Abstract
Accurate detection of brain tumors remains a significant challenge due to the diversity of tumor types along with human interventions during diagnostic process. This study proposes a novel ensemble deep learning system for accurate brain tumor classification using MRI data. The proposed system integrates fine-tuned Convolutional Neural Network (CNN), ResNet-50 and EfficientNet-B5 to create a dynamic ensemble framework that addresses existing challenges. An adaptive dynamic weight distribution strategy is employed during training to optimize the contribution of each networks in the framework. To address class imbalance and improve model generalization, a customized weighted cross-entropy loss function is incorporated. The model obtains improved interpretability through explainabile artificial intelligence (XAI) techniques, including Grad-CAM, SHAP, SmoothGrad, and LIME, providing deeper insights into prediction rationale. The proposed system achieves a classification accuracy of 99.4% on the test set, 99.48% on the validation set, and 99.31% in cross-dataset validation. Furthermore, entropy-based uncertainty analysis quantifies prediction confidence, yielding an average entropy of 0.3093 and effectively identifying uncertain predictions to mitigate diagnostic errors. Overall, the proposed framework demonstrates high accuracy, robustness, and interpretability, highlighting its potential for integration into automated brain tumor diagnosis systems.
Similar content being viewed by others
Introduction
Brain tumors are considered a significant global health concern due to their high mortality rates and the inherent complexity of their diagnosis and treatment. These may be classified as benign or malignant tumors, with both types capable of causing significant neurological diseases according to the tumor’s location within the brain or spinal cord. However, there is a lack of clear understanding of the precise cause of brain tumors, which limits diagnostic and, hence, treatment opportunities. Brain tumors are categorized as either primary—originating within the central nervous system (CNS)—or secondary, resulting from metastatic dissemination from extracranial malignancies. Among primary brain tumors, gliomas constitute the most prevalent and aggressive group, with Glioblastoma Multiforme (GBM) being the most lethal form, characterized by rapid proliferation and poor prognosis. In contrast, tumors such as meningiomas and pituitary adenomas are typically considered benign; however, due to their potential mass effect and involvement in critical neuroanatomical regions, they can still lead to significant clinical complications. Epidemiologically, brain tumors account for approximately 2.5% of all cancer-related mortalities. According to global estimates, the incidence of brain and other CNS tumors was projected to reach approximately 308,000 new cases in the year 2020. In India alone, the annual incidence ranges between 28,000 and 50,000 newly diagnosed cases1.
There are reasons why early diagnosis is often problematic, which include diverse symptoms that often mimic other diseases and the need to consult radiologists, which may add variability and human errors. Regarding the technologies discussed, it has been pointed out that deep learning technologies, particularly Convolutional Neural Networks (CNN), possess extensive potential in automating the detection and classification of brain tumors. Unlike the traditional situational diagnosis that often gives clinical judgment a clear win most of the time, deep learning frameworks have the ability to perform high-quality and even faster analysis of complicated medical images, trained with high levels of accuracy. Even though developing great accuracy for tumor classification of images, such as CNN-based designs like ResNet and EfficientNet designs through compounding minuscule image details, many models rely solely on one design and do not address the variability of brain tumor images2.
To eliminate this limitation, this research presents an ensemble approach, which combines three construction neural networks: EfficientNet-B5, ResNet-50, and an additional customized CNN. During training, this allows the ensemble method to more effectively train under image examples of brain tumors by redistributing the relative importance of each network among itself. In this work, a unique and powerful technique called the dynamic weighting strategy is introduced that enables the model to tune the significance of different designs at different training stages for better performance. This dynamic update raises the model’s performance level to meet the high standards expected in the analysis of complex medical images3,4. In addition, the ReduceLROnPlateau function is implemented, which helps to adjust the learning rate dynamically throughout the training phase to help avoid the overfitting problem and achieve better generalization performance on unseen data. Besides, it is also quite important that, apart from achieving high accuracy of the classification, the model’s comprehension is equally important with respect to this method5.
Several Explainable AI (XAI) approaches determine how well clinicians maintain their trust for medical applications. The decision-making process of the model receives evaluation through analysis tools that include Grad-CAM, Guided Grad-CAM, SHAP, SmoothGrad and LIME to determine which MRI scan areas contributed to model decisions. The supplementary parameter ‘uncertainty entropy’ adds to prediction reliability by displaying how certain the model remains about specific classifications. The identification of uncertain predictions occurs through this strategy which lets clinicians investigate questionable predictions when their confidence levels are low. The model exhibited robustness through its ability to accept different dataset formats during cross-dataset testing because this characteristic is essential for authentic clinical practice methods. A single strategy serves two purposes by speeding up and improving deep learning accuracy for brain tumor diagnosis and revealing how this model achieves its purpose6.
The key contributions of this work are listed as:
-
Dynamic ensemble model for Brain tumor classification: This research proposes a new strategy for combining the EfficientNet-B5 model, ResNet-50, and a CNN designed specifically for the task. It dynamically allows for the contribution rates of each of the models during learning, which enhances the reliability of the brain tumors classification.
-
Custom weighted cross-entropy loss function: A specific loss function is introduced to handle class imbalance and improve classification accuracy. The differentiation of the tumor classes is done by the model through the application of different odds to the tumor classes.
-
Explainability with multiple XAI techniques: To ensure that the constructed AI model is transparent and can be trusted when used in medical diagnosis, the study uses five XAI techniques including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM, and LIME.
-
Cross dataset validation for generalization: Cross-dataset validation is further performed to establish the generalization capability of the developed model, and the suitability of the model for real-world medical problem-solving is also ensured.
The study includes multiple main divisions which display both thorough assessments of brain tumor classification methods and the results achieved by ensemble deep learning modeling. An ensemble model consists of three main components, including EfficientNetB5 along with the ResNet50 and a self-made blank CNN. This investigation centres its analysis on three main aspects: dynamic weight distribution policy, novel loss function and explanation methods for XAI based on Grad-CAM, SHAP and SmoothGrad. The authors describe extensively the training procedure alongside model implementation details to solve the previously identified problems within this segment.
The experimental results segment provides detailed information about the result assessments and the finest domain models’ performance comparison. This result section presents accuracy together with precision of the model and its recall performance and F1 score measurements, and entropy-based uncertainty analysis. Additionally, it shows feature maps and XAI artefacts. The study contains cross-dataset validation results, which show how the model handles generalization abilities. The conclusion section summarizes the obtained findings, followed by a discussion of this study’s contributions which present future development possibilities for model improvement in medical applications.
Literature review
In7, authors examined the deadly nature of brain tumors affecting adults and children while emphasizing prompt and precise diagnosis to help prevent deaths. The authors studied the classification of brain tumors that included Glioma and meningioma, and pituitary neoplasms. They mentioned that although many papers covered brain tumor classification, there is minimal discussion about feature selection issues. Noreen et al. produced a deep-learning-based automatic diagnostic system that avoids the problems found in manual diagnosis and traditional feature-extraction approaches. Using the advanced models Inception-v3 and Xception for feature extraction produced a test accuracy level of 94.34% as they showed better performance than Inception-v3 alone. MRI scan interpretation through deep learning proved beneficial for radiologists as an analysis decision tool according to research which applied RF, support vector machine (SVM) and KNN classification techniques.
In 2024, Janidu Chathumina et al.8 explored the use of deep learning (DL) and computer vision in distinguishing anatomical features, particularly for brain tumor diagnosis. While DL methods show promising result in early detection, their integration into clinical workflows is hindered by explainability issues and the black-box nature of models. Despite achieving state-of-the-art (SOTA) performance, many models lack transparency, necessitating the use of XAI techniques to address these challenges. Chathumina’s review paper examines recent research on brain tumor diagnosis using DL, comparing studies that incorporate XAI techniques with those that do not. This study carries a comparative analysis of techniques and their performances, highlighting the strengths, weaknesses, and limitations of each. It identifies research gaps that need to be addressed and concludes with key findings, potential future directions, and recommendations for improving the integration of DL in clinical settings.
In9, the use of transfer learning for brain tumor classification using MRI images, applying three pre-trained CNN models—VGG-16, ResNet-50, and MobileNet has been explored. After preprocessing the MRI dataset with resizing, normalization, and augmentation, the models were fine-tuned and evaluated using accuracy, precision, recall, F1-score, and AUC. Among the models, VGG-16 achieved the best performance with 97.2% accuracy, an AUC of 0.974, and the shortest processing time (22%), making it the most efficient and accurate. ResNet-50 followed with 96% accuracy but higher computational cost (43%), while MobileNet had the lowest accuracy at 87% and moderate efficiency. The study concludes that VGG-16 is the most effective model for accurate and fast brain tumor classification in clinical settings.
The study in10 presents a novel ensemble approach combining ResNet50 and EfficientNet-B7 to improve brain tumor classification accuracy using over 22,000 high-resolution MRI images categorized into four classes: no tumor, glioma, meningioma, and pituitary tumor. The proposed model employs extensive preprocessing steps—including normalization, augmentation, and stratified splitting—followed by training both individual networks and integrating their predictions through a weighted average ensemble strategy (30% ResNet50, 70% EfficientNet-B7). The ensemble model significantly outperformed the standalone models, achieving a test accuracy of 99.53% compared to 97.4% for ResNet50 and 98.2% for EfficientNet-B7, with superior precision, recall, and F1-scores across all tumor classes. This dynamic ensemble model, enhanced by learning rate optimization and adaptive weight tuning, demonstrated exceptional generalization and robustness, making it a promising tool for reliable, real-time brain tumor detection in clinical environments.
Another study by Ghazanfar Latif et al.11 designed an automated approach to brain tumors, which was very important since the typical binary method of segregating tumourous and non-tumourous images was not very effective in the complete classification of the type of tumor. Their research focuses on classifying brain tumors into four types: necrotic lesions, edema lesions, enhancing lesions, and non-enhancing lesions. In this study, wavelet feature extraction of four MRI modalities, namely Flair, T1, T1c, and T2, has been applied to create the ensemble feature set for the binomial classification using the random forest trees. Once the tumorous images were identified, a region-growing segmentation algorithm was used to segment the tumor. In the final step, wavelet features were extracted from the segmented tumor regions for multiclass classification. The technique was investigated on 35 patients with 14 LGG, 21 HGG, and 21,700 MR images. The study came up with an average accuracy of 94.33% for binomial MR image classification and 96.08% for multiclass tumor classification which indicates the effectiveness of the deep learning method in enhancing the tumor diagnosis.
Shafi et al.12 presented a method for intracranial tumor classification—glioma, meningioma, pituitary adenoma, and multiple sclerosis which utilized MRI. Their complete system consists of pre-processing followed by feature extraction and feature selection before concluding with classification procedures. The first step adopts ROI extraction while applying Collewet normalization followed by quantization through Lloyd-max. The SVM operates as the fundamental classifier, whose final decision is reached by a voting mechanism. The proposed model achieved a weighted sensitivity of 97% and specificity of 97%. Its precision measurement reached 97% with overall accuracy at 97.5%, 98%, 83.8%, 98.011% and 98.719% respectively. Experimental data showed that training and testing accuracy reached approximately 97%. Furthermore, the evaluation demonstrated rates of 95.7% and 97.744%. The study demonstrated that the model displays an important capability to detect tumors and lesions which will advance neuro-medicine diagnosis for detecting tumors and multiple sclerosis in the brains.
Nahid Ferdous Aurna et al.13 proposed a novel approach for the pattern-based categorization of brain cancers through a two-layer feature CNN ensemble model. The study aims to perform CADx, an auxiliary to the human examination, which has limitations as a result of exhaustion and fatigue. The model uses three sets of MRI images, which consist of three types of brain tumors (meningioma, glioma, pituitary) and normal brain images. The authors used already existing five fixed models together with a specific CNN for feature extraction based on which the best models were concatenated in two stages. Using PCA, the principal components were obtained, and the optimum classifier was used to identify the important traits. The model yielded an average accuracy of 98% at the end of training. Also, a user interface (UI) was developed to assist with the real-time evaluation of the entire model.
In14, authros developed a novel CNN model applying XAI approaches for brain tumor diagnostics. While deep learning has shown amazing gains, it frequently lacks interpretability, making decision-making in key applications like medical diagnosis problematic. To overcome this, the authors presented a unique numerical Grad-CAM (numGrad-CAM) incorporated into a CNN model. This technique enhanced the classic Grad-CAM by adding numerical information, allowing clinicians to interpret and get clearer results. The model was assessed using technical assessments and human-centered (physician) evaluations, obtaining 97.11% accuracy, 95.58% sensitivity, and 96.81% specificity in brain tumor categorization. Additionally, the numGrad-CAM component achieved 90.11% accuracy compared to older CAM versions. Physicians responded favorably to the enhanced explainability features, indicating that the model has strong potential for clinical adoption by supporting safer and more interpretable diagnostic decisions.
The authors in15 addressed the problem of brain tumor identification and suggested to employ CNNs for MRI image classification. Early detection of brain cancer can greatly improve patient outcomes. CNN also provides a more thorough approach to deep learning techniques in the analysis of medical imaging. The study used two CNN models and carried out hyperparameter tuning and data augmentation work to boost performance. The model accuracy was up to 96%. Further, the XAI was deployed to examine models’ decision-making processes, enabling the discovery of patterns or factors that affected prediction results. This method enhanced the trustworthiness of diagnosis and provided transparency for clinical application. The study highlighted the potential of CNN models connected with XAI approaches in increasing brain tumor diagnostics, delivering an open-source library of code and images created during the testing for later research.
In16, authors employed deep learning (DL) for non-invasive diagnosis of adamantinoma Tous craniopharyngioma (ACP), which is currently diagnosed through biopsy. Though clinical experts can classify ACP via MRI with 86% accuracy, no study has explored the use of non-invasive DL classification because of predictive uncertainty. The authors used data from 86 suprasellar tumor patients collected from different centers to train a DL classifier in the preoperative MRI. Authors applyied a Bayesian DL framework to update their previously reported ACP classifier to yield predictive distributions rather than point values. Despite the poor performance of the calibrated model initially attributed to overfitting in the underlying model, an analytical estimate of predictive uncertainty was instrumental in proposing a mechanical classification abstention. This made the model achieve improved accuracies of up to 80%. These findings demonstrate the significance of model calibration for the predictive uncertainty of non-invasive brain tumor diagnosis in clinical applications.
In17, authors addressed the crucial demand for transparency in artificial intelligence (AI) models, specifically in medical applications, by adopting XAI approaches for brain tumor diagnosis. The lack of transparency among AI models raises questions about their trustworthiness, particularly in the field of healthcare. To address this challenge, this approach incorporates XAI to improve both the safety and accuracy of brain tumor detection. A four-class Kaggle brain tumor dataset (Dataset I) was used for experimentation purposes. The feature extractor was the pre-trained model, DenseNet201, and tumor areas were segmented using GradCAM. Features were acquired through the Iterative Neighborhood Component (INCA) technique and were classified using a SVM with a 10-fold cross-validation. The model obtained outstanding accuracy rates of 98.65% with Dataset I, surpassing state-of-the-art methods and suggesting future usefulness as a decision-support system for the identification of brain tumors by radiologists. A brief review is shown in Table 1.
Proposed methodology
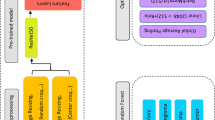
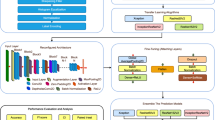
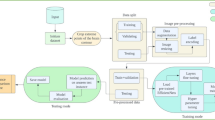
The proposed system offers an innovative approach for brain tumor classification by incorporating several state-of-the-art deep learning models. The ensemble model consists of three robust architectures, ResNet-50, EfficientNet-B5, and a custom CNN, where each architecture operates independently for feature extraction and classification. The dynamic weighting process maintains the appropriate ratio of outputs of the models by manipulating the influence of each model as training progresses, thus enhancing the performance and robustness of the models. Numerous strategies for data pre-processing, as well as augmentation, are also employed to improve the generalization of the proposed model. The method highlights the importance of model interpretability. The XAI methods, including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM, and LIME, are employed in order to provide clarity and understanding in the decision-making process, as demonstrated in Fig. 1. These methods can provide some insight that more traditional deep learning models do not always supply by illuminating the distinct areas of the MRI scan that lend the most credence to decisions from the model. Enhancing interpretability, model capabilities, and performance contributes to high classification accuracy, higher trust for decisions made by the model, and improved usefulness for medical diagnosis, all of which improve the model’s reliability for clinical application.
Proposed methodology.
Dataset
The study employs the Crystal Clean: Brain Tumors MRI Dataset18 for analysis which comprises about 22,000 high-quality MRI images separated into four categories: No Tumor, Glioma Tumor, Meningioma Tumor, and Pituitary Tumor as demonstrated in Fig. 2. Each class represents different tumor features so the dataset serves as an excellent basis for deep learning model training and testing.
The “No Tumor” class is defined by MRI images of normal brains showing clean, symmetrical brain structures devoid of abnormalities. Rising from glial cells, glioma tumors show up on MRI images as irregular lumps and usually induce asymmetry of surrounding brain tissue. Usually seen near the dura mater, meningioma tumors begin from the meninges and exhibit well-defined spherical or lobulated masses on MRI imaging. Finally, pituitary tumors—found in the pituitary gland—are small, limited to the sellar region, affecting little surrounding tissue.
Every class has a directory in the collection, and images reveal typical variations in orientation, size, and intensity seen in clinical settings. The dataset is separated into training (80%), validation (10%), and testing (10%) sets with balanced class representation. Each subset was generated independently with class-wise balancing, and no image overlap occurred between sets. Separate data generators were initialized for each split to prevent unintentional contamination during preprocessing or augmentation. Comprising 2456 No Tumor, 5048 Glioma, 5112 Meningioma, and 4721 Pituitary images, the training set matches validation and testing sets, further ensuring the robustness of the model.
Dataset example.
Preprocessing and augmentation
In this study, the images are preprocessed before feeding them to the model for training so that the model can easily extract features. Consistent scaling of the images using three color channels (RGB) yields a uniform 224 by 224 pixel size. Mathematically, the input image I ∈ R224 × 224 × 3 undergoes various transformations. Preprocessing enables basic normalization by use of a scalar function on the images.
The dataset is enlarged, and the model’s generalizing capability is increased using many methods of data augmentation. These include random rotations of up to 20 degrees, flips in both directions and variances in both width and height up to 20%. If (x,) is the pixel at coordinates (x,), horizontal flipping transforms it into I′(x,224−y), while vertical flipping results in I′(224−x,y). Also used to duplicate variations in image size is a 20% zoom range. Rotation is applied randomly within a range of 20 degrees19. The transformation matrix for rotation by an angle.
θ is given in Eq. (1):
This transforms the pixel coordinates (x,) into (x′,′) using (x′,y′)=Tθ(x,y).
Width and height shifts of up to 20% are also applied. For width shift,
x′=x+a⋅w, where w is the width and a∈[− 0.2,0.2]. Similarly, for height shift, y′=y+β⋅ℎ, where ℎ is the height and β∈[−0.2,0.2].
Zoom augmentation applies a zoom factor z∈[1 − 0.2,1 + 0.2], resulting in new pixel coordinates as shown in Eq. 2:
To enhance model robustness and generalization, various image augmentations are applied during training, as illustrated in Fig. 3. These augmentations simulate the diversity encountered in real-world scenarios. Both the training and validation data generators perform data shuffling to prevent overfitting by reducing model bias toward data order. However, the test data generator does not apply shuffling to ensure a consistent and unbiased evaluation. All generators process data in batches of size 16 to optimize memory usage and computational efficiency. For multi-class classification, the target labels are one-hot encoded, and the model is trained using categorical cross-entropy loss. This preprocessing pipeline ensures that the model is exposed to a broad range of visual variations, improving its generalization and performance during evaluation20.
Data augmentation steps.
Fine tuned EfficientNet-B5
The EfficientNetB5 model deployed in this research presents a high-performance deep learning system which adopts pre-trained ImageNet weights for transfer learning applications. The base model EfficientNetB5 functions as a CNN processing images without the fully connected layers due to its include_top = False functionality to accommodate specific classification tasks as shown in Fig. 4. Each input image requires processing with a 224 × 224 × 3 format because it carries three channels, which follow the RGB colour scheme. The mathematical framework behind the model depends on convolutional processes of the EfficientNet architecture, where the convolution operation is defined as shown in Eq. 3:
where I(i,j) represents the input image, and K(m,n) represents the convolutional kernel. EfficientNetB5 scales the width, depth, and resolution of the network uniformly through compound scaling, improving both accuracy and efficiency.
After the base model, the dropout layer randomly deactivates a fraction of neurons during training, which is represented as shown in Eq. (4):
Where r is a random variable, and p is the dropout rate (set to 0.5), ensuring that 50% of the neurons are dropped during training. Finally, the model outputs a softmax function for classification, defined as shown in Eq. (5):
This ensures that the final output is a probability distribution over the n classes, where \(\:{\sum\:}_{i=1}^{n}\sigma\:\left({z}_{i}\right)=1\) The model is optimized using Adamax, an adaptive learning algorithm, with categorical cross-entropy loss as shown in Eq. (6):
Where yi is the true label and \(\:{\widehat{y}}_{i}\)is the predicted probability. This setup ensures robust training and high accuracy for brain tumor classification tasks21,22.
Fine-Tuned EfficientNet-B5 architecture.
Fine tuned ResNet-50
The ResNet-50 model employed in this study uses transfer learning with pre-trained weights belonging to ImageNet and fine-tuned for the purpose of classifying brain tumors. ResNet-50 is a deep CNN that addresses the vanishing gradient issue through residual learning as depicted in Fig. 5, thereby allowing for increased depth. The key mathematical principle behind residual learning is as shown in Eqs. (7) and (8):
Where H(x) represents the desired mapping, F(x) is the residual function, and x is the input. The model learns the residuals rather than directly learning the mapping, enabling it to train deeper layers without degrading performance. In this model, the layers of ResNet-50 are frozen to preserve the learned features from the ImageNet dataset. The frozen layers are represented mathematically as shown in Eq. (9):
Where L is the loss function, and Wfrozen are the weights of the frozen layers. By freezing these layers, the pre-trained features remain intact, preventing them from being updated during backpropagation. The output of the last convolutional layer is pooled using global max pooling, represented as shown in Eq. (10):
Where yi represents the activation from the i-th feature map, this reduces the spatial dimensions while retaining the most important features for classification. The dropout layer is introduced after the ResNet-50 base to prevent overfitting. The dropout mechanism can be described in Eq. (11):
Where r is a random variable, and p is the dropout rate, set to 0.5, meaning 50% of the neurons are dropped out during each forward pass23.
Fine-Tuned ResNet-50 Architecture.
Base CNN architecture
The CNN-based model adopted in this study is categorized as the image categorization model for application in image diagnosis, including brain tumor detection. Its structure contains several convolution layers, partly followed up by pooling layers, attentions, and dense layers as demonstrated in Fig. 6. The convolutional operation applied to the input image is mathematically defined as in Eq. (12):
where I(i,j) is the input image, K(m,n) is the convolutional filter (or kernel), and Y(i,j) is the output of the convolution. The filter size in this model is 3 × 3, with 32, 64, 128, and 256 filters used in different layers to capture varying levels of detail. After each convolutional layer, the ReLU activation function is applied to introduce non-linearity as in Eq. (13):
This ensures that only positive values are propagated forward, making the model more capable of learning complex patterns. Max-pooling is applied to reduce the spatial dimensions of the feature maps. The max-pooling operation is defined as in Eq. (14):
.
Where x1,2,…,xn are the values in the pooling window, and ypooly is the maximum value selected. This operation reduces the dimensionality while retaining the most significant features.
An attention block is introduced in the middle of the architecture to enhance the focus on important regions of the image. The attention mechanism can be expressed as in Eq. (15):
Here, ai is the attention weight, and xi is the feature map. The attention mechanism helps the model focus on the most relevant features, such as tumor regions, by adjusting these weights.
Following the attention block, the output is flattened to a 1D vector using the Flatten operation, which is mathematically represented as reshaping the matrix into a single vector as in Eq. (16):
A dense layer with 256 units follows, where the output is given by Eq. (17):
Where Wi is the weight matrix, x is the input vector, and bi is the bias term. Another application of the ReLU activates all dense layer neurons. The following stage requires dropout techniques for overfitting prevention. During training, Dropout activates random neuron deactivation to ensure 50% neuron drop occurs during each training iteration. Classification happens through the application of softmax function at the end. MemoryWarning defines the output values as probabilities because the summation of all class probabilities equals one. The training of this model occurs with Adamax optimizer that involves categorical cross-entropy as loss mechanism.
This CNN architecture is designed for robust feature extraction and classification, enhanced by attention mechanisms and regularization techniques24,25.
CNN Architecture.
Attention block
The attention block in this model serves as a vital component which enables the model to detect important features throughout the input data especially required for complex applications such as brain tumor examination. Within the attention mechanism, the feature maps undergo a re-weighting process to help the model maintain focus on critical regions26.
The process begins with the application of global average pooling. For a feature map X∈RH×W×C, where H is the height, W is the width, and C is the number of channels, the global average pooling implemented, gives out an average of all the feature channels, effectively reducing the spatial dimensions as represented in Eq. (18):
This results in a vector of size C, which summarizes the most relevant information from each feature map across the entire spatial dimension.
Next, a Dense layer with 256 units is applied. The output z of this dense layer can be expressed as shown in Eq. (19):
where W1 is the weight matrix, X is the input from the previous global pooling layer, and b1 is the bias term. By using the ReLU activation function all negative values get moved to zero for achieving non-linearity.
The second Dense layer is employed with C being the number of channels found in the input feature map. Within this layer the sigmoid activation function creates attention weights through its output process, as shown in Eq. (20):
These attention weights αc scale each feature map, where 0 ≤ αc ≤ 10 for each channel. The sigmoid activation ensures that each channel is re-weighted based on its importance, with values closer to 1 indicating more important features and values closer to 0 indicating less important ones.
The next step involves reshaping these attention weights by expanding their dimensions, ensuring that they match the spatial dimensions of the original feature map H×W×C. This can be represented as shown in Eq. (21):
Where ⊗ denotes element-wise multiplication. This operation multiplies each channel of the original feature map by its corresponding attention weight, effectively re-weighting the feature maps to focus on the most relevant regions.
The output of this attention block is a feature map where important regions and channels are emphasized, allowing the model to focus on critical areas, such as regions where tumors are more likely to appear. Mathematically, the final output is shown in Eq. (22):
This mechanism improves the model’s capacity to capture fine-grained details by allowing it to selectively attend to relevant parts of the input, boosting its performance in classification tasks.
Ensemble model
The ensemble technique is applied in this study to combine EfficientNet-B5, ResNet-50, and a custom CNN model to enhance the classification system. Each of these models participates in their strength in the final classification output using a dynamic weight that varies depending on the current period of training. In this ensemble method, the learning process of the model is guaranteed to adjust the weights of the three network outputs to select the best-performing network (Fig. 7). The input to the ensemble model is an image X∈RH×W×C, where H is the height, W is the width, and C is the number of channels. The image is evaluated by three distinct modelsEfficientNet-B5, ResNet-50, and a CNN to extract high-level information. For each model, the output is represented as shown in Eq. (23):
where fEffNet(X), fResNet(X), and fCNN(X) represent the transformation of the input image X by the respective models. These outputs are feature vectors representing the image’s classification by each model.
The dynamic weighting mechanism applies weights to the outputs of each model based on the current epoch of training. At epoch t, the weights for EfficientNet, ResNet, and CNN are defined as shown in Eq. (24):
These weights are adjusted over time to give more importance to the CNN and ResNet models as the training progresses. The weighted output of the ensemble model is computed as shown in Eq. (25):
This combination ensures that the ensemble model adaptively leverages the strengths of each network. EfficientNet-B5’s strong feature extraction abilities are given higher weight in the early stages of training, while the weights shift toward ResNet-50 and CNN as training progresses. The final output of the ensemble model is passed through the softmax function for classification, as shown in Eq. (26):
where zi denotes the logit corresponding to class i, and the softmax function converts these logit values into probabilities for each of the classes. The model is trained via the use of the Adamax optimizer; this is a form of optimizer that adapts the learning rate during the training process. The loss function used is custom-weighted cross-entropy, where each class is assigned a different weight, as shown in Eq. (27):
Here, wi is the weight assigned to class i, yi is the true label, and \(\:{\widehat{y}}_{i}\) is the predicted probability for class i. This ensures that classes with higher importance are given more weight during optimization, reducing the likelihood of misclassification for these classes. By dynamically combining the outputs of multiple models and adjusting the loss function based on class importance, the ensemble model achieves greater robustness and accuracy in brain tumor classification tasks27. Algorithm 1 outlines the complete workflow of the proposed ensemble model, starting from the dataset input to the final model saving. To mitigate overfitting, the model incorporates various regularization techniques, including dropout layers, L2 regularization, and data augmentation. These strategies collectively enhance the model’s generalization ability and improve performance on unseen data. The three base model components (EfficientNet-B5, ResNet-50, and the custom CNN) have dropout layers set at a rate of 0.5 to randomly deactivate neurons for overfitting prevention. Weight decay as L2 regularization operates on fully connected layers by introducing penalties against large weight values to stop the model from developing complicated structures. The model’s generalization capability improved through the implementation of multiple augmentation techniques which included random rotations at 20 degrees and both horizontal and vertical flipping along with width and height shifts up to 20% and zoom augmentation at up to 20%. Training was halted by early stopping when the validation loss failed to improve for five consecutive epochs during the process.
Ensemble model architecture.
Ensemble model for brain tumor classification using explainability and cross-dataset validation.
Dynamic weights distribution
This research employs an ensemble model comprising three different networks, including EfficientNet-B5, ResNet-50, and a fine-tuned CNN. As the training continued, the weights attributed to each model changed dynamically as shown in Fig. 8. Primarily, due to the high feature extraction performance, the training weight of EfficientNet-B5 was initially larger than that of ResNet-50 and CNN. Again, when training continues until the fourth epoch, both CNN and ResNet-50 weights increase while the weight of EfficientNet-B5 decreases. With the method of adaptive weighting, the generalized ensemble model is warranted for proportioning the outputs of the three models in line with the accuracy achieved in each epoch28.
Mathematically, the dynamic weighting for each model is given as a function of the epoch t. For EfficientNet-B5, the weight decreases over time, ensuring it does not drop below a specified minimum, as shown in Eq. (28):
In contrast, the weights for ResNet-50 and the CNN increase with each epoch, as shown in Eq. (29):
These equations make sure that EfficientNet-B5 continues to dominate in the initial steps of the training process while the ResNet-50 and the CNN become more important throughout the training process. This shifting emphasis enables the ensemble to respond to various phases of the learning process, thus enabling each network to contribute in one way or the other. This dynamic weighting approach enhances the ensemble’s ability to generalize because the workability of each particular network at the optimal time during training is harnessed, enhancing efficiency in brain tumor classification missions.
Dynamic weight adjustment in ensemble model.
Dynamic Weights working.
Figure 9 shows how weight variations for every model change with increased training. Starting with 0.7 at epoch 0, EfficientNet-B5 is useful for early feature extraction. Its weight drops to 0.6 by the second epoch, so it stabilizes and has a negligible effect on the general ensemble output. On the other hand, in Epoch 2, the CNN model begins with a very low weight of 0.1 but increases pretty rapidly to 0.2. Consistent with its continuous contribution, the ResNet-50 model maintains a weight of 0.2 throughout the training run. After epoch 2, ResNet-50 and CNN both settle at 0.2; consequently, EfficientNet balances their respective contributions. This adaptive approach guarantees that, as training advances, the ensemble model relies more on EfficientNet’s strong feature extraction capabilities while in the early stages of training and progressively combines ResNet-50 and CNN; the same is shown in algorithm 2. As the strengths of every network are combined at many phases of the training process, the dynamic weighting lowers overfitting to any one model and boosts the generalizing ability of the model to unknown data.
Dynamic Weights over epochs.
To effectively combine the predictive strengths of the three constituent models including EfficientNet-B5, ResNet-50, and the custom CNN, a dynamic weighting strategy was adopted. This strategy assigns initial weights of 0.7, 0.2, and 0.1, respectively, to the output probabilities of the three models during the early training epochs, followed by linear adjustment at a fixed rate of 0.05 per epoch.
These initial weights were selected based on the representational capacity and architectural depth of the models. EfficientNet-B5, being deeper and computationally richer, was prioritized to leverage its powerful feature extraction ability. ResNet-50 and the CNN model, while relatively lighter, contribute complementary patterns and are progressively emphasized throughout training. This controlled redistribution encourages ensemble diversity and reduces early-stage overfitting to a dominant architecture.
Training details
The ensemble model received training with a purpose to enhance its performance alongside reinforcing generalizability capabilities. The model training necessary batch size of 16 was specified to prevent out-of-memory (OOM) errors which arose from running the high-resolution images combined with the complex architecture. The ImageDataGenerator class produced generators that served both training and validation and testing data from the dataset. The data augmentation strategies on these generators added variety to the dataset while reducing overfitting through techniques that involved image rotation, magnification, and vertical and horizontal flipping. Table 2 contains all the details of the experimental setup.
To mitigate overfitting, multiple regularization strategies were adopted including dropout (rate 0.5), L2 weight regularization in dense layers, extensive data augmentation, and early stopping based on validation loss. Additionally, the learning rate was dynamically reduced using the ReduceLROnPlateau scheduler to improve generalization performance.
During the training phase, three upgraded models were used: an adapted CNN, ResNet-50, and EfficientNet-B5, each of which gave distinct features to the collective. The CNN and ResNet-50 models had a learning rate of 1 × 10 − 3, but the EfficientNet-B5 model had a learning rate of 1 × 10 − 4 owing to its complexity. The Adamax optimizer, a version of the Adam optimizer with an infinite norm, was utilized to enable robust training with customizable learning rates. The optimization equation for updating model parameters θ can be written as shown in Eq. (30):
The algorithm uses mt as the first-moment estimate along with vt as the second moment estimate alongside learning rate η. Weighted cross-entropy loss was used to solve class imbalance issues. The custom loss function distributes weights to each class by using dataset prevalence data. The weighted cross-entropy can be expressed as shown in Eq. (31):
where wi is the weight assigned to class i, yi is the true label, and \(\:{\widehat{y}}_{i}\) is the predicted probability for class i. This enhances the model’s potential to generalize across all classes by making sure that classes with higher significance or fewer samples are given more weight during optimization. Moreover, the adaptive modification of the contribution of each sub-model (EfficientNet-B5, ResNet-50, CNN) over time was made straightforward by the dynamic weighting feature of the ensemble model. Each sub-model had its weights adjusted every epoch; CNN and ResNet-50 had growing weights over time, whereas EfficientNet’s weights slowly fell from higher initial levels. When the model did not improve after five consecutive epochs, training was halted early based on the validation loss being observed. By a factor of 0.2, the learning rate was also lowered by the ReduceLROnPlateau callback when the validation loss plateaued.
Feature maps of ensemble model
In deep learning, feature maps offer an accessible mechanism to illustrate the internal representations that a model learns during training. Feature maps show the output of each convolutional layer as it analyzes an input image, accumulating crucial attributes such as edges, textures, and patterns. In this study, feature maps are recovered from multiple layers of the CNN to explore how the model affects input images at each phase of the network.
To illustrate the feature maps, this study applies a trained CNN model for brain tumor classification. For any input image I∈RH×W×C, the model applies a series of convolutions that produce feature maps at each layer. The output of a convolutional layer Lk for a given filter F is defined as shown in Eq. (32):
where I(i, j) is the input image at position (i, j), F(m, n) is the filter, and YLk(i, j) represents the resulting feature map. These feature maps, also known as activations, are crucial for understanding which parts of the image the model focuses on at each stage of processing.
To analyze these activations, a custom function extracts and visualizes the feature maps from each convolutional layer. For each convolutional layer Lk, this study retrieves the set of feature maps Ak∈RHk×Wk×Dk , where Dk is the number of filters in the layer. Only a subset of filters (4 in this case) is selected for visualization to simplify the analysis.
For convolutional layers, the feature maps are displayed as 2D images using the following procedure:
-
Input Image Processing: The input image is passed through the network, and the activations are computed for each layer.
-
Feature Map Display: For each convolutional layer, the first few filters’ outputs are displayed as grayscale or color maps, providing insights into the regions of the image that are emphasized by the model.
For non-convolutional layers like Flatten and Dense, the output is either plotted as a line graph or a bar plot to show how features are transformed in fully connected layers. By visualizing these feature maps, this study can gain insights into how the network learns to identify critical patterns in brain MRI images, ultimately improving interpretability and explainability29.
Custom loss function
In the brain tumor classification challenge, the class imbalance is handled by applying a custom weighted cross-entropy loss function. The strategy makes sure that more difficult or underrepresented courses receive more attention by giving each class a varying weight according to the adjustable loss. By making up for class imbalances that may otherwise result in biased predictions, this approach boosts classification performance (Fig. 10).
A definition is provided for the categorical cross-entropy loss for a multi-class classification challenge, as shown in Eq. (33):
where yi represents the true label (one-hot encoded) for class i and \(\:\widehat{y}\)i is the predicted probability for class i. While this standard cross-entropy treats all classes equally, it may lead to poor performance in underrepresented classes. To mitigate this, the custom loss function introduces weights to the cross-entropy calculation.
The weighted cross-entropy modifies the standard cross-entropy by incorporating a weight vector w, which assigns different importance to each class. The weighted cross-entropy is given by Eq. (34):
In this implementation, the weight vector w=[1.0,2.0,2.0,3.0] assigns weights of 1.0, 2.0, 2.0, and 3.0 to the four different classes. The higher weight values are assigned to classes that are harder to classify or underrepresented in the dataset. Mathematically, the custom-weighted cross-entropy loss for a single sample can be written as shown in Eq. (35):
The weights are first multiplied element-wise with the true labels yi, and then the cross-entropy loss is scaled by the sum of the weighted labels. This ensures that the loss function gives more importance to misclassifications in the higher-weighted classes. The total loss across the batch is then averaged as shown in Eq. (36):
where m is the batch size and \(\:{L}_{sample}^{k}\) represents the loss for each sample in the batch.
Weighted cross-entropy loss serves as an incentive that guides the model to enhance accuracy rates for weightier classes thus producing balanced predictable results for all categories. The method proves useful in situations where datasets have unbalanced distribution as it frequently occurs within the context of medical image diagnosis30. Algorithm 3 for Custom Weighted Cross-Entropy Loss for Brain Tumor Classification is designed to calculate a weighted cross entropy for multi-class classification with a given weight to be associated with each class. The loss is derived from the standard form of cross-entropy loss modified by using weight factors that bring focus to some classes during learning. This approach assists in addressing class imbalance issues and enhancing the performance of the model, especially in under-represented classes.
Custom weighted cross-entropy loss for brain tumor classification.
Custom loss function workflow.
A weighted categorical cross-entropy loss function created specifically for balancing class distribution appeared in the framework for dealing with brain tumor dataset skew. A specific weight assignment mechanism within the model prevents training from ignoring minority classes during model training. The weights applied during training consisted of [1.0,2.0,2.0,3.0], which matched the classification categories starting with No Tumor then, followed by Glioma and Meningioma before ending with Pituitary.
At first, the weights established their value by applying inverse class frequency calculations from training data to emphasize infrequent classes in the loss function calculations. Between them, the pituitary class received higher weight values than the no tumor class, and the glioma class received lower weight values.
The procedure was followed by empirical weight adjustment by testing various near-weight combinations that allowed for the examination of validation accuracy with per-class precision/recall validation metrics. A configuration of [1.0,2.0,2.0,3.0] delivered maximum training stability together with optimal results in classification for all tumor categories.
Through this custom loss function, the predictive model acquired enhanced capability to detect fewer prevalent tumor types, which significantly improved its ensemble classifier robustness and fairness.
Entropy-based uncertainty estimation
For each model prediction, this study computed entropy as a measure of prediction uncertainty. Although the entropy values were not used to modify training or inference outcomes, study analyzed them post hoc to assess their potential as clinical triage signals. Specifically, study examined predictions exceeding an empirically determined entropy threshold (based on validation set distribution) as candidate low-confidence cases, envisioned for secondary review in a clinical workflow.
Explainability techniques
Multiple XAI approaches help this research demonstrate and evaluate the ensemble model’s decision-making behavior for brain tumor classification. Deep learning models provide outstanding predicted results but their complex multi-layer structure makes them known as “black boxes” in practice. XAI techniques enable better understanding between model performance and interpretability thus assisting doctors to accept model-generated predictions. The study utilizes five significant XAI techniques including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM and LIME for its investigations. The predictive process of the model varies with each distinct technique31.
Grad-CAM (gradient-weighted class activation mapping)
Grad-CAM stands as the most popular explanation method which identifies important image areas for model decisions. Using Grad-CAM produces a significance heatmap from the input image to identify important image regions while computing gradient values of the projected class score relative to the final convolution layer feature maps. The software uses this method to highlight specific MRI image areas which the model focuses on for different tumor identifications.
For an input image I, Grad-CAM computes a localization map LCGrad-CAM for class c as shown in Eq. (37):
Where Ak represents the feature maps, αkc are the gradients of the score for class c, and ReLU is used to focus on positive values, highlighting areas that contribute positively to the classification decision32.
SHapley additive explanations (SHAP)
The game-theoretic method SHAP gives each feature its important values through a heatmap overview of how all features affect the model outcome. The accuracy of results stands as a crucial convention since it depends on the features employed in classification models thus helping to interpret the importance of individual features throughout the classification steps. The computed SHAP values measure how prediction score values change between including and excluding a particular dataset feature.
The SHAP value for a feature i is given by Eq. (38):
Where S is a subset of all features F, and f is the model’s prediction function. In this study, SHAP helps us identify which image features (such as pixel intensity or specific regions) contribute most to the model’s predictions of various tumor types33.
SmoothGrad
SmoothGrad improves upon traditional gradient-based explanations by adding random noise to the input image and averaging the resulting gradient maps. This process smooths out noisy explanations and highlights consistent patterns across multiple noisy versions of the input. In mathematical terms, SmoothGrad modifies the input x by adding Gaussian noise N(0,σ2) and computes the average gradients over multiple samples as shown in Eq. (39):
Where \(\:{\stackrel{\sim}{x}}_{i}\) represents the noisy samples of the input image. SmoothGrad provides more refined and stable visualizations, helping us to interpret better the areas of the image that consistently influence the model’s decision34.
Guided Grad-CAM
The combination of Grad-CAM and guided backpropagation forms Grad-CAM. The areas of an image that prove useful for prediction become Grad-CAM’s main focus while guided backpropagation enhances these results through layer restriction on positive gradients. The approach leads to detailed visualization of image pixels which have maximum impact on the model.
The relevance score Rj for each pixel j is computed as shown in Eq. (40):
This approach provides a more fine-grained explanation of the model’s focus areas, making it easier to validate whether the model is relying on clinically relevant features in the MRI scans35,36.
LIME
LIME is a method to interpret black-box models by fitting a linear model locally in the space around the prediction. LIME modifies a small portion of the input and analyzes how the output changes; after doing so, a simpler, easier-to-understand model (for example, a linear model) is trained to mimic the ML model in the locally neighboring area of the modified ML.
The explanation model g is selected to minimize:
Where L(f, g, πx) measures the fidelity between the complex model f and the interpretable model g in the local neighborhood of the instance x, and Ω(g) is a regularization term to ensure simplicity37,38.
Experimental results
This section presents a detailed analysis of the proposed ensemble model for brain tumor classification. The model’s performance is evaluated using training, validation and testing on the same dataset, as well as on a different data to assess generazibility. During training, the dynamic weighting mechanisms are employed to optimize the contribution of CNN and ResNet-50 and EfficientNet-B5 by adaptively adjusting their weights. A weighted cross-entropy loss function is utilized to address class imbalance and ensure balanced detection across tumor types.
Model performance is quantitatively assessed using standard evaluation metrics, including accuracy, precision, recall, and F1 score. Additionally, the interpretability of the model is enhanced using XAI techniques such as Grad-CAM, SHAP, SmoothGrad, and LIME. These methods provide visual and feature-level insights into the model’s decision-making process.
An explanation of both model learning and network feature detection follows the presentation of the convolutional layer feature maps. The proposed model achieves higher accuracy and interpretation abilities compared to existing approaches when used for brain tumor diagnosis according to state-of-the-art analyses. The level of model confidence in making predictions is simulated by performing uncertainty analysis to strengthen critical medical diagnoses.
Training and validation results
The proposed model was trained over 20 epochs. At the end of the first epoch, it achieved an initial accuracy of 75%. Throughout training, precision and recall consistently remained above 98% across most epochs. All key performance metrics including accuracy, recall, and F1 score showed a steady improvement on both the training and validation datasets.
At the end of training during epoch 20, the model established paramount performance and attained the 99.11% accuracy, closely matched with the 99% validation accuracy, indicating strong generalization capability. The significant reduction in validation loss to 0.3766 guarantees no overfitting during the learning phase. As illustrated in Fig. 11, the model maintained robust performance across multiple evaluation parameters, confirming its effectiveness in brain tumor classification.
Training and validation results.
Testing results
The proposed model was evaluated using a test set of 2,168 MRI images classified into four categories, including normal, pituitary tumor, meningioma tumor, and glioma tumor. The high success rate of the system possibly lies in the fact that it can sort out various types of brain tumors.
For the ‘Normal’ class, recall, accuracy, F1-score, and recall were all 1.00, demonstrating extraordinary accuracy and almost remarkable genuine negative detection abilities. The model’s precision in identifying this kind of tumor is shown by its 0.98 accuracy rate, 0.99 recall rate, and 0.99 F1-score in the “Glioma Tumor” class. An F1-score of 0.99 was also attained by this class, “Meningioma Tumor,” with 0.98 for recall and 0.99 for accuracy. The model showed exceptional performance in the ‘Pituitary Tumor’ class, with 0.99 accuracy, 1.00 F1-score, and 0.99 recall, as shown in Table 3.
Weighted averages of accuracy, recall, and F1-score of 0.99 for each class indicate that the model performed consistently well across all tumor types and has a remarkable potential for generalization. The provided paradigm demonstrates exceptional accuracy together with well-controlled memory and precision requirements, suggesting its possible use in practical medical scenarios.
Confusion matrix.
A confusion matrix presented in Fig. 12 provides a complete evaluation of the performance of the proposed model on a test set for brain tumor classification. It illustrates the number of accurate and erroneous predictions for four separate classes: basal, glioma, meningioma, and pituitary.
These right predictions occur through the diagonal elements of this matrix which show 305 for normal classes, 626 for Glioma classes, 634 for Pituitary classes and 590 for Meningioma classes. Off-diagonal values relate to misclassifications. One normal image was misidentified by the model as both glioma tumor and pituitary tumor. Four Glioma Tumor images appeared to the model as Meningioma Tumors and it classified one image as a Pituitary Tumor together with the other misclassifications. There are three Meningioma Tumor cases classified as Glioma Tumor and one instance classified as Pituitary Tumor. The model’s solid performance can be observed through its accurate classifications of 305, 626, 634 and 590 samples. The relatively few misclassifications demonstrate the robustness of the model in distinguishing between these classes, as reflected in the overall accuracy of 99%.
Ablation study
Attention block
The attention block added to CNN demonstrates fundamental importance since it enables the network to target crucial imaging areas such as tumor boundary detection and intensity distribution tracking. The attention mechanism accomplishes channel-wise reweighting after global average pooling; thus, it enhances significant features and reduces less important features simultaneously.
The quantitative assessment of this component depends on an ablation study that trained two versions of the CNN model.
Pressing either of these two training configurations will enable the attention block functionality in operations.
(1) with the attention block.
(2) without the attention block.
The research design maintained all other elements of architectural and training design at a constant rate.
The results of the investigations are summarized in Table 4, evaluating the impact of incorporating an attention block into the CNN architecture. The inclusion of the attention mechanism led to notable improvements across all key performance metrics. Specifically, the F1-score increased by 2.3%, highlighting the attention block’s effectiveness in enhancing model performance, particularly in scenarios affected by class imbalance and image noise. These results confirm that the attention mechanism significantly contributes to more accurate tumor detection and classification.
Dynamic weights
The research implemented a weighting system that enabled the efficient combination of prediction capabilities of three models: EfficientNet-B5, ResNet-50, and the custom CNN. Initially, the ensemble assigns weights of 0.7 to EfficientNet-B5, 0.2 to ResNet-50, and 0.1 to the custom CNN. These weights are selected based on the architectural strengths of each model, with EfficientNet-B5 receiving the highest initial weight due to its superior feature extraction capabilities. To improve ensemble performance and avoid over-reliance on a single architecture, the model dynamically adjusts these weights during training. Specifically, a linear adjustment is applied at each epoch, modifying the weights by a factor of 0.05. Over time, this strategy increases the influence of ResNet-50 and the custom CNN, allowing the ensemble to leverage their complementary feature representations. This controlled redistribution enhances model diversity and mitigates early-stage overfitting caused by the dominance of EfficientNet-B5.
The defined boundary constraints stabilize the system but preserve its ability to adjust itself over prolonged periods. We conducted empirical checks using an ablation study that evaluated various weight initializations and weight adjustment protocols. The validation accuracy reached its peak when the proposed setting of (0.7, 0.2, 0.1 with 0.05 adjustment rate) was implemented for guiding ensemble training as summarized in Table 5.
Computational efficiency
Table 6 has a complete summary of the computational efficiency and major properties of the proposed ensemble brain tumor classification model. In total, there are 62,995,651 parameters in the model, of which 42,643,580 are trainable, and 20,352,071 are not trainable — pre-trained layers in some of the base models were used. Using an ensemble, the prediction accuracy is enhanced through the integration of three base models with dynamic weighting and summation layers. The model was specified with an input shape of (224, 224, 3), meaning that it takes color images of size 224 × 224 pixels with four classes of output. It uses approximately 240.31 MB of memory usage, as indicated by the model, due to its complexity as well as a need for resources. With respect to training performance, the model took 7,637 s over 20 epochs, with an average step time per batch of 0.3577 s. In addition, inference time was efficient, with the validation taking 69 s and the final test evaluation 31 s. The model behaves as having a high parameter count and high accuracy, with reasonable computation time, all of which is indicative of suitability for clinical applications. In general, the detailed metrics verify the model’s efficiency and strength, placing it as a very good fit for real-time deployment of brain tumor classification tasks.
The model demonstrates efficient computational performance. During training, the average step is approximately 0.3577 s. Validation is completed in 69 s, while testing takes only 31 s when executed on standard hardware. Memory requirements of around 240 MB are reasonable for modern clinical-grade systems, even without applying model compression techniques.
Given the critical importance of achieving high diagnostic accuracy in brain tumor detection, the model’s slightly increased complexity is justified to maximize clinical reliability and minimize diagnostic errors.
Cross-validation results
The proposed ensemble model obtained generalization ability and robustness through a 5-fold cross-validation assessment. Random shuffling, along with dataset stratification, occurred to preserve a balanced distribution of classes for every fold. The evaluation method measured each fold individually through independent training and testing until it calculated accuracy, sensitivity, specificity, precision, recall, F1-score, and AUC metrics for each split. The results appear in Table 7. The evaluation over 5 validation folds proves that the model delivers homogeneous and robust classification outcomes for every partition. The analysis includes these average values obtained from evaluating five cross-validation splits.
The model achieved strong performance metrics during evaluation, with an accuracy of 0.9906, sensitivity of 0.9848, and specificity of 0.9894. Additionally, it attained a precision of 0.9802, recall of 0.9837, an F1-score of 0.9831, and an area under the ROC curve (AUC) of 0.9914, indicating high diagnostic reliability and balanced classification performance. The model demonstrates robust performance through its ability to yield similar evaluation results across all validation folds, which confirms that it does not apply learnings only to specific data sections. The combination of ReduceLROnPlateau with data augmentation techniques, which included random rotation followed by horizontal/vertical flipping, zooming, and shifting, minimized overfitting in our model. Such strategies present the model with different visual patterns, which enhance its capability to handle unknown data. The data pipeline received a complete examination to avoid leakage incidents. The training validation and test splits contained no instances that matched with other data sections. Each data fold operated autonomously to defend against the contamination of evaluation results.
The analyzing process accounted for dataset bias possibilities. To minimize effects from a specific data distribution, we randomized and balanced all classes when the dataset was derived from a single source. The upcoming research will tackle this issue by including data from multiple sources. The evaluation results prove that the model achieves its performance levels because of its strong and regularized training methodology rather than because of bias, leakage, or overfitting effects.
Cross dataset validation results
The research employs a total dataset consisting of 13,351 MRI images from brain tumors fractured across four distinct classes: including Glioma and Meningioma and Pituitary Tumor as well as No Tumor images. The information combines publicly available resources that contain three different datasets.
-
Brain tumor dataset provides its MRI image collection through figshare to researchers for classification purposes.
-
The brain tumor classification MRI dataset obtained from Kaggle adds diverse image data by including various MRI scan variations in terms of size and direction.
-
Kaggle provides the brain tumor MRI dataset, which brings more high-quality tagged images to the collection.
Additionally, the dataset includes 4,204 images from the Crystal Clean: Brain Tumors MRI Dataset, which enhances its diversity and generalization. Each dataset is vast and well-balanced for trustworthy brain tumor classification because it includes MRI scans of different tumor forms in addition to images devoid of tumors. By merging these datasets, which give a variety of tumor characteristics, orientations, and image quality, deep learning models should be trained and evaluated more effectively. Based on the cross-dataset validation classification result, the model successfully diagnoses all four forms of brain tumors: pituitary, glioma, meningioma, and normal. With an overall accuracy rate of 99%, the model exhibited a remarkable aptitude for generalization on this heterogeneous dataset of 17,555 MRI images39,40. Precision, recall, and F1-score for the Normal and Pituitary Tumor classes reveal almost perfect values of 1.00; still, accuracy, memory, and F1-score are constantly excellent for each class. Using a recall of 1.00, an accuracy of 0.98, and an F1-score of 0.99, the tumor was certified to be a glioma. For meningioma tumors, the equivalent F1-score, accuracy, and recall were 0.99, 1.00, and 0.98 as presented in Table 8. As these data suggest, the system successfully recognizes images as either tumors or non-tumors with low rates of false positives and false negatives. A great cross-dataset validation performance demonstrates how long the model will endure in actual situations.
Cross dataset validation confusion matrix.
The confusion matrix in Fig. 13. demonstrates how well the model performs in classification across all four classes (Normal, Glioma, Meningioma, and Pituitary). Out of 2,484 normal images, the algorithm successfully predicted 2,467 with only 17 erroneous classifications. The program correctly diagnosed 5,092 out of 5,109 images associated with glioma tumors with fairly low mistakes.
With 82 erroneous classifications out of 5,177 images, Meningioma tumor had substantially more difficulties, showing that the accuracy of the model for this class is slightly worse than that of the other classes. Nevertheless, it fared well overall. Ultimately, the model demonstrated remarkable recall and accuracy, appropriately predicting 4,777 out of 4,785 cases of pituitary tumors. The confusion matrix illustrates the robustness of the approach, notably for Glioma and Pituitary Tumors, while also suggesting chances for future improvement in the classification of Meningioma. The few misclassifications show that the model generalizes effectively across diverse datasets and tumor types, displaying good performance even in cross-dataset scenarios.
Feature maps of ensemble model
The feature maps provided in Fig. 14 give a thorough view of how the model analyzes input MRI images of brain tumors across multiple levels. At the input layer, the images retain most of their original structure, highlighting the critical locations where the model starts processing. As the input proceeds through succeeding convolutional layers (Conv2D and MaxPooling layers), the feature maps get more abstract, concentrating on distinct portions of the image. In the early convolutional layers, this study observes obvious traces of the brain structures, showing the model’s attention on edges and significant features.
As the data proceeds to deeper layers like Conv2D_2 and Conv2D_3, the feature maps get increasingly pixelated and less interpretable to the human eye. These latter layers are responsible for spotting more complicated patterns that are not always visually intuitive but are crucial for categorization jobs. MaxPooling layers further lower the spatial resolution, conserving just the most relevant information. This approach highlights the model’s capacity to progressively distill vital information, concentrating on key portions of the image critical for tumor identification while abstracting away irrelevant features.
The activation values from the flatten layer are depicted in Fig. 15. The previous convolutional and pooling layers convert their multidimensional feature maps into one-dimensional vectors at this stage. Having thousands of activation values demonstrates that the layer maintains spatial information before moving it onto fully connected layers. The wide range of activation values shows that the model detects many distinct features that will later be used for classification. The Dense layer activation values appear in Fig. 16. The procedural part of this fully connected layer uses Flatten output while learning decision-making patterns for classification. The activation value spikes indicate specific neurons intensely respond to features extracted during previous network operations. The network processes the activations through additional layers to enhance the classification decision. Figure 17 shows the activation values for the Dropout layer, which is applied after the Dense layer to prevent overfitting. During training the Dropout mechanism chooses randomly to disable specific neurons because it improves generalization capabilities. Currently the Dense layer provides activations which will not be propagated through each cyclic process. The activation plot reveals spikes which indicate which neurons become active in that specific moment hence enabling the network to acquire more dependable characteristics. The network information spread is depicted through these graphs which also show why multiple layers help the system identify distinct brain tumor patterns. The research provides comprehensive understanding about model activity during training and evaluation sessions to assist in improving and solving models.
Layer-wise feature maps.
Activation values for the flatten layer.
Activation values for dense layer.
Activation values for the dropout layer.
Explainable AI (XAI) results
This study uses Grad-CAM brain MRI heatmaps to check model accuracy in visualizing tumor areas of importance. The Fig. 18(a) shows the original image. Figure 18(b) displays red and blue regions that relate to areas where the model determined the importance for its classification work. The model selects the tumor region in brain’s central area which proves its ability to detect essential tumor features within the image. The visual depiction of tumor-affected areas through this graphic indicates that the model’s decisions match medical assumptions and remain comprehensible.
The SHAP result presented in Fig. 19 demonstrates which image regions play a vital role in model classification. Model prediction contributors and detractors are shown with SHAP values in the form of red and blue colored areas. Regarded tumor border regions in this particular case carried higher SHAP values because they had a profound effect on classification outcomes. The SHAP values show that areas with lower predictions exist past the tumor region since they have negative values. The approach provides complete evidence regarding how model attributes behave and their contributions to the analysis.
The SmoothGrad result, which averages gradients across numerous noisy repetitions of the input image, provides the most trustworthy characteristics that influence the model’s prediction. The sites that contributed more often and considerably to the categorization decision are represented by the zones with brighter brightness. The highlighted area adjacent to the center of Fig. 20 illustrates that the model has concentrated on the tumor location. This technique promotes interpretability in the context of brain tumor classification by decreasing noise in the traditional gradient-based explanation and delivering a better representation of the model’s attention.
The technique outlined in this work, called Guided Grad-CAM, merges the benefits of guided backpropagation and Grad-CAM to give a full image of the locations leading to the model’s prediction. The overlay in Fig. 21 shows the specific locations in the brain MRI image that the model is concentrating on, particularly the area surrounding the tumor. The areas receiving the greatest focal point from the model are represented by patches of green and blue, and the overlay demonstrates how these areas impact the decision class. This method provides a more complete and accurate explanation for why a model is seeing what it sees to ensure that the model’s focal point is clinically aligned. In this study, LIME is used to identify the decision boundaries of the ensemble model as further evidence of the ways these different features drive one classification decision to the next. In Fig. 22, using LIME, this study provides an interpretable explanation for the model’s prediction of class 2. The red and green markings highlight which parts of the MRI image contributed most to the classification decision. Certain red sections are damaging to the prediction, while the green sections pose a positive increase to the prediction. As a byproduct of perturbing the original image and then testing how those perturbed images change the model’s decision, LIME enables, even approximates identification of the decision boundary by providing explanations of why this model predicted that class. This kind of openness gives physicians the ability to evaluate whether the model’s prediction is consistent or reasonable with medical expertise.
Original image (a) and Grad-CAM XAI heatmap (b).
SHAP XAI heatmap.
SmoothGrad XAI heatmap.
Guided Grad-CAM XAI heatmap.
LIMA XAI heatmap.
The combination of XAI techniques Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM, and LIME enables researchers to obtain essential information about how the ensemble model detects brain tumors. The prediction areas and properties in MRI images that the model relies on received useful analysis from these diagnostic tools. Model accuracy and interpretability were maintained through these solutions as they produced believable results. The solutions together with domain expertise allowed healthcare practitioners to understand the decision-making processes of the sog systems. The model achieves greater confidence regarding medical diagnostics through the incorporation of this approach which adds practicality to its functionality.
The integration of Grad-CAM with SHAP, SmoothGrad and LIME generates clear model interpretations through various approaches where Grad-CAM determines important scan regions which affected the model’s output decisions. Each pixel’s contribution levels to the classification result can be quantified through SHAP for more specific feature study. SmoothGrad improves gradient-based explanations through the process of gradient noise averaging which produces more clear and less noisy visual results. Each prediction from LIME receives surrounding explanation using a simpler interpretable model as a way to validate important regions. The approaches were used to study challenging misclassified images to uncover faint tumor areas that human readers may miss.
Uncertainty (entropy) results
The proposed model generates an uncertainty score through entropy assessment to determine brain tumor classifications at 0.3093. Entropy evaluates the randomness in predicted probability distributions through its values: low entropy values indicate strong prediction certainty, but high entropy values reveal uncertain outcomes. The measured entropy score stands at a low level compared to standard benchmarks, indicating precise predictions coming from the model.
The distribution of cases with entropy values greater than 0.5 is shown in Fig. 23 to study uncertain prediction behavior. The deployment process should implement automated safeguards for performing secondary verification on high-entropy prediction results before they become final approved results. The system’s resistance to errors improves through this strategy since uncertain outputs receive special monitoring that guards against classification risks.
Examples of low-confidence prediction.
State-of-the-art comparison
Research findings presented in Table 9 direct this state-of-the-art analysis, which evaluates various studies about XAI utilization in medical image analysis, especially brain tumor diagnosis. In14, the authors developed another CNN model for brain tumor detection by integrating numGrad-CAM as its explanation method. The model achieved 97.11% accuracy, together with 95.58% sensitivity and 96.81% specificity. Study results compared explainability against previous methods using CAM techniques but the latest technique contained only numGrad-CAM which enabled clinicians to understand model reasoning through 90.11% accurate sensitivity. However, by adding a custom loss function and dynamic weighting, the ensemble model reported in this study a mix of ResNet-50, EfficientNet-B5, and a custom CNN achieved a superior accuracy of 99.4% and 99.31% in cross-dataset validation. In41, authors designed the NeuroXAI framework to increase model transparency for medical practitioners adopting Grad-CAM, SHAP, and other XAI technologies, with a 98.62% accuracy rate. While NeuroXAI presents a wide range of interpretability, the proposed ensemble model trumps NeuroXAI in terms of generalization and accuracy and yields equal explainability techniques when cross-dataset validation is employed. After studying the treatment effectiveness of AI for glioma grading, the research in42 ultimately attained an accuracy of 88% with AI help. While AI helps enhance physicians’ understanding, the study suggested that applying SmoothGrad to explain explanations did not appreciably increase accuracy. On the other hand, a greater variety of XAI techniques and enhanced accuracy are indicated by the proposed ensemble model, delivering more helpful clinical decision assistance.
Unlike existing methods, our approach introduces a dynamic ensemble framework with adaptive weighting, which optimally combines multiple base models during training. Additionally, we developed a custom loss function specifically designed to handle class imbalance, improving the classification of minority classes. Our work also integrates multiple XAI techniques, including Grad-CAM, SHAP, SmoothGrad, and LIME, offering comprehensive interpretability to enhance clinical decision-making. Furthermore, cross-dataset validation was performed to demonstrate the model’s robustness across different datasets, and an entropy-based confidence score was incorporated to assess the certainty of predictions. These contributions collectively enhance the performance, generalizability, and explainability of our model, distinguishing it from existing approaches.
Results and discussion
This section presents a comprehensive analysis of the proposed model’s performance, including evaluation metrics, generalization strategies, explainability insights, and clinical relevance. We also discuss comparative benchmarks, ablation studies, and potential limitations, offering a balanced perspective on the system’s strengths and areas for future improvement.
Overall model performance
The performance of the proposed model was evaluated using standard metrics including accuracy, precision, recall, and F1-score, all of which indicated excellent outcomes. The model achieved a test accuracy of 99.4% and demonstrated strong generalization capability, achieving 99.31% accuracy in cross-dataset validation. To ensure the reliability of these results, rigorous safeguards were implemented to avoid overfitting or data leakage. Stratified sampling was employed to maintain class balance across training, validation, and testing splits. Data generators were initialized independently to prevent overlap or contamination, and the dataset was preprocessed through a robust pipeline ensuring no images were shared between partitions.
Strategies for generalization and robustness
To enhance generalization, the training process incorporated multiple regularization techniques, including extensive data augmentation, dropout layers, L2 weight regularization, early stopping, and dynamic learning rate scheduling (ReduceLROnPlateau). Importantly, the implementation of dynamic weights alongside a custom weighted cross-entropy loss function enabled the model to maintain balanced sensitivity toward underrepresented tumor classes, significantly improving classification performance. This configuration provided flexibility and allowed the ensemble to adapt throughout training—capitalizing on the strengths of EfficientNet-B5 during early epochs and progressively shifting emphasis to ResNet-50 and the custom CNN as training advanced. Moreover, the use of entropy-based uncertainty quantification further strengthened model reliability, yielding an average entropy score of 0.3093 across predictions. This metric was critical in identifying low-confidence cases, especially valuable in clinical contexts demanding high diagnostic safety.
Cross-validation and benchmarking
We acknowledge that the achieved performance is exceptionally high; however, its validity is supported by consistent outcomes across 5-fold cross-validation, internal test sets, and external validation datasets. Furthermore, our results align with or outperform those reported in recent literature on brain tumor classification using deep learning, where accuracies typically range between 97 and 99% on similar datasets. These comparisons confirm the competitive standing of our proposed system and highlight its robustness and reliability across partitions.
Model explainability and clinical trust
The integration of explainable AI (XAI) techniques, including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM, and LIME, enriched our understanding of model decisions. These techniques allowed visualization of critical MRI image regions contributing most to class-specific predictions, often aligning well with radiological features familiar to clinicians. Such transparency is crucial for ensuring that model outputs are interpretable and can be trusted by healthcare professionals.
Clinical relevance and risk mitigation
Automated brain tumor classification systems must address both performance and reliability under uncertainty. In real-world use, misdiagnoses—whether false positives or false negatives—carry serious consequences. False positives may lead to unnecessary emotional and financial stress, while false negatives could delay essential treatment. To mitigate such risks, we propose a confidence-aware safeguard mechanism integrated into the inference workflow. Predictions with high entropy values, signaling low confidence, can trigger additional review protocols, such as ensemble consensus verification or cross-model consistency checks. Envision the entropy-based low-confidence flagging as a triage mechanism, where predictions exceeding a predefined entropy threshold would be automatically highlighted within the clinical interface. These flagged cases could then be routed for secondary review by a senior radiologist or placed into a dedicated review queue within the hospital’s Picture Archiving and Communication System (PACS). This workflow aims to enhance diagnostic safety by ensuring that uncertain predictions receive appropriate human oversight. We also acknowledge that future work should include prospective user-testing with clinical teams to refine the practical deployment, integration, and usability of this safeguard mechanism in real-world clinical environments. Moreover, recording entropy values and prediction probabilities supports transparent auditing and enhances accountability. In future work, we aim to incorporate calibration techniques like temperature scaling and structured clinician feedback to further strengthen diagnostic safety.
Dynamic weighting limitation and future directions
While the dynamic weighting strategy was intentionally designed to assign higher initial weight to EfficientNet-B5 — leveraging its superior early-stage feature extraction — and progressively increase the influence of ResNet-50 and the custom CNN, we acknowledge an important limitation. The current approach relies on a fixed epoch-based adjustment schedule, which may not fully capture performance dynamics if sub-models improve or deteriorate at later stages of training. To address this, future work could explore adaptive, performance-driven weighting mechanisms, such as gradient-based dynamic weight updates, reinforcement learning controllers, or meta-learning strategies. These adaptive approaches would allow the ensemble to flexibly adjust the contribution of each sub-model in real time, based on ongoing performance metrics, rather than relying solely on a predefined adjustment scheme.
Limitations
While the outcomes of this research are promising, several limitations are acknowledged: The model has not yet been tested on real-time clinical patient data, and deployment will require further validation in such environments. The training process demands significant computational resources due to the model’s complexity and ensemble architecture, which may limit adoption in resource-constrained clinical settings. While the model size (~ 63 million parameters) and memory footprint (~ 240 MB) are reasonable for high-performance computing servers, we acknowledge that they may pose challenges for deployment on edge devices or in mobile health units. As a future direction, we suggest exploring model compression strategies such as pruning, quantization, and knowledge distillation to significantly reduce model size and computational load, thereby facilitating real-time inference and broader applicability in low-resource settings. Although the custom loss function and dynamic weight scheme provide notable improvements, additional enhancements may be needed to address highly unbalanced or rare tumor classes in larger and more diverse datasets.
Specifically, for the weighted loss function, the current study used initial class weights based on inverse class frequency, followed by empirical fine-tuning through iterative adjustments (in 0.5–1.0 steps) while monitoring validation precision and recall, with particular focus on underrepresented classes such as Pituitary tumors. While this pragmatic approach improved balance, we acknowledge that more systematic optimization methods—such as grid search, random search, or Bayesian optimization—could be explored in future work to identify optimal class weight configurations more rigorously and potentially enhance overall model performance and fairness.
While the integration of multiple explainable AI (XAI) techniques, including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM, and LIME, enriched the interpretability of model predictions, we acknowledge that this study did not include formal or informal clinical evaluation of these visualizations. Future work will involve systematic collaboration with clinical experts, such as radiologists or neuro-oncologists, to qualitatively and quantitatively assess the medical relevance of the highlighted regions. This will ensure that the explanations provided by the model align with clinically meaningful features and can be reliably used to support diagnostic decision-making in real-world settings.
Conclusion
The proposed method develops an ensemble deep learning system which uses EfficientNet-B5 along with ResNet-50, together with a newly developed CNN network to classify brain tumors. A dynamic weight assignment strategy forms the core innovation of this approach because it enables training models to change their contributions to achieve a more flexible ensemble model, which selects the best data at different learning stages. A weighted cross-entropy loss function was created to tackle the class imbalance issue together with its capability to improve tumor classification accuracy. The model achieved a successful accuracy rate of 99%. The precision measure reached 4% during the test while recall and F1-score values were found to be constantly significant across every class category. The analysis of combined MRI data from numerous healthcare institutions demonstrated 99.31% accuracy when testing data consisting of 17,555 images from various sources. The model exhibits broad applicability in real-world clinical practice because results from these tests demonstrate their general usefulness. The implementation of XAI techniques including Grad-CAM, SHAP, SmoothGrad, Guided Grad-CAM and LIME enabled interpretation along with transparency. The identification methods based on XAI techniques determined which parts of the MRI scans the model relied on for classification, thus delivering information that could assist clinicians when they validate prediction results. The incorporation of uncertainty entropy improves model reliability since it tracks prediction confidence but also marks unforeseen situations that need attention with further investigation.
Data availability
The datasets used in this study are publically available at the Kaggle repository: https://www.kaggle.com/datasets/sabersakin/brainmri; https://www.kaggle.com/datasets/mohammadhossein77/brain-tumors-dataset; https://www.kaggle.com/datasets/masoudnickparvar/brain-tumor-mri-dataset; https://github.com/Retinderdeepsingh/Brain-tumor-classification/blob/main/ensemble-model.ipynb.
References
Price, M. et al. Childhood, adolescent, and adult primary brain and central nervous system tumor statistics for practicing healthcare providers in neuro-oncology, CBTRUS 2015–2019. Neurooncol. Pract. 11 (1), 5–25 (2024).
Fernando, K. R. M. & Tsokos, C. P. Deep and statistical learning in biomedical imaging: state of the Art in 3D MRI brain tumor segmentation. Inf. Fusion. 92, 450–465 (2023).
Srinivasan, S. et al. Multimodal biomedical image segmentation using Multi-Dimensional U-CNN. BMC Med. Imaging. 24 (1), 38 (2024).
Rahman, M. A. et al. GliomaCNN: an effective lightweight CNN model in assessment of classifying brain tumor from magnetic resonance images using explainable AI. Comput. Model. Eng. Sci. 140 (3), 2425–2448 (2024).
Li, C. et al. U-KAN makes strong backbone for medical image segmentation and generation. Proc. Conf. AAAI Artif. Intell.140, 4652–4660 (2025).
Padmapriya, S. T. & Devi, M. S. G. Computer-aided diagnostic system for brain tumor classification using explainable AI. In 2024 IEEE International Conference on Interdisciplinary Approaches in Technology and Management for Social Innovation (IATMSI) (2024).
Noreen, N., Palaniappan, S., Qayyum, A., Ahmad, I. & Alassafi, M. O. Brain tumor classification based on fine-tuned models and the ensemble method. Comput. Mater. Contin. 67 (3), 3967–3982 (2021).
Chathumina, J. & Jayakumar, K. Enhancing brain tumor diagnosis: A comparative review of systems with and without eXplainable AI. In 2024 5th Information Communication Technologies Conference (ICTC), vol. 14, 309–313 (2024).
Shamshad, N. et al. Enhancing brain tumor classification by a comprehensive study on transfer learning techniques and model efficiency using MRI datasets. IEEE Access 12, 100407–100418. https://doi.org/10.1109/ACCESS.2024.3430109 (2024).
Singh, R. et al. Ensemble deep learning models for enhanced brain tumor classification by leveraging ResNet50 and EfficientNet-B7 on high-resolution MRI images. IEEE Access 12, 178623–178641. https://doi.org/10.1109/ACCESS.2024.3494232 (2024).
Latif, G., Iskandar, D. N. F. A. & Alghazo, J. Multiclass brain tumor classification using region growing based tumor segmentation and ensemble wavelet features. In Proceedings of the International Conference on Computing and Big Data, 2018 (2018).
Shafi, A. S. M., Rahman, M. B., Anwar, T., Halder, R. S. & Kays, H. M. E. Classification of brain tumors and auto-immune disease using ensemble learning. Inf. Med. Unlocked. 24 (100608), 100608 (2021).
Aurna, N. F., Yousuf, M. A., Taher, K. A., Azad, A. K. M. & Moni, M. A. A classification of MRI brain tumor based on two stage feature level ensemble of deep CNN models. Comput. Biol. Med. 146 (105539), 105539 (2022).
Marmolejo-Saucedo, J. A. & Kose, U. Numerical grad-cam based explainable CNN for brain tumor diagnosis. Mob. Netw. Appl. (2022).
Schiavon, D. E. B., Becker, C. D. L., Botelho, V. R. & Pianoski, T. A. Interpreting convolutional neural networks for brain tumor classification: an explainable artificial intelligence approach. In Intelligent Systems 77–91 (Springer, 2023).
Prince, E. W., Ghosh, D., Görg, C. & Hankinson, T. C. Uncertainty-aware deep learning classification of adamantinomatous craniopharyngioma from preoperative MRI. Diagnostics (Basel) 13, 6 (2023).
Taşcı, B. Attention deep feature extraction from brain MRIs in explainable mode: DGXAINet. Diagnostics (Basel) 13, 5 (2023).
Hashemi, S. M. H. Crystal clean: brain tumors MRI dataset. Kaggle (2023).
Islam, M. M., Talukder, M. A., Uddin, M. A., Akhter, A. & Khalid, M. BrainNet: Precision brain tumor classification with optimized EfficientNet architecture. Int. J. Intell. Syst. 2024, 1 (2024).
Ullah, N., Khan, J. A., Falco, D. & Sannino, I. Explainable artificial intelligence: Importance, use domains, stages, output shapes, and challenges .ACM Comput. Surv. 57, 1–36 (2024).
Sharma, A. K. et al. Brain tumor classification using the modified ResNet50 model based on transfer learning. Biomed. Signal. Process. Control. 86 (105299), 105299 (2023).
Zheng, S. et al. Asymmetric adaptive heterogeneous network for multi-modality medical image segmentation. IEEE Trans. Med. Imaging. 44 (4), 1836–1852 (2025).
Zahoor, M. M. et al. Brain tumor MRI classification using a novel deep residual and regional CNN. Biomedicines 12 (7), 1395 (2024).
Raza, A. et al. A hybrid deep Learning-Based approach for brain tumor classification. Electronics 11 (7), 1146. https://doi.org/10.3390/electronics11071146 (2022).
Zhang, Z. et al. A novel deep learning model for medical image segmentation with convolutional neural network and transformer. Interdiscip Sci. 15 (4), 663–677 (2023).
Ullah, N., Khan, M. S., Khan, J. A., Choi, A. & Anwar, M. S. A robust end-to-end deep learning-based approach for effective and reliable BTD using MR images. Sens. (Basel). 22 (19), 7575 (2022).
Xiao, G. et al. Contrastive learning with dynamic weighting and Jigsaw augmentation for brain tumor classification in mri. Neural Process. Lett. 55 (4), 3733–3761 (2023).
Srinivasan, S. et al. A hybrid deep CNN model for brain tumor image multi-classification. BMC Med. Imaging. 24 (1), 21 (2024).
Francisco, P. D., Antuono, K., Robbelein, N., Noppe, W. & Weijtjens, C. Long-term fatigue Estimation on offshore wind turbines interface loads through loss function physics-guided learning of neural networks. Renew. Energy. 205, 461–474 (2023).
de Vries, B. M. et al. Explainable artificial intelligence (XAI) in radiology and nuclear medicine: a literature review. Front. Med. (Lausanne) 10, 1180773 (2023).
Özbay, F. A. & Özbay, E. Brain tumor detection with mRMR-based multimodal fusion of deep learning from MR images using Grad-CAM. Iran. J. Comput. Sci. 6 (3), 245–259 (2023).
Hossain, S., Chakrabarty, A., Gadekallu, T. R., Alazab, M. & Piran, M. J. Vision transformers, ensemble model, and transfer learning leveraging explainable AI for brain tumor detection and classification. IEEE J. Biomed. Health Inf. 28 (3), 1261–1272 (2024).
Snehal, R. A., Kapdi, M. S. & Raval, M. Interpretable machine learning model to predict survival days of malignant brain tumor patients. Mach. Learn. Sci. Technol. 4, 2 (2023).
Spagnolo, F. et al. Instance-level quantitative saliency in multiple sclerosis lesion segmentation. ArXiv [eess IV] (2024).
Nhlapho, W., Atemkeng, M., Brima, Y. & Ndogmo, J. C. Bridging the gap: exploring interpretability in deep learning models for brain tumor detection and diagnosis from MRI images. Information 15, 4 (2024).
Yu, W. et al. Liver vessels segmentation based on 3d residual U-NET. In 2019 IEEE International Conference on Image Processing (ICIP) (2019).
Zhang, Z. et al. Deep learning and radiomics-based approach to meningioma grading: exploring the potential value of peritumoral edema regions. Phys. Med. Biol. 69, 10 (2024).
Ullah, N., Hassan, M., Khan, J. A., Anwar, M. S. & Aurangzeb, K. Enhancing explainability in brain tumor detection: A novel DeepEBTDNet model with LIME on MRI images. Int. J. Imaging Syst. Technol. 34, 1 (2024).
Sakin, S. Brain Tumor MRI Dataset Merged (2023).
Nickparvar, M. Brain Tumor MRI Dataset (2021).
Zeineldin, R. Deep Multimodality Image-Guided System for Assisting Neurosurgery. Dissertation, Karlsruhe: Karlsruher Institut für Technologie.
Jin, W., Fatehi, M., Guo, R. & Hamarneh, G. Evaluating the clinical utility of artificial intelligence assistance and its explanation on the glioma grading task. Artif. Intell. Med. 148 (102751), 102751 (2024).
Guha, S., Kodipalli, A., Fernandes, S. L. & Dasar, S. Explainable AI for interpretation of ovarian tumor classification using enhanced ResNet50. Diagnostics (Basel). 14 (14), 1567 (2024).
Wang, Y. et al. A gradient mapping guided explainable deep neural network for extracapsular extension identification in 3D head and neck cancer computed tomography images. Med. Phys. 51 (3), 2007–2019 (2024).
Acknowledgements
Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2025R178), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Retinderdeep Singh: Conceptualization, Methodology, Software development, Formal analysis, Writing—original draft preparation.Sheifali Gupta: Data curation, Model training, Experimental setup, Writing—review and editing.Ashraf Osman Ibrahim: Investigation, Data analysis, Visualization, Writing—review and editing.Lubna A. Gabralla: Data curation, Model training, Experimental setup, Writing—review and editing.Salil Bharany: Supervision, Project administration, Validation, Resources, Writing—review and editing.Ateeq Ur Rehman: Investigation, Data analysis, Visualization, Writing—review and editing.Seada Hussen: Conceptualization, Supervision, Resources, Writing—review and editing, Final approval of the version to be submitted.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Singh, R., Gupta, S., Ibrahim, A.O. et al. Advanced dynamic ensemble framework with explainability driven insights for precision brain tumor classification across datasets. Sci Rep 15, 29090 (2025). https://doi.org/10.1038/s41598-025-14917-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14917-w