Chun, J. et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 68, 461–466 (2018).
Ahmad, F., Ahmad, F., Ahmad, I. & Khan, M. S. Indole acetic acid production by the Indigenous isolates of Azotobacter and fluorescent Pseudomonas in the presence and absence of Tryptophan. Turk. J. Biol. 29, 29–34 (2005).
Ivanchenko, M. G., Napsucialy-Mendivil, S. & Dubrovsky, J. G. Auxin-induced Inhibition of lateral root initiation contributes to root system shaping in Arabidopsis Thaliana. Plant. J. 64, 740–752 (2010).
Ryu, C. M. et al. Bacterial volatiles promote growth in Arabidopsis. Proc. Natl. Acad. Sci. U. S. A. 100, 4927–4932 (2003).
Burg, S. P. & Burg, E. A. The interaction between auxin and ethylene and its role in plant growth. Proc. Natl. Acad. Sci. U.S.A. 55, 262–269 (1966).
De La Alarcón, C. & Villegas, I. Resveratrol as an antioxidant and pro-oxidant agent: mechanisms and clinical implications. Biochem. Soc. Trans. 35, 1156–1160 (2007).
Kang, H. I. et al. Antioxidative effects of peanut sprout extracts. J. Korean Soc. Food Sci. Nutr. 39, 941–946 (2010).
Overvoorde, P. J. et al. Functional genomic analysis of the AUXIN/INDOLE-3-ACETIC ACID gene family members in Arabidopsis Thaliana. Plant. Cell. 17, 3282–3300 (2005).
Wilmoth, J. C. et al. NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lateral root formation. Plant. J. 43, 118–130 (2005).
Okushima, Y., Fukaki, H., Onoda, M., Theologis, A. & Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant. Cell. 19, 118–130 (2007).
Pérez-Alonso, M. M. et al. Endogenous indole-3-acetamide levels contribute to the crosstalk between auxin and abscisic acid, and trigger plant stress responses in Arabidopsis. J. Exp. Bot. 72, 459–475 (2021).
Lehmann, T. et al. Arabidopsis NITRILASE 1 contributes to the regulation of root growth and development through modulation of auxin biosynthesis in seedlings. Front. Plant. Sci. 8, 1–15 (2017).
Marchant, A. et al. AUX1 promotes lateral root formation by facilitating indole-3-acetic acid distribution between sink and source tissues in the Arabidopsis seedling. Plant. Cell. 14, 589–597 (2002).
De Smet, I. et al. Auxin-dependent regulation of lateral root positioning in the basal meristem of Arabidopsis. Development 134, 681–690 (2007).
Atkinson, J. A. et al. Branching out in roots: Uncovering form, function, and regulation. Plant. Physiol. 166, 538–550 (2014).
Omelyanchuk, N. A. et al. A detailed expression map of the PIN1 auxin transporter in Arabidopsis Thaliana root. BMC Plant. Biol. 16, 1–12 (2016).
Gelová, Z. et al. Developmental roles of auxin binding protein 1 in Arabidopsis Thaliana. Plant. Sci. 303, 110750 (2021).
Zhang, W., Swarup, R., Bennett, M., Schaller, G. E. & Kieber, J. J. Cytokinin induces cell division in the quiescent center of the Arabidopsis root apical meristem. Curr. Biol. 23, 1979–1989 (2013).
Damodaran, S. & Strader, L. C. Factors governing cellular reprogramming competence in Arabidopsis adventitious root formation. Dev. Cell. 59, 1–14 (2024).
Michniewicz, M. et al. Transportar of IBA1 links auxin and cytokinin to influence root architecture. Dev. Cell. 50, 599–609 (2019).
Zolman, B. K., Yoder, A. & Bartel, B. Genetic analysis of Indole-3-butyric acid responses in. Genet 156, 1323–1337 (2000).
Zolman, B. K., Nyberg, M. & Bartel, B. IBR3, a novel peroxisomal acyl-CoA dehydrogenase-like protein required for indole-3-butyric acid response. Plant. Mol. Biol. 64, 59–72 (2007).
Zolman, B. K., Martinez, N., Millius, A., Adham, A. R. & Bartel, B. Identification and characterization of Arabidopsis indole-3-butyric acid response mutants defective in novel peroxisomal enzymes. Genetics 180, 237–251 (2008).
Strader, L. C. et al. Multiple facets of Arabidopsis seedling development require indole-3-butyric acid-derived auxin. Plant. Cell. 23, 984–999 (2011).
Chelikani, P., Fita, I. & Loewen, P. C. Diversity of structures and properties among catalases. Cell. Mol. Life Sci. 61, 192–108 (2004).
Meyer, Y., Reichheld, J. P. & Vignols, F. Thioredoxins in Arabidopsis and other plants. Photosynth Res. 86, 419–433 (2005).
Kemf, B. & Bremer, E. OpuA, an osmotically regulated binding protein-dependent transport system for the osmoprotectant Glycine betaine in Bacillus subtilis. J. Biol. Chem. 270, 16701–16713 (1995).
Boch, J., Kempf, B., Schmid, R. & Bremer, E. Synthesis of the omsoprotectant Glycine betaine in Bacillus subtilis: characterization of the GbsAB genes. J. Bacteriol. 178, 5121–5129 (1996).
Stock, A. M., Robinson, V. L. & Goudreau, P. N. Two-component signal transduction. Annu. Rev. Biochem. 69, 183–215 (2000).
Martínez-Hidalgo, P. & Hirsch, A. M. The nodule microbiome: N2-fixing rhizobia do not live alone. Phytobiomes J. 1, 70–82 (2017).
Msanne, J. et al. Characterization of abiotic stress-responsive Arabidopsis Thaliana RD29A and RD29B genes and evaluation of transgenes. Planta 234, 97–107 (2011).
Peng, Y. et al. Pseudoxanthomonas sp. JBR18, a halotolerant endophytic bacterium, improves the salt tolerance of Arabidopsis seedlings. Plant. Physiol. Biochem. 207, 108–415 (2024).
Acknowledgements
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (2023R1A2C1006404, RS-2024-00440009, RS-2024-00400556) to A.Y.S., and the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Research Initiative Program (KGM9942522 and KGM1002521) to A.Y.S. and S.Y.K.
Funding
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (2023R1A2C1006404, RS-2024-00440009, RS-2024-00400556) to A.Y.S., and the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Research Initiative Program (KGM9942522 and KGM1002521) to A.Y.S. and S.Y.K.
Author information
Authors and Affiliations
Contributions
AYS, SYK, and YKK conceived the project, designed the analysis, and organized the manuscript. SL isolated bacterial strain, performed the root formation, and abiotic stress experiment. HN and NK performed the bacteria data analysis. JA measured IAA concentration from bacteria. AHC, HK, SL, and SL performed the gene expression experiment and grew the plant material. YP performed the production of AAI and dDDH figures. AYS, SL, and SYK wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the name of the author Hualin Nie, which was incorrectly given as Haulin Nie.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, S., Kim, Y.K., Nie, H. et al. Functional characterization of a novel plant growth-promoting rhizobacterium enhancing root growth and salt stress tolerance. Sci Rep 15, 30405 (2025). https://doi.org/10.1038/s41598-025-14065-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14065-1