Abstract
We develop an AI system that pairs engineering problems with biology-inspired solutions at a large scale, by analyzing over 101 million abstracts to identify thematic links between engineering and biology. We detect coherent themes in each domain with transformer-based embeddings and BERTopic, then link them in a topic graph that quantifies their co-occurrence. We use TRIZ (Theory of Inventive Problem Solving) analysis to show how biological principles can overcome specific engineering limitations. By integrating language models, topic modeling, and contradiction analysis, the approach highlights latent thematic overlaps. Our methodology is demonstrated in four distinct case examples—including adhesive mechanisms for robotic climbing and thermal insulation inspired by dental bonding—validating our approach. This systematic approach can accelerate the discovery of new bio-inspired innovations.
Similar content being viewed by others
Introduction
Interdisciplinary research advances science by tackling complex real-world problems through the integration of diverse fields. However, collaboration between engineering and biology is challenging due to epistemological and methodological disparities. Engineering emphasizes quantitative, deterministic methods, while biology focuses on qualitative, adaptive approaches to complex systems1.
Recent advancements in Artificial Intelligence (AI), particularly in Natural Language Processing (NLP), offer promising solutions for bridging these gaps by identifying semantic commonalities across large text corpora. Yet, NLP techniques alone often lack a structured method for interpreting and resolving fundamental contradictions or trade-offs that arise when merging diverse domains.
TRIZ (Theory of Inventive Problem Solving) is a systematic innovation methodology developed by Genrich Altshuller that resolves contradictions through structured approaches, including 40 inventive principles and a contradiction matrix2. This heuristics-based methodology, distilled from over 40,000 patents, frames every engineering problem as a contradiction between two performance attributes and recommends one of 40 inventive principles to resolve it. By systematically targeting inherent contradictions within problem statements, TRIZ addresses gaps in traditional problem-solving approaches. Building upon Altshuller’s foundational work, recent research3 has investigated TRIZ’s application in bioinspired innovation, demonstrating its potential for guiding nature-inspired design solutions.
In TRIZ, a “contradiction” is a pair of performance attributes where improving one traditionally degrades the other (e.g., higher mechanical resistance vs. lower weight). This research systematically aligns engineering and biology articles by integrating TRIZ contradiction formalism with NLP and graph embedding techniques.
At the intersection of engineering and biology lies an active area of innovation, shaped by their contrasting yet complementary epistemologies. Engineering thrives on quantitative rigor, developing precise solutions under well-defined constraints, while biology navigates the emergent complexity of living systems, embracing adaptability and resilience. These differences, often seen as barriers, may be reframed as complementary strengths: biological systems offer solutions to challenges that defy deterministic approaches, while engineering provides the tools to formalize and scale biological insights. Bridging these domains requires not only technological sophistication but also a framework that respects their unique paradigms while facilitating mutual enrichment.
This study addresses the following research question: How can Natural Language Processing techniques and graph embedding, augmented by TRIZ contradiction formalism, effectively pair engineering and biology articles to reveal cross-domain innovation opportunities?
We make three key contributions:
-
1.
We develop a three-stage classification methodology that enables curation of a large engineering–biology corpus (126 k papers) built with the express intent of enabling inference of thematic associations across the two disciplines;
-
2.
We introduce a novel graph-based approach that generates a 544-topic knowledge network with 46,362 weighted cross-links, helping uncover bio-inspired solutions;
-
3.
We demonstrate the practical value of our approach through four detailed case studies that reveal cross-domain innovation opportunities.
Literature review
Previous studies address isolated components of the problem, but none provide an end-to-end pairing pipeline.
Natural Language Processing (NLP) has become a widely used tool for analyzing, processing, and extracting insights from unstructured textual data. Transformer-based models like BERT4 and GPT5 have significantly advanced semantic analysis by capturing contextual nuances in text. These models have been employed in various domains, including biomedical data mining6, patent discovery7, and engineering knowledge analysis8,9. Despite these advancements, the application of NLP to interdisciplinary pairing remains underexplored, particularly for domains with contrasting problem-solving approaches like engineering and biology.
Recent studies use machine learning and topic modeling (e.g., BERTopic) to uncover latent themes, enabling automated detection of cross-domain documents9,10. Generative AI models can further assist in abstracting and summarizing complex knowledge, enabling the creation of high-level ‘knowledge maps’ that guide researchers toward innovative cross-domain solutions.
Recent studies have demonstrated the utility of semantic similarity measures, such as cosine similarity and word embeddings, for linking concepts across fields. For example, researchers in bioinformatics have employed these techniques to predict protein structures11,12. However, such approaches often lack a structured framework for identifying and resolving the inherent contradictions between the problem statements of these domains.
Graph embedding techniques have facilitated the representation of complex relationships within datasets. Algorithms such as node2vec13, DeepWalk14, and GraphSAGE15 encode graph structures into vector spaces, enabling efficient clustering and visualization of semantically related concepts. These methods create knowledge graphs that visualize interdisciplinary research evolution and uncover hidden connections16.
In scientific literature, graph embeddings have been used to map relationships between publications, authors, and keywords, facilitating the discovery of novel research directions. Despite their success, existing implementations primarily focus on single-domain analyses or broad interdisciplinary frameworks. Existing approaches rarely tackle the specific challenge of pairing engineering and biology articles, where thematic overlaps need to reconcile distinct epistemological paradigms. Additionally, these approaches often lack a structured problem-solving framework.
TRIZ, a systematic problem-solving methodology developed by Genrich Altshuller through extensive patent analysis, provides a structured approach to innovation by identifying and resolving the fundamental contradictions within problem statements2. TRIZ tools such as the Contradiction Matrix and 40 Inventive Principles have been widely adopted in engineering and manufacturing to identify creative solutions to technical challenges. Conversely, research on applicability of TRIZ or TRIZ-inspired principles in biology has been limited—except in notable areas such as biomimicry and ecological modeling3,17,18—supporting its role in balancing the synergies and conflicts between engineering and biology.
By transforming problem statements into generalized contradictions, TRIZ enables the identification of complementary challenges across domains. For example, the engineering problem of “increasing material strength without increasing weight” aligns with the biological problem of “enhancing cellular resilience without compromising flexibility.” These generalized formulations pave the way towards a common language for multidisciplinary collaboration.
It is important to note that our application of the TRIZ lens in this framework is conceptual—serving as an organizing principle that highlights potential contradictions and synergies between engineering and biological problem statements. While we do not yet provide quantitative data to validate these pairings, this approach backed by literature paves the way for future studies that will directly leverage TRIZ principles to develop data-driven pairing strategies. Our proposed NLP-based framework also constitutes the first layer towards a broader framework to be augmented by TRIZ, as suggested in prior studies10.
Cross-domain research is essential for addressing complex global challenges, such as climate change, healthcare innovation, and sustainable development. Traditional approaches, such as expert consultations and collaborative workshops, rely heavily on human intuition and expertise, making them resource-intensive and difficult to scale. AI-assisted frameworks provide an approach that scales to large corpora, enabling systematic knowledge integration across domains19.
Despite these advancements, few frameworks explicitly focus on pairing articles from disciplines as distinct as engineering and biology. Existing approaches often overlook the nuanced contradictions that define these fields, limiting their ability to uncover meaningful synergies. This study addresses this gap by integrating NLP, graph embedding, and TRIZ methodologies into a unified framework.
Materials and methods
This research employs a mixed-methods approach, integrating computational tools and theoretical frameworks to systematically pair engineering and biology articles.
We employ a supervised machine learning classifier trained to identify articles that span multiple fields of study20,21. This ensures that we focus on genuinely interdisciplinary documents rather than relying solely on domain metadata. Building on recent frameworks, our approach also leverages topic modeling22 to identify thematic structures. Finally, large language models5,23 can then assist in summarizing and contextualizing these insights, refining the pairings identified through NLP and graph embeddings.
The aim is a maintainable workflow that can be updated regularly. The methodology unfolds in three distinct phases: data collection and preprocessing, model creation, and evaluation.
The study begins with data collection from Semantic Scholar, which encompasses reputable scientific databases, including IEEE Xplore, PubMed, and Scopus, among others24. Our initial dataset dated 2023-08-01 comprises approximately 101 million articles of all disciplines. We employed the Semantic Scholar datasets API to retrieve a collection of JSONL files, which were subsequently processed in batches. Although we retrieved many articles, the dataset is moderate in size (\(\approx\)128 GB) because we store only abstracts and metadata, not full text.
To ensure we focus on genuinely interdisciplinary articles, a supervised classifier is employed to filter and identify documents that span multiple domains, rather than relying solely on metadata-based categorization20,21. This interdisciplinary classifier is trained on Byte-Pair Encoding (BPE) sequences \(\le\) 300 tokens with post-padding and uses a two-layer Text-CNN (256 filters each; kernel sizes 2 & 3) with max-pooling and PReLU activations in the convolutional layers, sigmoid for the output layer. Embeddings are trainable; dropout = 0.2 (normal); optimizer = Adam (learning rate \(1\times 10^{-3}\)); batch = 64; loss = binary cross-entropy. The model ran up to six epochs with early stopping (patience 2), He-uniform initializer and zero biases, and the output bias set to the label mean.
This is complemented with a discipline-specific filtering stage (trained on Semantic Scholar metadata as well) which enables to narrow down the pool of interdisciplinary article to those focused on engineering and biology solely. Specifically, the engineering classifier stage is followed by a biology classifier stage. Engineering encompasses a broader definition, inclusive of physics, material sciences, and computer science. The different classifier stages attribute, to each abstract, a probability of (a) being interdisciplinary, (b) being engineering-related, and (c) being biology-related. For further implementation details of the interdisciplinary articles classifier, see20. The methodology employed to build the discipline-specific classifiers is analogous.
Finally, the resulting dataset is equally divided between (a) interdisciplinary articles that combine engineering and biology on one hand and (b) engineering-only articles (that is, articles classified as not biology-related by the biology classifier) on the other. The motivation for an equal split between articles combining engineering and biology and engineering-only articles is to enable the generation of topics sufficiently representative of engineering per se. Engineering articles primarily focus on material science, robotics, and energy systems, while biology articles emphasize genomics, systems biology, and biomaterials.
We define \(p_{\textrm{ID}}\), \(p_{\textrm{ENG}}\), and \(p_{\textrm{BIO}}\) as the classifier-predicted probabilities that an abstract is interdisciplinary, engineering-related, and biology-related, respectively. The raw S2ORC dump (101 M abstracts) is trimmed using the following thresholds:
-
1.
Interdisciplinarity. Text-CNN20 \(\rightarrow\) keep abstracts with \(p_{\textrm{ID}}\ge 0.50\).
-
2.
Engineering. Boosted Trees, similar approach to 20 \(\rightarrow\) \(p_{\textrm{ENG}}\ge 0.47\).
-
3.
Biology. Boosted Trees, similar approach to 20 \(\rightarrow\) \(p_{\textrm{BIO}}\ge 0.37\).
The three classifiers employed exhibit F1 scores or (1) 0.82, (2) 0.86, and (3) 0.84 respectively. Threshold values were set to maximize Matthews correlation coefficient25. We excluded abstracts that were empty or contained unreadable characters; all other records were kept if (and only if) they passed the three classifier thresholds and the engineering-only down-sampling step.
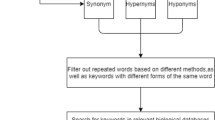
Following preprocessing, we apply BERTopic22 to the filtered corpus. BERTopic is a transformer-based topic-modelling pipeline that clusters documents with UMAP + HDBSCAN and names each cluster with class-based TF-IDF, enabling the extraction of coherent topics from the collection of engineering and biology articles. The overall process is visually summarized in Fig. 1.
Overview of the article-to-topic pipeline, showing filtering, preprocessing, and classification steps for engineering and biology articles.
Each identified topic is labeled according to its association with either engineering or biology. By evaluating article abstracts against these topics, each article is assigned a primary and a secondary topic (based on probability of belonging to a given topic), capturing its strongest thematic alignment and reinforcing its interdisciplinary character. Topic model training is the most computationally expensive step in this process and was performed using an NVIDIA A100 GPU.
We assessed topic quality using the \(C_V\) measure of Mimno et al.26, which has been shown to robustly reflect human judgments across varied corpora. Additionally, Stevens et al.27 demonstrate that average coherence tends to plateau or decline as topic numbers increase in large datasets. We compute \(C_V\) coherence for all topics to ensure our model balances interpretability and coverage.
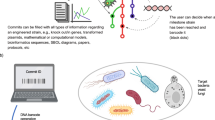
Subsequently, we construct an interdisciplinary graph where nodes represent topics and edges indicate the frequency of co-occurrences derived from the training corpus. Figure 2 shows the step-by-step process that leads to the creation of this graph. In this graph, edges connecting engineering and biology topics serve as interdisciplinary “weights,” revealing meaningful cross-domain links. Frequency thresholds are applied to highlight balanced, non-trivial connections that guide the identification of novel cross-domain opportunities.
Construction of an interdisciplinary topic graph. Each node represents a discovered topic, and edges reflect significant co-occurrences between topics in the filtered set of engineering and biology articles. This graph structure provides a foundation for further embedding, clustering, and identification of meaningful cross-domain linkages.
In our implementation, we begin by constructing a topic association table that links each document’s primary topic with every secondary topic present in the document. When a secondary topic meets our criteria, an edge is established between the primary and secondary topics. The edge weight is determined by counting each occurrence of their co-occurrence—so if Topic A and Topic B appear together in five documents, their edge weight is 5. This straightforward, count-based method effectively normalizes the association frequency across our dataset.
Topic modeling relied on BERTopic, driven by all-MiniLM-L6-v2 sentence-transformer embeddings and a CountVectorizer (English stop-words; vocabulary terms appearing \(\ge 15\) times). Dimensionality reduction used UMAP (\(n_{\text {neighbors}}=15\), \(n_{\text {components}}=5\), metric = cosine, random_state = 42), and clustering followed with HDBSCAN (min_cluster_size = 10). Topics were constrained to min_topic_size = 50, extracted with top_n_words = 10, and accompanied by document–topic probability estimates; a global seed of 42 ensured reproducibility.
The model was configured with a minimum topic size of 10 and a maximum number of topics capped at 1000 to balance granularity with interpretability.
We then inspected the topic–edge-weight histogram and trimmed any topic (or topic pair) whose weight lay outside one standard deviation of the mean—here, \(76 \le w \le 308\)—thereby attempting to suppress dominant “catch-all” clusters.
As a concrete example, consider that our function iterates over the topic association table, assigning each valid topic as a node in the graph and computing edge weights. In one instance, the topic labeled ’Topic 109’ was found to have an edge with ’Topic 327’ with a weight of 5—illustrating how recurring co-occurrences across documents are captured in the graph structure.
Having developed a topic model and calculated the article probabilities, the next step is to create an interdisciplinary model represented as a graph. The framework integrates NLP techniques with graph embedding to analyze and visualize relationships between articles. Before semantic analysis, we employ advanced topic modeling techniques (e.g., BERTopic) to identify latent thematic structures, associating each article with a set of primary and secondary topics. This enhances our understanding of how engineering and biology themes intersect and guides more precise semantic pairing.
For example, an engineering article on “Optimizing material composites for enhanced durability” aligns with a biology article on “Mechanisms of cellular resilience under stress.”. Graph embedding techniques enable visualizing the relationships between topics and associated articles. Frequency-based thresholds are also applied to avoid trivial or overly generic linkages and to filter out rare, potentially noisy associations. By calibrating these thresholds, we aim to retain balanced, actionable links that reveal non-obvious synergies between engineering and biology.
To illustrate, the first step involves associating to each interdisciplinary abstract a primary topic and a secondary topic, using the previously created topic model. A conceptual example is shown in Table 1. Each row is one abstract. “Topic 1” is the highest-probability topic assigned by BERTopic; “Topic 2” is the runner-up. Reading across therefore shows the two strongest thematic anchors for every document.
We next count how often each Topic 1–Topic 2 pair appears across the corpus. The resulting frequency is the edge-weight in the interdisciplinary graph. This is illustrated in Table 2.
This process leads to the creation of a comprehensive interdisciplinary topic network, as illustrated in Fig. 3. Each node in the network represents a topic derived from engineering or biology articles, while the edges capture meaningful co-occurrences that link these topics together. By examining this network, researchers can discover both direct and subtle connections between the two domains. Such insights may highlight promising areas for collaboration, guide the exploration of complementary themes, and ultimately pave the way for more integrated, innovative research.
Subgraph extracted from the interdisciplinary topic network. In this example, nodes represent topics derived from engineering and biology articles, and edges denote semantic co-occurrence frequencies (’weights’), illustrating the framework’s ability to detect cross-domain linkages.
The effectiveness of the framework is assessed using expert validation that provides qualitative insights into the relevance of the proposed pairings. The classifiers employed to narrow down the unfiltered pool of articles to a subset focused on biology and engineering can be evaluated through quantitative metrics such as precision, recall, and F1-score, as detailed in Douard et al.21.
During expert-validation we qualified every engineering–biology pairing as either directly or indirectly relevant. Direct relevance was assigned when the biological insight pointed to an immediately deployable mechanism, material, or design (e.g., gecko feet for robotic adhesion). Indirect relevance covered cases where the biological finding served chiefly as a guiding analogy—valuable, but still requiring abstraction or intermediate engineering steps. For instance, ant foraging behaviour—ants iteratively reinforce the shortest paths with pheromone trails—has been abstracted into ant-colony-optimization heuristics for network routing and project-scheduling; the principle guides the design, but nothing physical is transferred, so its relevance is indirect28,29.
This distinction is well documented in the literature: direct analogies are typically literal form-level transfers, whereas indirect analogies require abstraction of deeper principles before application30. TRIZ-based needs-analysis frameworks make the same split, labeling resources that touch the product directly as “direct-relevance” and those drawn from the broader super-system as “indirect-relevance”31.
Our expert validation phase involved domain experts (primarily with general, multidisciplinary engineering backgrounds) qualitatively reviewing a representative sample of engineering–biology topic pairings to confirm thematic relevance and practical plausibility. Conceptually, they evaluated whether each pairing shared clear thematic overlap, suggested a novel insight, and appeared implementable, providing informal calibration for our pairing heuristics without prescribing rigid criteria.
Results and case examples
Our pipeline filtered 101 million papers to a 126,012 engineering-biology corpus, generating 46,362 cross-domain connections, as summarized in Fig. 4. This analysis identified 544 distinct topic nodes and identified candidate biological analogues, such as gecko-foot adhesion applications for robotic climbing systems, which are studied in this section.
The filtering outcome for the 101 M abstracts dataset illustrates the pipeline’s step-by-step approach. The figure visualizes each stage of dataset refinement—from preprocessing through domain-specific classification—to arrive at a final subset of 126,012 abstracts focused on engineering and biology.
While additional validation may be needed to fully quantify performance, our three-pass filtering helps isolate thematically relevant, cross-domain content: we chose each threshold by optimizing the Matthews correlation coefficient via k-fold cross-validation on held-out abstracts and conducted qualitative spot-checks on random samples at each stage to confirm correct category alignment. Future work may include developing a representative gold-standard validation set. This gold-standard set would be independently annotated by more than one domain experts, enabling to compute inter-annotator agreement (e.g., Cohen’s \(\kappa\))32.
The exact article count for each filtering stage towards the 126,012 abstracts dataset is shown in Table 3. Numbers indicate abstracts remaining after each filter is applied. Interdisciplinarity filtering alone shrinks the corpus by one order of magnitude. The three-stage filtering process reduces the initial 101M abstracts to a final subset of 63,006 abstracts focused on biology, isolating a manageable pool for topic modeling. This is complemented by an equal proportion of 63,006 engineering-only abstracts to arrive to the 126,012 abstract dataset used for topic modeling. As alluded to, the resulting graph included 544 topic nodes and 46,362 edges.
Our BERTopic model achieved a mean \(C_V\) coherence of 0.53 across 544 topics. This value aligns with Mimno et al.’s findings that large, heterogeneous corpora yield moderate coherence scores26. Moreover, Stevens et al. show that as the number of topics grows—here capped at 1000—coherence typically plateaus or declines27. A mean of 0.53 thus reflects a reasonable trade-off between thematic clarity and the preservation of low-frequency, potentially innovative clusters.
Our goal is to generate innovative solutions, and two primary criteria determine the relevance of our pipeline’s outputs: (a) whether they solve the engineering problem at hand in a novel manner, and (b) whether they are feasible in practice. Indeed, when approached with creative insight, even seemingly remote concepts can represent meaningful associations. As these ideas extend beyond the realm of well-established, documented cases, determining their precise relevance becomes increasingly challenging. Nonetheless, it is precisely within these less conventional associations that the highest potential for innovation often resides.
Case examples
Adhesive mechanism for robotic climbing
To assess our framework’s applicability and get an early sense of the relevance of interdisciplinary pairings, we examined well-known cases of biomimicry to determine whether our approach can effectively identify natural analogs that inform engineering design. As a case study, we considered the research question: ’How can we design a high-strength adhesive mechanism for a robotic climbing system?’. From this question, we extracted the following keywords: Robotic Climbing System, High-Strength Adhesive, and Robotic Manipulator Kinematics. Querying our interdisciplinary graph with these keywords yielded several engineering concepts, notably Bipedal Robot Locomotion and Robotic Manipulator Kinematics. Further navigation toward high-weight biological nodes highlighted Biomechanics as a central biological concept. Biomechanics examines the physical forces and mechanical properties underlying biological functions and provides insight into how microscopic structures on arthropod feet generate sufficient adhesive forces—whether through van der Waals interactions, capillary forces, or mechanical interlocking—and manage these forces during attachment and detachment. Biomechanics is particularly relevant in the context of arthropod adhesion, which is a well-known bioinspired solution to the research question33.
To enrich this biomimetic case study, we applied the TRIZ contradiction matrix, setting “Strength” (14) as the parameter to improve and “Adaptability or Versatility” (35) as the one at risk. The matrix points to Principle 3–Local Quality, which recommends localized adjustments to the adhesive surface so it adapts to varying conditions. This strategy echoes arthropod feet, whose microstructural variations maximize grip, simultaneously boosting both strength and adaptability.
Self-organizing coordination in multi-agent robotics
Consider the research question, “How to achieve self-organizing coordination in multi-agent robotic systems?”. Applying our methodology, we identified Swarm Intelligence Optimization Algorithms as the pertinent engineering concept and Animal Locomotion and Spinal Coordination as an associated biological concept. The latter is particularly compelling, as it provides valuable insights into decentralized control mechanisms and the coordination of complex movements in biological systems34. These insights can, in turn, inspire novel approaches to enabling self-organizing coordination in multi-agent robotic systems. This analysis also reveals a methodological challenge: while some identified topics are broad, others are highly specific. Future research should focus on refining the granularity of our topic identification process to achieve an optimal balance between breadth and specificity in interdisciplinary mapping.
For the multi-agent robotics case, we framed a TRIZ contradiction: increase “Extent of Automation” (38) without diminishing “Ease of Operation” (33). The matrix points to Principle 1, Segmentation, advocating modular, autonomous yet cooperative agents. Echoing spinal coordination in animal locomotion, this partitioned design distributes control while keeping operator interaction simple.
Rapid-motion imaging inspired by neuromorphic vision
To further demonstrate the framework’s practical applicability and innovation potential, Douard et al.’s case study35 on bioinspired imaging explores a cross-domain pairing that addresses the research question: ’How can image sensors track rapid motion without information loss?’. By translating biological principles—such as decentralized coordination observed in spinalized cats, adaptive triggering inspired by echolocation, and selective attention modeled after visual cortex processing—this study identifies key design elements that enhance rapid motion tracking in imaging sensors. Furthermore, the asynchronous operation of neuromorphic sensors enables each pixel to independently trigger events upon detecting significant changes in light intensity. This event-driven mechanism, which mirrors the biological process of selective attention, minimizes latency and reduces redundant data capture. These findings illustrate the potential of bioinspired methodologies and showcase the framework’s capacity to bridge diverse knowledge domains.
Extending this bio-inspired imaging example, we framed the challenge in the TRIZ contradiction matrix with “Loss of Information” (24) as the improving feature and “Loss of Time” (25) as the degrading one. This reflects the challenge of enhancing motion tracking without increasing latency. The matrix points to Principle 10, Prior Action, which advocates preparing the system in advance. Neuromorphic sensors exemplify this: each pixel asynchronously pre-conditions itself to fire the moment it detects a luminance change, mirroring biological pre-activation and preventing information loss.
Thermal insulation informed by dental bonding
At times, the biological strategies may appear distant at first glance. For instance, the query “How to insulate an injection molding barrel to avoid heat losses?” identifies Structural Analysis and Materials Engineering in the engineering domain and Dental Bonding and Adhesive Effectiveness in the biological domain. Dental adhesives are engineered to secure strong bonds while reducing thermal transfer, a balance that could inspire innovative material formulations or structural designs for improved insulation of injection molding barrels36. These associations often warrant further investigation. There is a fine distinction between genuinely transformative innovation inspired by uncommon pairings and ideas that ultimately lack practical merit—a challenge inherent to our work.
For thermal-insulating injection-molding barrels, we framed the problem as a TRIZ contradiction between “Temperature” (17) and “Energy Loss” (22): retaining heat without raising power demand. The matrix recommends Principle 22, Blessing in Disguise–repurposing an apparent drawback as an asset. Drawing on dental adhesives, which achieve strong bonding while curbing heat transfer, we transform a “limitation” into a passive-insulation strategy.
The analysis revealed a key limitation: specific topics were overrepresented in pairings, likely due to their broad relevance or generic framing. To address this imbalance, future iterations could incorporate diversity constraints, temporal parameters, and frequency limitations on pairings, ensuring that less-explored yet promising connections are adequately prioritized. These findings also emphasize the inherent subjectivity of pairing relevancy assessments and the need for further refinement. Directly relevant pairings can be defined as those in which the biological concept immediately suggests a tangible strategy or solution for the engineering challenge. In contrast, indirectly relevant pairings can be defined as those where the biological insight, while not directly applicable, provides an underlying principle or analogy that can inspire innovative approaches upon further exploration.
Assignments to “direct” or “indirect” relevance were made during our expert validation phase based on whether the biological insight immediately suggested a concrete engineering mechanism (direct) or a broader guiding principle requiring analogy (indirect). Conceptually, this distinction hinges on the level of specificity with which a pairing can be mapped to an engineering solution. In principle, experts could review examples, define clear criteria, and develop an initial annotation rubric. Future work can potentially incorporate data-derived quantitative measures such as citation-overlap analysis—to systematically validate and refine the direct/indirect classification.
Preliminary quantitative signal
To obtain an early quantitative indication of relevance, we randomly sampled 50 engineering–biology pairings from the graph and asked a domain expert with a mechatronics engineering background to judge each as relevant (positive) or not relevant (negative). Thirty-two pairings were deemed relevant, while 18 were not. Because the sample is small, we report only the aggregated proportion of direct and indirect pairings as a single positive class, with a fuller disaggregated analysis planned for a multi-rater follow-up study. The split provides a first quantitative signal that the pipeline surfaces thematically plausible links more often than chance, motivating a forthcoming evaluation.
Discussion
Interdisciplinary topic graphs can reveal latent connections between engineering and biology, encouraging shared understanding and complementary solutions. This methodology enables a systematic exploration of problem spaces, such as integrating structural optimization techniques with biological principles of resilience, or aligning energy efficiency strategies with adaptive mechanisms observed in nature. Such connections often remain hidden due to the siloed nature of scientific literature. Additionally, the visualization tools examined can effectively convert extensive textual data into actionable insights, enabling researchers to pinpoint opportunities for innovation. This approach provides guidance for cross-domain dialogue, integrating computational tools with frameworks like TRIZ to transform knowledge synthesis.
Another parallel: biological systems achieve high functionality while minimizing resource use. TRIZ echoes this imperative by favoring inventions that maximize performance while lowering the cost of attaining it. Consequently, the “natural” character of bio-inspired solutions aligns with TRIZ’s foundational principle of reducing reliance on resources external to the system under study.
Limitations remain, including the tendency to surface well-known, broadly framed articles more frequently than lesser-known but potentially transformative ones. Addressing this will require refining our selection metrics, introducing mechanisms that promote thematic diversity, and fine-tuning the language models on domain-specific corpora for greater semantic precision. Moreover, further integrating graph embeddings and contradiction resolution strategies promises a richer, more balanced exploration of interdisciplinary opportunities. By striving to diversify the articles drawn into these pairings, we can cultivate a more equitable and innovative research ecosystem that effectively merges the rigor of engineering methods with the adaptive ingenuity of biological systems.
Future work may probe potential biases by performing stratified Cohen’s \(\kappa\) agreement32 analyses across subfields (e.g., materials vs. robotics) and threshold-sensitivity tests to pinpoint where our filtering pipeline is most sensitive.
Recommendations for future research
We recommend future work prioritize: (1) hierarchical topic modeling for enhanced topic granularity, (2) diversity constraints to surface uncommon yet valuable pairings, and (3) explainable-AI components showing pairing rationale.
Hierarchical topic modeling may help progressively refine broad clusters while maintaining coherence. Cross-domain expansion to other scientific disciplines will validate scalability. Class-imbalance countermeasures through diverse sampling may address topic over-representation. Success metrics should include practical outcomes to establish meaningful real-world benchmarks.
Conclusion
We describe an engineering–biology pairing method applied to a 100 M-paper corpus and demonstrate that it can generate bio-inspired pairings. Our end-to-end framework (1) filters a 101 M-paper corpus to a 126,012 high-confidence engineering–biology training corpus, (2) maps it into 544 coherent topics, and (3) surfaces 46,362 data-backed cross-links–demonstrating, at scale, how TRIZ-inspired NLP can expose actionable synergies.
The proposed framework combines transformer-based language models with topic modeling techniques to identify thematic clusters in research literature. This approach goes beyond simple keyword searches to discover meaningful connections between engineering challenges—such as structural optimization—and potentially relevant biological strategies like adaptive behaviors.
Expert assessment and case study evidence indicate that the proposed pairings effectively orient researchers toward conceptually coherent areas of investigation. To advance system effectiveness, future research should emphasize developing sophisticated topic granularity, establishing diversity constraints to expand recommendation scope, and integrating user feedback loops with explainability frameworks to enhance both system reliability and utility.
Data availability
Researchers who are interested in accessing the data may contact the corresponding author for further details. Sample pairing results or analysis snippets are available on request. Access to the data is granted on a case-by-case basis, subject to review and agreement on appropriate data use policies.
References
C.D. Manning, Raghavan, P. & Schütze, H. “Introduction to Information Retrieval: Probabilistic information retrieval.” (2008).
Altshuller, G. S.. “The Theory of Inventive Problem Solving.” (1996).
Nikolay, B. & Bogatyreva, O. “BioTRIZ: A Win-Win Methodology for Eco-innovation.” (2014).
Jacob, D., Chang, M.-W., Lee, K. & Toutanova, K. “BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding.” North American Chapter of the Association for Computational Linguistics (2019).
T.B. Brown et al. “Language Models are Few-Shot Learners.” ArXiv abs/2005.14165 (2020).
Lee, J. et al. BioBERT: A pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2019).
Jieh-Sheng, L. & Hsiang, J. “Patent classification by fine-tuning BERT language model.” World Patent Information (2020).
Fabio, P., Rocktäschel, T., Lewis, P., Bakhtin, A., Wu, Y., Miller, A.H. & Riedel, S. “Language Models as Knowledge Bases?” Conference on Empirical Methods in Natural Language Processing (2019).
Nicolas, D., Samet, A., Giakos, G.C. & Cavallucci, D. “Navigating the Knowledge Network: How Inter-Domain Information Pairing and Generative AI Can Enable Rapid Problem-Solving.” TFC (2023).
Nicolas, D., Samet, A., Giakos, G.C. & Cavallucci, D. “Bridging Two Different Domains to Pair Their Inherent Problem-Solution Text Contents: Applications to Quantum Sensing and Biology.” TFC (2022).
Park, J., Kim, Kwangmin, Hwang, W. & Lee, D. Concept embedding to measure semantic relatedness for biomedical information ontologies. J. Biomed. Inform. 94, 103182 (2019).
Jumper, J. M. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Aditya, G. & Leskovec, J. “node2vec: Scalable Feature Learning for Networks.” Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (2016).
Bryan, P., Al-Rfou, R. & Skiena, S.S. “DeepWalk: online learning of social representations.” Proceedings of the 20th ACM SIGKDD international conference on Knowledge discovery and data mining (2014).
W.L. Hamilton, Ying, Z. & Leskovec, J. “Inductive Representation Learning on Large Graphs.” Neural Information Processing Systems (2017).
Xin, X., Hu, J., Lyu, X., Huang, H. & Cheng, X. “Exploring the Interdisciplinary Nature of Precision Medicine: Network Analysis and Visualization.” JMIR Medical Informatics 9 (2020).
J.F.V. Vincent & Cavallucci, D. “Development of an Ontology of Biomimetics Based on Altshuller’s Matrix.” TFC (2018).
Vincent, J.F.V. “Biomimetics with Trade-Offs.” Biomimetics 8 (2023).
Armands, S., Osis, J. & Donins, U. “Knowledge Integration for Domain Modeling.” MDA/MDSD (2017).
Nicolas, D., Samet, A., Giakos, G.C. & Cavallucci, D. “Quantifying Interdisciplinarity in Scientific Articles Using Deep Learning Toward a TRIZ-Based Framework for Cross-Disciplinary Innovation.” Machine Learning and Knowledge Extraction (2025).
Nicolas, D., Samet, A., Giakos, G.C. & Cavallucci, D. “A Novel Interdisciplinarity Model Towards Inter-domain Information Pairing.” TFC (2024).
Grootendorst, M.R. “BERTopic: Neural topic modeling with a class-based TF-IDF procedure.” ArXiv abs/2203.05794 (2022).
Zhang, T. et al. Benchmarking large language models for news summarization. Trans. Assoc. Comput. Linguistics 12, 39–57 (2023).
Kyle, L. Wang, L.L., Neumann, M., Kinney, R.M. & Weld, D.S. “S2ORC: The Semantic Scholar Open Research Corpus.” Annual Meeting of the Association for Computational Linguistics (2020).
Davide, C. & Jurman, G. “The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation.” BMC Genomics 21 (2020).
David, M., Wallach, H.M., Talley, E.M., Leenders, M. & McCallum, A. “Optimizing Semantic Coherence in Topic Models.” Conference on Empirical Methods in Natural Language Processing (2011).
Keith, S., Kegelmeyer, W.P., Andrzejewski, D. & Buttler, D.J. “Exploring Topic Coherence over Many Models and Many Topics.” Conference on Empirical Methods in Natural Language Processing (2012).
Dorigo, M. & Gambardella, L. M. Ant colony system: A cooperative learning approach to the traveling salesman problem. IEEE Trans. Evol. Comput. 1, 53–66 (1997).
Di, C., Gianni, A. & Dorigo, M. AntNet: Distributed Stigmergetic Control for Communications Networks. J. Artif. Intell. Res. 9, 317–365 (1998).
Vincent, J. F. V. & Mann, D. L. Systematic technology transfer from biology to engineering. Philos. Trans. R. Soc. Lond. Series A Math. Phys. Eng. Sci. 360, 159–173 (2002).
Guo, J. et al. A needs analysis approach to product innovation driven by design. Proc. CIRP 39, 39–44 (2016).
Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 20, 37–46 (1960).
Persson, B. N. J. Biological adhesion for locomotion on rough surfaces: Basic principles and a theorist’s view. MRS Bull. 32, 486–490 (2007).
Webb, Barbara. Swarm intelligence: From natural to artificial systems. Connection Sci. 14, 163–164 (2002).
Nicolas, D., Samet, A., Cavallucci, D. & Giakos, G.C. “An Interdisciplinary Framework for Systematic Innovation: A Case Study on Nature-Inspired Imaging Solutions.” 2024 IEEE International Conference on Imaging Systems and Techniques (IST) (2024) 1-5.
Meerbeek, V. et al. Buonocore memorial lecture. Adhesion to enamel and dentin: current status and future challenges. Oper. Dentistry 28(3), 215–35 (2003).
Acknowledgements
This study was carried out within the framework of the Collaborative Doctoral Program in Applied Artificial Intelligence for Industry, a joint initiative between the University of Strasbourg and the National Institute of Applied Sciences (INSA) in France, in partnership with the Laboratory for Quantum Cognitive Imaging and Neuromorphic Engineering q(CINE) and Bioinspired Space Systems at Manhattan University in New York, USA. We sincerely thank the Conception, Système d’Information et Processus inventifs (CSIP) group at the ICube Laboratory, INSA Strasbourg, as well as our colleagues, for their steadfast support, collaborative efforts, and insightful contributions throughout this project.
Funding
This work was completed as part of a PhD contract funded by the AIARD (Artificial Intelligence Aided Research and Development) industrial chair. The AIARD industrial chair is co-financed by the Grand Est region, the Eurométropole de Strasbourg, and ten collaborating companies. These partners provide financial support to advance research and development in artificial intelligence. A complete list of the participating companies, along with further details about the industrial chair, is available on the Chair’s website: www.aiard.eu.
Author information
Authors and Affiliations
Contributions
Conceptualization, N.D. and D.C.; methodology, N.D. and A.S.; software, N.D.; formal analysis, N.D.; investigation, N.D.; resources, G.G.; data curation, N.D.; writing—original draft preparation, N.D.; writing—review and editing, G.G., D.C., and A.S.; visualization, N.D.; supervision, D.C. and G.G.; project administration, G.G. and D.C.; funding acquisition, D.C. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors. Hence, institutional ethics approval was not required.
Ethical considerations
We complied with all ethical standards and licensing requirements associated with the use of the Semantic Scholar Open Research Corpus (S2ORC). The dataset, made available under the ODC-By 1.0 license, was utilized strictly for non-commercial research purposes in accordance with the licensing terms. To ensure adherence to data privacy regulations and the protection of intellectual property rights, no personally identifiable information or sensitive data was included in our analysis.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Douard, N., Cavallucci, D., Samet, A. et al. Large-scale transformer-based topic graphs identify thematic links between engineering and biology. Sci Rep 15, 29256 (2025). https://doi.org/10.1038/s41598-025-15067-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15067-9