Abstract
Alzheimer’s disease (AD) presents a pressing global health challenge, demanding improved strategies for early detection and understanding its progression. In this study, we address this need by employing survival analysis techniques to predict transition time from Cognitive Normal (CN) to Mild Cognitive Impairment (MCI) in elderly individuals, considering the predictive value of baseline comorbidities. Leveraging data from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) and Australian Imaging, Biomarker & Lifestyle Flagship Study of Ageing (AIBL) databases, we construct feature sets encompassing demographics, cognitive scores, and comorbidities. Various machine learning and deep learning methods for survival analysis are employed. Our top-performing model, fast random forest, achieves a concordance index of 0.84 when considering all feature modalities, with comorbidity data emerging as a significant predictor. The top features identified by the best-performing model include one demographic feature (age), seven cognitive scores (ADAS13, RAVLT learning, FAQ, ADAS11, RAVLT immediate, CDRSB, ADASQ4), and two comorbidities (Endocrine & Metabolic, Renal & Genitourinary). Age is highlighted as the most influential predictor, while cognitive scores are crucial indicators of Alzheimer’s disease. External validation against the AIBL dataset affirms the robustness of our approach. Overall, our study contributes to a deeper understanding of the role of baseline comorbidities in AD risk prediction and emphasizes the importance of incorporating comprehensive feature assessment in clinical practice for early diagnosis and personalized treatment planning.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is a degenerative brain disorder that causes a steady deterioration in cognitive function1. It is the primary cause of dementia in the aging population, and is typically marked by a gradual onset of cognitive decline preceding observable clinical symptoms by approximately 20 years which presents a substantial window of opportunity for potential intervention in disease progression2. A global study estimating the prevalence of dementia worldwide reveals that the number of affected individuals is projected to increase from 57.4 million cases in 2019 to 152.8 million cases by 20503. While treatments, including medications, may offer some mitigation of AD progression, a definitive cure remains elusive4. This escalating global prevalence, coupled with the absence of a definitive treatment, highlights the urgent need for deeper insights into disease characteristics and effective strategies for the detection of the disease, specially in the pre-symptomatic/early stage.
The progression of Alzheimer’s disease can be divided into three main stages: preclinical/cognitive normal (CN)5, preclinical stage where brain changes occur without obvious symptoms; mild cognitive impairment (MCI)6, where cognitive decline and mild memory problems are present but do not yet significantly interfere with daily functioning; and dementia due to AD7, where symptoms become more prominent and cause significant disruptions to daily functioning, such as confusion, memory loss, and behavioral and personality changes. Usually, patients develop mild cognitive impairment, and are at an increased risk of gradually progressing to Alzheimer’s dementia. Early detection of AD relies on identifying CN individuals who are at risk of progressing to MCI.
There is no shortage of studies employing machine learning and deep learning techniques for the classification of Alzheimer’s disease stages and achieving great levels of accuracy 8,9,10. In the literature, survival analysis has been compared to binary classification methods, highlighting its advantages, particularly in the context of medical prognosis and disease progression prediction. It provides valuable insights into the time until an event, such as disease progression or death, rather than merely predicting whether an event will occur. This helps in understanding the temporal dynamics of disease progression11. Additionally, survival analysis allows for continuous risk assessment over time, enabling the prediction of survival probabilities at various time points. These features make survival analysis a powerful tool for predicting and managing disease progression in clinical settings. Moreover, its major advantage over traditional regression techniques lies in its capability to handle censored data — individuals who remain asymptomatic or lost to follow up—thereby leveraging the entirety of available data11. In recent years, survival analysis has garnered increased attention within the machine learning literature focused on predicting Alzheimer’s disease progression, with many studies utilizing these techniques to investigate the transition from Mild Cognitive Impairment to AD across various data modalities. In 2020, a study investigated MCI to AD conversion prediction using longitudinal multivariate predictive models12. They compared stable and progressive MCI groups and employed survival analysis with the extended Cox model and logistic regression on ADNI data. The dataset included structural magnetic resonance imaging (sMRI) and neuropsychological measures of 321 participants at baseline and follow-up points up to 36 months, comprising 51 features. Several other studies combined information from various modalidties such as sMRI, Fluorodeoxyglucose-Positron Emission Tomography (FDG-PET), genetics, and neuropsychological tests together or in various combinations 13,14,15,16. The progression of CN to AD has also been studied with survival analysis techniques. Notably, one particular study compared machine learning techniques across two datasets using many modalities (demographics, apolipoprotein E (APOE) status, cognitive scores (CS), medical history, family history, medical examination, blood test and adverse events)17. Deep learning survival analysis approaches have also garnered considerable attention in recent studies, which have been applied to examine both CN to AD progression18,19 and MCI to AD progression19,20 scenarios using multiple data modalities.
However, there has been notably limited exploration of survival analysis in studying the conversion from CN to MCI. To the best of our knowledge, the only previous work that employed machine learning survival analysis techniques from CN to MCI conversion using the ADNI dataset was presented in14, where researchers utilized the ADNI dataset to investigate the duration of conversion from CN to MCI and from MCI to AD, employing various data modalities such as MRI, PET, blood biomarkers, clinical data, and demographics, yielding a total of 1,085 features. The study incorporated survival analysis techniques and reported a concordance index of 0.66, denoting the model’s predictive accuracy in determining CN to MCI conversion times. Previous studies in the literature predominantly focus on survival analysis of the progression from CN or MCI to AD, considering dementia as the event of interest. However, there exists a notable gap in research concerning the survival analysis of CN to MCI, with MCI as the event. Addressing this gap is crucial as early detection of MCI presents a pivotal opportunity for intervention and therapeutic strategies, ultimately aiding in the management and potentially delaying the onset of Alzheimer’s disease.
Furthermore, this limited number of studies investigating CN to MCI conversion or early-stage AD progression is accompanied by the predominant focus of existing survival analysis studies on some specific types of data, including neuroimaging, cognitive scores,and genetics. While neuroimaging is pivotal in confirming structural alterations linked to AD, its utility in early detection is hindered by drawbacks like high costs, limited specificity, and the need for testing at single time points. This is often due to delayed concerns from patients or their families, which prompt assessments at specific instances rather than at multiple intervals. Although multiple time point testing is feasible and may provide a more comprehensive view of disease progression, it requires additional resources and may not always be practical in clinical settings. In contrast, Electronic Health Records (EHR) offer a promising avenue for AD prediction, providing comprehensive patient information, longitudinal assessment, accessibility, cost-effectiveness, and a holistic approach to care, without the necessity for specialized imaging equipment or procedures. Comorbidities and sociodemographic traits have been linked to an increased risk of death in AD and dementia patients. Nonetheless, the findings have often been inconsistent with respect to the exact impact of comorbidities and sociodemographic variables 21,22,23,24,25. Comorbidity is defined as the coexistence of multiple diseases alongside the primary condition being considered within an individual26. The incorporation of comorbidity data within structured EHR presents an exciting opportunity for data mining through machine learning algorithms, potentially leading to more precise and efficient methods for AD classification, and thereby enabling earlier diagnosis and intervention. Comorbidity data encompasses the documentation of multiple coexisting medical conditions or diseases in a patient’s health record, and has been correlated with an elevated risk of mortality in AD and dementia patients, offering a comprehensive and multidimensional approach to AD prediction27. The underlying hypothesis posits that individuals with similar historical disease experiences are more likely to develop similar diseases in the future28.
Comorbidities, or the simultaneous presence of multiple medical conditions in an individual, are prevalent in AD patients and exert a substantial influence on disease trajectory 28,29,30. The interplay between AD and comorbidities represents a complex yet potentially revealing aspect of the disease’s progression. Understanding these interactions can provide crucial insights into the predictive associations, temporal patterns, and clinical outcomes associated with AD. In previous survival analysis studies, the patients’ medical history was included among the selected features for AD prediction17. However, there was a lack of detailed examination into the individual impact of these specific medical data on the prediction of AD.
The contributions of this work are multi-faceted. Firstly, it expands the scope of existing research by focusing on the CN to MCI conversion, a stage that has received limited attention in survival analysis studies as compared to CN to AD conversion. By specifically targeting this transition, the study aims to provide a more comprehensive understanding of early-stage AD progression and identify important predictors. Secondly, the study incorporates baseline comorbidities alongside demographics and cognitive scores, introducing a novel approach to feature selection that considers the holistic health profile of individuals. This integration of comorbidity data from EHRs offers a promising avenue by investigating the potential predictive role of comorbidities in AD progression. Moreover, the study contributes to methodological advancements by evaluating the performance of various machine learning and deep learning algorithms in predicting survival time to early-stage Alzheimer’s, considering the challenges posed by heterogeneous and censored data. By comparing and contrasting the efficacy of these algorithms, the study aims to identify the most suitable model for AD risk prediction, enhancing the accuracy and reliability of early detection methods. Additionally, the study employs Partial Dependence Plots (PDPs) to gain deeper insights into the impact of selected features on model predictions, providing a systematic approach to understanding the predictive relationship between individual features and progression to MCI.
Overall, this work addresses critical gaps in the literature regarding the application of ML survival analysis techniques in studying early-stage AD progression, introduces a novel approach to feature selection incorporating baseline comorbidities, and contributes to methodological advancements in AD prediction using machine learning and deep learning algorithms. By advancing our understanding of early-stage AD progression and enhancing the accuracy of AD prediction models, this study aims to pave the way for earlier diagnosis and intervention strategies, ultimately improving patient outcomes and quality of life.
Organization of the paper
The remainder of this paper is organized as follows: Section 2 presents the cohort characteristics and the performance results of the proposed models. Section 3 provides a discussion of the findings, the value of incorporating comorbidities as predictive features in Alzheimer’s disease modeling. Section 4 outlines the materials and methods, including dataset details, model selection, and evaluation procedures. Finally, Section 5 concludes the study and proposes directions for future research.
Results
Cohort characteristics
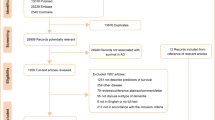
In this work, we use data from the publicly available Alzheimer’s Disease Neuroimaging Initiative (ADNI)3132 for model development and Australian Imaging, Biomarker & Lifestyle Flagship Study of Ageing (AIBL)33 for external validation. All participants with a baseline of Cognitive Normal, at least one follow-up visit, and documented medical history were selected. In ADNI, there are 389 censored and 105 transitioned individuals, while the AIBL dataset consists of 290 censored and 30 transitioned individuals. All their baseline variables are summarized in Table 1. In ADNI, we notice a statistically significant difference in age between censored (mean = 73.84, SD = 5.84) and transitioned (mean = 75.80, SD = 5.65) groups (p = 0.002). Additionally, although not significant, a higher percentage of males transitioned compared to censored individuals (56.2% vs. 47.0%, p = 0.120). In terms of cognitive scores, significant differences were observed in several measures between censored and transitioned groups. For instance, the Clinical Dementia Rating Scale Sum of Boxes (CDRSB), Alzheimer’s Disease Assessment Scale (ADAS), Rey Auditory Verbal Learning Test (RAVLT), and Functional Activities Questionnaire (FAQ) showed statistically significant differences between the two groups. Regarding comorbidity features, the presence of certain conditions differed significantly between censored and transitioned individuals. Notably, there was a higher prevalence of Endocrine Metabolic (46.3% vs. 57.1%, p = 0.062) and Renal Genitourinary (42.2% vs. 57.1%, p = 0.009) conditions in transitioned individuals compared to censored individuals. In comparison to ADNI, AIBL participants tend to be slightly younger (mean age: 72.32 vs. 73.84 years) but exhibit comparable levels of educational attainment and racial distribution. Notably, AIBL has a higher proportion of males in the censored group compared to ADNI (55.9% vs. 47.0%). AIBL also demonstrates similar trends in cognitive scores, albeit with variations in specific measures. Regarding comorbidities, AIBL exhibits differences in the prevalence of certain conditions compared to ADNI, suggesting potential variations in health profiles between the two cohorts. For instance, in endocrine and metabolic conditions, in ADNI, both censored and transitioned participants had over 45% prevalence, whereas in AIBL, both transitioned and censored individuals were below 17%.
Performance of the machine and deep learning models
In this study, we conducted an extensive evaluation of machine learning and deep learning survival analysis models to predict early stage Alzheimer’s disease progression. The models—Cox proportional hazards (CoxPH), recursive partitioning for survival trees (Rpart), random survival forest (RSF), fast random survival forest (FastRSF), cross-validated generalized linear model via penalized maximum likelihood (CVGlmnet), DeepSurv, DeepHit, and CoxTime—were compared across four distinct feature sets (FS) (FS1, FS2, FS3, and FS4), each combining demographics, cognitive scores, and comorbidities in varying ways. The workflow of our models is detailed in the Methods section.
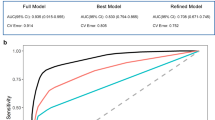
The performance of our machine learning models on the ADNI dataset across the four different feature sets (FS1, FS2, FS3, and FS4) is presented in Fig. 1 in the form of a heatmap that shows the mean value of the C-index. All reported results are based on the unseen testing data. As a benchmark, the Cox proportional hazards model (first column) is included. Among the evaluated models, both the fast random survival forest machine learning model and the DeepSurv deep learning model excelled, achieving a C-index of 0.84 when applied to FS1. The CoxPH model, Rpart, and RSF achieved C-indices of 0.82, 0.75, and 0.76, respectively. CVGlmnet and CoxTime yielded moderate results, with C-indices of 0.67 and 0.75. The Deephit model demonstrated a slightly lower C-index of 0.66. This feature set incorporates all data modalities, resulting in superior performance compared to the other feature sets. To complement these findings, we conducted bootstrapping (1,000 resamples) on FS1 to obtain 95% confidence intervals for the C-index (Table 3). Fast RSF achieved the highest mean C-index of 0.8607 [0.8107–0.9107], confirming its strong and stable predictive performance. DeepSurv and RSF followed closely with 0.8226 [0.7726–0.8726] and 0.7844 [0.7344–0.8344], respectively. We conducted Kruskal–Wallis test which revealed significant global differences (\(\chi ^2\) = 60.56, df = 7, \(p < 0.0001\)), and Dunn’s post-hoc comparisons with Holm correction confirmed that Fast RSF significantly outperformed DeepSurv (p = 0.041), as well as other baseline models such as CoxPH (p = 0.002) and DeepHit (\(\text {p} < 0.001\)). These findings provide statistical support for selecting Fast RSF as the top-performing model.
Heatmap displaying the mean concordance index-measured performance of each machine learning algorithm with each feature set (FS) on the ADNI dataset. Mean C-index values are computed over outer CV folds, without bootstrapping. Confidence intervals are reported separately for final model comparisons. Abbreviations: Dem = demographics, CS = cognitive scores, Com = comorbidities.
Excluding comorbidities (FS3) led to a significant performance drop, with the C-index falling to 0.76 for the fast random survival forest and 0.33 for DeepSurv. This decrease highlights the pivotal role of comorbidity information in enhancing the model’s predictive power, especially evident in complex models like Deepsurv. The CoxPH and RSF models attained C-indices of 0.75 and 0.76, respectively. Rpart and CVGlmnet achieved scores of 0.5 and 0.68, while Deephit showed a moderate value of 0.61. CoxTime delivered 0.66. Similarly, excluding cognitive scores (FS2) resulted in a C-index drop to 0.59 for the fast random survival forest and 0.74 for DeepSurv, which is an expected decrease knowing the importance of cognitive scores in predicting outcomes. CoxPH and CoxTime dropped to 0.62 and 0.59, respectively. Rpart and RSF showed scores of 0.5 and 0.59, while CVGlmnet and Deephit reached 0.58 and 0.53, respectively. When using FS4, which only includes demographics, the performance of most models declined. The fast random survival forest model’s c-index dropped sharply to 0.48, indicating a significant loss in predictive accuracy. Other models, such as CVGlment and Rpart, also showed lower c-index values of 0.50 and 0.56, respectively. Interestingly, the Cox model maintained a more stable performance compared to other feature sets, also, Deephit maintained a relatively stable performance with a slight decrease to 0.65. Overall, the inclusion of comorbidities and cognitive scores (FS1) significantly enhances the predictive accuracy of most models compared to using only demographic features (FS4).
The average C-index across all eight models was 0.76 for FS1, decreasing to 0.67 for FS3 and even lower for FS2. Notably, the fast random survival forest consistently outperformed other models when all data modalities were included.
Our top models, achieving a c-index of 0.84, have surpassed previous survival analysis studies conducted on the same dataset (ADNI) and cohort (CN to MCI), which attained c-index scores of 0.6614 and 0.6816, respectively. Further details of the comparison, including the features and models used in each study, are provided in Table 2. Our approach yields a significant enhancement in the early prediction of Alzheimer’s disease progression. By leveraging three cost-effective and non-invasive modalities, we contrast favorably with previous approaches that relied on expensive and invasive techniques such as MRI, PET scans, and blood biomarkers. This deliberate selection of features and models underscores our ability to achieve highly promising outcomes, demonstrating that readily available clinical data can be sufficient for accurate prediction.
Predictive features
Having identified the two top-performing models with a c-index of 0.84 using FS1, which included demographic information, comorbidities, and cognitive scores, we now proceed to perform a statistical test of significance to determine if these top two performing models are significantly different in performance compared to other models and possibly between themselves. As described in the methodology, we employed the Kruskal–Wallis test to assess whether there were statistically significant differences in the predicted risk scores across the eight models trained on the FS1 feature set. This non-parametric test evaluates the global differences in model predictions without assuming normality. The resulting chi-squared statistic was 5.623 (df = 7, p = 0.5844), indicating no significant difference in median risk rankings among the models.
Despite the lack of statistical significance, we proceeded with model selection based on both predictive performance and practical relevance. Fast Random Survival Forest and DeepSurv emerged as the top-performing models by C-index, with RSF achieving the highest overall score (0.84). To further support this choice, we conducted a second Kruskal–Wallis test on the bootstrapped C-index distributions across models, followed by Dunn’s post-hoc test with Holm correction. These additional analyses revealed that Fast RSF performed significantly better than most baseline models, reinforcing its suitability for downstream interpretation. Ultimately, we selected Fast RSF as the model of focus due to its strong performance, model stability across imputations, and interpretability in clinical contexts. This selection reflects a balance between statistical evidence and the practical needs of transparent decision-making in medical applications.
The features influencing the outcomes of the top-performing model (fast random survival forest on FS1) will undergo further analysis. Initially, the model ranks these features using a “permutation” method. Subsequently, we identify and select the top 10 features based on this ranking which were : ADAS13, AGE, RAVLT learning, FAQ, ADAS11, RAVLT immediate, Comorbidity Renal& Genitourinary, CDRSB, ADASQ4, Comorbidity Endocrine & Metabolic. The features selected consist of one demographic feature: Age, 7 cognitive scores: ADAS13, RAVLT learning, FAQ, ADAS11, RAVLT immediate, CDRSB, ADASQ4, and 2 comorbidities: Endocrine & Metabolic and Renal & Genitourinary. The selection of features aligns with existing literature, where age emerges as the most significant risk factor in Alzheimer’s disease, reflecting its well-established association with disease progression34. Additionally, cognitive scores serve as crucial indicators of AD, further supporting their inclusion in the predictive feature.
For our exploratory analysis we employ Partial Dependence Plots (PDPs) to offer a visual representation of the relationship between these individual features and the target variable while holding other features constant. The PDPs for the top 10 selected features by our fast random survival forest models are shown in Fig. 2. Specifically, Fig. 2a corresponds to ADAS13, Fig. 2b to AGE, Fig. 2c to RAVLT learning, Fig. 2d to FAQ, Fig. 2e to ADAS11, Fig. 2f to RAVLT immediate, Fig. 2g to Comorbidity (Renal & Genitourinary), Fig. 2h to CDRSB, Fig. 2i to ADASQ4, and Fig. 2j to Comorbidity (Endocrine & Metabolic). The blue shading in the plot, which is darker for higher values and lighter for smaller values, indicates the varying impact of the feature on the predicted survival function. For example, in the case of “AGE” feature, as age increases, the shading becomes darker and the darker survival function is lower, suggesting a stronger negative relationship between age and survival probability. Conversely, for younger ages, the shading is lighter and the lighter survival function is elevated, indicating a weaker impact of age on survival probability. The trend observed in the survival function value aligns with the shading pattern: as age increases, the survival function value decreases, indicating a higher predicted probability of experiencing the event of interest (conversion to MCI). Conversely, for younger ages, the survival function value increases, suggesting a lower risk of experiencing the event.
Partial Dependence Plots of top 10 selected features by the best performing machine learning model. Time x-axis is in months.
Kaplan–Meier survival curves stratified by (a) presence of endocrine/metabolic comorbidities (e.g., diabetes), (b) renal/genitourinary comorbidities, and (c) age group (\(\le 70\) vs. \(>70\) years).
When examining the partial dependence survival profiles of the comorbidity features “Renal&Genitourinary” and “Endocrine&Metabolic”, we observe two distinct survival function lines: a light blue line representing when the feature is absent (0) or does not exist, and a dark blue line representing when the feature is present (1) or the condition exists. Across both features, the darker line consistently depicts a lower survival function compared to the lighter line, suggesting that the presence of these comorbidities is associated with decreased survival probabilities.
To further evaluate the clinical relevance of these observations, Kaplan–Meier curves were generated for subgroups defined by the key features. Individuals with renal comorbidities exhibited significantly poorer survival compared to those without (Fig. 3a, log-rank p=0.04), supporting the trend observed in the partial dependence plot. In contrast, while individuals with endocrine/metabolic comorbidities, including diabetes, also showed a lower survival trend, this difference was not statistically significant (Fig. 3b, log-rank p=0.06). Additionally, age-stratified survival curves revealed that individuals aged above 70 years had significantly worse outcomes compared to those aged 70 or younger (Fig. 3c, log-rank p=0.003). These results emphasize the predictive value of age and renal health in survival outcomes and reinforce the interpretability of the model’s stratification capabilities.
External validation
Our best performing model, fast random survival forest, underwent external validation against the AIBL dataset to assess its generalizability. To strengthen this validation strategy, we adopted a two-stage approach. In the first stage, we applied the original top-performing model—trained on the full ADNI feature set (FS1)—to the AIBL dataset after aligning feature dimensions by adding missing variables with neutral values. This resulted in a C-index of 0.73, demonstrating robust predictive performance and highlighting the model’s consistent capabilities across datasets. In the second stage, we repeated the process using only the features common to both ADNI and AIBL. The fast random survival forest model was retrained using these shared features and achieved a C-index of 0.79 on the ADNI test set and 0.75 on the AIBL dataset. Next, because zero-imputation may introduce bias, we performed a sensitivity analysis using multiple imputation on the AIBL dataset. Across five imputed versions, the model achieved a pooled C-index of 0.77, indicating improved generalizability under less biased conditions and highlights variability due to missing data. Internal performance remained stable, reinforcing the model’s reliability. The outcomes of all three validation steps are summarized in Table 4.
Discussion
The integration of survival analysis techniques with machine learning presents a promising avenue for advancing early-stage Alzheimer’s disease prediction. By leveraging demographic, cognitive, and comorbidity data, this study shows the feasibility of using readily available clinical features for accurately identifying early-stage AD progression from CN to MCI. Our findings highlight the critical role of age and specific comorbidities as predictive features. The superior performance of the fast random survival forest model demonstrates the feasibility of using readily available clinical data, with external validation on the AIBL dataset demonstrating the robustness and generalizability of the proposed approach.
When examining the performance of the ML and DL survival analysis models, we observe distinct patterns about the importance of including both cognitive scores and comorbidity data. Models like the fast random survival forest and DeepSurv models demonstrated outstanding performance, achieving a C-index of 0.84 when applied to FS1, which incorporated all data modalities. This outcome is in line with expectations, given that FS1 incorporates all data modalities. The exclusion of either comorbidities (FS3) or cognitive scores (FS2) resulted in notable declines in performance, with C-index scores dropping to 0.33 and 0.74 for the DeepSurv model, and dropping to 0.76 and 0.59 for the fast-RSF respectively. The sharp drop in DeepSurv’s performance on FS3 (from 0.84 to 0.33) likely reflects its reliance on high-dimensional, informative input and its tendency to overfit when such complexity is reduced. Comorbidity features provided critical variance for pattern recognition, and their exclusion substantially limited the model’s ability to generalize. This drop shows how DeepSurv’s is sensitive to feature richness and highlights the challenges of using deep learning models when high-quality, diverse inputs are not available. In contrast, simpler models like CoxPH exhibited more stable performance under reduced feature sets, suggesting greater robustness to overfitting and stronger reliance on structured, lower-dimensional data. Additionally, the cognitive scores at baseline may have been more similar across individuals, further reducing discriminative signal. This divergence in performance patterns between Cox models and machine learning models highlights the complexity of Alzheimer’s disease prediction and the necessity of tailoring modeling approaches to specific data characteristics.
Incorporating various feature sets, each comprising different combinations of demographics, cognitive scores, and comorbidities, serves to explore the individual and combined effects of these factors on the outcome of interest. The trend observed in the top performing model was also clearly evident in the other models. Firstly, with the feature set FS1, which includes demographics, cognitive scores, and comorbidities, the eight models exhibited an average performance of 0.76. By selectively excluding specific data modalities, its contribution to the predictive performance of the model is highlighted. For instance, when comorbidity data is excluded, FS3, the average performance of eight models notably decreased to 0.67. Additionally, the models demonstrated their lowest average performance when utilizing FS2 without cognitive scores. This decline in performance aligns with expectations, as cognitive scores are essential components for accurate predictions in this context.
Moreover, compared to prior studies, which achieved maximum C-index values of 0.66 and 0.68, our models showcased significant improvements. This shows the efficacy of utilizing cost-effective and non-invasive data modalities, such as demographics, cognitive scores, and comorbidities, as opposed to resource-intensive techniques like MRI and PET imaging. This approach not only enhances predictive accuracy but also broadens the accessibility of AD progression modeling in clinical settings. These findings advocate for the comprehensive consideration of cognitive and comorbidity data in future predictive modeling efforts. Expanding this approach to integrate longitudinal data and exploring additional feature combinations could further refine prediction models and advance the early diagnosis and management of Alzheimer’s disease.
The lack of statistically significant differences in the Kruskal-Wallis test emphasizes the comparable performance among the eight models when using FS1. However, the interpretability and highest C-index performance of the fast random survival forest model made it the choice for predictive feature analysis. Its ability to handle complex relationships while maintaining clarity in feature importance makes it a valuable tool in exploring the association of key predictive features such as age, cognitive scores, and comorbidities. This approach facilitates a deeper understanding of Alzheimer’s disease progression and provides actionable insights for early detection and intervention strategies.
The selection of features was performed using the permutation method, which systematically evaluates the contribution of each feature on the model’s predictive capability by measuring the change in performance when the feature is randomly shuffled. The identified features match well with established clinical findings, reinforcing the validity of the results. Age emerged as the most critical predictor, reflecting its well-documented association with Alzheimer’s disease progression34. Similarly, cognitive scores, such as ADAS13 and RAVLT, are essential measures for assessing cognitive decline and AD status35. Their inclusion underscores their value in early-stage prediction models. Comorbidities, while not direct causes of Alzheimer’s disease, may have indirect associations. Conditions like metabolic disorders (e.g., diabetes, obesity) and renal diseases (e.g., chronic kidney disease) have been linked to cognitive decline and dementia36,37. While these conditions are included as predictors, their clinical interpretation should be approached with caution, as associations do not imply causation. These findings emphasize the importance of including comorbidities in predictive models, as they might exacerbate underlying pathological processes or contribute to increased cognitive decline risk.
The use of Partial Dependence Plots provided crucial insights into how individual features influence survival predictions in the fast random survival forest model. This method enabled the visualization of the direct impact of each feature while keeping other variables constant, ensuring clarity in interpreting the model’s behavior. Age emerged as a critical predictor, with a consistent trend of lower survival probabilities at higher ages, reinforcing its well-established predictive association in Alzheimer’s disease progression. This pattern was further validated through Kaplan–Meier survival curves, which revealed significantly poorer survival in individuals over 70 years of age compared to those 70 or younger (log-rank p = 0.003). The detailed representation of the relationship between age and survival function further substantiates its importance in predictive modeling. The analysis of comorbidity features, such as Renal & Genitourinary and Endocrine & Metabolic, highlighted their significant association on survival probabilities. The presence of these conditions is associated with reduced survival probabilities, a trend that aligned with survival curve analyses. Specifically, individuals with renal issues showed significantly worse outcomes (log-rank p = 0.04), while differences related to endocrine/metabolic issues—including diabetes—did not reach statistical significance (log-rank p = 0.06). This convergence between model-based interpretation and classical survival analysis reaffirms the robustness of the feature selection and the model’s ability to leverage clinically relevant features for accurate risk stratification. This can extend to clinical decision-making, where understanding the interplay of age, cognitive scores, and comorbidities can support risk stratification and inform early clinical decision-making. It is important to note that while these features were predictive of conversion risk, the associations observed do not imply causality.
The external validation of the fast random survival forest model using the AIBL dataset, with a concordance index (C-index) of 0.73, highlights the model’s robust predictive capabilities across diverse datasets. The model has shown to be consistent and reliable when applied to real-world scenarios, showcasing its potential for broader applicability. To further ensure the validity of this external validation, we conducted a secondary evaluation using only the features shared between ADNI and AIBL. The model retained high performance with a C-index of 0.79 on ADNI and 0.75 on AIBL, reinforcing its robustness even under reduced input dimensionality. The observed reduction in model performance on AIBL compared to ADNI may be partly explained by differences in cohort characteristics, as detailed in Table 1. Nonetheless, both strategies rely on assumptions regarding how missing features in AIBL are handled—either through zero imputation or through exclusion via common feature alignment. To probe this further, we incorporated a sensitivity analysis using multiple imputation to fill in the unavailable AIBL features with plausible values, rather than assuming their absence. This led to an improved external validation performance with a pooled C-index of 0.77, suggesting that imputing missing variables yields more realistic risk estimates and enhances model generalizability. The individual C-index scores across the five imputed datasets varied from 66.5 to 82.7, therefore, there is the uncertainty introduced by different imputation pathways. These results support the notion that while the model remains stable under different input assumptions, the choice of imputation strategy materially affects external performance estimates and should be carefully considered in cross-cohort deployment.
Another plausible contributor to the performance drop observed in the AIBL validation is the absence of several key predictive features. In our top-performing FS1-ADNI model, five of the highest-ranked cognitive features (ADAS13, ADAS11, ADASQ4, RAVLT learning, and FAQ) were not available in the AIBL dataset. These features were among the top 10 based on feature importance rankings and significantly contributed to model performance in ADNI. Their absence in AIBL, due to non-collection rather than biological variation, likely constrained the model’s predictive power during external validation. The multiple imputation results further highlight this issue—when these features were recovered using statistically informed methods, the model’s ability to generalize improved. This reinforces the importance of feature completeness in external validation and supports the need for harmonized data collection protocols across studies to minimize biases introduced by structural missingness.
Material & methods
This study investigates the impact of baseline comorbidities in elderly individuals by incorporating them into various feature sets. These sets, along with additional demographic data, are employed to assess the efficacy of diverse machine learning algorithms in predicting survival time to early-stage Alzheimer’s (from CN to MCI), considering the challenges posed by heterogeneous and censored data. The methodology will follow the flow shown in Fig. 4. An interpretation of the inserved relation of features selected by the models and their predictive relevance to AD development is attempted through partial dependency plots.
Overview of the pipeline of the survival analysis techniques machine learning system.
Dataset
In this work, we use data from the publicly available Alzheimer’s Disease Neuroimaging Initiative32 and Australian Imaging, Biomarker & Lifestyle Flagship Study of Ageing33 databases. ADNI was launched as a collaborative effort between public and private sectors, aiming to assess the efficacy of longitudinal MRI and PET imaging alongside additional biomarkers, clinical evaluations, and neuropsychological assessments for monitoring the advancement of Mild Cognitive Impairment and early Alzheimer’s Disease. ADNI provides a wide variety of datasets and data modalities covering many facets of Alzheimer’s disease, and for this study we are utlizing the two datasets: “ADNIMERGE”, which includes a heterogeneous collection of demographics and cognitive scores, and “RECMHIST”, which contains information on comorbidities. All participants with a baseline of Cognitive Normal, at least one follow-up visit (as single visits lack relevance in survival analysis), and documented medical history were selected. AIBL,founded in 2006, comprises similar data modalities to ADNI as it adopted its key elements from ADNI, but not as extensive. Following the same flow as earlier, we selected the demographics, cognitive scores, and comorbidities information. This selection yielded a total of 494 ADNI participants and 320 AIBL participants who met the criteria. The use of baseline data indicates that the survival analysis took into account the interval between the study’s start and the MCI diagnosis or censoring. Comorbidity data from the RECMHIST dataset were mapped into 19 binary variables indicating the presence or absence of each condition (e.g., hypertension, diabetes, depression). If a condition was undocumented, it was assumed to be absent, following standard practice in ADNI-based studies. To align the AIBL dataset with the ADNI-trained models, features missing in AIBL were added and imputed with neutral values (i.e., zeros). While this allowed for external evaluation using the trained models, we acknowledge that this may not fully capture the underlying biological or clinical signal. To address concerns regarding feature mismatch across cohorts, we also constructed a version of the dataset restricted to the intersection of available features in both ADNI and AIBL. This common feature set consisted of 17 variables shared by both datasets, including one demographic feature (AGE), one categorical variable (PTGENDER), four cognitive scores (CDRSB, MMSE, RAVLT immediate, LDELTOTAL), and eleven comorbidity indicators. The comorbidity features include: Cardiovascular, Endocrine/Metabolic, Gastrointestinal, Hepatic, Malignancy, Musculoskeletal, Neurologic, Psychiatric, Renal/Genitourinary, and Smoking history. These common features were used to retrain the top-performing model on the ADNI training data, followed by internal testing and external validation on AIBL, allowing for a more direct and unbiased comparison across datasets. This approach avoids reliance on imputing missing data in AIBL and serves as a more conservative and reliable test of external generalizability.
Dataset preprocessing
The combined demographics, cognitive scores, and comorbidity data comprise both categorical and numerical features, along with missing values. Categorical features underwent one-hot encoding, generating a new binary feature for each level of every categorical feature. Given the presence of missing values common in real-world datasets, data imputation technique is applied to estimate these missing values. To prevent data leakage, preprocessing steps—including imputation—were carried out within each fold of the cross-validation. Preprocessing parameters were derived from the training set and applied to the test set. Empirical sampling from a histogram was chosen due to the subset’s small proportion of missing values (none exceeding 5%). This decision is supported by findings that highlight the benefits of methods like Predictive Mean Matching (PMM) for multiple imputation38. The paper emphasizes its ability to preserve data features, avoid impossible values, and remain robust against model misspecification. This imputation occurred within the cross-validation loop, initially on the training set and subsequently on the test set, using the prediction matrix developed from the training set. Overall, the complete ADNI feature set post-processing consisted of 36 features (43 after one-hot encoding), of which 6 were demographic (12 after one-hot encoding), 11 were cognitive scores, and 19 were comorbidities. The complete AIBL feature set post-processing consisted of 16 features of which 2 were demographic, 4 cognitive scores, and 10 comorbidities.
In order to explore the individual and combined effects of each data modality we decided to incorporate various feature sets, each comprising different combinations of the demographics, cognitive scores, and comorbidities features. In total, four feature sets were constructed: FS1 encompassed demographics, cognitive scores, and comorbidities; FS2 comprised demographics and comorbidities; FS3 comprised demographics and cognitive scores; and finally, FS4 solely consisted of demographics.
Model selection
In our study, we adopted a multifaceted approach to analyze survival data by leveraging various machine learning techniques available in the MLR3 (Machine Learning in R) version 0.16.139 package in R version 4.3.240. A total of five machine learning and three deep learning algorithms, equipped to handle high-dimensional, heterogeneous, censored data, were selected for evaluation. Given the limited number of features (up to 36 features), no feature selection methods were employed. The machine learning for survival analysis models used are: the Cox proportional hazards (CoxPH) model, the penalised Cox regression cross-Validated Group Lasso Regularization (CVGLmnet), random forest survival analysis (RSF), fast random forest survival analysis (Ranger), recursive partitioning and regression trees (Rpart). Additionally, the deep learning for survival analysis models used are: DeepSurv, DeepHit, and the Cox-Time model. These models were chosen for their ability to capture complex relationships in survival data and provide transparent decision rules in most,, offering a balanced comparison across traditional, ensemble-based, and deep learning approaches.
The cox proportional hazards model is one of the most common regression modeling frameworks for survival analysis41 and is explored with its penalized regression variant as a useful benchmark in survival analysis, providing a reliable reference point against which the performance of other models can be evaluated. The Cox model can be expressed by hazard function \(h(t)\), representing the risk of an event occurring. The following estimate can be made:
where,
-
\(t\) represents the survival time.
-
\(h(t)\) is the hazard function determined by a set of \(p\) covariates (\(x_1, x_2, \ldots , x_p\)).
-
The coefficients (\(b_1, b_2, \ldots , b_p\)) measure the impact (i.e., the effect size) of covariates.
-
The term \(h_0\) is called the baseline hazard.
The regularized regression form of Cox implements the elastic net regularization technique, which combines both L1 (lasso) and L2 (ridge) penalties42. The penalties are applied to the coefficients of the predictor variables. Cross-validation is performed to select the optimal value of the regularization parameter (lambda) and potentially other tuning parameters. Cross-validation helps prevent over-fitting by assessing the model’s performance on a validation set that is separate from the training data. After cross-validation, the model with the optimal regularization parameter(s) is selected.
The two ensemble learning methods implementing random forests: Random Survival Forest43 and Fast Random Survival Forest44 survival techniques introduce randomization to the survival model and excel at capturing intricate relationships within the data by aggregating predictions from multiple decision trees. With \(X\) as the feature matrix and \(T\) as the survival time,
where \(h_i(t | X)\) is the survival function of the \(i\)-th tree. We also integrated the tree-based method recursive partitioning and regression trees into our analysis pipeline45. This method recursively partitions the data based on co-variates, providing transparent decision rules that are easily interpretable. Additionally, there is the penalized regression variant of Cox, Cross-Validated Group Lasso Regularization. It is a method for fitting Cox proportional hazards models with group lasso regularization. It extends the glmnet package by incorporating cross-validated group lasso regularization, which is beneficial for handling high-dimensional data with correlated covariates in survival analysis. To show the cross-validated Cox models with L1 and L2 regularization. Let \(\lambda\) be the regularization parameter, \(\beta\) the vector of model coefficients, \(\ell (\beta )\) the partial log-likelihood of the Cox model, and \(P(\beta )\) the regularization penalty term (e.g., group lasso). The optimization objective is given by:
In the realm of deep learning, we employed state-of-the-art models such as DeepSurv and DeepHit46,47. These neural network architectures are specifically designed for survival analysis tasks, capable of capturing complex nonlinear relationships between covariates and survival outcomes. DeepSurv, for example, utilizes deep neural networks to learn hierarchical representations of the data, while DeepHit combines recurrent neural networks with multi-task learning to account for censoring and event occurrences simultaneously. Finally, we explored the Cox-Time model, an extension of the Cox proportional hazards model that accommodates time-varying covariates48. This model allows for more dynamic modeling of survival data, where the effects of covariates may change over time. All deep learning models were implemented using the pycox library48, which builds on torchtuples. DeepSurv followed the default architecture proposed by Katzman et al.46, consisting of a 2-layer fully connected feedforward neural network with ReLU activation functions. DeepHit and Cox-Time similarly used the default architectures from Kvamme et al.48, with ReLU activations in hidden layers and appropriate output activations (softmax for DeepHit and linear for Cox-based models). Dropout regularization was applied to mitigate overfitting, and both dropout and learning rate were included in the hyperparameter search space, as detailed in Table 5. The choice to retain these default configurations ensured model comparability and stability given the moderate input dimensionality and sample size.
Model tuning
To maximize model performance, one or more of the hyper-parameters of some of the selected algorithms needs to be tuned. In an ideal scenario with unlimited resources, tuning all hyper-parameters jointly would be preferable, but in practice, only a subset can be tuned, constituting the search space or tuning space. The search space for each model is shown in Table 5. The hyper-parameters were optimized through nested resampling (nested cross-validation), which helps reduce optimistic bias and provides a more reliable estimate of model performance compared to simple cross-validation. The dataset is first divided into training and testing sets in a stratified manner, with a ratio of 90% for training and 10% for testing as shown in Fig. 4. The test data remains entirely unseen, ensuring unbiased evaluation. In each iteration of the 10-fold outer cross-validation, the remaining 9 folds are used for inner 5-fold cross-validation to optimize the hyper-parameters, while the held-out outer fold is used for performance evaluation. Random search was used as the tuning strategy, and the concordance index (C-index) was employed as the objective function to guide hyperparameter selection. This inner cross-validation loop was responsible for fine-tuning hyper-parameters, with each configuration being evaluated across 100 iterations to ensure comprehensive exploration of the parameter space. We acknowledge that this approach may lead to variability in the reported performance due to the relatively small test sample. After the hyperparameters are optimized using the inner loop, the model is trained on the entire training set (outer loop training folds) with the selected hyperparameters. Then, it’s evaluated on the held-out test set from the outer loop to obtain an unbiased estimate of its performance. Certain parameters of the machine learning models were predetermined with a fixed value and not subjected to optimization. Specifically, the importance of features in both random survival forest and fast random survival forest models was set to “permutation”. Additionally, the optimizer for all deep learning models was fixed as ’Adadelta’, an optimization method that dynamically adjusts learning rates based on recent gradient updates to ensure stable and efficient convergence during training. This way the configurations were consistent across models and focused the optimization efforts on the parameters with a wider range of variation and an impact on performance enhancement.
Model evaluation and interpretation
The performance of the models is evaluated using the concordance index (C-index) metric49, the most widely used metric for the global evaluation of models in survival analysis. A good model according to the C-index (C = 1 is perfect, \(\hbox {C} > 0.7\) is good) is one that always assigns higher scores to the subjects who survive longer, experience the event (MCI diagnosis) later or even never. Essentially, the higher the C-index, the better the model’s ability to correctly order participants based on their predicted probabilities. After evaluating all eight models, the highest C-index is selected as the top-performing model. To further quantify the uncertainty around model performance, we estimated 95% confidence intervals for the C-index using non-parametric bootstrapping. Specifically, for each model, the test set predictions were resampled with replacement 1,000 times, and the C-index was computed on each sample. The confidence intervals were then defined as the 2.5th and 97.5th percentiles of the resulting distribution. To support the statistical validity of model comparison, we additionally applied Dunn’s post-hoc test to the bootstrapped C-index distributions. This allowed us to identify pairwise differences in model performance and determine whether the observed superiority of the top-performing model was statistically significant rather than due to sampling variability.
Survival models are known as “risk prediction models”, and therefore, for a more comprehensive understanding of the differences between models we not only consider the absolute performance with C-index but also the relative performance among all models with the risk scores.The risk scores provided by the machine learning survival models are expressed as continuous rankings, providing valuable insights into the relative risk levels of each patient. These rankings inform us not only about who is predicted to have higher or lower risk but also quantify the magnitude of this predicted difference. While C-index serves as the primary metric for evaluating model accuracy—accounting for both event times and censoring—we also performed Kruskal-Wallis and post-hoc tests as supplementary analysis to explore the distributional differences in estimated risk scores across models. These statistical comparisons provide additional, but not performance-determining, insights into how models differentiate risk across individuals. We employ the Kruskal-Wallis test50, a non-parametric statistical method specifically designed for comparing distributions of model outcomes, in our case, the risk score ranking of the eight models on test subjects are compared. If the test indicates a significant difference among the models, we proceed with post hoc analysis (Dunn’s test) to determine which specific models differ from each other.
The top-performing model will be further evaluated using the external validation dataset AIBL. This external validation step is crucial for assessing the generalizability and robustness of our model beyond the dataset used for training and testing (ADNI). To address potential bias from zero-imputation of missing comorbidity features in the AIBL dataset, we performed a sensitivity analysis using multiple imputation. We first aligned the ADNI and AIBL datasets by introducing missing columns to AIBL with NA values, ensuring feature correspondence. Multiple imputation was performed using predictive mean matching (PMM) via the mice package in R51, generating five imputed datasets. Each was then evaluated using the top-performing model trained on ADNI. The C-index was computed for each imputed version, and results were pooled using Rubin’s rules to obtain the final performance estimate on AIBL.
Partial Dependence Plots (PDPs) will be used to gain deeper insights into the impact of selected features by the top performing model on its predictions52. These plots offer a systematic approach to understanding the marginal effect of individual features on model output by intervening on one feature while keeping others constant and observing the resulting changes in predictions. Through PDP analysis, we aim to elucidate the relative importance of features and their associations with the model’s predictions. By analyzing the shape and direction of PDP curves, we can discern how each feature influences the model’s predictions and derive meaningful interpretations regarding its behavior. Additionally, Kaplan-Meier curves will be generated to visualize survival differences between key subgroups, enhancing the clinical interpretability of the model’s predictions. A statistical log-rank test will also be performed to assess whether the survival distributions between these subgroups are significantly different. Lastly, the most frequently selected features across models will be analyzed, where possible, and evaluate feature importance scores to quantify each feature’s contribution to model predictions.
Conclusion
In this study, we investigated the impact of baseline comorbidities on predicting survival time to early-stage Alzheimer’s (from CN to MCI) in elderly individuals. Utilizing data from the ADNI and AIBL databases, we constructed feature sets incorporating demographics, cognitive scores, and comorbidities. We applied various machine learning algorithms, including Cox proportional hazards, random forest survival analysis, and deep learning models, to assess predictive efficacy. Our top-performing model, fast random survival forest, achieved a c-index of 0.84 when considering all feature modalities. Notably, exclusion of comorbidity data led to a significant decrease in predictive performance, highlighting its importance. Feature analysis revealed age, cognitive scores, and certain comorbidities as key predictors. External validation was conducted in two steps. First, the full ADNI-trained model was applied to the AIBL dataset after aligning features, which yielded a c-index of 0.73. Second, to address dataset mismatch concerns, we retrained the fast random survival forest model on ADNI using only the common features shared with AIBL. This version achieved a c-index of 0.79 on ADNI and 0.75 on AIBL, reinforcing the model’s generalizability even under feature alignment constraints. While nested cross-validation was used during model development for hyperparameter tuning and internal validation, the final performance metrics were derived from a 10% independent test set. Given the relatively small size of this test subset, we acknowledge that reported results may be sensitive to partitioning. This study is also limited by the use of only baseline features, without accounting for longitudinal changes in comorbidities or cognitive scores. Additionally, comorbidity groupings were broad and may benefit from more granular classifications. Future work should consider dynamic modeling with longitudinal data to capture temporal trends in disease progression, as well as evaluate model performance in more diverse and real-world clinical populations. Expanding the feature space to include lifestyle or socioeconomic factors may also enhance model generalizability and clinical relevance. Overall, our approach demonstrates promising potential for early Alzheimer’s prediction using readily available clinical data.
Data availability
The dataset analyzed in this study can be accessed from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) repository at adni.loni.usc.edu (Accession Number: sa000002). Data from the Australian Imaging Biomarkers and Lifestyle flagship study of ageing (AIBL) are also available at the ADNI database (www.aibl.csiro.au/adni/index.html). Access to both datasets requires a simple application procedure that is readily granted to researchers. For any further inquiries or to request data used in this study, please contact the corresponding author.
References
Holtzman, D. M., Morris, J. C. & Goate, A. M. Alzheimer’s disease: the challenge of the second century. Sci. Transl. Med. 3, 77sr1–77sr1 (2011).
Castellani, R. J., Rolston, R. K. & Smith, M. A. Alzheimer disease. Disease-a-month: DM 56, 484 (2010).
Nichols, E. et al. Estimation of the global prevalence of dementia in 2019 and forecasted prevalence in 2050: an analysis for the global burden of disease study 2019. The Lancet Public Heal. 7, e105–e125 (2022).
Cummings, J. L. & Cole, G. Alzheimer disease. Jama 287, 2335–2338 (2002).
Sperling, R. A. et al. Toward defining the preclinical stages of alzheimer’s disease: Recommendations from the national institute on aging-alzheimer’s association workgroups on diagnostic guidelines for alzheimer’s disease. Alzheimer’s & dementia 7, 280–292 (2011).
Albert, M. S. et al. The diagnosis of mild cognitive impairment due to alzheimer’s disease: recommendations from the national institute on aging-alzheimer’s association workgroups on diagnostic guidelines for alzheimer’s disease. Alzheimer’s & dementia 7, 270–279 (2011).
McKhann, G. M. et al. The diagnosis of dementia due to alzheimer’s disease: Recommendations from the national institute on aging-alzheimer’s association workgroups on diagnostic guidelines for alzheimer’s disease. Alzheimer’s & dementia 7, 263–269 (2011).
Tanveer, M. et al. Machine learning techniques for the diagnosis of alzheimer’s disease: A review. ACM Transactions onMultimed. Comput. Commun. Appl. (TOMM) 16, 1–35 (2020).
Pellegrini, E. et al. Machine learning of neuroimaging for assisted diagnosis of cognitive impairment and dementia: a systematic review. Alzheimer‘s & Dementia: Diagn. Assess. & Dis. Monit. 10, 519–535 (2018).
Mirzaei, G. & Adeli, H. Machine learning techniques for diagnosis of alzheimer disease, mild cognitive disorder, and other types of dementia. Biomed. Signal Process. Control. 72, 103293 (2022).
Ohno-Machado, L. Modeling medical prognosis: survival analysis techniques. J. Biomed. Inform. 34, 428–439 (2001).
Platero, C. & Tobar, M. C. Longitudinal survival analysis and two-group comparison for predicting the progression of mild cognitive impairment to alzheimer’s disease. J. Neurosci. Methods 341, 108698 (2020).
Orozco-Sanchez, J., Trevino, V., Martinez-Ledesma, E., Farber, J. & Tamez-Peña, J. Exploring survival models associated with mci to ad conversion: A machine learning approach. bioRxiv 836510 (2019).
Li, Y., Wang, L., Zhou, J. & Ye, J. Multi-task learning based survival analysis for multi-source block-wise missing data. Neurocomputing 364, 95–107 (2019).
Lu, P. & Colliot, O. Multilevel survival modeling with structured penalties for disease prediction from imaging genetics data. IEEE J. Biomed. Health Inform. 26, 798–808 (2021).
Liu, K., Chen, K., Yao, L. & Guo, X. Prediction of mild cognitive impairment conversion using a combination of independent component analysis and the cox model. Front. Hum. Neurosci. 11, 33 (2017).
Spooner, A. et al. A comparison of machine learning methods for survival analysis of high-dimensional clinical data for dementia prediction. Scientific reports 10, 1–10 (2020).
Mirabnahrazam, G. et al. Predicting time-to-conversion for dementia of alzheimer’s type using multi-modal deep survival analysis. Neurobiol. Aging 121, 139–156 (2023).
Nakagawa, T. et al. Prediction of conversion to alzheimer’s disease using deep survival analysis of mri images. Brain Commun. 2, fcaa057 (2020).
Pölsterl, S., Sarasua, I., Gutiérrez-Becker, B. & Wachinger, C. A wide and deep neural network for survival analysis from anatomical shape and tabular clinical data. In Machine Learning and Knowledge Discovery in Databases: International Workshops of ECML PKDD 2019, Würzburg, Germany, September 16–20, 2019, Proceedings, Part I, 453–464 (Springer, 2020).
Wattmo, C., Londos, E. & Minthon, L. Risk factors that affect life expectancy in alzheimer’s disease: a 15-year follow-up. Dement. Geriatr. Cogn. Disord. 38, 286–299 (2014).
Rountree, S. D., Chan, W., Pavlik, V. N., Darby, E. J. & Doody, R. S. Factors that influence survival in a probable alzheimer disease cohort. Alzheimer’s research & therapy 4, 1–6 (2012).
Steenland, K., MacNeil, J., Seals, R. & Levey, A. Factors affecting survival of patients with neurodegenerative disease. Neuroepidemiology 35, 28–35 (2010).
Walsh, J. S., Welch, H. G. & Larson, E. B. Survival of outpatients with alzheimer-type dementia. Ann. Intern. Med. 113, 429–434 (1990).
Brookmeyer, R., Corrada, M. M., Curriero, F. C. & Kawas, C. Survival following a diagnosis of alzheimer disease. Arch. Neurol. 59, 1764–1767 (2002).
Valderas, J. M., Starfield, B., Sibbald, B., Salisbury, C. & Roland, M. Defining comorbidity: implications for understanding health and health services. Ann. Fam. Med. 7, 357–363 (2009).
Todd, S., Barr, S., Roberts, M. & Passmore, A. P. Survival in dementia and predictors of mortality: a review. Int. J. Geriatr. Psychiatry 28, 1109–1124 (2013).
Davis, D. A., Chawla, N. V., Blumm, N., Christakis, N. & Barabási, A.-L. Predicting individual disease risk based on medical history. In Proceedings of the 17th ACM conference on Information and knowledge management, 769–778 (2008).
Santiago, J. A. & Potashkin, J. A. The impact of disease comorbidities in alzheimer’s disease. Front. Aging Neurosci. 13, 631770 (2021).
Rajamaki, B., Hartikainen, S. & Tolppanen, A.-M. The effect of comorbidities on survival in persons with alzheimer’s disease: a matched cohort study. BMC Geriatr. 21, 1–9 (2021).
Sun, H. et al. Altered motor activity patterns within 10-minute timescale predict incident clinical alzheimer’s disease. J. Alzheimers Dis. 98, 209–220 (2024).
Mueller, S. G. et al. The alzheimer’s disease neuroimaging initiative. Neuroimaging Clin. 15, 869–877 (2005).
Ellis, K. A. et al. The australian imaging, biomarkers and lifestyle (aibl) study of aging: methodology and baseline characteristics of 1112 individuals recruited for a longitudinal study of alzheimer’s disease. Int. Psychogeriatr. 21, 672–687 (2009).
Guerreiro, R. & Bras, J. The age factor in alzheimer’s disease. Genome Med. 7, 1–3 (2015).
Jacobs, D. M. et al. Neuropsychological detection and characterization of preclinical alzheimer’s disease. Neurology 45, 957–962 (1995).
Cai, H. et al. Metabolic dysfunction in alzheimer’s disease and related neurodegenerative disorders. Curr. Alzheimer Res. 9, 5–17 (2012).
Zhang, C.-Y., He, F.-F., Su, H., Zhang, C. & Meng, X.-F. Association between chronic kidney disease and alzheimer’s disease: an update. Metab. Brain Dis. 35, 883–894 (2020).
Austin, P. C., White, I. R., Lee, D. S. & van Buuren, S. Missing data in clinical research: a tutorial on multiple imputation. Can. J. Cardiol. 37, 1322–1331 (2021).
Lang, M. et al. mlr3: A modern object-oriented machine learning framework in r. J. Open Source Softw. 4, 1903 (2019).
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Cox, D. R. Regression models and life-tables. J. R. Stat. Soc. Series B Stat. Methodol. 34, 187–202 (1972).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1 (2010).
Ishwaran, H. & Kogalur, U. B. Random survival forests for r. R news 7, 25–31 (2007).
Wright, M. N. & Ziegler, A. ranger: A fast implementation of random forests for high dimensional data in c++ and r. arXiv preprint arXiv:1508.04409 (2015).
Therneau, T., Atkinson, B., Ripley, B. & Ripley, M. B. Package ‘rpart’. Available online: cran. ma. ic. ac. uk/web/packages/rpart/rpart. pdf (accessed on 20 April 2016) (2015).
Katzman, J. L. et al. Deepsurv: personalized treatment recommender system using a cox proportional hazards deep neural network. BMC Med. Res. Methodol. 18, 1–12 (2018).
Lee, C., Zame, W., Yoon, J. & Van Der Schaar, M. Deephit: A deep learning approach to survival analysis with competing risks. In Proceedings of the AAAI conference on artificial intelligence, vol. 32 (2018).
Kvamme, H., Borgan, Ø. & Scheel, I. Time-to-event prediction with neural networks and cox regression. J. Mach. Learn. Res. 20, 1–30 (2019).
Harrell, F. E., Califf, R. M., Pryor, D. B., Lee, K. L. & Rosati, R. A. Evaluating the yield of medical tests. Jama 247, 2543–2546 (1982).
Kruskal, W. H. & Wallis, W. A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 47, 583–621 (1952).
Van Buuren, S. & Groothuis-Oudshoorn, K. mice: Multivariate imputation by chained equations in r. J. Stat. Softw. 45, 1–67 (2011).
Friedman, J. H. Greedy function approximation: a gradient boosting machine. Ann. Stat. 1189–1232 (2001).
Acknowledgements
This work is supported by Khalifa University under Award no. FSU-2021-005. ADNI data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative database adni.loni.usc.edu. Data collection and sharing for this project were funded by the ADNI(National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). For a full list of funding sources and contributors, please visit adni.loni.usc.edu. As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: ADNI Acknowledgement List. AIBL data used in this study was obtained from the Australian Imaging Biomarkers and Lifestyle flagship study of ageing (AIBL) funded by the Commonwealth Scientific and Industrial Research Organisation (CSIRO) which was made available at the ADNI database (www.loni.usc.edu/ADNI). The AIBL researchers contributed data but did not participate in analysis or writing of this report. AIBL researchers are listed at www.aibl.csiro.au.
Author information
Authors and Affiliations
Contributions
Aamna AlShehhi (AAS) led the project by providing the overarching idea, securing funding, and supervising the research. Ferial Abuhantash (FA) conceptualized and conducted the study, performed the literature review, developed and implemented the code, built the models, analyzed the data, generated visualizations, and wrote the initial manuscript. Roy Welsch (RW) and Stan Finkelstein (SF) contributed critical feedback, offered expert opinions, and reviewed the manuscript. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abuhantash, F., Welsch, R., Finkelstein, S. et al. Alzheimer’s disease risk prediction using machine learning for survival analysis with a comorbidity-based approach. Sci Rep 15, 28723 (2025). https://doi.org/10.1038/s41598-025-14406-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14406-0
This article is cited by
-
Longitudinal multi-modal data prediction model for mild cognitive impairment by deep survival analysis
BMC Medical Informatics and Decision Making (2026)