Abstract
Amphibians, particularly salamanders, are facing dramatic population declines worldwide; therefore, investigating their role as predators through diet analysis could inform targeted conservation programs. DNA metabarcoding significantly outperforms morphology-based identification of prey taxa in other vertebrates, but has rarely been applied to amphibians, and then only for faecal samples. Here, we chose the golden Alpine salamander Salamandra atra aurorae, an endangered endemism of the Italian Alps, to (i) develop a new metabarcoding protocol for amphibian stomach flushing samples; (ii) investigate the diet of this subspecies, including differences in the Frequency of Occurrence (%FO) of prey taxa between sexes; and (iii) compare these results with a recently published morphology-based identification of diet for the same samples. From 53 golden Alpine salamanders from the Province of Trento (Italy) population, targeting two COI fragments with DNA metabarcoding, we detected an extraordinary 177 prey taxa (103 to species), obtaining a much higher proportion of positive samples and taxonomic resolution compared to the morphological study. The FO of these taxa showed a strong positive correlation between sexes. In amphibian diet studies, the inclusion of metabarcoding protocols for stomach flushes could accelerate our understanding of their trophic ecology and improve future conservation actions.
Similar content being viewed by others
Introduction
According to the latest Global Amphibian Assessment1, amphibians are the most threatened Vertebrate Class globally due to the spread of new pathogens, the climate crisis, habitat loss, and overexploitation. In particular, the conservation status of salamanders is worsening rapidly worldwide, as the Red List Index of the Order Caudata is consistently the lowest1. In this context, studies focused on their feeding behavior and preferred prey have significantly advanced the understanding of trophic interactions2, ecosystem network dynamics3, predator-prey dynamics4, and the overall impact of species on their environment5. In terms of habitat conservation, diet studies help to identify critical foraging areas and preferred food sources6. This information is essential for the development of targeted conservation plans aimed at preserving ecosystems and mitigating the threats posed by habitat loss, degradation, and overexploitation7, as well as the presence of parasitic8 and invasive species9,10.
The study of the amphibian diet is traditionally based on morphological identification of their prey from stomach flushing samples, which is considered a minimally invasive, non-lethal method in amphibians (e.g.11,12). Compared to fecal samples, which are often the preferred sample type for other species as they can be collected without disturbing or harming individuals (e.g., lizards:13; felids:14), stomach flushes appear to provide more reliable diet composition than faeces in the case of amphibians15,16.
Morphological-based identification of taxa detected in both stomach flush and faecal samples, however, has its own limitations when compared to molecular approaches such as DNA metabarcoding. For instance, DNA metabarcoding protocols of these sample types can identify species even when morphologically diagnostic characters are absent or unrecognizable due to partial or complete digestion17. If comprehensive DNA barcodes reference databases are available, prey identification with this method could potentially reach higher taxonomic resolution with respect to the morphological approach17. In fact, numerous studies on non-amphibian species have confirmed that DNA metabarcoding has significantly greater resolution and better accuracy than morphological identification (e.g., Natterer’s bat Myotis nattereri18; European hake Merluccius merluccius7; Selvagens gecko Tarentola (boettgeri) bischoffi19; Blue Catfish Ictalurus furcatus20). This is especially the case for samples from animals with generalist diets, such as some insectivores21. Very few DNA metabarcoding-based studies of amphibian diet have been published thus far, and all of these use faecal samples8,22,23,24. To the best of our knowledge, this is the first study to use DNA metabarcoding to analyze amphibian stomach flushing samples.
Therefore, here we aimed to (i) develop a new DNA metabarcoding protocol for the analysis of stomach flushing contents in amphibians; (ii) apply this protocol to the golden Alpine salamander Salamandra atra aurorae (Trevisan, 1982) as a case study; (iii) compare the efficiency of our new protocol with available morphological identification25, in order to highlight advantages and drawbacks of each method. This work aimed at providing useful information for conservation planning of the target taxon by investigating its dietary habits, including potential sex differences.
Methods
Study species and study area
The golden Alpine salamander (S. a. aurorae), a subspecies of the Alpine salamander S. atra (Laurenti, 1768), is endemic to the Italian Alps with a very limited distribution of only 26 km². Two main populations are known, located on the Vezzena Plateau in the Autonomous Province of Trento (Trentino-Alto Adige/Südtirol Region) and the Sette Comuni Plateau in the neighboring Province of Vicenza (Veneto Region;26). In the latest IUCN Red List assessment for Italy (2022), the conservation status of this subspecies was reclassified from Vulnerable (VU) to Endangered (EN), due to habitat degradation and its declining distribution and occupancy area27.
The sampling area for this study was located in the Municipality of Levico Terme on the Vezzena Plateau (Fig. 1). This plateau has a mean elevation of 1450 m a.s.l. and a typically Alpine climate, but with humidity levels similar to the neighboring pre-Alpine Veneto Region28. The vegetation consists mainly of silver fir (Abies alba Mill., 1759), beech (Fagus sylvatica L., 1753), and spruce (Picea abies (L.) H. Karst., 188129). Of note, on 29 October 2018, the ‘Vaia’ windstorm felled millions of trees in the northeastern Alps, including a large number in the study area, causing significant damage to the forest and possibly threatening this species’ survival25.
Map of the general study area, with the Vezzena Plateau (Trentino-Alto Adige/Südtirol Region, Italy) outlined in red. Image created with QGIS.org (2024) QGIS Geographic Information System. Open Source Geospatial Foundation Project http://qgis.org.
Sample collection
The sampling design and protocol are detailed in25. Briefly, the study area was divided into 29 square plots (20 × 20 m), which were surveyed for adult salamanders on the 26, 28 and 29 July 2022 (during or immediately after rainfall). All experimental procedures were approved by the Italian Ministry of Ecological Transition (authorization MiTE-0014200 of the 7 February 2022) and were performed in accordance with relevant guidelines and regulations. Sampling was performed as follows: 53 S. a. aurorae adult individuals (defined as total length > 90 mm;30) were captured in the wild by hand using sterile gloves, placed in a sterile container, and kept at 4 °C to reduce metabolic rate and slow down digestion31. Stomach flushing samples were obtained using 10–30 ml water to flush the stomach into sterile 50 ml Falcon tubes using the protocol established by Fraser32. Sex and reproductive state were noted for each individual. Nine of the 53 samples did not contain visible prey remains. No mortality or stress reactions were observed from the animals during the entire sampling process. Samples were kept at room temperature in 70% ethanol until further analysis, while sampled individuals were released in their respective capture plots within 6 h after capture.
Sample processing
First, morphological analysis was carried out by MUSE - Science Museum of Trento, and results were published in25. To minimize sample DNA contamination at this step, researchers wore sterile gloves (changed between each sample) and worked with sterile Petri dishes and forceps. Then, after morphological identification, stomach contents and liquid (see below) were immediately transferred to new sterile Falcon tubes and transferred at room temperature to the Animal, Environmental and Antique DNA Platform of the Conservation Genomics Research Unit (FEM, S. Michele all’Adige, Italy), where metabarcoding analysis was carried out.
Since each sample contained up to 100 ml of 70% ethanol (added during the stomach flushing process), in order to concentrate the DNA, before DNA extraction, each sample was mechanically fractured and homogenized in liquid nitrogen using a sterile ceramic pestle and mortar, and the homogenate was filtered through a Sterivex-GP 0.22 μm Filter unit (Millipore, Burlington, Massachusetts, USA). All the following procedures were performed under BSL2 biological hoods. Total DNA extraction was carried out from the filters using the DNeasy PowerWater Sterivex Kit (Qiagen®), following manufacturer’s instructions except for modifications noted in33. At least one negative control was included in each extraction batch. Extracted DNA was eluted in 100µL EB buffer.
PCR amplification targeted the Cytochrome Oxidase I (COI) gene, which is commonly used for DNA metabarcoding of invertebrates (e.g.24,34,35,36). In order to broaden the taxonomic scope of this study, two primer pairs were used here, each including standard Illumina sequencing adapters: ZBJ-ArtF1c and ZBJ-ArtR2c (hereafter PRIMER_A) targeting a 157 bp (bp) fragment37, and mlCOIintF-XT with jgHCO2198, as modified by Wangensteen et al.38, targeting a 313 bp sequence (here, PRIMER_B). PCR reaction mix for PRIMER_A consisted of 10 µl of Promega Flexi Buffer 5X, 6 µl of MgCl₂ 25µM, 2 µl of each forward and reverse primers 10ρmol/µl, 0.25 µl of dNTP’s 10mM each, 0.25 µl of Promega - GoTaq HS G2 5U/µL, 5 µl of template DNA and sterile H₂O up to a volume of 50 µl. The reaction mix for PRIMER_B was the same, except that 1 µl of dNTP’s 10mM each. The PCR reaction programs for both primers are detailed in Figure S1 (Supplementary Information). One PCR negative control (reaction mix only), as well as the extraction negative controls, were included for each PCR reaction.
After amplification success was confirmed on a QIAxcel Advanced System (QIAGEN, Germany), all samples were purified with the MinElute PCR Purification Kit (QIAGEN, Germany). Library preparation followed guidelines of the Nextera XT Index Kit (FC-131‐1001/FC‐131‐1002; Illumina, Inc., San Diego, California, USA). All samples were quantified and normalized, before being pooled together to a final concentration of 13ρM. Libraries were then sequenced on an Illumina MiSeq platform with the 2 × 300 bp paired-end approach and a sequencing depth of 30,000 reads per sample.
Data analysis
Bioinformatic analysis was carried out with MICCA39. Briefly, raw paired-end reads were merged with a minimum overlap length of 50 bp, and a maximum of 10 mismatches. After primer trimming, the sequences were filtered by quality and length allowing an expected error rate of 0.1%, and minimum lengths of 155 bp for PRIMER_A and 300 bp for PRIMER_B. Sequences were dereplicated, denoised, and filtered for any PCR/sequencing errors or chimeras with the denovo_unoise MICCA command, which implements the UNOISE3 algorithm40, to obtain Amplicon Sequence Variants (ASVs). Finally, to eliminate potential contamination from the dataset, the corresponding number of reads of each ASV found in either extraction or PCR negative controls was subtracted from each sample24. Singletons were also removed. Details on the commands and parameters used for this bioinformatic analysis are shown in Figure S2 (Supplementary Information).
Since a reference database of COI sequences for local invertebrate fauna was not available, ASVs detected by both PRIMER_A and PRIMER_B were identified with BLAST41, only considering the top result (i.e. the results with lowest e-value). ASVs were considered assigned only if they met the following criteria: (i) they belonged to the Phyla Annelida, Arthropoda, or Mollusca; (ii) they shared at least 99% of coverage with the matching GenBank reference; and (iii) they passed the minimum identity thresholds of 90% for family, 95% for genus, and 98% for species. Any unassigned ASV was discarded. To further validate these results, the plausibility that each species could be found in the sampling area was checked with presence records from the Global Biodiversity Information Facility (GBIF; https://www.gbif.org/) and the Checklist of Species of the Italian Fauna (https://www.faunaitalia.it/checklist/). In addition, the ecological role of each taxon was noted after consulting expert taxonomists at FEM and peer-reviewed literature42,43,44.
The dietary preference of S. a. aurorae was quantified by calculating the percent Frequency of Occurrence (%FO) for each prey taxon as the percentage of samples in which the taxon was detected out of the total number of samples, considering the combined results obtained with both primer pairs45,46. The correlation between %FO of the main prey taxa found in the stomach of males (34 individuals) versus females (19), and of pregnant (11) versus non-pregnant (8) females, was tested with the Spearman’s rank correlation coefficient in RStudio using the cor.test function47. Scatterplots of these correlations were produced with ggplot248 and edited with Inkscape49. %FOs, represented with barplots, was also used to qualitatively compare results between metabarcoding analysis (this study) and morphology-based identifications (published in25).
Results
All 53 stomach flushing samples were successfully amplified on the Qiaxcel Screening System and, therefore, were sequenced together with the negative controls, obtaining a total of 32,761,400 raw reads. Following bioinformatic merging and denoising, the total dataset consisted of 3,199,813 reads, including 1,233,061 reads of PRIMER_A and 1,966,752 of PRIMER_B. Overall, 320 ASVs were identified for PRIMER_A and 1,267 for PRIMER_B. Of these ASVs, 201 for PRIMER_A and 618 for PRIMER_B were taxonomically classified with BLAST. Our DNA metabarcoding protocol was able to identify from 2 to 29 prey taxa in all 53 samples (mean: 9.4 ± 2.0 taxa). Overall, 177 taxa, including 103 species, were identified; 52 (29.4%) were detected by both primers, 65 (36.7%) only by PRIMER_A, and 60 (33.9%) only by PRIMER_B (details shown in Table S1, Supplementary Materials).
The most common taxa in the diet of S. a. aurorae were Diptera (%FO: 84.9%), Myriapoda (76%), Arachnida (77.4%; including Aranea with %FO: 58.5%, Opiliones 35.8%, and Acarina 5.7%), Coleoptera (56.6%), and Psocoptera (50.9%), followed by Collembola (45.3%), Lepidoptera (39.6%), Clitellata (34%), Hemiptera (24.5%), and Gastropoda (22.6%). The least common were Neuroptera (13.2%), Dermaptera (3.8%), and Malacostraca (1.9%). The complete list of taxa detected with DNA metabarcoding in this study and %FO for each are shown in Table 1. Of the 177 identified taxa, six species (the house fly Helina troene Walker, 1849; the Highlands thricops Thricops albibasalis Zetterstedt, 1849; the parasitoid wasps Dusona angustifrons Forster, 1868, Enizemum ornatum Gravenhorst, 1829, and Syrphoctonus longiventris Thomson, 1890; and the booklouse Stenopsocus lachlani Kolbe, 1880), four genera (Leptomias, Muscopterix, Metylophorus and Amphigerontia), and one family (Pemphigidae) had a known distribution limited to the Americas, Asia, or northern Europe, and were therefore not expected to be found in our study area (Table 1).
The proportion of taxa detected showed a very strong positive correlation between the diet of male and female salamanders (Spearman’s rho: 0.923; p-value < 0.001; Fig. 2A). The proportion of prey found in samples also showed a positive correlation between females with different reproductive status (pregnant or non-pregnant; Spearman’s rho: 0.797; p-value < 0.001; Fig. 2B).
Scatterplots showing the correlations between the Frequency of Occurrence (FO) of each prey taxon in Salamandra atra aurorae (A) male and female diet: x-axis: females; y-axis: males, and (B) pregnant and non-pregnant female diets: x-axis: non-pregnant females; y-axis: pregnant females. Black lines represent the correlation trend line, and Spearman’s rho values are also reported. Plots created with the R package ggplot2 and edited with Inkscape.
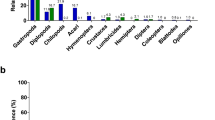
Using the same samples as in Centomo et al.25, our metabarcoding protocol identified at least one prey item in a higher number of samples compared to the morphological analysis (53 here, versus 44 as reported in25), and more total taxa (177 versus 16 in25), and achieved a higher taxonomic resolution (103 species here, compared to 16 higher-level taxa, Family at best, in25)and Diptera; Neuroptera and Psocoptera. In addition, of the taxa detected by both methods, metabarcoding showed a higher frequency of occurrence (%FO), meaning that when a taxon was detected it was generally found in a higher proportion of samples with respect to the morphological study (as shown in Fig. 3), with two exceptions (Acarina and Isopoda). On the other hand, DNA metabarcoding failed to detect two taxa which were found by Centomo et al.25, albeit at low %FO, namely Anoplura, represented in Fig. 3, and Pseudoscorpiones, included in the Arachnida Class for morphological data in Fig. 3.
Comparison of the Frequency of Occurrence (%FO) of each prey taxon found by DNA metabarcoding (blue; data from this study) or morphology (dark red; data from25) in Salamandra atra aurorae stomach flushing samples.
Discussion
As amphibians are rapidly declining worldwide, studies investigating their feeding behavior and diet preferences are fundamental to understanding trophic interactions, ecosystem network and predator-prey dynamics, in addition to identifying critical foraging areas and preferred food sources (e.g.4). More in general, this information is fundamental for a better understanding of the biology and behavior of these elusive species17 and has great conservation implications in terms of explaining populations fluctuations, the impact of habitat degradation and the climate crisis, or for establishing successful Conservation Breeding Programs (CBP)50. Although our target species is not involved in any CBP at present, these programs are, more in general, becoming increasingly relevant to support amphibian conservation in light of their ongoing population decline51.
The DNA metabarcoding protocol developed here aimed, for the first time, at rapidly identifying prey taxa from stomach flushing samples, a minimally invasive sampling method for amphibians. This approach obtained a detailed list of prey from all samples collected during this study, and the comparison of these results with a traditional taxonomic study of the same samples proved our DNA metabarcoding protocol to be a fast and reliable alternative, although a few taxon-specific limitations still exist. The application of this method provided insights into the diet of the endangered subspecies used here as a case study, the golden Alpine salamander S. a. aurorae, which was shown to ingest a remarkable 177 taxa, 103 of which we were able to identify to species. As salamanders are currently threatened worldwide, the development of minimally invasive protocols such as the one presented here is fundamental for comprehensive conservation-oriented programs to identify the resources required by target species. The application of metabarcoding to S. a. aurorae stomach flushes offers valuable ecological insights that can inform conservation strategies. As an endangered taxon with a restricted distribution, understanding its trophic interactions is crucial for habitat management and protection. Our findings indicate that S. a. aurorae consumes a diverse range of invertebrates, reflecting a flexible feeding strategy. This adaptability may provide resilience in the face of certain environmental changes, but it also underscores the importance of maintaining prey biodiversity within its habitat, especially that of soil-dwelling invertebrates. The main threats to S. a. aurorae include illegal collection for pet trade, habitat alteration due to forestry practices, and potential climate-driven changes in microhabitat conditions52. A precise identification of the taxonomic composition of the golden Alpine salamander diet, as provided by our study, can establish an important baseline for monitoring shifts in trophic ecology and for assessing the compatibility of land use practices with foraging requirements. Furthermore, the notable presence, in the salamander’s diet, of taxa associated with dead wood on the forest floor53 underscores the importance of appropriate forest management, as previously highlighted25,54. In addition, diet studies of generalist species such as S. a. aurorae potentially provide information on smaller, more elusive invertebrates, shedding light on their diversity within forest ecosystems and helping to assess habitat integrity55. Therefore, our approach should be extended to other amphibian species and habitats. The knowledge gained from this research, being relevant for both predator diet and soil invertebrate biodiversity, could assist in the rapid development of integrated conservation and management strategies, which are essential for the protection of the world’s most threatened vertebrate group and their habitats.
Overall, our results align with previous research on other Salamandra species8,24, which predominantly prey on Diptera, Myriapoda, Aranea, Coleoptera, and Psocoptera. However, with 177 taxa detected, including 103 species, the golden Alpine salamander’s diet appears to be richer with respect to the fire salamander S. salamandra, for which Marques et al.24 found 58 prey taxa (48 genera), and Wang et al.8 20 prey taxa. Even lower numbers were detected in three sympatric Desmognathus species in another recent metabarcoding study (11 genera or species for D. ocoee, 11 for D. amphileucus, and 22 for D. monticola23). However, it should be noted that these discrepancies might be due to the different sample type used in these previous studies (faecal samples) and/or to marker choice (18S8; COI, but different single set of primers23). In fact, the effect of prey digestion on morphological identification of prey items in amphibian species has been reported before15,16. As for DNA metabarcoding studies, stomach content samples are known to contain better quality DNA compared to fecal samples56, although to the best of our knowledge no studies comparing DNA metabarcoding data from stomach flushes and fecal samples in amphibians have been published so far.
Based on our results, a strong correlation was observed between the dietary composition of both males and females, suggesting that S. a. aurorae does not exhibit strong sexual dietary dimorphism and has a generalist trophic strategy. Similarly, no substantial dietary divergence was detected between pregnant and non-pregnant females, indicating that reproductive status does not significantly influence prey selection in this taxon.
Although promising, the DNA metabarcoding approach still presents technical challenges. For instance, the need for using multiple primer pairs in DNA metabarcoding diet studies is currently debated. Although it has been argued that choosing appropriate primers according to each study design can effectively eliminate the need for multiple primer sets, at least in studies focusing on insect species (e.g., insect mock community57), combinations of different primers are still used where the expected targets encompass a broader taxonomic range (e.g., diet of three horseshoe bats58; of three insectivore mammals59; of S. salamandra24). In this study, which targeted Annelida and Mollusca in addition to Arthropoda, 29.4% of the 177 taxa identified were detected by both primers, 36.7% only by PRIMER_A, and 33.9% only by PRIMER_B. Therefore, the use of two primers greatly improved the overall dataset, and we conclude that a combination of primer sets should be used when studying the diet of generalist species like the golden Alpine salamander.
Overall, our results confirm the potential of metabarcoding studies to reach a much higher taxonomic resolution and accuracy with respect to morphological analysis (e.g.7,60). The comparison of results from the DNA metabarcoding protocol presented here and those from the morphological analysis published in Centomo et al.25 showed that the former performed better in terms of number of positive samples, number of total taxa identified, taxonomic resolution, and %FO obtained, for taxa that were found by both methods. The false negatives detected in the DNA metabarcoding approach (i.e., Pseudoscorpiones and Anoplura, which were found by25 but not in the present study) may be attributed to the lack of comprehensive reference COI sequences for these taxa (e.g.61); to differential amplification biases of the primers chosen59; or DNA degradation of certain prey items. However, as the two missing taxa were found in less than 5% of the samples in25, our overall knowledge of the golden Alpine salamander’s diet was not changed significantly by using a molecular approach. Importantly, however, it should be noted that DNA metabarcoding cannot at present distinguish between prey life stages, providing less ecological information than morphology-based identifications62,63. Furthermore, metabarcoding does not provide other data, such as the abundance of each prey taxon64, which is relevant to some aspects of trophic ecology such as trophic strategy, prey selectivity and inter-individual diet variation. Given the limitations of both methods, we conclude that an integrated approach represents the most effective way to investigate the diet of amphibians.
The detection of a few taxa with geographical distributions inconsistent with our study area suggests that some sequences may have originated from contaminated reference databases or sequencing errors, a challenge inherent to DNA metabarcoding studies. According to GBIF, however, two species and two genera found here have been detected in areas geographically close to the study area (i.e., central Italy, eastern France, Austria, and southern Germany: E. ornatum; Austria: S. lachlani; northern Italy and in the Veneto Prealps: Metylophorus; southern Europe: Amphigerontia). Additionally, it should be noted that Pemphigidae are currently considered a subfamily under the name Eriosomatinae65,66, and the latter has also been recorded in northern Italy and Slovenia. The identification of the remaining taxa by metabarcoding could be explained as misassignments, due to undetected PCR/sequencing errors, or to errors or biases linked to the reference database67,68,69. Nonetheless, considering the current rate of alien species translocation, further investigation into the invertebrate fauna of the study area would help to confirm, or rule out, the presence of these ‘unlikely’ taxa.
Most prey taxa found in the stomach flushing samples are known to dwell in the soil litter for at least one life stage, and could therefore plausibly become prey of the target species42,43,44,70. However, as some of the taxa detected are known parasites or predators (i.e., Hymenoptera: Braconidae, Ichneumonidae, and Chrysididae parasitize Lepidoptera and Diptera; Neuroptera and Psocoptera prey on aphids43), part of the biodiversity identified by metabarcoding could be secondary prey. While this possibility needs to be considered when drawing conclusions on the target species’ ecology and behavior, nonetheless the DNA metabarcoding approach represents a powerful survey methodology to investigate the complexity of the trophic network of ground dwelling animals. In this scenario, combining morphological and metabarcoding approaches could help distinguish between primary and secondary prey.
While our study confirms the advantages of DNA metabarcoding in dietary studies, certain methodological challenges must also be acknowledged. The use of COI primers, although effective for a wide range of invertebrates, introduces inherent biases due to differential amplification efficiencies across taxa: some prey groups may be underrepresented or entirely missed if they possess a lower primer affinity59. Future studies could mitigate this limitation by incorporating additional genetic markers or employing shotgun sequencing approaches to increase taxonomic coverage. Furthermore, the reliability of prey identification is contingent upon the completeness of reference databases61. Our manual verification of species distributions helped reduce potential misidentifications, but continued efforts to expand DNA barcode libraries for regional invertebrate fauna would further enhance accuracy.
Data availability
The dataset generated and analyzed in this study is publicly accessible on the EMBL Database, Project Accession: PRJEB86487 (https://www.ebi.ac.uk/ena/browser/view/ERP169832).
References
Luedtke, J. A. et al. Ongoing declines for the world’s amphibians in the face of emerging threats. Nature 622 (7982), 308–314 (2023).
Clare, E. L. Molecular detection of trophic interactions: emerging trends, distinct advantages, significant considerations and conservation applications. Evol. Appl. 7 (9), 1144–1157 (2014).
Rodríguez-Barreras, R., Godoy-Vitorino, F., Præbel, K. & Wangensteen, O. DNA metabarcoding unveils niche overlapping and competition among Caribbean sea urchins. Reg. Stud. Mar. Sci. 40, 101537 (2020).
Urban, P., Præbel, K., Bhat, S., Dierking, J. & Wangensteen, O. S. DNA metabarcoding reveals the importance of gelatinous zooplankton in the diet of Pandalus borealis, a keystone species in the Arctic. Mol. Ecol. 31 (5), 1562–1576 (2022).
Siegenthaler, A. et al. Metabarcoding of shrimp stomach content: Harnessing a natural sampler for fish biodiversity monitoring. Mol. Ecol. Resour. 19 (1), 206–220 (2019).
Krebs, C. J. in Ecology: the Experimental Analysis of Distribution and Abundance. 6 edn (eds Pearson International) (Cummings, 2009).
Riccioni, G., Stagioni, M., Piccinetti, C. & Libralato, S. A metabarcoding approach for the feeding habits of European Hake in the Adriatic sea. Ecol. Evol. 8 (21), 10435–10447 (2018).
Wang, Y. et al. Diet diversity and environment determine the intestinal Microbiome and bacterial pathogen load of fire salamanders. Sci. Rep. 11 (1), 20493 (2021).
Robeson, M. S. et al. Assessing the utility of metabarcoding for diet analyses of the omnivorous wild pig (Sus scrofa). Ecol. Evol. 8, 185–196 (2018).
Mychek-Londer, J. G., Chaganti, S. R. & Heath, D. D. Metabarcoding of native and invasive species in stomach contents of great lakes fishes. PLoS ONE. 15, e0236077 (2020).
Romano, A., Salvidio, S., Palozzi, R. & Sbordoni, V. Diet of the newt Triturus carnifex (Laurenti, 1768), in the flooded karst sinkhole Pozzo del Merro, central Italy. J. Cave Karst Stud. 74 (3), 271–277 (2012).
Roner, L. et al. A midsummer night’s diet: snapshot on trophic strategy of the Alpine salamander Salamandra atra. Diversity 12, 202 (2020).
Pincheira-Donoso, D. Testing the accuracy of fecal-based analyses in studies of trophic ecology in lizards. Copeia 2008(2), 322–325 (2008).
Rodgers, T. W. & Janečka, J. E. Applications and techniques for non-invasive faecal genetics research in felid conservation. Eur. J. Wildl. Res. 59, 1–16 (2013).
Crovetto, F., Romano, A. & Salvidio, S. Comparison of two non-lethal methods for dietary studies in terrestrial salamanders. Wildl. Res. 39 (3), 266–270 (2012).
Costa, A. et al. What goes in does not come out: different non-lethal dietary methods give contradictory interpretation of prey selectivity in amphibians. Amphibia-Reptilia 35 (2), 255–262 (2014).
Ficetola, G. F., Manenti, R., Taberlet, P. & Environmental DNA and metabarcoding for the study of amphibians and reptiles: species distribution, the microbiome, and much more. Amphibia-Reptilia 40 (2), 129–148 (2019).
Hope, P. R. et al. Second generation sequencing and morphological faecal analysis reveal unexpected foraging behaviour by Myotis Nattereri (Chiroptera, Vespertilionidae) in winter. Front. Zool. 11, 1–15 (2014).
Gil, V. et al. Questioning the proverb ‘more haste, less speed’: classic versus metabarcoding approaches for the diet study of a remote Island endemic gecko. PeerJ 8, e8084 (2020).
Evans, H. K., Bunch, A. J., Schmitt, J. D., Hoogakker, F. J. & Carlson, K. B. High-throughput sequencing outperforms traditional morphological methods in blue catfish diet analysis and reveals novel insights into diet ecology. Ecol. Evol. 11 (10), 5584–5597 (2021).
Jusino, M. A. et al. An improved method for utilizing high-throughput amplicon sequencing to determine the diets of insectivorous animals. Mol. Ecol. Resour. 19 (1), 176–190 (2019).
Pereira, A., Samlali, M. A., S’Khifa, A., Slimani, T. & Harris, D. J. A pilot study on the use of DNA metabarcoding for diet analysis in a montane amphibian population from North Africa. AJH 70(1), 68–74 (2021). https://doi.org/10.1080/21564574.2021.1921058.
Funk, A. T., Holt, B. H. & Pierson, T. W. Evaluation of fecal metabarcoding for studying the diets of sympatric dusky salamanders (Desmognathus). Herpetol Conserv. Biol. 17 (3), 478–488 (2022).
Marques, A. J., Mata, V. A. & Velo-Antón, G. COI metabarcoding provides insights into the highly diverse diet of a generalist salamander, Salamandra salamandra (Caudata: Salamandridae). Diversity 14 (2), 89 (2022).
Centomo, E., Roner, L., Salvatori, M., Pedrini, P. & Romano, A. Rare and hungry: feeding ecology of the golden Alpine salamander, an endangered amphibian in the Alps. Animals 13 (13), 2135 (2023).
Romanazzi, E. & Bonato, L. Updating the range of the narrowly distributed endemites Salamandra atra aurorae and S. atra pasubiensis. Amphibia-Reptilia 35, 123–128 (2014).
Rondinini, C., Battistoni, A. & Teofili, C. (eds) Lista Rossa IUCN Dei Vertebrati Italiani 2022 (Comitato Italiano IUCN e Ministero dell’Ambiente e della Sicurezza Energetica, 2022).
Eccel, E. & Saibanti, S. Inquadramento climatico dell’altopiano di Lavarone-Vezzena nel Contesto generale Trentino. Studi Trentini Di Scienze Naturali Acta Biologica. 82 (2005), 111–121 (2007).
Beukema, W. & Brakels, P. Discovery of Salamandra Atra aurorae (Trevisan, 1982) on the Altopiano Di vezzena, Trentino (Northeastern Italy). Acta Herpetol. 3 (1), 77–81 (2008).
Klewen, R. F. Die Landsalamander Europas 1: Die Gattungen Salamandra Und Mertensiella (A. Ziemsen: Wittenberg Lutherstadt,, 1988).
Trzcionka, M., Withers, K. W., Klingenspor, M. & Jastroch, M. The effects of fasting and cold exposure on metabolic rate and mitochondrial proton leak in liver and skeletal muscle of an amphibian, the cane toad Bufo Marinus. J. Exp. Biol. 211, 1911–1918 (2008).
Fraser, D. F. Coexistence of salamanders in the genus Plethodon: A variation of the Santa Rosalia theme. Ecol 55, 238–251 (1976).
Zanovello, L. et al. A validated protocol for eDNA-based monitoring of within-species genetic diversity in a pond-breeding amphibian. Sci. Rep. 13, 4346 (2023).
Curran, T. G. et al. One Bat’s waste is another man’s treasure: a DNA metabarcoding approach for the assessment of biodiversity and ecosystem services in Ireland using Bat faeces. Biodivers. Conserv. 31, 2699–2722 (2022).
Fonseca, M. L. et al. Dietary preferences and feeding strategies of Colombian Highland woolly monkeys. Sci. Rep. 12, 14364 (2022).
Wilson Rankin, E. E., Knowlton, J. L., Shmerling, A. J. & Hoey-Chamberlain, R. Diets of two non-native praying mantids (Tenodera sinensis and Mantis religiosa) show consumption of arthropods across all ecological roles. Food Webs. 35, e00280 (2023).
Zeale, M. R. K., Butlin, R. K., Barker, G. L. A., Lees, D. C. & Jones, G. Taxon-specific PCR for DNA barcoding arthropod prey in Bat faeces. Mol. Ecol. Resour. 11, 236–244 (2011).
Wangensteen, O. S., Palacín, C., Guardiola, M. & Turon, X. DNA metabarcoding of Littoral hard-bottom communities: high diversity and database gaps revealed by two molecular markers. PeerJ 6, e4705 (2018).
Albanese, D., Fontana, P., De Filippo, C., Cavalieri, D. & Donati, C. MICCA: a complete and accurate software for taxonomic profiling of metagenomic data. Sci. Rep. 5, 9743 (2015).
Edgar, R. C. UNOISE2: improved error-correction for illumina 16S and ITS amplicon sequencing. BioRxiv 081257 (2016).
Madden, T., The, B. L. A. S. T. & Sequence Analysis, T. Oct 9 [Updated 2003 Aug 13]. In: McEntyre J, Ostell J, editors. The NCBI Handbook [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2002-. Chapter 16. (2002). Available from: http://www.ncbi.nlm.nih.gov/books/NBK21097/.
Pozzi, G. Insetti d’Italia E d’Europa: Conoscerli E Riconoscerli (Le guide di airone, Mondadori, 1988).
Chinery, M. Guida Degli Insetti d’Europa: Atlante Illustrato a Colori (Nuova edizione aggiornata, Franco Muzzio ed., 2010).
Bellmann, H. Guida Ai Ragni d’Europa Oltre 400 Specie Europee (Franco Muzzio ed., 2011).
Granquist, S. M., Esparza-Salas, R., Hauksson, E., Karlsson, O. & Angerbjörn, A. Fish consumption of harbour seals (Phoca vitulina) in North Western Iceland assessed by DNA metabarcoding and morphological analysis. Polar Biol. 41 (11), 2199–2210 (2018).
Lee, C. I., Wang, F. Y., Liu, M. Y., Chou, T. K. & Liao, T. Y. DNA metabarcoding for dietary analysis of holland’s carp (Spinibarbus hollandi) to evaluate the threat to native fishes in Taiwan. J. Fish. Biol. 99 (5), 1668–1676 (2021).
RStudio, T. & RStudio RStudio: Integrated Development for R. PBC, Boston, MA URL. http://www.rstudio.com/ (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. ISBN 978-3-319-24277-4. https://ggplot2.tidyverse.org (2016).
Inkscape Project. Inkscape. (2020). Retrieved from https://inkscape.org.
Ferrie, G. M. et al. Nutrition and health in amphibian husbandry. Zoo Biol. 33 (6), 485–501 (2014).
Della Togna, G. et al. Evaluating amphibian biobanking and reproduction for captive breeding programs according to the amphibian conservation action plan objectives. Theriogenology 150, 412–431 (2020).
IUCN SSC Amphibian Specialist Group. Salamandra atra. The IUCN Red List of Threatened Species 2022: e.T19843A89706038. (2022).
Parisi, F. et al. Linking Deadwood traits with saproxylic invertebrates and fungi in European forests—A review. iForest 11, 423–436 (2018).
Romano, A. et al. Forest management and conservation of an elusive amphibian in the alps: habitat selection by the golden alpine salamander reveals the importance of fine Woody debris. Ecol. Manag. 424, 338–344 (2018).
Boyer, S., Cruickshank, R. H. & Wratten, S. D. Faeces of generalist predators as ‘biodiversity capsules’: A new tool for biodiversity assessment in remote and inaccessible habitats. Food Webs. 3, 1–6 (2015).
de Sousa, L. L., Silva, S. M. & Xavier, R. DNA metabarcoding in diet studies: unveiling ecological aspects in aquatic and terrestrial ecosystems. Environ. DNA. 1 (3), 199–214 (2019).
Elbrecht, V. et al. Validation of COI metabarcoding primers for terrestrial arthropods. PeerJ 7, e7745 (2019).
Aldasoro, M. et al. Gaining ecological insight on dietary allocation among horseshoe bats through molecular primer combination. PLoS One. 14 (7), e0220081 (2019).
Browett, S. S. et al. Primer biases in the molecular assessment of diet in multiple insectivorous mammals. Mamm. Biol. 101, 293–304 (2021).
Yoon, T. H. et al. Metabarcoding analysis of the stomach contents of the Antarctic Toothfish (Dissostichus mawsoni) collected in the Antarctic Ocean. PeerJ 5, e3977 (2017).
Muster, C. et al. The dark side of pseudoscorpion diversity: the German barcode of life campaign reveals high levels of undocumented diversity in European false scorpions. Ecol. Evol. 11, 13815–13829 (2021).
Harper, L. R. et al. Using DNA metabarcoding to investigate diet and niche partitioning in the native European otter (Lutra lutra) and invasive American mink (Neovison vison). Ecol. Evol. 9, 8181–8193 (2019).
Nielsen, J. M., Clare, E. L., Hayden, B., Brett, M. T. & Kratina, P. Diet tracing in ecology: method comparison and selection. Methods Ecol. Evol. 9, 278–291 (2018).
Lamb, P. D. et al. How quantitative is metabarcoding: a meta-analytical approach. Mol. Ecol. 28, 420–430 (2019).
Sano, M. & Akimoto, S. I. Morphological phylogeny of gall-forming aphids of the tribe Eriosomatini (Aphididae: Eriosomatinae). Syst. Entomol. 36 (4), 607–627 (2011).
Li, X., Jiang, L. & Qiao, G. Is the subfamily Eriosomatinae (Hemiptera: Aphididae) monophyletic? Turk. J. Zool. 38 (3), 285–297 (2014).
Mioduchowska, M., Czyż, M. J., Gołdyn, B., Kur, J. & Sell, J. Instances of erroneous DNA barcoding of metazoan invertebrates: are universal COX1 gene primers too universal? PLoS One. 13 (6), e0199609 (2018).
Alberdi, A. et al. Promises and pitfalls of using high-throughput sequencing for diet analysis. Mol. Ecol. Resour. 19 (2), 327–348 (2019).
Pentinsaari, M., Ratnasingham, S., Miller, S. E. & Hebert, P. D. BOLD and GenBank revisited–Do identification errors arise in the lab or in the sequence libraries? PLOS One 15(4), e0231814 (2020).
Bonato, L., Peretti, E., Sandionigi, A. & Bortolin, F. The diet of major predators of forest soils: a first analysis on syntopic species of chilopoda through DNA metabarcoding. Soil. Biol. Biochem. 158, 108264 (2021).
Acknowledgements
This study was partially carried out with funding to HCH at the Fondazione E. Mach (Project BIOALPEC) under the National Biodiversity Future Centre (NBFC) Project (code CN_00000033, Concession Decree No. 1034 of 17 June 2022 adopted by the Italian Ministry of University and Research, CUPD43C22001280006), funded under the National Recovery and Resilience Plan (NRRP), Mission 4 Component 2 Investment 1.4 - Call for tender No. 3138 of 16 December 2021, rectified by Decree n.3175 of 18 December 2021 of Italian Ministry of University and Research funded by the European Union – NextGenerationEU. This manuscript reflects only the authors’ views and opinions, neither the European Union nor the European Commission can be considered responsible for them. This study was also financially supported by the Autonomous Province of Trento Natura 2000 Network. The authors would like to thank the Fondazione E. Mach for access to facilities, the forestry staff of the Municipality of Levico and the PAT Forest Service for facilities and field support, and Emma Centomo for fieldwork support. We would like to thank Paolo Fontana and Lucio Bonato for their expert advice on, respectively, invertebrate species distributions and S. a. aurorae biology.
Author information
Authors and Affiliations
Contributions
Conceived and designed the study: all authors; acquired the funding: M.G., A.R., P.P., H.C.H.; collected the samples: F.P., L.R., A.R.; designed the molecular protocol: L.Z, F.P., M.G., H.C.H.; performed laboratory analyses: F.P., M.G.; completed bioinformatic and statistical analyses: L.Z., F.P.; drafted the manuscript: L.Z., H.C.H; read, commented and approved the manuscript: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zanovello, L., Polli, F., Girardi, M. et al. DNA metabarcoding analysis of stomach flushing contents reveals the exceptionally diverse diet of the golden alpine salamander. Sci Rep 15, 34080 (2025). https://doi.org/10.1038/s41598-025-14757-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14757-8