Abstract
Glioblastoma is an aggressive, malignant primary brain tumour and the most prevalent histological type of glioma. Our study attempted to investigate the independent predictors of overall survival (OS) and cancer-specific survival (CSS) in Asian patients with glioblastoma and establish predictive models for the OS and CSS of Asian patients with glioblastoma based on the machine learning algorithms. Data from Asian patients with glioblastoma in the SEER database were retrieved and stochastically grouped into a training set (n = 845) and a validation set (n = 362), and patients in our centre were assigned to the test set (n = 172). Univariate and multivariate Cox regression analyses were performed to evaluate the prognostic factors. Predictive models for OS and CSS were established based on eight machine learning algorithms, including Lasso Cox, random survival forest, CoxBoost, generalized boosted regression modelling (GBM), stepwise Cox and survival support vector machine, eXtreme Gradient Boosting, supervised principal component and partial least squares regression for Cox, and the selected predictive models were evaluated by the area under the ROC curves (AUC) and 95% confidence interval (CI), calibration curves and decision curve analyses in the training set, validation set and test set. In our retrospective study, age, tumour history, histologic type, surgery and chemotherapy were confirmed to be predictors of OS (p < 0.05); age, tumour history, histologic type, surgery and chemotherapy were identified as independent factors for CSS (p < 0.05). The predictive model for OS based on the GBM algorithm exhibited excellent predictive performance at 6 months (AUC = 0.837, 95% CI: 0.803–0.870), 12 months (AUC = 0.809, 95% CI: 0.780–0.839) and 24 months (AUC = 0.750, 95% CI: 0.717–0.783) in the training set, and the powerful predictive performance of the GBM model was confirmed in the validation and test sets, with good concordance between the predicted and observed OS rates demonstrated by calibration curves and clinical decision making performance suggested by the decision curve analyses curves. The predictive model based on the GBM algorithm for CSS also performed best = in the training set at 6 months (AUC = 0.808, 95% CI: 0.770–0.847), 12 months (AUC = 0.755, 95% CI: 0.721–0.789) and 24 months (AUC = 0.692, 95% CI: 0.657–0.728) in the training set, and convincing predictive effectiveness was also confirmed in the validation and test sets with good calibration and clinical utility. Age, tumour history, histologic type, surgery and chemotherapy were confirmed to be independent factors for OS; and age, tumour history, histologic type, surgery and chemotherapy were identified as prognostic factors for CSS in our retrospective study. The predictive model constructed for OS and CSS based on the GBM algorithm in Asian patients with glioblastoma can be used to accurately predict OS and CSS in clinical practice, which may help tailor personalized treatment regimens and provide significant benefits for these patients.
Similar content being viewed by others
Introduction
Glioblastoma is the most prevalent histological type of glioma, accounting for more than half of the total number of gliomas, with a frequency of approximately four incidences among 100,000 people per year1,2,3. Glioblastoma is more likely to occur in people aged 55–85 years, with a median age of 64 years. According to the WHO grading system, glioblastoma represents a highly invasive and fast-growing neoplasm originating in brain tissue and is categorized under the most severe classification level of central nervous system tumours4. Patients diagnosed with glioblastoma face challenging clinical outcomes, typically exhibiting limited life expectancy with a median survival of only 15 months and a median progression-free survival of 6.2–7.5 months despite standard therapies, including maximal safe surgical resection followed by concurrent radiotherapy and adjuvant chemotherapy5,6,7. The identification of prognostic indicators and the development of reliable prediction tools for patients with glioblastoma have significant clinical value, as they enable timely therapeutic strategies and personalized care approaches, ultimately leading to improved treatment outcomes. Such advancements in risk stratification and predictive analytics may contribute to improved mortality rates through optimized clinical decision-making processes.
Machine learning (ML), which demonstrates superior capabilities in pattern recognition and predictive analytics and enables more robust data modelling and precise parameter estimation through advanced computational algorithms, has been widely used in various areas of clinical research8,9 and practice. Compared with traditional survival prediction models built employing multivariate Cox regression, ML enables the extraction of meaningful patterns from complex datasets, facilitates the identification of underlying correlations among variables, and supports the development of sophisticated forecasting frameworks through diverse algorithmic approaches10. Thus, our study aimed to investigate the independent predictors of overall survival (OS) and cancer-specific survival (CSS) and establish ML model to OS and CSS of Asian patients with glioblastoma. The model was trained and validated using data from the SEER database and tested through the data in our centre and can be used to provide guidance for clinical treatment decisions.
Materials and methods
Patient information acquisition
We searched the publicly accessible SEER database (https://seer.cancer.gov/), which consists of 17 registries covering more than 26.5% of the United States population between 2000 and 2021, to obtain the majority of information on patients with glioblastoma available in February 2025 through SEER*Stat (version 8.4.3). The inclusion criteria were as follows: (1) patients were diagnosed with glioblastoma between 2010 and 2021; (2) the race of the patient was Asian; (3) the histologic type ICD-O-3 site code was 9440/3, 9441/3, 9442/3 or 9445/3; (4) the pathology was confirmed; and (5) the surgery codes were “00”, “20”, “21”, “30”, or “55”. The exclusion criteria consisted of the following conditions: (1) patient age less than 18 years; (2) unknown tumour size; (3) unknown laterality; (4) unknown survival time or status; and (5) survival time of 0 months. The main outcome measures in our study were OS and CSS. OS was defined as the time span from the initial diagnosis of PUC to the time of death or the terminal follow-up. CSS was measured as the time from the diagnosis of glioblastoma to glioblastoma-specific death. Moreover, we collected the above patient information as a test set from the Second Affiliated Hospital of Dalian Medical University. This study obtained the approval of the Institutional Research Ethics Committees of the Second Affiliated Hospital of Dalian Medical University and the informed consent of all the enrolled patients and/or their legal guardians. All studies were performed in accordance with the Declaration of Helsinki and relevant guidelines.
Statistical analysis
Counting data were converted into categorical variables. Univariate and multivariate Cox analyses of the independent predictors of OS and CSS, presented as hazard ratios (HRs) and 95% confidence intervals (CIs), were performed, and a p value < 0.05 was considered to indicate statistical significance. Inverse probability of treatment weighting (IPTW) uses standardized methods to control and minimize potential biases and confounding effects on survival analyses by assigning appropriate weights to each observation through propensity score values. Cox regression analysis was subsequently performed to confirm the independent factors after IPTW. We randomly divided the SEER database data into training (n = 845) and validation (n = 362) sets at a proportion of 7:3, and the patients in our centre were assigned to the test set (n = 172). Univariate Cox analyses in the training set were performed to screen variables to construct the machine learning predictive model. Eight machine learning algorithms were implemented, including random survival forest (RSF), generalized boosted regression modelling (GBM), Lasso Cox, CoxBoost, stepwise Cox and survival support vector machine (Survival-SVM), eXtreme Gradient Boosting (XGBoost), supervised principal component (SuperPC) and partial least squares regression for Cox (plsRcox) to establish predictive models for OS and CSS. OS and CSS were evaluated at discrete time points (6, 12, and 24 months) to assess short-, medium-, and long-term prognostic performance. Discrimination was assessed using Harrell’s concordance index (C-index) to present global measure and time-dependent ROC curves using the timeROC and riskRegression packages with AUCs calculated separately for each time point. The 95% confidence intervals for AUC values were estimated using bootstrap resampling (500 repetitions) to account for variability in model performance. To further validate the models, calibration curves were utilized to examine the alignment between predicted probabilities and actual outcomes. Decision curve analyses (DCAs) were used to evaluate the practical clinical value of these methods. Using the best-performing model, patients were stratified into high-risk and low-risk groups based on the median predicted risk score. This cutoff ensured balanced group sizes and maximized discriminative power for survival outcomes. Kaplan‒Meier (KM) survival curves and log-rank tests were performed to confirm the predictive model discrimination capacity between high-risk and low-risk groups. All the above analyses were performed in R version 4.4.2.
Results
Baseline characteristics
The baseline characteristics of glioblastoma patients in SEER set and test set were presented in the Table 1, key differences emerged in age, tumor history prevalence, and histologic type (p <0.05) which may implying the data heterogeneity, while other demographic and clinical features remained comparable between cohorts.
Identification of the risk factors for patients with glioblastoma
Identification of the risk factors for OS
Univariate and multivariate Cox regression analyses were performed to screen for significant predictors of OS in the SEER set (n = 1207) and test set (n = 172). As shown in Table 2, age, histologic type, combined summary stage, surgery, radiotherapy and chemotherapy were confirmed as independent prognostic factors for OS in the SEER set (p <0.05). In the test set, age, tumour history, histologic type, surgery and chemotherapy were screened as statistically significant prognostic factors for patients with glioblastoma (p <0.05). To address potential selection bias of the tumor history on OS, we employed IPTW strategy to generate weighted cohorts using year of diagnosis, age, gender, histologic type, primary site, laterality, tumor size, combined summary stage, surgery, radiotherapy, chemotherapy. As shown in Table 3, tumor history continued to be an independent predictor of worse OS after employing IPTW (HR: 2.06, 95% CI: 1.30–3.25, p =0.002), reinforcing the robustness of the association, while no significant difference was explored after IPTW in SEER set.
Identification of the risk factors for CSS
The detailed univariate and multivariate Cox regression analysis results for CSS in the SEER set and test set are presented in Table 4. Age, histologic type, primary site, combined summary stage, surgery, radiotherapy and chemotherapy were identified as statistically significant factors after univariate and multivariate Cox regression analyses in the SEER set (p <0.05). In the test set, age, tumour history, histologic type, surgery and chemotherapy were confirmed to be independently significant factors for CSS (p <0.05). We used the IPTW technique to create weighted cohorts based on year of diagnosis, age, gender, histologic type, primary site, laterality, tumor size, combined summary stage, surgery, radiotherapy, and chemotherapy to address the potential selection bias of tumor history on CSS. As demonstrated in Table 5, tumor history remained an independent predictor of worse OS after IPTW (HR: 1.93, 95% CI: 1.30–2.86, p =0.001) in the test set, confirming the association strength.
Machine learning based predictive model for OS
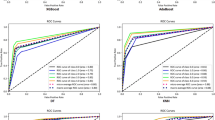
We used univariate Cox regression to select features in the training set. As presented in Table6,nine features, including age, tumour history, histologic type, laterality, tumour size, combined summary stage, surgery, radiotherapy and chemotherapy, were statistically significant and were selected for the construction of predictive models. Multiple machine learning algorithms, including RSF, GBM, Lasso Cox, CoxBoost, Survival-SVM, XGBoost, SuperPC and plsRcox, were employed to develop prognostic frameworks capable of estimating OS probabilities at 6-month, 12-month, and 24-month intervals. As the ROC curves in Fig. 1; Table 7 show, the GBM model exhibited excellent predictive performance at 6 months (AUC = 0.837, 95% CI: 0.803–0.870), 12 months (AUC = 0.809, 95% CI: 0.780–0.839) and 24 months (AUC = 0.750, 95% CI: 0.717–0.783) in the training set, and the powerful predictive performance of the GBM model was confirmed in the validation and test sets. The DCA curve also suggested that the GBM model holds considerable utility in making clinical decisions (Fig. 2). The GBM model showed good agreement between the predicted and observed OS rates at 6 months, 12 months and 24 months in the training, validation, and test sets, as shown by the calibration curves (Fig. 3). The survival curves in Fig. 4 demonstrate the differentiation capability between low-risk and high-risk OS patients in the training, validation and test sets (p <.05).
Receiver operating characteristic curves of the 6-month, 12-month, and 24-month OS prediction models for glioblastoma patients in the training (A-C), validation (D-F), and test (G-I) set.
Decision curve analyses (DCAs) of the 6-month, 12-month, and 24-month OS prediction models for glioblastoma patients in the training (A-C), validation (D-F), and test (G-I) set.
Calibration curves of 6-month, 12-month, and 24-month OS prediction in the training (A), validation (B), and test (C) set.
The survival curves between low-risk and high-risk glioblastoma patients for OS.
Machine learning based predictive model for CSS
As shown in Table 8, seven features, including age, primary site, laterality, combined summary stage, surgery, radiotherapy and chemotherapy, were statistically significant, as confirmed by univariate Cox regression, and were incorporated to establish the predictive models. Among the eight machine learning algorithm-based models, RSF, GBM, Lasso Cox, CoxBoost, Survival-SVM, XGBoost, SuperPC and plsRcox, the GBM model exhibited the best predictive performance in the training set at 6 months (AUC = 0.808, 95% CI: 0.770–0.847), 12 months (AUC = 0.755, 95% CI: 0.721–0.789) and 24 months (AUC = 0.692, 95% CI: 0.657–0.728) in the training set, as shown in Table 9; Fig. 5, and convincing predictive effectiveness was also confirmed in the validation and test sets. The utility of the GBM model for clinical decision making was suggested by the DCA curve (Fig. 6). The calibration curves of the GBM model in the training, validation, and test sets showed good concordance between the predicted and observed overall survival rates at 6 months, 12 months and 24 months (Fig. 7). As illustrated in Fig. 8, Kaplan-Meier analysis revealed a statistically significant stratification (p <0.05) of CSS outcomes between the low-risk and high-risk subgroups in the training, validation, and test sets (p <0.05).
Receiver operating characteristic curves of the 6-month, 12-month, and 24-month CSS prediction models for glioblastoma patients in the training (A-C), validation (D-F), and test (G-I) set.
Decision curve analyses (DCAs) of the 6-month, 12-month, and 24-month CSS prediction models for glioblastoma patients in the training (A-C), validation (D-F), and test (G-I) set.
Calibration curves of 6-month, 12-month, and 24-month CSS prediction in the training (A), validation (B), and test (C) set.
The survival curves between low-risk and high-risk glioblastoma patients for CSS.
Discussion
This study clarified the survival-associated factors of Asian patients with glioblastoma in the SEER database and our retrospective cohort and developed a novel machine learning predictive model based on the data from the SEER database, which was validated with retrospective cohort data. Age, histologic type, surgery and chemotherapy were identified as independent factors for OS and CSS in the SEER database and retrospective cohort, which was also validated by other studies11,12. Maximal surgical resection followed by simultaneous radiotherapy and temozolomide chemotherapy is the recommended standard for the treatment of patients with glioblastoma12,13but not all patients receive standard treatment for several reasons. Moreover, we found that tumour history was an important risk factor for the OS and CSS of patients with glioblastoma in our retrospective cohort before and after IPTW, which has not yet been widely recognized. In the research of Ebad Ur Rehman et al., tumour history was an independent OS-associated factor, and the most frequent prior cancers reported were genitourinary, breast, haematologic and lymphatic, and gastrointestinal malignancies; however, the exact mechanism underlying the connection was unclear14. A common hypothesis is a shared genetic pathway, and Turcotte et al. reported that primary cancers can give rise to syndromes that show an autosomal dominant pattern of inheritance, such as gliomas15. Another hypothesis is that an important risk factor for the development of CNS tumours in cancer survivors is prior cranial radiotherapy and chemotherapy, which has been validated by various studies16,17,18,19,20. In the retrospective study for the second primary gliomas conducted by Maluf et al., most had received chemotherapy and/or radiation before their glioma diagnosis21. What’s more, both intravenous and intrathecal methotrexate administration have been found to be associated to an increased risk of meningioma development22,23. Diagnostic delays in some Chinese patient populations often result in initial cancer detection at advanced stages, consequently strengthening the observed correlation between prior tumor history and adverse prognosis. The relevant connections and underlying mechanisms still need to be explored in depth.
We were the first to apply machine learning models in the construction of predictive models for patients with glioblastoma based on clinical data. The predictive performance of the generalized boosted regression modelling algorithm was more stable and better than those of the other algorithms, and the overall survival of patients with glioblastoma reached AUROCs of 0.856, 0.777 and 0.691 for 6-month, 12-month and 24-month survival, respectively, in the test set. The predictive model for cancer-specific survival attained AOROCs of 0.805, 0.736 and 0.703 for 6-month, 12-month and 24-month survival, respectively, in the test set. The performance of our machine learning-based predictive models was slightly preferable to that of the prognostic nomogram models, which were developed on the clinical data of Niu et al. using traditional Cox regression analysis24. A variety of predictive models have been created as artificial intelligence continues to evolve. The radiomics-based machine learning prediction model using pretreatment multiparametric magnetic resonance imaging achieved AUROCs of 0.791 and 0.708 in internal and external validation, respectively25. The SVM and LR pathomics models based on glioblastoma tissue and normal brain tissue were constructed with AUC values of 0.779 and 0.785, respectively26. The effectiveness of multimodal prediction tends to be better than that of a single type of data. The comprehensive clinical-radiomics-TME model for predicting OS constructed by Zhou et al. exhibited satisfactory survival prediction ability, with AUROCs of 1-, 2-, and 3-year OS prediction of 0.842, 0.844, and 0.795, respectively, in the test set27. The combination of clinical data and genetic information through artificial neural networks accurately predicts 15-month survival, with an accuracy rate of 83.3% and an AUROC of 0.8128. Combining multiple types of data, including clinical, radiomics, pathomics and genetic data, may help further improve the predictive efficacy of prognostic models.
There are several limitations to our study. Since the patient data in the training and validation sets were collected from the SEER database, the retrospective nature and missing data were unavoidable, and the single-centre data of patients with glioblastoma also limited the sample size of the test set, which may have led to bias or heterogeneity. Thus, large-scale prospective studies and detailed characteristic analyses, such as analyses of chemotherapeutic regimens, are needed to explore and verify the prognostic factors in our centre and construct a predictive model with greater predictive effectiveness and the capability for further survival prediction. Moreover, the combination of additional data, including radiomics, pathomics and genetic data, may contribute to further enhancing the predictive power of the model.
Conclusion
Age, tumour history, histologic type, surgery and chemotherapy were confirmed to be independent factors for OS, and age, tumour history, histologic type, surgery and chemotherapy were identified as prognostic factors for CSS in our retrospective study. The predictive model constructed for OS and CSS based on the generalized boosted regression modelling algorithm in Asian patients with glioblastoma can be used to accurately predict OS and CSS in clinical practice, which may help tailor personalized treatment regimens and provide significant benefits for Asian patients with glioblastoma.
Data availability
For patient privacy reasons, datasets generated and/or analyzed in this study are not publicly available, but are available from the corresponding authors upon request.
References
Ostrom, Q. T., Gittleman, H., Kruchko, C. & Barnholtz-Sloan, J. S. Primary brain and other central nervous system tumors in appalachia: regional differences in incidence, mortality, and survival. J. Neurooncol. 142, 27–38 (2019).
Lapointe, S., Perry, A. & Butowski, N. A. Primary brain tumours in adults. Lancet 392, 432–446 (2018).
Grochans, S. et al. Baranowska-Bosiacka, I. Epidemiology of glioblastoma Multiforme-Literature review. Cancers (Basel). 14, 2412 (2022).
Louis, D. N. et al. The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Stupp, R. et al. Cilengitide combined with standard treatment for patients with newly diagnosed glioblastoma with methylated MGMT promoter (CENTRIC EORTC 26071 – 22072 study): a multicentre, randomised, open-label, phase 3 trial. Lancet Oncol. 15, 1100–1108 (2014).
Weller, M. et al. EANO guideline for the diagnosis and treatment of anaplastic gliomas and glioblastoma. Lancet Oncol. 15, e395–403 (2014).
Stupp, R. et al. Maintenance therapy with Tumor-Treating fields plus Temozolomide vs Temozolomide alone for glioblastoma: A randomized clinical trial. JAMA 314, 2535–2543 (2015).
Skrede, O. J. et al. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet 395, 350–360 (2020).
Le Berre, C. et al. Application of artificial intelligence to gastroenterology and hepatology. Gastroenterology 158, 76–94e2 (2020).
Achilonu, O. J. et al. Predicting colorectal cancer recurrence and patient survival using supervised machine learning approach: A South African Population-Based study. Front. Public. Health. 9, 694306 (2021).
Kim, M. et al. Glioblastoma as an age-related neurological disorder in adults. Neurooncol Adv. 3, vdab125 (2021).
Gilbert, M. R. & Loghin, M. The treatment of malignant gliomas. Curr. Treat. Options Neurol. 7, 293–303 (2005).
Zhao, M. J., Lu, T., Ma, C., Wang, Z. F. & Li, Z. Q. A narrative review on the management of glioblastoma in China. Chin. Clin. Oncol. 11, 29 (2022).
Ebad Ur Rehman, M. et al. Impact of prior cancer history on survival in brain malignancy: A propensity score-adjusted, population-based study. Cancer Rep (Hoboken) 7, e (2024). (1984).
Turcotte, L. M. et al. Risk factors, and surveillance of subsequent malignant neoplasms in survivors of childhood cancer: A review. J. Clin. Oncol. 36, 2145–2152 (2018). Risk.
Bowers, D. C. et al. Subsequent neoplasms of the CNS among survivors of childhood cancer: a systematic review. Lancet Oncol. 14, e321–328 (2013).
Yamanaka, R. & Hayano, A. Secondary glioma following acute lymphocytic leukemia: therapeutic implications. Neurosurg. Rev. 40, 549–557 (2017).
Pettorini, B. L. et al. Radiation-induced brain tumours after central nervous system irradiation in childhood: a review. Childs Nerv. Syst. 24, 793–805 (2008).
Relling, M. V. et al. High incidence of secondary brain tumours after radiotherapy and antimetabolites. Lancet 354, 34–39 (1999).
Little, M. P. et al. Risks of brain tumour following treatment for cancer in childhood: modification by genetic factors, radiotherapy and chemotherapy. Int. J. Cancer. 78, 269–275 (1998).
Maluf, F. C., DeAngelis, L. M., Raizer, J. J. & Abrey, L. E. High-grade gliomas in patients with prior systemic malignancies. Cancer 94, 3219–3224 (2002).
Taylor, A. J. et al. Population-based risks of CNS tumors in survivors of childhood cancer: the British childhood cancer survivor study. J. Clin. Oncol. 28, 5287–5293 (2010).
Chojnacka, M. et al. Second brain tumors following central nervous system radiotherapy in childhood. Br. J. Radiol. 87, 20140211 (2014).
Niu, X., Chang, T., Yang, Y. & Mao, Q. Prognostic nomogram models for predicting survival probability in elderly glioblastoma patients. J. Cancer Res. Clin. Oncol. 149, 14145–14157 (2023).
Karabacak, M., Patil, S., Gersey, Z. C., Komotar, R. J. & Margetis, K. Radiomics-Based machine learning with natural gradient boosting for continuous survival prediction in glioblastoma. Cancers (Basel). 16, 3614 (2024).
Li, W. et al. Pathomics models for CD40LG expression and prognosis prediction in glioblastoma. Sci. Rep. 14, 24350 (2024).
Zhou, Q. et al. Integrated MRI radiomics, tumor microenvironment, and clinical risk factors for improving survival prediction in patients with glioblastomas. Strahlenther Onkol. 201, 398-410 (2024).
Azimi, P., Yazdanian, T., Zohrevand, A. & Ahmadiani, A. Predicting survival in glioblastoma using gene expression databases: A neural network analysis. Int. J. Mol. Cell. Med. 13, 79–90 (2024).
Funding
The present study was supported by Dalian Institute of Chemical Physics-Dalian University of Medical Science and Technology Joint Innovation Fund (DMU-2&DICP UN202309) and Liaoning Provincial Science and Technology Department (2022-MS-15, 2022-MS-16).
Author information
Authors and Affiliations
Contributions
YTH and NWC designed the study, revised the manuscript and was responsible to undertake project leadership and guarantee this study. DLL, LXZ and LFX contributed to write this manuscript and analyzed the data of patient, RHZ, HYG, JLG and MHW collected the data of patients. All authors contributed to the article and approved the final version of manuscript for submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The Ethics Committee of the Second Affiliated Hospital of Dalian Medical University approved the research.
Patient consent for publication
The patient provided informed consent.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, D., Zhang, L., Xu, L. et al. Development and validation of a machine learning-based survival prediction model for Asian glioblastoma patients using the SEER database and Chinese data. Sci Rep 15, 31114 (2025). https://doi.org/10.1038/s41598-025-15553-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15553-0