Abstract
Mesophotic sponge grounds (MSG), which are known as local biodiversity hotspots along the Eastern Mediterranean, have been suggested to serve as refugia for threatened species from shallower habitats. However, to understand their role as a refuge, their connectivity to shallow habitats should be assessed. This study evaluates the genetic connectivity of the broadcasting sponge Axinella polypoides, which inhabits both shallow and mesophotic zones, across two scales: (1) geographically, between Eastern and Western Mediterranean populations, and (2) bathymetrically, between shallow and MSG populations along the Israeli coast. Using ddRADseq (N = 106, 2,295 SNPs), significant genetic differentiation was observed between Eastern and Western Mediterranean populations, whereas panmixia was identified in Eastern populations. A reduced dataset (N = 95, 5,400 SNPs) revealed subtle genetic substructure differences between shallow and mesophotic populations, particularly at Achziv. Moreover, shallow populations were generally less genetically divergent than mesophotic ones, except for Rosh-Carmel mesophotic population, which showed higher gene flow with shallow populations. Herzliya shallow population was identified as a key gene source. These results underscore the Eastern Mediterranean MSG’s role as refugia for A. polypoides, and possibly for species with similar life-history traits. Our findings emphasize the importance of protecting these habitats, especially given the increased anthropogenic and climate-driven pressures.
Similar content being viewed by others
Introduction
The Mediterranean Sea, a hotspot of biodiversity, hosts at least 17,000 species with high level (ca. 20%) of endemism1to which climate change poses a threat2,3,4. Sponges are one of the most prevalent phyla worldwide5and in the Mediterranean Sea in particular, being a dominant phylum in benthic ecosystems6,7. The Mediterranean sponge community is diverse and rich, with over 730 recorded species, nearly half (48%) are endemic8. The ecological importance of the sessile sponges is mainly due to two reasons: First, sponges are very active filter feeder organisms, that greatly contribute to the food web by mediating the transfer of organic matter and nutrients between the pelagic zone and the benthos, the so-called bentho-pelagic coupling9,10,11; and second, they are often considered as “environmental engineers” since they create complex structures, thus serving as habitats for other organisms12. However, like the general Mediterranean biodiversity, sponge diversity has been described as non-homogeneous, with a decreasing gradient of diversity and abundance from the northwest to the southeast, which contributed to the perception of the Levant basin as poor in species diversity and richness compared to the rest of this sea1,8,13.
During the last decade, rich sponge grounds have been discovered along the Israeli Mediterranean shore at mesophotic depths of 80–130 m14. Subsequent studies of these mesophotic sponge grounds (MSG) discovered they are hotspots of local biodiversity: 35 species were new to the known local fauna, many of them new to the Levant Basin, and three new to science. Moreover, at least 70 additional species have yet to be identified. However, among this relatively high richness of sponges, there is only a small overlap of species found in shallow and mesophotic communities14,15.
The MSG is dominated mainly by massive sponges such: Agelas oroides (Schmidt, 1864) and Irciniinds, as well as branched sponges such as Axinellids. Whereas the shallow habitats are mostly covered by algae and to a lesser extent by encrusting sponges (e.g. Crambe crambe (Schmidt, 1862), Phorbas spp.) and Axinella spp.14,16.
Compared to shallow areas, these MSG are characterized by more stable abiotic conditions, which may contribute to the observed increase in biodiversity in these habitats9,17. Located below the seasonal thermocline, the MSG, experience consistently lower and more stable temperatures- typically averaging around 18˚C18,19which are more favorable for many sponge species, as higher temperatures can limit their survival20,21,22. Additionally, light intensity is much lower in these areas (0.0005% at the lower mesophotic limit: ~198 m) compared to shallow habitats (13.3% up to 40 m), resulting in reduced competition with algae on the rocky substrate23,24and therefore the mesophotic habitats became biodiversity hotspot for benthic fauna, dominated by sponges (95% of the total live coverage14. Furthermore, compared to shallow habitats, the MSG are more distant from the terrestrial land and thus less exposed to anthropogenic disturbances14. Moreover, recently it was suggested that these MSG may serve as a refuge for some shallow-water species that suffer from the rising seawater temperature and possibly be a source of larvae for re-settlement in shallow waters once environmental conditions are favorable20,25.
To understand the role of these Eastern Mediterranean MSG as a refuge, it is necessary to provide baseline data regarding the level of genetic connectivity between the MSG and the shallow coast. In sponges the gene flow heavily depends on the reproductive strategy of the species, and it is also greatly influenced by prevalent oceanographic currents, as well as by the distance between the suitable substrates26,27. For example, released gametes and/or long-lived larvae, theoretically, can enable dispersal over greater distances compared to crawling larvae that live for a few hours and have limited mobility28. They are usually limited by larger-scale barriers. Specifically, the Almeria-Oran front, for instance, is a major driver of genetic separation between populations from the Atlantic Ocean and the Mediterranean Sea. Other fronts such as the Sicily Strait, the Otranto Channel, and the southern margin of the Aegean Sea, act as genetic breaks for several sessile animals27,29,30including sponges31,32. In contrast, sessile species with short pelagic larval duration phase, such as brooding corals, gorgonians or sponges, have more significant genetic breaks between localities, than species with higher motility abilities. However, their genetic differentiation is less influenced by the presence of oceanography discontinuities (i.e. fronts or physical barriers) than those who have highly mobile larvae or lifestyle27. Besides, the distribution of such species, in which the larval dispersal distance is shorter, is usually accomplished by the “stepping stones” pattern33,34,35. Importantly, different oceanographic fronts have been identified in the Mediterranean Sea playing a crucial role in isolating populations27. At a smaller scale along the Eastern Mediterranean Israeli shore, only a few studies examined connectivity of benthic shallow macrofaunal populations (e.g. cnidarians, bivalves36,37. Yet, none of them evaluated connectivity patterns with respect to deeper (MSG) populations. Two of these Israeli MSG have already been declared as Marine Reserves (MR): Rosh HaCarmel Sea and Rosh Hanikra-Akhziv Reserves, while several others are in various stages of approval or designation as MR (https://en.parks.org.il/). Nevertheless, the connectivity patterns between the MSG populations along the Israeli shore, and their connectivity to the shallow habitats, have not yet been assessed. Understanding these patterns is crucial for conservation, as it may determine whether these areas can serve as refuge.
In the current research we studied the connectivity and the genetic structure of wild populations of the Mediterranean widespread protected demosponge Axinella polypoides Schmidt, 1862, across the sea, from the Western (Spain and France) to the Eastern Mediterranean Israeli locations. The latter included much more samples taken from a wide depth range (15–130 m) along ~ 120 km (Fig. 1). Axinella polypoides is a common Mediterranean and East Atlantic Ocean species, and its populations along the Israeli shore represent the eastern edge of its documented distribution38. In Israel, it is one of only 12 species, that can be found both in the mesophotic and shallow habitats (~ 15–130 m23, and it is the third most abundant species in the MSG14. The reproductive strategy of A. polypoides has been recently described25. It is a gonochoristic broadcaster (i.e. oviparous) species, showing seasonality along the shallow depths of the Israeli shore, whereas it reproduces all year-round (with high fecundity levels) in the MSG25. Based on this reproductive strategy, we hypothesized that the extended duration of the planktonic embryonic development will lead to relatively long offsprings dispersal, resulting in high connectivity between spatially close populations, regardless of depth26.
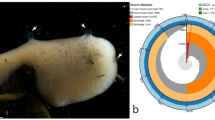
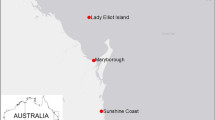
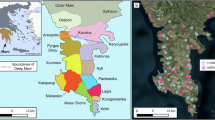
Sampling sites of A. polypoides. (A) Mediterranean Sea sampling sites: Catalonia, Marseille (each from one site, < 30 m depth) and the Israeli sites. (B) Map of the sampling sites along the Israeli coast. Rocky areas along the coastline are indicated as border-lined shadowed areas: Achziv - AC; Rosh-Carmel - R; Atlit – A; Herzliya - H. Small letters and color indicate the depth: shallow (< 30 m) – sh (blue); Medium-depth (~ 40 m) – m (yellow); and MSG - deep (80–130 m) – d (red). (C) An in situ picture of A. polypoides from Herzliya MSG (Israel).
Materials and methods
a. Study sites and sampling
A total of 116 A. polypoides specimens were collected from four sites along the Eastern Mediterranean Israeli coast (Fig. 1 and Table S1), and three depth zones at each of the site: mesophotic (80–130 m), middle-depth (35–40 m), and shallow (< 30 m). Atlit and Herzliya sites are the peaks of submerged sandstone ridges that stretch parallel to the Israeli Mediterranean shore and are characterized as patchy rocky habitats. Rosh-Carmel and Achziv are more continuous rocky sites, being extensions of terrestrial mountain ridges that span from the shore to deeper than 100 m depth39. The conservation status of the shallow sites at Herzliya and Atlit as marine reserves is currently ‘proposed’ and has not been approved yet. The Atlit and Herzliya mesophotic sites have been approved as MR but not declared yet. The only sampling sites from declared MR are the Achziv and Rosh-Carmel sites in both depths (shallow and mesophotic).
Samples of sponges from mesophotic sponge grounds were sampled by a Remotely Operated Vehicle (ROV; ECA-Robotics H800), on board the R/V Mediterranean Explorer. Middle-depth and shallow samples (~ 40 and 25 m, respectively) were collected by SCUBA diving. Tissue sections from the samples collected by SCUBA diving were small (up to 3 cm3, and individuals Collected by ROV were relatively small. To avoid sampling individuals that might originate from the same clone, all Collected specimens were at least 4 m apart.
In addition to the Eastern Mediterranean (Israel) sponges, we also included samples (n = 16) from the Western Mediterranean coasts of Catalonia (Spain) and Marseille (France) Collected by SCUBA diving down to 30 m depth (Fig. 1 and Table S1). The latter samples were used to analyze gene flow and infer long distance connectivity patterns between the Eastern and the Western parts of the Mediterranean Sea.
In all cases, the samples were preserved in 100% EtOH, replaced twice, and were kept at −20 °C until further processing. The morphological identification was done based on spicules composition and 3D arrangement within the sponge, studied with a stereo microscope (Olympus KL 200).
b. DNA extractions and ddRADseq library Preparation and sequencing
To assess the genetic connectivity between A. polypoidespopulations we employed “double digest Restricted site-associated DNA sequencing” (ddRADseq40;. DNA was extracted from all 116 samples, using the DNeasy Blood & Tissue kit (Qiagen, www.qiagen.com) following the manufacturer’s protocol, with minor modification in the final DNA elution step (eluted using 100ul elution buffer). Double-stranded DNA (dsDNA) was quantified using Qubit dsDNA HS assay (Life Technologies). ddRADseq libraries were performed for all samples40with some modifications following Combosch et al. (2017). All the protocol steps were conducted on random and heterogenic batches of samples to avoid batch effect. Genomic DNA was standardized to 500 ng and was digested using two high-fidelity restriction enzymes: Eco-RI and BfaI (New England Biolabs) for 6 h at 37 °C. Digested fragments were cleaned by manually pipetting with Agencourt AMPure beads (1.5x volume ratio, Beckam Coulter) and subsequently quantified afterwards with Qubit dsDNA HS assay (Life Technology). The resulting genomic fragments were standardized to 200 ng and ligated to custom-made P1 and P2 adaptors that contained sample-specific barcodes and primer annealing sites. Barcoded individuals were pooled into libraries and were cleaned by manual pipetting using AMPure beads (1.5x volume ratio). Using a Blue Pippin Prep (Sage Science) the libraries were size-selected to a range of 200–400 bp. PCR amplification was performed to each library, using the Phusion HF polymerase (Thermo Scientific), with a different set of PCR primers for each library. The PCR program was: 98 °C/30 s – (98 °C/10 s – 65 °C/30 s – 72 °C/1.5 min) x 12 cycles – 72 °C/10 min. The PCR products were cleaned by manually pipetting with Agencourt AMPure beads (1.5x volume ratio) and quantified using a Qubit dsDNA HS assay. Quality checks were conducted to all libraries with a TapeStation 2200 for genomic DNA (Agilent Technologies). Libraries were normalized based on their concentration and two pools were created. Libraries were pair-end sequenced (2 × 150 bp) on the Illumina NovaSeq 6000 at Novagene Europe (Cambridge, UK).
c. Locus assembly and outlier filtration
Sequences quality was examined using Fastqc42. Filtering by quality and locus assembly were conducted with the Stacks pipeline version 2.61 45. The resulting sequences were DNA fragments with two appropriate restriction enzyme cut sites that were selected, amplified, and sequenced (RAD-tags). These reads were processed using process_radtags, which refines raw RAD-tags by trimming low-quality reads, eliminating those with uncalled bases, and discarding reads lacking complete barcodes or restriction cut sites. The rescue feature (-r) in process_radtags was used to recover minimally diverged barcodes and RAD-tags, with only two mismatches allowed (-adapter_mm = 2). The trimming feature (-t) was used to truncate low quality reads at the 3’-end, to a read length of 145 bp, so that to increase confidence level in the Single Nucleotide Polymorphism (SNP) calling. After these primary filtrations, 703,175,678 loci were retained, out of the initial 877,673,690 raw reads. Because five individuals (originated from different studied sites) were found to have low quantity of reads, they were filtered out from further analyses, and a total of 111 individuals were included, with 645,901,476 remaining loci (74% from the initial reads), ranging from 337,653 to 39,294,555 with an average of 5,988,963 loci per individual.
Optimization tests were conducted following previous work44,45to determine the optimal Stacks parameters. In brief, each test was performed on five sets of three randomly selected individuals. Parameters that were not tested were set to their default values, with r = 0.8 for all tests. Based on the number of assembled loci, polymorphic loci, SNPs, and the coverage, the chosen parameters were: M = 2, m = 3 and n = 1.
We used the Matrix condenser interface to visualize and assess missing data at the individual level (https://bmedeiros.shinyapps.io/matrix_condenser/)46. Individuals with more than 35% missing data were removed, resulting in the exclusion of five additional samples and a final dataset of 106 samples for downstream analysis. We considered two separate datasets: (A) The entire dataset including 106 individuals from all 14 sites (Catalonia, Marseille and the 12 Israeli sites), and (B) That included 95 samples, all from the Israeli coast, in order to increase the number of shared SNPs in the specimens from the Eastern Mediterranean. The populations included for each dataset are presented in Table S1. In dataset B, shallow and medium depths were both considered as shallow depth (sh) after the results observed for dataset A, since the physical distance between them was very small both horizontally (average of 4.3 km) and vertically (average of 10 m). This pooling increased the sample size of some sites (Table S1).
The populations module of Stacks was used with each dataset to conduct several filtration steps (Table S2). SNPs common among a minimum percentage of individuals were filtered using (-r), choosing shared SNPs between 80% (-r 0.8) and 90% (-r 0.9) of individuals for datasets A and B, respectively. Using -write_single_snp, only the first SNP from each locus was chosen to reduce the linkage disequilibrium among different loci. SNPs with minor allele frequency (- minmaf < 0.05) were removed to avoid artifacts derived from PCR or sequencing errors, that might lead to bias in tests for selection47,48. SNPs departing from Hardy-Weinberg equilibrium (p = 0.05) that were present in at least two populations and SNPs that showed excess heterozygosity (Ho > 0.5) were removed as well49.
The sponge A. polypoides is a low microbial abundance (LMA) sponge50,51. Indeed, only a relatively small number of bacteria and archaea hits were found using -blastn, comparing the Stacks output to a nr database extracted from NCBI (accessioned on 22/06/2022), with e-value of 1e-6 or lower. The summarization of filtered SNPs from each step is presented in Table S2.
To assess genetic connectivity between populations, SNPs potentially under selection should be removed from the neutral dataset52,53. To achieve that we used a combination of Bayescan version 2.1 56 and Arlequin version 3.5.2 57. For Bayescan, default parameters were used based on the bayesian approach that calculated the coefficient of a specific population’s Fst, and compared each locus’ Fst coefficient to all the other populations. The chosen outliers were defined by a q-value > 0.05 (FDR is an analogue of the p-value56;. For Arlequin, p-values were calculated based on 100,000 coalescent simulations of samples to compare the created null distribution to the population differentiations (Fst), with 100 demes per group and on the observed heterozygosity across all loci55. The ‘Allowed missing level per site’ was set to 0.05 and the ‘Non-hierarchical Island Model’ was chosen. The obtained p-values were corrected using the p.adjust function in R with fdr method57. Overall, 76 SNPs were identified for dataset A as under selection: 72 SNPs for the ARLEQUIN analysis and 14 SNPs for BAYESCAN (10 overlapped SNPs between both analyses). As for dataset B, 32 SNPs were identified as under selection, all of which were detected by ARLEQUIN except two that were inferred in both methods (Table S2). No clones were found using the function mlg from the package popr in R58. The final number of neutral SNPs retained for datasets A and B, were 2,295 and 5,400, respectively.
d. Population genetics analyses
All the population genetic analyses were done separately to datasets A and B. Genetic diversity and demographic statistics were calculated within each population using Stacks. Expected (He) and observed (Ho) heterozygosity and inbreeding coefficients (Fis) were calculated for the whole dataset and for each population separately. To assess the genetic differentiation between pairs of populations pairwise-Fst values were calculated using ARLEQUIN version 3.5.2, with the default parameters (0.05 allowed level of missing data) and with 20,000 permutations. Heatmaps were created using the R package pheatmap59. Population structure was assessed using STRUCTURE version 2.360 ran for 100,000 MCMC iterations using the admixture model, with a burn-in of 50,000 iterations, setting a putative K from 1 to 8, with 15 repetitions for each run. We determined the most likely number of clusters using STRUCTURE HARVESTER61 (ln(Pr) and delta-k) and averaged each individual’s proportional coefficient across a k value, using Clustering Markov Packager Across K (CLUMPAK;62). Based on the STRUCTURE analysis, using the average proportion of membership of each pre-defined population in each of the three clusters, the Eastern Mediterranean Israeli populations were averaged and presented altogether as one group, using the MAPMIXTURE package. Pie charts were created and presented on a map63. Population structure was assessed with the usage of discriminant analysis of principal components (DAPC; adegenet R package,64), with the function snapclust.choose.k to choose the optimal value of k based on Akaike Information Criterion (AIC), using the k-means algorithm (pop.ini = “kmeans”), allowing a maximum k (number of clusters) of 8 (max = 8), and a maximum number of iterations of 100 (max.iter = 100). By comparing different clustering solutions that resulted from increasing k, this function calculates the AIC rates for each solution and the one with the lowest value is considered as the optimal one. The function xvalDAPC from the adegenet R package was used to select the most appropriate number of Principal Components (PCs) for the analysis, with 1,000 replicates (n.rep = 1,000), based on the lowest mean of squares error (MSE). After, using the function assignplot we graphically identified the genetic clusters assigned to each individual, and produced scatterplots of a given number of PCs and eigenvalues using scatter.dapc. The program fineRADstructure was used as another approach to infer the populations’ structure. This software considers nearest neighbor haplotypes and linkage information, and provides high resolution co-ancestry outputs65. We ran the analysis with both datasets deselecting the --write_single_SNP option which allowed the inclusion of linked SNPs in the different RAD-tags, resulting in a final dataset of 16,680 and 35,486 SNPs (for datasets A and B respectively). The fineRADstructure analysis utilized default settings with parameters set to -x 100,000, -y 100,000, -z 1,000 for individual population assignment. Additionally, tree construction was executed with -x 10,000. The obtained results were visually interpreted using the FineRADstructure R Library and the fineRADstructurePlot R script, both of which are included in the fineRADstructure package.
We computed a pairwise distance matrix by standard AMOVA computations (1,000 permutations) in ARLEQUIN version 3.5.2.2 to obtain the significance of the difference between the following groups: (1) individuals from Catalonia and Marseille were grouped and tested versus individuals from Israel using dataset A; (2) Using dataset B (only specimens from Israel) individuals from shallow depths were grouped and individuals from MSG were grouped; and (3) Individuals from Israel (dataset B) were grouped regardless their depth, separating them into four groups according to the site they were sampled from: Achziv, Rosh-Carmel, Atlit, and Herzliya.
To identify the gene flow patterns between the Israeli populations, relative migration analysis based on Nei’s Gst was calculated using the divMigrate function of the diveRsity R package to dataset B, comparing between sampling sites and depth, supported by bootstrap repetitions (boot= 100066,67;. Finally, a population assignment analysis test was performed, calculating the likelihood ratio thresholds for the eight Israeli populations (dataset B), based on Monte Carlo Likelihood ratio test with an α of 0.002 and 5,000 replicates using Genodive version 3.0668. This approach involved assigning or excluding reference populations as potential sources of individuals based on their multilocus genotypes. It achieves this by assessing the likelihood of an individual’s genotype occurring within a population, considering the allele frequencies present. Additionally, it identifies recent migrants between distinct populations. Genetic assignment techniques facilitate the inference of individuals’ origin, especially beneficial, in scenarios where population-level genetic distinctions are minimal. All maps were created using QGIS version 3.26 (http://www.qgis.org).
Results
Population genetics statistics
Population genetics statistics are found in Table 1. Catalonia and Marseille populations (dataset A) exhibited higher number of private alleles per individual (21 and 30, respectively) than Israeli populations, which ranged between 4 and 8 (Table 1, dataset A). Overall, the genetic diversity (expected heterozygosity; He) in each dataset was quite similar for the whole dataset and between sites, ranging from 0.046 ± 0.02 in Achziv Medium depth (ACm) population, to 0.079 ± 0.02 in Marseille population. The genetic diversity was slightly higher and even in dataset B, The Israeli samples ranged from 0.096 ± 0.01 in Atlit mesophotic (Ad) population, to 0.111 ± 0.01 in Herzliya shallow (Hsh) population. Similarly, in dataset A, the lowest observed heterozygosity (Ho) value was found in ACm population (0.050 ± 0.02), and the highest Ho in Marseille population (0.072 ± 0.02). The Ho in dataset B, showed also similarity with the populations found with the extreme values for He, ranging between 0.080 ± 0.01 in Ad population to 0.093 ± 0.01 in Hsh population. The inbreeding coefficient (Fis) was also similar for the entire dataset and between all sites, with slightly positive values. The same populations that were found with the minimum Ho and He values, also had the lowest Fis values in both datasets. In dataset A, the lowest Fis was 0.018 ± 0.02 (in ACm) and in dataset B it was 0.060 ± 0.06 (in Ad). However, the highest Fis values were found in Catalonia (for dataset A, 0.055 ± 0.02) and in Hsh and ACd (for dataset B, 0.090 ± 0.06, 0.090 ± 0.07 respectively) (Table 1).
Genetic structure between Eastern and Western mediterranean populations (Dataset A)
A clear genetic structure was detected in dataset A, separating the Western Mediterranean A. polypoides populations (Catalonia and Marseille) from those at the Eastern Mediterranean (Fig. 2 and S3). In the STRUCTURE analysis, despite the optimal number of clusters (three, k = 3; Fig. S1), the genetic structure suggested the separation of the samples into two main genetic groups: the individuals from Catalonia and Marseille (orange-blue cluster, Fig. 2A) and individuals from the Israeli coast (blue-green cluster, Fig. 2A). Samples from the Eastern Mediterranean were assigned to the blue cluster with a percentage ca. 90%, and the rest being assigned to the green genetic cluster (Fig. 2A, B). On the other hand, samples from the Western Mediterranean had a genetic assignment to the orange genetic cluster always higher than 50%, with some introgression of the blue cluster (Fig. 2A, B). DAPC and assignplot results were like those from STRUCTURE, grouping samples into two separated clusters: Cluster 1 with only individuals from both Catalonia and Marseille, and Cluster 2 with Israeli individuals (Fig. 2C, Fig. S3). The pairwise Fst values also differentiated A. polypoides populations into two major clusters of the populations mentioned above. The highest Fst values were observed between Catalonia and Marseille populations (Western Mediterranean) and the Israeli populations ranging between 0.185 and 0.305. Moreover, all the calculated pairwise-Fst values between Eastern and Western Mediterranean samples were significant (pv < 0.05, Fig. 3A). This grouping was also supported by AMOVA, that showed significant genetic differences between the grouping of all samples from the Western Mediterranean to all samples from Israel, which explained 28.54% of the total variance (p = 0.009, Table 2 (. The lowest pairwise-Fst values were found among Israeli populations comparisons, ranging between − 0.020 and 0.022, but 14 of the Fst’s were slightly negative and only some were significant. Nevertheless, higher and significant genetic differentiation was found between Catalonia and Marseille (Fst = 0.047, pv < 0.05, Fig. 3A). The fineRADstructure results were also in line with the STRUCTURE and DAPC analyses, showing a clear differentiation between the samples from Catalonia and Marseille with respect to the Israeli ones (Fig. S4A).
Individual genotype assignment of A. polypoides to clusters (K) for dataset A: 106 individuals and 14 sites. (A) As inferred by STRUCTURE with k = 3; (B) Pie frequency charts showing percentage of populations assigned to each of the three clusters represented in (A) for Catalonia - C, Marseille - F, and the averages of all the Israeli sites: Hsh, Hm, Hd, Ash, Am, Ad, Rsh, Rm, Rd, ACsh, ACm, and ACd; (C) Density distribution of individual genotypes along Discriminant Function 1 from the DAPC, illustrating population clustering patterns based on genetic similarity with k=2, 65PCs and 1 eigenvalue (clusters separation is presented in Figure S3A).
Genetic differentiation of A. polypoides measured as pairwise Fst. Significant values (pv < 0.05) are presented in bold and marked with asterisk (*). (A) For dataset A: 106 individuals and 14 sites. (B) For dataset B, including 95 individuals and 8 sites along the Israeli shore. Labels on the axes representing the source population: Catalonia - C, Marseille - F, and the averages of all the Israeli sites: Achziv - AC; Rosh-Carmel - R; Atlit – A; Herzliya – H. Small letters indicate the depth: shallow (< 30 m) – sh; Medium-depth (40 m) – m; and MSG (80–130 m) – d.
Genetic structure of Israeli coast populations (Dataset B, only Israel)
No genetic structure was found in the populations along the Israeli coast as referred from the reduced dataset (dataset B). The optimal k using Structure Harvester was two (k = 2; Fig. S5), showing no clear difference in the genetic structure among the Israeli populations, with all individuals showing a similar pattern of genetic assignment (Fig. 4A). According to AIC the optimal k for DAPC analysis was one (Fig. S6). Thus, we decided to run DAPC grouping samples by sampling area, resulting in Achziv mesophotic samples (ACd) being relatively isolated from the other individuals, with most individuals scattered separately from the rest (Fig. 4B). Achziv shallow individuals (ACsh) showed some isolation as well, both in the DAPC and fineRADstructure (Fig. 4B and S4B). On the other hand, in the DAPC analysis, Rosh-Carmel mesophotic individuals (Rd) were located mixed with the individuals from the shallow depths. However, in general, the shallow populations showed some sort of overlap in the center of the plot, while the mesophotic populations were slightly separated from them (Fig. 4B). The fineRADstructure (haplotypic analysis) showed no clear structure between the Israeli populations, and high genetic homogeneity between all individuals. We detected, though, a separated clade of samples from Herzliya site, with a mixture between shallow and mesophotic individuals (Fig. S4B, black clade). In addition, Rosh-Carmel mesophotic individuals appeared dispersed between most of the clades (Fig. S4B).
Although all Fst values were relatively low, and only some were significant for the different pairwise comparisons between Israeli sites (ranging from 0.005 to 0.023), we detected a clear separation between mesophotic samples and the shallow ones, except for Rd that grouped with the shallow samples. The only significant comparison with Rd was observed with the Hsh population (Fst = 0.017, p = 0.0004). The highest genetic differentiation was observed between Achziv’s shallow and mesophotic populations, with a significant Fst of 0.023 (p = 0.01; Fig. 3B).
The AMOVA analysis comparing samples from MSG with shallow samples, found significant differences between these bathymetric groups, although it represented only 0.2% of the overall explained variance (Table 2).
The migration analysis indicated bidirectional connections among all Israeli populations. Specifically, migration from Herzliya and Achziv’s mesophotic populations (Hd and ACd) to all the other populations, including direction opposite to the major currents, with relatively low Gst values (ranging from 0.23 to 0.45 with an average of 0.3). However, shallow populations displayed higher migration rates between them, with Gst values ranging from 0.32 to 1 (average of 0.6, Fig. 5A). An increase of the migration threshold showed the highest migration rates between all the shallow-water sampling stations. The mesophotic-depth sampling sites presented much lower migration rates from the shallow-water ones and from each other, except for Rosh Carmel mesophotic (Rd), which showed a relatively high migration value towards Hsh (Gst = 0.8, Fig. 5B). Based on population assignment, the Herzliya shallow site (Hsh; light blue) emerged as the only source population for most of the other populations. Rosh-Carmel shallow (Rsh) was the only exception, that displayed a 7% contribution from Atlit shallow (Ash), and Achziv shallow (ACsh) that had 10% of individuals assigned to Rosh-Carmel shallow. Additionally, the population of Achziv mesophotic (ACd; red) exhibited a self-assignment of 50% apart from the assignment to Hsh (Fig. 5C).
Individual genotype assignment of A. polypoides to clusters (K) for dataset B, including 95 individuals and eight sites from Israel (dataset B), (A) as inferred by STRUCTURE with k = 2 and by (B) DAPC grouping samples per site retaining 11PCs.
Populations migration and assignment of A. polypoides between eight sites from dataset B (red = MSG, blue = shallow). (A) Contemporary migration depicting the relative strength of gene flow in Israel. Each direction is based on source population numbers on the nodes, using Nei’s GST method; (B) With a filter threshold of 0.75; (C) Population assignment providing the proportion of individuals assigned to each site.
Discussion
In this study, we describe the genetic connectivity and gene flow of the sponge A. polypoides at two different scales. The long-distance analyses compared samples from the Western and the Eastern Mediterranean. It detected a clear genetic structure between the two basins (Fig. 2). On the other hand, the small-scale analysis within Eastern Mediterranean, revealed low genetic differentiation and high connectivity among all examined populations along the Israeli coast (Figs. 4 and 5). However, a subtle genetic substructure was found between the Israeli populations, probably driven mainly by bathymetric features (summarized in Fig. 6).
Map illustrating A. polypoides connectivity patterns along the north of the Israeli coast. Yellow polygons represent the approved Marine Reserves (MR) and green polygons represent the proposed MR, using data obtained from Ecological Geographic data center, ICT, Israel Nature and Parks Authority. Gene flow of A. polypoides populations is illustrated by the white arrows. The red arrow indicates the major current along the shore. The shapes represent the sampling sites: Achziv (Rosh-Hanikra nature reserve) - AC; Rosh-Carmel - R; Atlit – A; Herzliya (Yam-Poleg) – H. Small letters indicate the depth: shallow (up to 40 m) – sh (blue); and MSG (80–130 m) – d (red).
The genetic difference between Western and the Eastern Mediterranean samples, resembles other studies with a similar spatial separation between samples (Fig. 269,70,71,72;. This is consistent with the results from a recent study of the genetic connectivity of the shallow-water Atlanto-Mediterranean sponge Petrosia ficiformis that found a marked genetic structure between samples distributed across the entire Mediterranean Sea31. In that study, the Israeli samples were separated (by microsatellite markers) from the rest, mainly due to isolation by distance and to the presence of prevalent oceanographic fronts31. Such oceanographic fronts play an important role in separating internal basins in the Mediterranean, like the Sicily Strait, that separates the Eastern and Western Mediterranean basins27,69,72,73,74,75. However, it was suggested that in sponges and other sessile organisms, high differentiation between populations is mostly caused by reduced larval dispersal capabilities27as it is the case for the above-mentioned P. ficiformis, which has a crawling larva with a very limited dispersal ability31. For organisms with greater dispersal ability, either through longer lifespan of the pelagic larvae or by motility of both adult and larvae, it was mainly suggested that the genetic differences are consequence of the combination of hydrodynamics processes, and the limitations of the biological dispersal27. These results contrast, though, to what has been reported for highly motile organisms such as the fish Serranus cabrilla and Dicentrarchus labrax, where no structure was detected between populations in the Eastern and Western Mediterranean Sea76,77. Despite the fact that information on the biology of A. polypoides larvae and gametes is still limited, the genetic difference we found between the Eastern and Western Mediterranean (Israeli versus both Catalonia and Marseille) populations, is in congruence with what is expected of sessile invertebrates with limited dispersal abilities16,28,78. However, to better understand the connectivity and gene flow patterns of A. polypoides between and within basins, further studies should investigate additional sampling areas throughout the species distribution range, and Western Mediterranean sample size should be increased.
Our smaller-scale analysis within the Levant basin, demonstrated that A. polypoides populations along the Israeli Mediterranean coast exhibited low genetic differentiation and high connectivity. This could have been expected because of the relative proximity of the study sites (along ~ 120 km coast, Fig. 1). Nevertheless, this connectivity pattern contrasts with other studies on sponge populations, using similar distance scales regardless of the type of larvae the species have. Such is the case of two Mediterranean viviparous sponge species, with assumed limited dispersal ability (Scopalina lophyropoda and Ircinia fasciculata, using microsatellites), and another Southern Ocean and species (Dendrilla antarctica, using ddRADseq), in which clear genetic breaks were detected at a reduced scale32,79,80. A similar separation pattern has been detected even for oviparous sponges, with assumed higher dispersal ability, such as Cliona delitrix, Chondrosia reniformis, and Petrosia ficiformis31,73,81using COI and microsatellites. The moderate-high connectivity observed in the oviparous A. polypoides25 we studied, seems not to follow this trend, at least for the Israeli coast. Similarly, high connectivity in small spatial scale was recently reported for the deep-sea glass-sponge Vazella pourtalesii in the northwest Atlantic using ddRADseq82. Notably, compared with microsatellites, ddRADseq provides thousands of genome-wide SNPs, offering higher resolution for fine-scale structure, but requires high-quality DNA and has a higher per-sample cost83.
Remote biodiversity hotspots, such as the Israeli MSG, require special conservation attention. The assessment of the connectivity within MSG, as well as between them and shallow habitats, are of utmost importance because they are considered as potential refugia for species that would not survive if environmental conditions on shallow depths worsened33,84. Such a detrimental change is already happening in the Levant Basin. For example, the temperature-driven disappearance of Agelas oroides from the shallow-water habitats along Israel20and additional local population extinctions in other regions in the Mediterranean Sea21,22,85. For a mesophotic habitat to be considered as a refuge, it is not enough for its populations to simply overlap in distribution with the shallow populations, but also their vertical connectivity must be determined86. The low differentiation we detected between all Israeli A. polypoides populations, indicates a moderate-high gene flow between shallow and mesophotic populations, hence supporting the second requirement of a refuge. The clustering analyses together with the relatively low pairwise Fst we detected (Table 2; Fig. 3B), suggest that all Israeli samples should be indeed regarded as a unique panmictic population, although with some subtle substructure detected for several mesophotic sites (Figs. 4B and 5). Additional support for this statement is the low level of population structure between all Israeli populations (Fig. 4A). The MSG populations of A. polypoides, reproduce year-round and invest more in sexual reproduction compared to the shallow-water populations, which reproduce seasonally25. Therefore, due to the detected bidirectional vertical connectivity (Fig. 5A), propagules could theoretically disperse between shallow and mesophotic habitats. Thus, if the current trend of conditions escalates at the shallow habitats, the MSG A. polypoides populations could serve as a gene source for propagules resettlement in the shallow depths, when environmental conditions will allow it. However, partial overlaps in timing of reproduction between these habitats may serve as a barrier leading to maintenance of genetically distinct populations. Furthermore, the Depth Differentiation Hypothesis (DDH) identifies depth as a major driver of ecological and genetic differentiation in marine systems, driven by environmental shifts and distinct water masses that promote local adaptation17,87,88.
Using the reduced dataset that included only the samples from Israeli sites (dataset B), we found a sub-structure among this finer-scale area89. The difference between all shallow and mesophotic populations was significant (p = 0.01, Table 2), but it explained only 0.2% of the total variation. This is in line with the clustering analyses results of high gene flow among shallow populations, and some sort of genetic isolation of Achziv, Atlit, and Herzliya mesophotic sites from the rest (Figs. 3B, 4 and 5B). The higher similarities and the mixture of the samples from the MSG of Rosh-Carmel with the shallow populations found in the pairwise Fst, migration and clustering analyses (Figs. 3B, 4 and 5 and S4B), can be attributed to bathymetric properties. Rosh-Carmel mesophotic site is a continuous extension of the Carmel terrestrial mountain that creates uninterrupted rocky substrate dropping from sea surface down to 300 m39,90. These bathymetric conditions provide higher potential for larvae settlement throughout the entire depth range. In accordance, it also allows greater connectivity with the other shallow populations that were already shown to have lower differentiations among them. Interestingly, the migration analysis and population assignment showed that the specimens from Herzliya shallow population are the main gene source for the other populations (Fig. 5). The explanation to this finding could be attributed to the fact that the long-shore currents along the Israeli continental shelf prevail mainly northwards18,90. These currents could carry larvae originated from Herzliya to all other sites that are located further north. It was also suggested that Achziv’s water mass is more affected by coastal dynamics than by the major currents, probably because the bay of Haifa can constitute a barrier to the water mass flowing from the south18. In turn, this may explain the results of Achziv’s populations which showed the highest differentiation from all other populations, in addition to the lowest signatures of migration. Remarkably, despite the small distance between mesophotic and shallow sites in Achziv (~ 6 km), we detected a subtle genetic differentiation between them as well. The spatial distances between samples from shallow to MSG depths within the sites Atlit, Rosh-Carmel, and Achziv are 6–8 km, while the Herzliya shallow site is 16 km away from its nearest MSG. The high gene flow between shallow and mesophotic areas in Herzliya, despite being the most distant among the shallow-MSG populations, also might be due to the site’s bathymetric properties. Compared to the other sites, Herzliya has a higher number of pinnacles (on which MSG occur), and its shallow rocky substrate expands over a greater area than that at the other sites14,91. The Herzliya populations represent only the edges of their depth range, which contains suitable habitats for settlement between them (at 60 and 80 m depth). As such, propagules can potentially disperse in a wider area and spread more efficiently.
Mesophotic sponge grounds are particularly vulnerable and require attention and legislation to promote their preservation11,21. This is generally true for the entire Mediterranean, but especially for the Levant Basin. The biodiversity of this Basin is severely threatened due to the extreme rise in seawater temperatures coupled with increasing local anthropologic threats. Such threats include the damming of the Nile, enlargement of the Suez Canal, shipping, fishing, and more recently, gas exploration and construction of marine infrastructure1,9,21,92,93. MR could help minimize threats and damages. However, establishing such reserves effectively requires the assessment of their connectivity, as carried out in this study94,95. This work contributed to our understanding of the biology and distribution of a key Atlanto-Mediterranean sponge A. polypoides in shallow and mesophotic habitats, an important yet vulnerable species50,96in the list of SPA-BD97. We showed that all A. polypoides populations studied along the Israeli shore, are reasonably well connected, and that MSG habitats, that have more stable conditions than the shallow habitats, might serve as a refuge for shallower populations. The subtle substructure detected for some mesophotic sites points to a relative isolation of A. polypoides in several MSG sites, except for Rosh Carmel mesophotic population. Therefore, the study indicates occurrence of site-specific differences among mesophotic ecosystems in agreement with previous studies (e.g17., and the relevance of determining the areas with the greatest gene flow between shallow and mesophotic habitats whose preservation should be prioritized. However, to ensure successful performance of populations at a regional level in a fast-changing environment, the study highlights the importance of protecting each MSG area individually, rather than focusing on just one33,95,98. These findings should be considered when designing future MR (Fig. 6, green polygons) or managing current ones (Fig. 6, yellow polygons).
Data availability
RADseq data for each individual sample are deposited in the NCBI SRA database, BioProject PRJNA1201738.
References
Coll, M. et al. The biodiversity of the mediterranean sea: estimates, patterns, and threats. PLoS One. 5, 8 (2010).
Albano, P. G. et al. The dawn of the tropical Atlantic invasion into the mediterranean sea. Proc Natl. Acad. Sci. U S A 121, (2024).
Chatzimentor, A., Doxa, A., Katsanevakis, S. & Mazaris, A. D. Are mediterranean marine threatened species at high risk by climate change? Glob Chang. Biol. 29, 1809–1821 (2023).
Lejeusne, C., Chevaldonné, P., Pergent-Martini, C., Boudouresque, C. F. & Pérez, T. Climate change effects on a miniature ocean: the highly diverse, highly impacted Mediterranean Sea. Trends Ecol Evol 25, 250–260 Preprint at (2010). https://doi.org/10.1016/j.tree.2009.10.009
Van Soest, R. W. M. et al. Global diversity of sponges (Porifera). (2012). https://doi.org/10.1371/journal.pone.0035105
Strano, F., Micaroni, V., Costa, G., Bertocci, I. & Bertolino, M. Shallow-water sponge grounds along the Apulian Coast (central mediterranean Sea). Mar. Biodivers. 50, 7 (2020).
Díaz, J. A., Ordinas, F., Farriols, M. T., Melo-Aguilar, C. & Massutí, E. Sponge assemblages in fishing grounds and seamounts of the Balearic Islands (western Mediterranean). Deep Sea Res. Part. I Oceanogr. Res. Pap. 203, 104211 (2024).
Xavier, J. R. & van Soest, R. W. M. Diversity patterns and zoogeography of the Northeast Atlantic and mediterranean shallow-water sponge fauna. Hydrobiologia 687, 107–125 (2012).
Bell, J. J. et al. Academic Press,. Sponge functional roles in a changing world. In Advances in Marine Biology 95, 27–89 (2023).
Bell, J. J. The functional roles of marine sponges. Estuar. Coast Shelf Sci. 79, 341–353 (2008).
Maldonado, M. et al. Sponge grounds as key marine habitats: a synthetic review of types, structure, functional roles, and conservation concerns. Mar. Anim. Forests. 1–39. https://doi.org/10.1007/978-3-319-17001-5_24-1 (2015). Springer International Publishing.
Hogg, M. et al. Deep-sea sponge grounds seas regional. (2010). http://hdl.handle.net/20.500.11822/8579
Voultsiadou, E. Reevaluating sponge diversity and distribution in the mediterranean sea. Hydrobiologia 628, 1–12 (2009).
Idan, T. et al. Shedding light on an East-Mediterranean mesophotic sponge ground community and the regional sponge fauna. Mediterr. Mar. Sci. 19, 84–106 (2018).
Goren, L., Idan, T., Shefer, S. & Ilan, M. Macrofauna inhabiting massive demosponges from shallow and mesophotic habitats along the Israeli Mediterranean Coast. Front Mar. Sci 7, (2021).
Idan, T. Mesophotic sponge grounds in the Levant basin: community composition and dynamics. PhD dissertation. (Tel-Aviv University, (2020).
Slattery, M., Lesser, M. P., Rocha, L. A., Spalding, H. L. & Smith, T. B. Function and stability of mesophotic coral reefs. Trends Ecol Evol 39, 585–598 Preprint at (2024). https://doi.org/10.1016/j.tree.2024.01.011
Ozer, T., Gertman, I., Gildor, H. & Herut, B. Thermohaline Temporal variability of the SE mediterranean coastal waters (Israel) – long-term trends, seasonality, and connectivity. Front Mar. Sci 8, (2022).
Ozer, T., Gertman, I., Kress, N., Silverman, J. & Herut, B. Interannual thermohaline (1979–2014) and nutrient (2002–2014) dynamics in the Levantine surface and intermediate water masses, SE Mediterranean sea. Glob Planet. Change. 151, 60–67 (2017).
Idan, T., Goren, L., Shefer, S. & Ilan, M. Sponges in a changing climate: survival of Agelas oroides in a warming mediterranean sea. Front Mar. Sci 7, (2020).
Garrabou, J. et al. Collaborative database to track mass mortality events in the mediterranean sea. Front Mar. Sci 6, (2019).
Grenier, M., Idan, T., Chevaldonné, P. & Pérez, T. Mediterranean marine keystone species on the Brink of extinction. Glob Chang. Biol. 29, 1681–1683 (2023).
Kress, N., Gertman, I. & Herut, B. Temporal evolution of physical and chemical characteristics of the water column in the Easternmost Levantine basin (Eastern mediterranean Sea) from 2002 to 2010. J. Mar. Syst. 135, 6–13 (2014).
Castellan, G., Angeletti, L., Montagna, P. & Taviani, M. Drawing the borders of the mesophotic zone of the Mediterranean sea using satellite data. Sci Rep 12, 5585 (2022).
Idan, T., Shefer, S., Goren, L., Brickner, I. & Ilan, M. Does depth matter? Reproduction pattern plasticity in two common sponge species found in both mesophotic and shallow waters. Front Mar. Sci 7, (2020).
Pérez-Portela, R. & Riesgo, A. Population genomics of early-splitting lineages of metazoans. In 103–137 (2018). https://doi.org/10.1007/13836_2018_13
Pascual, M., Rives, B., Schunter, C. & Macpherson, E. Impact of life history traits on gene flow: a multispecies systematic review across oceanographic barriers in the mediterranean sea. PLoS One 12, (2017).
Maldonado, M. The ecology of the sponge larva. Can. J. Zool. 84, 175–194 (2006).
Villamor, A., Costantini, F. & Abbiati, M. Genetic structuring across marine biogeographic boundaries in rocky shore invertebrates. PLoS One 9, 7 (2014).
Patarnello, T., Volckaert, F. A. M. J. & Castilho, R. Pillars of hercules: is the Atlantic-Mediterranean transition a phylogeographical break? Mol. Ecol. 16, 4426–4444 (2007).
Riesgo, A. et al. Genetic diversity, connectivity and gene flow along the distribution of the emblematic Atlanto-Mediterranean sponge Petrosia ficiformis (Haplosclerida, Demospongiae). BMC Evol. Biol 19, 24 (2019).
Riesgo, A. et al. Population structure and connectivity in the mediterranean sponge Ircinia fasciculata are affected by mass mortalities and hybridization. Nat. Commun. 117, 427–439 (2016).
Magris, R. A., Pressey, R. L., Weeks, R. & Ban, N. C. Integrating connectivity and climate change into marine conservation planning. Biol. Conserv. 170, 207–221 (2014).
Lesser, M. P., Slattery, M. & Leichter, J. J. Ecology of mesophotic coral reefs. J. Exp. Mar. Biol. Ecol. 375, 1–8 (2009).
Bell, J. J. et al. Society for conservation biology global conservation status of sponges. Biology (Basel). 29, 42–53 (2015).
Chomsky, O., Douek, J., Chadwick, N. E., Dubinsky, Z. & Rinkevich, B. Biological and population-genetic aspects of the sea anemone Actinia equina (Cnidaria: Anthozoa) along the mediterranean Coast of Israel. J. Exp. Mar. Biol. Ecol. 375, 16–20 (2009).
Shabtay, A., Tikochinski, Y., Benayahu, Y. & Rilov, G. Preliminary data on the genetic structure of a highly successful invading population of oyster suggesting its establishment dynamics in the levant. Mar. Biol. Res. 10, 407–415 (2014).
de Voogd, N. J. et al. World Porifera database. Axinella Polypoides Schmidt (2023).
Sade, A. R. et al. The Israel National bathymetric survey - a new look at the seafloor off Israel. Isr. J. Earth Sci. 55, 185–187 (2006).
Peterson, B. K., Weber, J. N., Kay, E. H., Fisher, H. S. & Hoekstra, H. E. Double digest radseq: an inexpensive method for de Novo SNP discovery and genotyping in model and non-model species. PLoS One 7, (2012).
Combosch, D. J., Lemer, S., Ward, P. D., Landman, N. H. & Giribet, G. Genomic signatures of evolution in Nautilus—an endangered living fossil. Mol. Ecol. 26, 5923–5938 (2017).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Available online at (2010). https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Catchen, J., Hohenlohe, P. A., Bassham, S., Amores, A. & Cresko, W. A. Stacks: an analysis tool set for population genomics. Mol. Ecol. 22, 3124–3140 (2013).
Paris, J. R., Stevens, J. R. & Catchen, J. M. Lost in parameter space: a road map for stacks. Methods Ecol. Evol. 8, 1360–1373 (2017).
Jeffries, D. L. et al. Comparing RADseq and microsatellites to infer complex phylogeographic patterns, an empirical perspective in the crucian carp, Carassius carassius, L. Mol. Ecol. 25, 2997–3018 (2016).
de Medeiros, B. A. S. & Farrell, B. D. Whole-genome amplification in double-digest RADseq results in adequate libraries but fewer sequenced loci. PeerJ 6, e5089 (2018). https://doi.org/10.7717/peerj.5089
Benestan, L. et al. RAD genotyping reveals fine-scale genetic structuring and provides powerful population assignment in a widely distributed marine species, the American Lobster (Homarus americanus). Mol. Ecol. 24, 3299–3315 (2015).
Roesti, M., Salzburger, W. & Berner, D. Uninformative polymorphisms bias genome scans for signatures of selection. BMC Evol. Biol 12, 94 (2012).
Hohenlohe, P. A., Amish, S. J., Catchen, J. M., Allendorf, F. W. & Luikart, G. Next-generation RAD sequencing identifies thousands of SNPs for assessing hybridization between rainbow and Westslope cutthroat trout. Mol. Ecol. Resour. 11, 117–122 (2011).
Coppari, M. et al. The role of mediterranean sponges in benthic-pelagic coupling processes: Aplysina aerophoba and Axinella polypoides case studies. J. Exp. Mar. Biol. Ecol. 477, 57–68 (2016).
Wehrl, M., Bakterielle & Aufnahme Selektivität und interne Prozessierung bei marinen Schwämmen (Porifera). Doctoral dissertation, Universität Würzburg. (2006).
Casanova, A. et al. Low impact of different SNP panels from two building-loci pipelines on RAD-Seq population genomic metrics: case study on five diverse aquatic species. BMC Genomics 22, (2021).
Luikart, G., England, P. R., Tallmon, D., Jordan, S. & Taberlet, P. The power and promise of population genomics: from genotyping to genome typing. Nat. Rev. Genet. 4, 981–994 (2003).
Foll, M. & Gaggiotti, O. A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a bayesian perspective. Genetics 180, 977–993 (2008).
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Storey, J. D. The positive false discovery rate: a bayesian interpretation and the Q-Value1. Ann Stat 31, 2013-2035 (2003).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R Stat. Soc. B. 57, 289–300 (1995).
Kamvar, Z. N., Brooks, J. C. & Grünwald, N. J. Novel R tools for analysis of genome-wide population genetic data with emphasis on clonality. Front Genet 6, 208 (2015).
Kolde, R. pheatmap (RRID:SCR_016418). Preprint at (2018).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. (2000). http://www.stats.ox.ac.uk/pritch/home.html
Earl, D. A. & vonHoldt, B. M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour. Preprint at (2011).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Jenkins, T. L. mapmixture: an R package and web app for Spatial visualisation of admixture and population structure. Mol Ecol. Resour 24, e13943 (2024).
Jombart, T. Adegenet: an R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Malinsky, M., Trucchi, E., Lawson, D. J. & Falush, D. RADpainter and fineradstructure: population inference from RADseq data. Mol. Biol. Evol. 35, 1284–1290 (2018).
Sundqvist, L., Keenan, K., Zackrisson, M., Prodöhl, P. & Kleinhans, D. Directional genetic differentiation and asymmetric migration. (2013). https://doi.org/10.1002/ece3.2096
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. DiveRsity: an R package for the Estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Meirmans, P. G. GENODIVE version 3.0: easy-to-use software for the analysis of genetic data of diploids and polyploids. Mol Ecol Resour, 1126–1131. Preprint at (2020).
Valente, S., Serrão, E. A. & González-Wangüemert, M. West versus East mediterranean sea: origin and genetic differentiation of the sea cucumber Holothuria polii. Mar. Ecol. 36, 485–495 (2015).
Sá-Pinto, A., Branco, M. S., Alexandrino, P. B., Fontaine, M. C. & Baird, S. J. E. Barriers to gene flow in the marine environment: insights from two common intertidal limpet species of the Atlantic and mediterranean. PLoS One 7, e50330 (2012).
Dailianis, T., Tsigenopoulos, C. S., Dounas, C. & Voultsiadou, E. Genetic diversity of the imperilled bath sponge Spongia officinalis linnaeus, 1759 across the mediterranean sea: patterns of population differentiation and implications for taxonomy and conservation. Mol. Ecol. 20, 3757–3772 (2011).
López-Márquez, V. et al. Genetic connectivity of two marine gastropods in the mediterranean sea: seascape genetics reveals species-specific oceanographic drivers of gene flow. Mol. Ecol. 30, 4608–4629 (2021).
Moussa, M., Choulak, S., Rhouma-Chatti, S., Chatti, N. & Said, K. First insight of genetic diversity, phylogeographic relationships, and population structure of marine sponge Chondrosia reniformis from the Eastern and Western Mediterranean Coasts of Tunisia. Ecol Evol 12, e8494 (2022).
Bergamasco, A. & Malanotte-Rizzoli, P. The circulation of the mediterranean sea: a historical review of experimental investigations. Adv. Oceanogr. Limnol. 1, 11–28 (2010).
Paterno, M. et al. Population genomics Meet lagrangian simulations: oceanographic patterns and long larval duration ensure connectivity among Paracentrotus lividus populations in the Adriatic and ionian seas. Ecol. Evol. 7, 2463–2479 (2017).
Schunter, C. et al. Matching genetics with oceanography: directional gene flow in a mediterranean fish species. Mol. Ecol. 20, 5167–5181 (2011).
Quéré, N. et al. Gene flow at major transitional areas in sea bass (Dicentrarchus labrax) and the possible emergence of a hybrid swarm. Ecol. Evol. 2, 3061–3078 (2012).
Maldonado, M. & Bergquist, P. Atlas of Marine Invertebrate Larvae (Academic, 2002). Chp-2.
Blanquer, A. & Uriz, M. Population genetics at three Spatial scales of a rare sponge living in fragmented habitats. BMC Evol. Biol. 10, 1–9 (2010).
Leiva, C. et al. Population substructure and signals of divergent adaptive selection despite admixture in the sponge Dendrilla Antarctica from shallow waters surrounding the Antarctic Peninsula. Mol. Ecol. 28, 3151–3170 (2019).
Chaves-Fonnegra, A., Feldheim, K. A., Secord, J. & Lopez, J. V. Population structure and dispersal of the coral-excavating sponge Cliona delitrix. Mol. Ecol. 24, 1447–1466 (2015).
Patova, A. et al. Population genomics and connectivity of Vazella pourtalesii sponge grounds of the Northwest Atlantic with conservation implications of deep sea vulnerable marine ecosystems. Sci. Rep. 15, 1540 (2025).
Sunde, J., Yıldırım, Y., Tibblin, P. & Forsman, A. Comparing the performance of microsatellites and RADseq in population genetic studies: analysis of data for Pike (Esox lucius) and a synthesis of previous studies. Front Genet 11, 218 (2020).
Slattery, M., Lesser, M. P., Brazeau, D., Stokes, M. D. & Leichter, J. J. Connectivity and stability of mesophotic coral reefs. J. Exp. Mar. Biol. Ecol. 408, 32–41 (2011).
Dinçtürk, E., Öndes, F., Leria, L. & Maldonado, M. Mass mortality of the keratose sponge Sarcotragus foetidus in the Aegean sea (Eastern Mediterranean) correlates with proliferation of Vibrio bacteria in the tissues. Front Microbiol 14, (2023).
Lesser, M. P., Slattery, M. & Mobley, C. D. Biodiversity and functional ecology of mesophotic coral reefs. Annu. Rev. Ecol. Evol. Syst. 49, 49–71 (2018).
Etter, R. J. & Rex, M. A. Deep-Sea Biodiversity (Harvard University Press, 2010).
Monteiro, C. A., Paulino, C., Jacinto, R., Serraõ, E. A. & Pearson, G. A. Temporal windows of reproductive opportunity reinforce species barriers in a marine broadcast spawning assemblage. Sci Rep 6, 29198 (2016).
Vendrami, D. L. J. et al. RAD sequencing resolves fine-scale population structure in a benthic invertebrate: implications for Understanding phenotypic plasticity. R Soc. Open. Sci 4, (2017).
Ery, K. O. E., Neev, D., Bento, Y. K. & Emery, K. O. Mediterranean Beaches of Israel. (1960).
Zviely, D. & Polinov, S. & Reich-Roman, A. Maritime policy for israel’s mediterranean Waters. 1–213 (2016).
Corrales, X. et al. Future scenarios of marine resources and ecosystem conditions in the Eastern mediterranean under the impacts of fishing, alien species and sea warming. Sci Rep 8, (2018).
Ibrahim, O., Mohamed, B. & Nagy, H. Spatial variability and trends of marine heat waves in the Eastern mediterranean sea over 39 years. J Mar. Sci. Eng 9, (2021).
Andrello, M., Mouillot, D., Somot, S., Thuiller, W. & Manel, S. Additive effects of climate change on connectivity between marine protected areas and larval supply to fished areas. Divers. Distrib. 21, 139–150 (2015).
Carr, M. H. et al. The central importance of ecological Spatial connectivity to effective coastal marine protected areas and to meeting the challenges of climate change in the marine environment. Aquat. Conserv. 27, 6–29 (2017).
Azzola, A. et al. You cannot conserve a species that has not been found: the case of the marine sponge Axinella polypoides in liguria, Italy. Aquat. Conserv. 31, 737–747 (2021).
UNEP/MAP-SPA/RAC. SPA-BD Protocol - ANNEX II: list of endangered or threatened species. (2018).
Lowe, W. H. & Allendorf, F. W. What can genetics tell Us about population connectivity? Mol. Ecol. 19, 3038–3051 (2010).
Acknowledgements
We thank Ana Riesgo and Gilberto Bergamo for their support. Fieldwork was conducted aboard the R/V Mediterranean Explorer, with the assistance of EcoOcean, Itay Katzman and Oded Ezra.
Funding
This work was supported by the Merci Foundation; the Israel Ministry of Science (grant #2197); the EU Biodiversa + project “SPONBiodiv” (ANR-22-EBIP-0004-06, France); the AMiDEX international project “MARRIO” funded by Aix Marseille University (France); and the Sponge-Pump project (PID2021-124856NB-100) funded by MCIN/AEI and ERDF (Spain).
Author information
Authors and Affiliations
Contributions
T.M., L.G., T.I., Y.T. and M.I. conceived and designed the study; T.M., S.S., L.G., T.I., A.R., R.C. and T.P. conducted fieldwork and collected samples; T.M., C.S. and S.T., conducted laboratory work; T.M. and S.T. performed statistical analyses and interpreted the results; T.M. wrote the first version of the manuscript, and all co-authors contributed to the current version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Morav, T., Taboada, S., Goren, L. et al. Deep connections: exploring the genetic connectivity of mesophotic and shallow-water populations of the sponge Axinella polypoides. Sci Rep 15, 39183 (2025). https://doi.org/10.1038/s41598-025-18563-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-18563-0