Abstract
Glioblastoma (GBM) displays marked intratumoral heterogeneity and high invasiveness, making en bloc resection difficult. In particular, total/subtotal resection is challenging for lesions near the diencephalon, midbrain, or brainstem or in motor areas, as damage to these sites can compromise postoperative function and reduce quality of life. Resection of GBM, especially deep-seated tumors, rarely achieves circumferential margins; pathological examination can only be performed on a portion of the tumor. Notably, traditional random intraoperative sampling introduces bias, limiting molecular profiling. The use of imaging-based biomarkers may mitigate this sampling bias, but critical molecular–region signatures remain undefined. In this study, we identified imaging parameters on structural magnetic resonance perfusion imaging to facilitate targeted sampling of representative glioma regions. Whole-exome sequencing of 29 samples from 6 patients revealed significant intratumoral heterogeneity in gene mutations across sampling sites, with edge regions showing the highest heterogeneity. Key cancer driver and drug-target gene mutations also exhibited heterogeneous spatial distributions. Critically, regions with elevated perfusion-weighted imaging cerebral blood volume (PWI-CBV) values demonstrated greater mutation heterogeneity and enrichment of key mutation events. To ensure comprehensive detection of heterogeneous mutations intraoperatively, we propose a three-site sampling strategy within high PWI-CBV regions. This approach may increase the accuracy and completeness of molecular pathological data, ultimately supporting personalized therapeutic planning.
Similar content being viewed by others
Introduction
Gliomas are the most prevalent primary malignant tumors of the central nervous system in adults1; glioblastoma (GBM)2 accounts for more than 50% of all gliomas3,4,5. However, effective treatment options for GBM remain limited, and surgical intervention continues to be the preferred therapeutic approach6,7,8,9. On the basis of findings from tumor morphopathology and molecular pathology studies, the 2021 edition of the WHO classification of central nervous system tumors specifies distinct molecular pathological events in GBM10,11,12. These characteristic molecular pathological events offer important guidance for diagnosis and treatment planning13. Molecular pathological feature-based diagnosis would enable more accurate and effective treatment planning13,14; however, GBM exhibits heterogeneous gene mutations both among patients and within individual tumors15,16limiting the accuracy and comprehensiveness of intraoperative sampling and thus making it difficult to obtain reliable subsequent pathological results, make precise diagnoses and develop effective treatment plans17,18,19. Therefore, surgeons need to fully consider intratumoral gene mutation heterogeneity (GMH) by adopting evidence-based methods during surgery to ensure that the sample accurately and comprehensively reflects the characteristics of the entire GBM tumor. The Response Assessment in Neuro-Oncology (RANO) consortium has suggested adopting intraoperative multisite uniform sampling (RANO plan) to reduce sampling bias; however, verification is needed to determine whether a more efficient and targeted optimization scheme for multisite sampling can be adopted for GBM20.
Significant intratumoral heterogeneity can be observed via multimodal magnetic resonance imaging (MRI) of GBM21,22. This heterogeneity manifests as evident differences in imaging characteristics across different intratumoral regions, which can be attributed to factors such as the degree of tumor cell differentiation, the distribution of intratumoral blood vessels, and the metabolic status of the tumor cells23,24. Our concern lies in whether the MRI features of GBM can be utilized to optimize the RANO criteria. To address this, we obtained 29 tumor samples from 6 GBM patients via intraoperative neuronavigation under general anesthesia and characterized the intratumoral GMH of the samples via whole-exon sequencing (WES). We then integrated multimodal data such as brain-tumor 3D spatial reconstruction, MRI scans, and WES results to optimize the RANO plan (Fig. 1A).
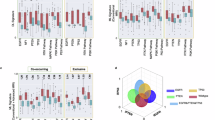
The intratumoral genetic–imaging feature heterogeneity of GBM is significant and differs across different tumor anatomical subregions. (A), Schematic diagram of the MRI feature-guided intraoperative multisite sampling and follow-up study. (B), Proportions of SGMs, CGMs and UGMs in different samples from different patients. (C), Correlation analysis between PGMs and variance in the ROGs of the original CE-T1W images over time (n = 6). (D), Correlation analysis between the number of gene mutations in individual samples and the distance from the tumor centroid in patients whose tumors tended to show sphere-like growth on imaging (n = 14 of 3 patients). (E), Correlation analysis between the proportion of PGMs and the distance from the tumor centroid in patients whose tumors tended to show sphere-like growth on imaging (n = 14 of 3 patients). GBM, glioblastoma; PnSn, sample no. n of patient no. n, SGM, shared gene mutation; CGM, crossover gene mutation; UGM, unique gene mutation; PGM, private gene mutation.
Materials and methods
Sample acquisition and statistics
We recruited patients who were diagnosed with GBM and underwent intraoperative multisite sampling between July 2017 and December 2018. The study protocol was approved by the Ethics Committee of DaPing Hospital (Project number: Medical Research Ethical Review (2024) NO.217) and registered in the Chinese Medical Research Registration and Filing Information System (CMR, medicalresearch.org.cn) (Registration No: MR-50-24-029389) and China Clinical Trial Registry (chictr.org.cn) (Registration No: ChiCTR2500103340). All procedures strictly adhered to the Declaration of Helsinki principles and relevant national ethical guidelines, with written informed consent obtained from all participants or their legally authorized representatives. All operations were rigorously executed in strict compliance with the pre-established study protocol. The inclusion criteria were as follows: (a) a diagnosis of IDH-wildtype glioblastoma; (b) provision of informed consent for intraoperative multisite sampling for research purposes; and (c) postoperative pathologically confirmed GBM. The exclusion criteria were as follows: (a) non-glioma-related injuries or disease affecting survival during the disease course up to May 2024; (b) refusal to allow the samples to be used for scientific studies; and (c) unclear postoperative diagnosis or missing imaging data. Ultimately, 6 patients aged between 48 and 74 years were enrolled. Additional details are presented in the Supplementary Table 1.
Multisite sampling
The enrolled patients underwent preoperative MRI to acquire contrast-enhanced T1-weighted (CE-T1W)25 and CBV-weighted images26. Additionally, a computed tomography scan was performed for 3D reconstruction. Radiologists manually delineated and modeled glioma regions using PWI and CE-T1W imaging data to define regions of interest (ROIs), from which CBV and CE-T1W signal intensity values were extracted. During this process, 4–6 sampling sites were selected within the solid tumor regions of GBM (areas exhibiting varying CE-T1W or CBV signal intensities), with the following four principles: (a) Regions with high or low signal intensities on CE-T1W and CBV images were selected, categorizing areas into four types: T1C-high/CBV-high, T1C-high/CBV-low, T1C-low/CBV-high, and T1C-low/CBV-low (some patients had only two types of regions); (b) solid tumor tissue regions were identified on the basis of PWI sequences, avoiding cystic cavities; (c) when feasible, sites were selected to include both central and peripheral regions of the tumor; and (d) on the basis of the surgical approach and craniotomy method, puncture trajectories were planned to avoid interference between sampling sites, minimize surgical duration, and prioritize superficial-first accessibility. We tried to avoid the undesirable effect of reduced tumor purity by sampling from the lateral portion of the radiographically visible tumor rather than from the invasive edge. The signal characteristics used in the study were extracted from the entire ROI, covering the whole tumor. However, surgical feasibility and the need to minimize interference between multiple sampling sites limit the number of obtainable samples per tumor. All imaging data are legally available upon request. Using intraoperative neuronavigation under general anesthesia, we collected samples through puncture biopsy. Multisite sampling was conducted from shallow to deep based on spatial position while ensuring an intact endocranium to minimize navigation errors caused by tissue drift. The sampling time for each patient was kept within 20 min. The samples obtained were then immersed in RNA Stabilization Reagent for Animal Tissue (Beyotime R0118). Peripheral blood samples were simultaneously taken as normal controls. A total of 35 samples were obtained from 6 patients (6 whole blood control samples and 29 tumor samples). All participating patients recovered well after surgery.
Whole-exome sequencing
Sample DNA was extracted from tissues and peripheral blood with an AllPrep DNA/RNA Mini Kit (Qiagen 80204). The fragment size, mass and total concentration of the DNA were subsequently measured. Library preparation, exon capture and sequencing were performed by Genetron Health (Beijing, China). The raw data were subjected to quality filtering to remove low-quality reads, and adapter sequences were sequenced with Trimmomatic27 to obtain clean data. The clean data were then aligned to the human reference genome (hg19/GRCh37) with the Burrows–Wheeler Alignment (BWA) (http://broadinstitute.github.io/picard/), followed by base correction and local alignment via GATK v4.1 to increase the overall alignment quality. Single nucleotide variants (SNVs) were identified with Mutect2 v4.128,29, whereas short fragment insertions and deletions (InDels) were identified with Strelka2 v2.9.1029. Variant effect predictor (VEP v95) was employed for annotating SNVs and InDels30, which then underwent functionality filtering with the following criteria: (a) Retained variants: missense, nonsense, frameshift, splice-site (± 2 bp), and start/stop-loss mutations; (b) excluded variants: synonymous, intronic (non-splice), intergenic, and benign variants (PolyPhen-2 score < 0.85); and (c) database filtering: variants with gnomAD v2.1 MAF ≥ 0.1% and prioritized COSMIC v90 cancer-associated mutations were removed. Finally, the Integrative Genomics Viewer (IGV v2.8.2) (https://github.com/igvteam/igv/issues) was used to confirm the remaining SNVs and InDels to obtain a final list of mutations.
Image processing and identification of clusters on the basis of the CBV and CE-T1W images
The CE-T1W brain MR images of 29 samples were imported into 3D Slicer 5.5.0 software31. Expert neuroimaging specialists delineated the regions of glioma (ROGs), with intraoperative neuronavigation-guided sampling sites (positioning error ≤ 2 mm). Using the PyRadiomics package (version 3.0.1), histogram features were extracted from CBV and CE-T1W images within the defined ROGs32. The input features used for consensus clustering were only the average signal intensity value (CBV_mean and T1C_mean) of each sampling ROG in the CBV images and the CE-T1W images. These radiomics features were subjected to unsupervised consensus clustering using the CBV and CE-T1W images via the R package ConsensusClusterPlus (v1.66.0)33:
-
(a)
number of repetitions = 50 bootstraps;
-
(b)
pItem = 0.8 (resampling 80% of any sample);
-
(c)
pFeature = 1 (resampling 100% of each sample of the input radiomics features);
-
(d)
clusterAlg = “km”;
-
(e)
Distance = “euclidean”.
Three distinct clusters were identified on the basis of the following aspects: (a) average pairwise consensus matrix within clusters; (b) delta plot of the relative change in the area under the CDF curve; and (c) average silhouette distance. Additionally, the ROG volume, centroid coordinates, and statistical features of the ROGs were calculated with the segment statistics package in 3D Slicer.
Quantification of imaging feature heterogeneity and GMH
CE-T1WI can clearly display the shape, size, boundaries, and complex information of the internal tissue structure and cell composition of the tumor. We quantified imaging feature heterogeneity using the variance in signal intensity on CE-T1WI within the sampling site ROI. This metric was chosen because CE-T1WI is clinically pivotal for GBM diagnosis and treatment assessment and exhibits characteristic patterns that reflect intratumoral heterogeneity. Critically, CE-T1WI signal variance correlates with biological heterogeneity: high signal areas indicate blood‒brain barrier disruption due to immature vasculature from angiogenic tumor subclones, whereas low signal areas may represent invasive tumor cells along mature vessels or quiescent stem-like populations34. Therefore, we use the CE-T1WI variance within the sampling site range guided by neuronavigation to quantify the imaging feature heterogeneity of this sampling site; these features are considered to be loosely positively correlated.
Defining intratumor gene mutational heterogeneity
The distribution of gene mutations in the samples of the subjects differed. We categorized the types of gene mutations into three groups: shared gene mutations (SGMs), crossover gene mutations (CGMs), and unique gene mutations (UGMs). CGMs are found in more than one but not all samples, and UGMs are found in only one sample from the same subject. These two types of mutations show different features and steps of the heterogeneous evolution of GBMs; therefore, they are called private gene mutations (PGMs). GMH can be calculated as PGM/(PGM + SGM).
Defining gene mutation coverage (GMC)
The gene mutation detected in a sample represents only a proportion of the overall gene mutations present in the subject. To measure the efficiency of detecting genetic mutations in a single sample of the subject, we defined the GMC of the subjects as the proportion of the number of genetic mutations detected in one or more samples of the subject to the total number of genetic mutations detected in all samples of the subject.
Cancer driver gene mutation (CDGM) analysis
CDGMs were initially identified from an integrated glioma driver gene database (n = 687 genes) derived from prior pan-glioma studies35. From this pool, 59 genes harboring mutations in our cohort were classified as CDGMs. To assess clonality, the variant allele frequency (VAF) was calculated for each mutation. On the basis of established thresholds36mutations with VAF ≥ 30% were designated early clonal events, whereas those with VAF < 30% were considered late subclonal events or passenger mutations. Early clonal events were considered to indicate CDGMs. All VAF distributions are provided in Supplementary Table 2.
Evaluation of the sparsity of multisite sampling
We measure sparsity in terms of the size of the sphere that can cover all the sampling sites; a larger sphere (the larger diameter of the sphere) indicates a sparser distribution of sampling sites. For two-site sampling, sparsity was quantified by calculating the distance between two sites via 3D Slicer 5.5.0 software. For three-site sampling, sparsity was determined on the basis of the diameter of the minimum coverage circle, which was calculated with 3D Slicer 5.5.0 software. After mathematical analysis, in cases where the three sites formed acute or right triangles, the diameter of the minimum coverage circle corresponded to the diameter of its circumcircle. However, if an obtuse triangle was formed or the three sites were collinear, then the diameter of the minimum coverage circle was set to the longest distance between three points after connecting them in pairs. A schematic diagram is provided to illustrate this concept (Figure S4F).
Statistical analysis
All the data were analyzed with IBM SPSS version 26, R software version 4.3.3, and all the graphs were prepared in GraphPad Prism version 9.5.1. The threshold for statistical significance was set at P < 0.05. Correlation analysis was performed with simple linear regression. The normality of the data was assessed with the Shapiro‒Wilk normality test. Paired or unpaired data between two groups were compared with paired t tests or independent-samples t tests, respectively. Multiple comparisons with normally distributed data were conducted with one-way ANOVA followed by Tukey’s multiple comparison test.
Results
GBM demonstrates significant intratumoral genetic feature heterogeneity according to imaging features
A total of 29 qualified samples were obtained from 6 patients (Supplementary Table 1), and a comprehensive analysis of the results of WES data revealed 596 gene mutations (Supplementary Table 2). Significant GMH was observed among patients and among samples from the same patient (Supplementary Table 3). On the basis of the presence of GMH in patient samples, gene mutations were categorized into SGMs, CGMs, and UGMs and further classified into PGMs and SGMs. MUC16, RPTN, and TP53 were the three genes that are most frequently mutated in GBM. The total number of gene mutations differed significantly across different patient samples (Supplementary Fig. 1A), ranging from 77 to 178, with an average of 103.5 ± 37.59. The GMC of all the samples ranged from 5.83 to 82.93%, with an average of 54.87 ± 18.02% (Supplementary Fig. 1B). The proportions of SGMs in individual patients ranged from 1.95 to 52.27%, with an average of 26.73 ± 20.82%. Similarly, the proportions of SGMs in individual samples varied from 3.57 to 86.79%, with an average of 45.83 ± 28.86%. The proportions of CGMs and UGMs in individual samples exhibited similar trends, ranging from 0.00 to 88.46% and averaging 38.18 ± 27.23% for CGMs and ranging from 1.85 to 58.97% and averaging 15.98 ± 11.51% for UGMs (Fig. 1B). Notably, both PGMs and SGMs demonstrated interpatient as well as intratumoral heterogeneity. We calculated the proportion of PGMs of patients (Supplementary Fig. 1C) and extracted the variance values within the ROG on CE-T1WI (Supplementary Fig. 1D). Importantly, our study revealed a significant positive correlation between the proportion of PGMs and the variance measurements (R = 0.86, P = 0.028), suggesting a potential relationship between genetic heterogeneity and imaging feature heterogeneity (Fig. 1C). These results indicate that there is high genetic heterogeneity within GBM, and this heterogeneity is highly correlated with imaging feature heterogeneity.
Characterization of the intratumoral GMH in different anatomical subregions
To explore potential intratumoral genetic mutation heterogeneity (GMH) in central versus marginal regions of GBM, we selected patients whose tumor imaging profiles, analyzed via a simplified model approximating tumor growth dynamics, suggested spherical growth patterns37 (Supplementary Table 4). Molecular phylogenetic analysis revealed no significant correlation between the number of gene mutations in individual samples and the distance from the tumor centroid to each sample (R = -0.28, P = 0.34) (Fig. 1D). Notably, we observed a positive correlation between the proportion of PGMs and the distance from the tumor centroid to each sample (R = 0.71, P = 0.0045) (Fig. 1E). Furthermore, observations from GBM sampling sites suggested a tendency toward increased PGM frequency in marginal samples (Supplementary Fig. 1E). These findings might tentatively suggest the presence of a high level of GMH in the marginal region, which may align with the invasive and infiltrating phenotype observed in GBM38. Notably, this observed spatial heterogeneity might not strictly mirror evolutionary hierarchies, as emerging evidence points toward possible discordance between anatomical zonation and clonal evolution in GBM39,40,41.
Many key gene mutation events are present among PGMs
We conducted a comprehensive screening and analysis of CDGMs35 across all samples, resulting in the detection of a total of 30 CDGMs. In terms of gene mutation types, missense variants constituted the majority of CDGMs. Specifically, 15 CDGMs were SGMs, whereas 11 were PGMs (including CGMs and UGMs) within any given sample. Additionally, MUC16, SETD2, TP53 and WDR63 presented with either SGM or PGM patterns across different samples (Fig. 2A). Notably, there was considerable variation in the specific CDGM profiles of individual patients, with an average PGM rate of 47.10% (Supplementary Fig. 2A). We also investigated drug target gene mutations (DTGMs) and detected a total of 4 genes, including PIK3CA, PTEN, TP53, and EGFR. Among them, PIK3CA and PTEN mutations were SGMs (50%), EGFR mutations were PGMs (25%), and TP53 mutations were either SGMs or PGMs across different samples. Molecular phylogenetic analysis revealed that most CDGMs clustered on distinct branches within the molecular phylogenetic tree, indicating their heterogeneity (Supplementary Fig. 2B). These results indicate that high GMH was observed in the subclonal development of GBM, emphasizing the importance of multisite sampling.
High PWI-CBV value regions cover more key mutation events from PGMs. (A), Overview of the gene mutation profiles grouped by patients. (B), Difference in the mean CBV value according to unsupervised consensus clustering based on the CBV and CE-T1W images (n = 29). (C), Difference in the number of UGMs according to unsupervised consensus clustering based on the CBV and CE-T1W images (n = 29). (D), CDGM distribution according to unsupervised consensus clustering based on the CBV and CE-T1W images. E, DTGM distribution according to unsupervised consensus clustering based on the CBV and CE-T1W images. CBV, cerebral blood volume; CE-T1W contrast-enhanced T1-weighted; UGM, unique gene mutation; CDGM, cancer driver gene mutation; DTGM, drug target gene mutation.
High PWI-CBV regions included more key gene mutation events than low PWI-CBV regions
To further explore the associations between imaging features and intratumoral GMH in GBM, unsupervised consensus clustering was performed on the 29 samples via CBV and CE-T1W images (Supplementary Table 5). The clustering result, with k = 3, was visualized with a heatmap (Supplementary Fig. 3). Three distinct GBM sample clusters were identified on the basis of their CBV values and named cluster CBVhigh, cluster CBVmedium and cluster CBVlow (Fig. 2B). The number of UGMs in the samples in cluster CBVhigh was significantly greater than that in the samples in cluster CBVmedium (P = 0.041) and cluster CBVlow (P = 0.0091) (Fig. 2C), indicating that elevated CBV is associated with heightened intratumoral genetic heterogeneity. On the basis of established glioma evolution theory, we propose that this phenomenon may be driven by increased independent subclonal development and differentiation propensity within vascular-rich niches42,43. Furthermore, we analyzed the distributions of CDGMs and DTGMs among the three clusters of GBM samples. Cluster CBVhigh accounted for 29 CDGMs, cluster CBVmedium accounted for 29 CDGMs, and cluster CBVlow accounted for 23 CDGMs. Collectively, 71.60% of CDGMs were detected in clusters CBVhigh and CBVmedium (Fig. 2D). This suggests that molecular pathological events linked to dominant subclones in later GBM development are concentrated in regions with higher PWI-CBV values. Additionally, among all the DTGMs identified in our analysis, those in the CBVhigh cluster and the CBVmedium cluster included p53 mutations, which are common in GBM (Fig. 2E). These findings suggest that neurosurgeons can utilize preoperative multimodal MRI to target regions displaying higher PWI-CBV values during surgery to potentially obtain more DTGM-rich tissue samples for improved treatment guidance.
Multisite sampling of GBM impacts the GMC and the detection of PGMs
GMC is a very important feature assessed in GBM sampling and affects the quality of the sampling results. After identifying the relationships between imaging features and intratumoral GMH, to further optimize the intraoperative sample acquisition strategy for GBM, we compared the GMC values for single-site and four-site sampling (Fig. 3A and Supplementary Table 6). As the number of sampling sites increased, extremely significant differences were observed in the GMC among the single-site, two-site and three-site sampling methods (one vs. two sites: P = 1.1 × 10− 6; two vs. three sites: P = 4.4 × 10−10; one vs. three sites: P = 4.8 × 10− 12); however, the difference between the GMC values for the three-site and four-site sampling methods was relatively small. For single-site sampling, we observed a Gaussian distribution pattern in this cohort of GBMs (61.7 ± 10.82%), with nearly 58.62% of the sites exhibiting a GMC between 60% and 70% (Supplementary Fig. 4A, B).
Nevertheless, no significant difference in the GMC was found among the different multimodal imaging feature clusters (Fig. 3B), and there was no correlation between the GMC and the distance from the tumor centroid to the sample (Supplementary Fig. 4C). These findings confirm that the intratumoral variance in the GMC is high and that no guidance from morphological or imaging evidence can significantly improve the GMC with single-site sampling.
The GMC values of the two-site and three-site sampling methods conformed to Gaussian distributions (75.50 ± 5.20% for two-site sampling and 86.44 ± 9.99% for three-site sampling) (Supplementary Fig. 4D, E). The GMC obtained with four-site sampling did not follow a Gaussian distribution and was deemed inadequate for further analysis. In contrast to single-site sampling, the absolute and relative positions of the sampling sites in intraoperative multisite sampling are often crucial considerations for surgeons.
Therefore, in this study, a method was developed to measure the absolute sparsity between sampling points: the sparsity of two-site sampling was quantified by calculating the distance between the two sites, and the sparsity of three-site sampling was determined on the basis of the minimum coverage circle diameter (Figure S4F). Interestingly, we found that the sparsity of two-site and three-site sampling was not correlated with the GMC (two: R = -0.18, P = 0.19; three: R = -0.067, P = 0.63) (Fig. 3C, D). However, we did identify a positive correlation between the sparsity of two-site and three-site sampling and the number of PGMs (two: R = 0.38, P = 0.0042; three: R = 0.46, P = 0.00046) (Fig. 3E, F). These findings suggest that increasing the sparsity between two or three sites during sampling can improve the detection of PGMs.
“Vegetation” hypothesis conceptualize the detection of shared (SGMs) and private (PGMs) mutations during sampling
We propose a “vegetation” hypothesis to conceptualize the detection of shared (SGMs) and private (PGMs) mutations during sampling (Fig. 3G). This model analogizes SGMs to uniform “soil” present throughout the tumor, while PGMs are akin to variably distributed “vegetation” growing within that soil. In highly heterogeneous GBM (depicted as thin soil with dense, patchy vegetation), sampling any single site yields high PGM detection but low GMC (limited soil coverage). Conversely, in less heterogeneous GBM (thick soil with sparse vegetation), single-site sampling yields high GMC but low PGM detection. Thus, the model illustrates the inherent trade-off between GMC and PGM detection efficiency in a single sample, driven by the underlying heterogeneity. Therefore, multisite sampling is necessary to accurately identify SGMs and fully detect PGMs in patients with highly heterogeneous GBM.
Multisite sampling in GBM does not affect the GMC but aids in the detection of PGMs. (A), Differences in gene mutation coverage in multisite sampling. (B), Difference in gene mutation coverage following unsupervised consensus clustering on the basis of the CBV and CE-T1W images (n = 29). (C), Correlation analysis between gene mutation coverage and the distance between the sites in two-site sampling (n = 56). (D), Correlation analysis between the gene mutation coverage and the radius of the minimum covering circle (n = 54). (E), Correlation analysis between the number of PGMs and the distance between the sites of two-site sampling (n = 56). (F), Correlation analysis between the number of PGMs and the radius of the minimum covering circle (n = 54). G, Schematic diagram of the “vegetation” hypothesis for detecting homogeneity and heterogeneity in an intraoperative sample; the soil (brown) refers to SGMs, and vegetation refers to PGMs (specifically, trees, bushes and grasses refer to PGMs with high, moderate and low frequencies, respectively). The amount of soil and vegetation represents the number of corresponding gene mutations. An uneven “vegetation” distribution means that the detection of PGM will be different for each sample. GMC, gene mutation coverage; UGM, unique gene mutation; PGM, private gene mutation; GBM, glioblastoma; GMH, gene mutation heterogeneity.
Discussion
In our study, WES was integrated with MRI to map the intratumoral distribution of genetic-imaging feature heterogeneity in GBM patients. The abundance of gene mutations was not correlated with the centripetal structure of the selected samples, which exhibited symmetrical and simple developmental morphology, suggesting that the evolution and development of glioma cells do not occur solely from an anatomical perspective. Additionally, we observed a significant presence of PGMs at the edge (regions distal to the glioma centroid) of the gliomas, indicating that this particular subregion may exhibit heterogeneity. Coincidentally, a previous report also demonstrated that nonenhanced regions of GBM lesions presented with a greater burden of unique mutations, similar to our UGMs44. This conclusion further supports the existence of high GMH at the invasive front of GBM, suggesting that glioma cells in this region may be more malignant. At the molecular biology level, genetic mutations can reflect intratumoral heterogeneity in GBM. During the heterogeneous development of GBM, distinct subregions exhibit unique CDGM and DTGM profiles. Intratumoral heterogeneity can lead to diversity in tumor cell subclones, which facilitates the selection and generation of dominant clones in GBM under therapeutic pressure, resulting in GBM treatment resistance. While extensive research has focused on understanding GBM heterogeneity, insufficient attention has been given to its homogeneity. From a clinical perspective, it is preferable to identify drug targets within SGMs whenever possible. Targeting PGMs (present only in subclones) is related to a risk of selecting resistant subclones lacking the mutation, leading to treatment failure. Although current molecular targeted therapies for GBM show limited efficacy and are primarily used as second- or third-line treatments following the failure of first-line therapy, we recommend incorporating pathological information from multiple sampling sites to identify DTGMs within SGMs and detecting PGMs to help identify subclone-specific targets, avoiding futile broad-spectrum therapies or intensified combination therapy; thus, the two approaches are complementary.
Building on this “vegetation” hypothesis, we further evaluate its clinical implications for optimizing intraoperative sampling. The primary objective of intraoperative biopsy for GBM is twofold: to comprehensively identify SGMs (for pan-tumor targets) and to maximize PGM detection (for subclone-specific targets), thereby supporting stratified therapeutic strategies. Through a reasonable and evidence-based sampling plan, we can improve the efficiency of identifying SGMs to select targeted drugs more accurately, increase the number of detected PGMs to inform subclone-targeted therapies, and minimize the trade-off between GMC and PGM detection to maximize the benefits of sampling. Standardizing the location and quantity of intraoperative samples is crucial for managing GBM with high intratumoral heterogeneity. Future studies with larger cohorts should quantitatively compare CBV-targeted and RANO-standardized sampling in terms of GMC, PGM detection rate, and sampling bias reduction. This will clarify the optimal scenarios for each approach: RANO for homogeneous tumors and CBV guidance for heterogeneously enhancing lesions. Notably, our CBV-guided three-site sampling strategy does not contradict but complements the RANO recommendations. The RANO approach emphasizes systematic sampling across perpendicular axes covering core and edge regions—a robust method for gliomas with low imaging feature heterogeneity. However, for tumors exhibiting marked PWI-CBV heterogeneity (as in our cohort), initiating sampling in high-CBV regions may enhance the detection of spatially restricted mutations without deviating from the spatial distribution principles of RANO.
In our study, we considered the GMC and detection of PGMs to be the two most important sampling guidance factors. We found that the detection of PGMs depends on the sampling sites with high PWI-CBV values and spatial separation between the sampling sites. The GMC is related only to the number of sampling sites, and increasing the number of sampling sites is necessary for improving GMC detection; however, this should be limited in minimizing patient injury. Three-site sampling achieved a GMC of 86.44 ± 9.99% in our study; the difference in the GMC between the three-site and four-site sampling approaches was minimal; and three-site sampling resulted in substantially reduced injury to the patient. Considering these factors, we propose a three-site strategy to obtain samples from three intratumoral regions with the highest PWI-CBV values. If feasible, a relatively greater spatial separation between the sampling sites may enhance the detection of PGMs.
Limitations
This study has several limitations that warrant consideration. First, the small cohort size (6 patients, 29 samples) restricts the statistical power for subgroup analyses (e.g., molecular subtypes or growth patterns) and necessitates the validation of our CBV-guided tripartite sampling strategy in larger cohorts. Second, while MRI-based targeting has improved actionable mutation detection, the clinical utility remains constrained by the limited efficacy of contemporary targeted therapies in first-line GBM management; consequently, our approach primarily serves recurrence treatment planning rather than immediate therapeutic changes. Third, the proposed optimal sampling site number (n = 3) was inferred from diminishing GMC returns but requires verification through denser spatial profiling, as maximal 4-site sampling precluded saturation analysis. Finally, the absence of transcriptome and single-cell sequencing data limits our ability to characterize expression-level heterogeneity and cellular-resolution features. Future studies should expand cohort sizes and integrate multiomics protocols to address these gaps.
Conclusions
In this original research, we observed a positive correlation between the intratumoral GMH of GBM and sampling sparsity as well as the PWI-CBV value. These findings indicate that the PWI-CBV can serve as a reliable marker for guiding intraoperative sampling to assess intratumoral GMH comprehensively and increase sampling quality. On the basis of these conclusions, we propose a PWI-CBV-guided optimization strategy for the RANO sampling framework. This strategy prioritizes high-CBV regions within systematically distributed sites, potentially enhancing mutation detection in radiologically heterogeneous GBMs; however, this approach needs to be validated in clinical practice.
Data availability
Sequence data that support the findings of this study have been deposited in the National Genomics Data Center of China (https://ngdc.cncb.ac.cn/) with the primary accession code PRJCA027955. Other data included in this study are available upon request by contact with the corresponding author.
Abbreviations
- CBV:
-
Cerebral blood volume
- CDGM:
-
Cancer driver gene mutation
- CE-T1W:
-
Contrast-enhanced T1-weighted
- CGM:
-
Crossover gene mutation
- DTGMs:
-
Drug target gene mutations
- GMC:
-
Gene mutation coverage
- GMH:
-
Gene mutation heterogeneity
- PGM:
-
Private gene mutation
- GBM:
-
Glioblastoma
- MRI:
-
Magnetic resonance imaging
- PWI:
-
Perfusion-weighted imaging
- ROGs:
-
Regions of glioma
- SGM:
-
Shared gene mutation
- UGM:
-
Unique gene mutation
- WES:
-
Whole-exon sequencing
References
Lapointe, S., Perry, A. & Butowski, N. A. Primary brain tumours in adults. Lancet 392, 432–446. https://doi.org/10.1016/s0140-6736(18)30990-5 (2018).
Berger, T. R., Wen, P. Y., Lang-Orsini, M. & Chukwueke, U. N. World health organization 2021 classification of central nervous system tumors and implications for therapy for Adult-Type gliomas: A review. JAMA Oncol. 8, 1493–1501. https://doi.org/10.1001/jamaoncol.2022.2844 (2022).
Miller, K. D. et al. Brain and other central nervous system tumor statistics, 2021. CA Cancer J. Clin. 71, 381–406. https://doi.org/10.3322/caac.21693 (2021).
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the united States in 2012–2016. Neuro Oncol. 21, v1–v100. https://doi.org/10.1093/neuonc/noz150 (2019).
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the united States in 2013–2017. Neuro Oncol. 22, iv1–iv96. https://doi.org/10.1093/neuonc/noaa200 (2020).
Stummer, W. & Kamp, M. A. The importance of surgical resection in malignant glioma. Curr. Opin. Neurol. 22, 645–649. https://doi.org/10.1097/WCO.0b013e3283320165 (2009).
McGirt, M. J. et al. Independent association of extent of resection with survival in patients with malignant brain Astrocytoma. J. Neurosurg. 110, 156–162. https://doi.org/10.3171/2008.4.17536 (2009).
Ma, R., Taphoorn, M. J. B. & Plaha, P. Advances in the management of glioblastoma. J. Neurol. Neurosurg. Psychiatry. 92, 1103–1111. https://doi.org/10.1136/jnnp-2020-325334 (2021).
Wu, J. S. et al. Current status of cerebral glioma surgery in China. Chin. Med. J. (Engl). 124, 2569–2577 (2011).
Horbinski, C., Berger, T., Packer, R. J. & Wen, P. Y. Clinical implications of the 2021 edition of the WHO classification of central nervous system tumours. Nat. Rev. Neurol. 18, 515–529. https://doi.org/10.1038/s41582-022-00679-w (2022).
van der Meulen, M., Ramos, R. C., Mason, W. P., Von Deimling, A. & Maas, S. L. N. Opinion & special article: glioma classification: how to interpret molecular markers in a diffuse glioma pathology report. Neurology 99, 903–908. https://doi.org/10.1212/wnl.0000000000201262 (2022).
Chai, R. et al. Molecular pathology and clinical implications of diffuse glioma. Chin. Med. J. (Engl). 135, 2914–2925. https://doi.org/10.1097/cm9.0000000000002446 (2022).
Reifenberger, G., Wirsching, H. G., Knobbe-Thomsen, C. B. & Weller, M. Advances in the molecular genetics of gliomas - implications for classification and therapy. Nat. Rev. Clin. Oncol. 14, 434–452. https://doi.org/10.1038/nrclinonc.2016.204 (2017).
Huse, J. T. & Aldape, K. D. The evolving role of molecular markers in the diagnosis and management of diffuse glioma. Clin. Cancer Res. 20, 5601–5611. https://doi.org/10.1158/1078-0432.Ccr-14-0831 (2014).
Huse, J. T. & Holland, E. C. Targeting brain cancer: advances in the molecular pathology of malignant glioma and Medulloblastoma. Nat. Rev. Cancer. 10, 319–331. https://doi.org/10.1038/nrc2818 (2010).
Nicholson, J. G. & Fine, H. A. Diffuse glioma heterogeneity and its therapeutic implications. Cancer Discov. 11, 575–590. https://doi.org/10.1158/2159-8290.Cd-20-1474 (2021).
Colbassani, H. J., Nishio, S., Sweeney, K. M., Bakay, R. A. & Takei, Y. CT-assisted stereotactic brain biopsy: value of intraoperative frozen section diagnosis. J. Neurol. Neurosurg. Psychiatry. 51, 332–341. https://doi.org/10.1136/jnnp.51.3.332 (1988).
Yu, K. et al. Surveying brain tumor heterogeneity by single-cell RNA-sequencing of multi-sector biopsies. Natl. Sci. Rev. 7, 1306–1318. https://doi.org/10.1093/nsr/nwaa099 (2020).
Bex, A. & Mathon, B. Advances, technological innovations, and future prospects in stereotactic brain biopsies. Neurosurg. Rev. 46, 5. https://doi.org/10.1007/s10143-022-01918-w (2022).
Karschnia, P. et al. A framework for standardised tissue sampling and processing during resection of diffuse intracranial glioma: joint recommendations from four RANO groups. Lancet Oncol. 24, e438–e450. https://doi.org/10.1016/s1470-2045(23)00453-9 (2023).
Hu, L. S., Hawkins-Daarud, A., Wang, L., Li, J. & Swanson, K. R. Imaging of intratumoral heterogeneity in high-grade glioma. Cancer Lett. 477, 97–106. https://doi.org/10.1016/j.canlet.2020.02.025 (2020).
Fan, H. et al. Artificial intelligence-based MRI radiomics and radiogenomics in glioma. Cancer Imaging. 24, 36. https://doi.org/10.1186/s40644-024-00682-y (2024).
Shiroishi, M. S., Boxerman, J. L. & Pope, W. B. Physiologic MRI for assessment of response to therapy and prognosis in glioblastoma. Neuro Oncol. 18, 467–478. https://doi.org/10.1093/neuonc/nov179 (2016).
Roodakker, K. R. et al. Region-by-region analysis of PET, MRI, and histology in En bloc-resected oligodendrogliomas reveals intra-tumoral heterogeneity. Eur. J. Nucl. Med. Mol. Imaging. 46, 569–579. https://doi.org/10.1007/s00259-018-4107-z (2019).
Zhang, L. et al. Multicenter clinical radiomics-integrated model based on [(18)F]FDG PET and multi-modal MRI predict ATRX mutation status in IDH-mutant lower-grade gliomas. Eur. Radiol. 33, 872–883. https://doi.org/10.1007/s00330-022-09043-4 (2023).
van Santwijk, L., Kouwenberg, V., Meijer, F., Smits, M. & Henssen, D. A systematic review and meta-analysis on the differentiation of glioma grade and mutational status by use of perfusion-based magnetic resonance imaging. Insights Imaging. 13, 102. https://doi.org/10.1186/s13244-022-01230-7 (2022).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219. https://doi.org/10.1038/nbt.2514 (2013).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods. 15, 591–594. https://doi.org/10.1038/s41592-018-0051-x (2018).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17 https://doi.org/10.1186/s13059-016-0974-4 (2016).
Zhou, L. et al. Clinical application of 3D slicer combined with sina/mosocam multimodal system in preoperative planning of brain lesions surgery. Sci. Rep. 12, 19258. https://doi.org/10.1038/s41598-022-22549-7 (2022).
van Griethuysen, J. J. M. et al. Computational radiomics system to Decode the radiographic phenotype. Cancer Res. 77, e104–e107. https://doi.org/10.1158/0008-5472.Can-17-0339 (2017).
Wilkerson, M. D. & Hayes, D. N. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573. https://doi.org/10.1093/bioinformatics/btq170 (2010).
Kunimatsu, A. et al. Texture analysis in brain tumor MR imaging. Magn. Reson. Med. Sci. 21, 95–109. https://doi.org/10.2463/mrms.rev.2020-0159 (2022).
Yang, K. et al. Glioma targeted therapy: insight into future of molecular approaches. Mol. Cancer. 21, 39. https://doi.org/10.1186/s12943-022-01513-z (2022).
Boscolo Bielo, L. et al. Variant allele frequency: a decision-making tool in precision oncology? Trends Cancer. 9, 1058–1068. https://doi.org/10.1016/j.trecan.2023.08.011 (2023).
Ellingson, B. M. et al. Contrast-enhancing tumor growth dynamics of preoperative, treatment-naive human glioblastoma. Cancer 122, 1718–1727. https://doi.org/10.1002/cncr.29957 (2016).
Gillies, R. J. Cancer heterogeneity and metastasis: life at the edge. Clin. Exp. Metastasis. 39, 15–19. https://doi.org/10.1007/s10585-021-10101-2 (2022).
Percuoco, V. et al. Glioblastoma invasion patterns from a clinical perspective-a systematic review. Neurosurg. Rev. 47, 864. https://doi.org/10.1007/s10143-024-02944-6 (2024).
Rabah, N. & Mohand, A. Kravchenko-Balasha, N. Understanding glioblastoma signaling, heterogeneity, invasiveness, and drug delivery barriers. Int. J. Mol. Sci. 24 https://doi.org/10.3390/ijms241814256 (2023).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401. https://doi.org/10.1126/science.1254257 (2014).
O’Connor, J. P. et al. Imaging intratumor heterogeneity: role in therapy response, resistance, and clinical outcome. Clin. Cancer Res. 21, 249–257. https://doi.org/10.1158/1078-0432.Ccr-14-0990 (2015).
Frosina, G. Advancements in Image-Based models for High-Grade gliomas might be accelerated. Cancers (Basel). 16. https://doi.org/10.3390/cancers16081566 (2024).
Hu, L. S. et al. Integrated molecular and multiparametric MRI mapping of high-grade glioma identifies regional biologic signatures. Nat. Commun. 14, 6066. https://doi.org/10.1038/s41467-023-41559-1 (2023).
Acknowledgements
We would like to extend our heartfelt thanks to all the scholars who contributed to this study. This study was supported by the National Natural Science Foundation of China (No. 92059204 and 82272895) and the Natural Science Foundation of Chongqing Municipality (CSTB2022NSCQ-LZX0061). We thank Professor Zhong Peng of the Department of Pathology, Daping Hospital, for reviewing and further improving the pathological diagnosis of the patient. We also thank Qian Yu for her language assistance.
Author information
Authors and Affiliations
Contributions
Jun-han Wang: Conceptualization, Data curation, Formal analysis, Methodology, Visualization, and Writing – original draft. Ji Xia: Data curation, Formal analysis, Investigation, Methodology, Software, Visualization. Peng-fei Wu: Data curation, Investigation, Software. Liang Yi: Conceptualization, Funding acquisition, Project administration, Resources, Supervision, and Writing –review & editing. Shi-cang Yu: Conceptualization, Funding acquisition, Project administration, Resources, Supervision, and Writing –review & editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The study protocol was approved by the Ethics Committee of Daping Hospital (Project number: Medical Research Ethical Review (2024) NO.217) and registered in the Chinese Medical Research Registration and Filing Information System (CMR, medicalresearch.org.cn) (Registration No: MR-50-24-029389) and China Clinical Trial Registry (chictr.org.cn) (Registration No: ChiCTR2500103340). All procedures strictly adhered to the Declaration of Helsinki principles and relevant national ethical guidelines, with written informed consent obtained from all participants or their legally authorized representatives. All operations were rigorously executed in strict compliance with the pre-established study protocol.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.




Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Jh., Xia, J., Wu, Pf. et al. An optimized tissue sampling scheme guided by MRI features reveals intratumoral heterogeneity in glioblastoma. Sci Rep 15, 33194 (2025). https://doi.org/10.1038/s41598-025-17539-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-17539-4