Abstract
Periodontal disease and dental caries are two oral conditions that have been associated with atherosclerotic cardiovascular disease (ASCVD). However, it is unclear if one of the key mechanisms involved in this association could be a shared genetic susceptibility. The goal of this study was to explore whether there is an intersection of genetic loci among individuals with comprehensive oral examinations and subclinical ASCVD screenings. We leveraged data from oral and medical examinations obtained from the Dental and Heart Strategies Concentrating on Risk Evaluation (Dental/Heart SCORE) projects. Genome-wide association studies (GWASs) were performed independently in 552 participants (aged 45–75 years). The decayed, missing, or filled teeth index (DMFT) and periodontal disease indices were used to reflect oral conditions; coronary artery calcium scores (CAC) and carotid intima media thickness (CIMT) were analyzed as subclinical ASCVD traits. Single nucleotide variant (SNV) associations with oral and ASCVD traits were found; however, there were only a few regions of suggestive genetic loci overlap between these conditions. The most robust associations found for each phenotype are as follows: DMFT with rs79198416 (near CDC73/KCNT2; p = 7.57E-07), periodontal disease with rs73870587 (DIPK2A, p = 7.38E-08); CIMT with rs113152669 (LRP1B p = 4.07E-07), and CAC with rs76676138 (CNTNAP2; p = 2.47E-19). Although genetic associations were identified for each of the phenotypes of interest in the GWASs, there were no regions of shared genetic loci that significantly intersected across phenotypes. Thus, our results suggest that incorporation of environmental, behavioral, microbiome-related factors, and larger sample sizes, are warranted in future studies between oral and cardiovascular health.
Similar content being viewed by others
Introduction
Despite being highly preventable1dental caries is the most prevalent chronic disease in both children and adults2 with more than 1 in 5 people having untreated dental caries in the US population3. Different dental diseases have been suggested to affect the risk for systemic inflammatory conditions1, including atherosclerotic cardiovascular disease (ASCVD), the leading cause of death worldwide4. Subclinical ASCVD markers, such as the carotid intima-media thickness (CIMT) and coronary artery calcification (CAC), along with risk factors such as inflammatory dysregulation and lifestyle stressors, can predict future risk of cardiovascular outcomes5.
A suggested mechanism that may play a role in the correlation between oral disease and ASCVD is genetic susceptibility, determined in part by single nucleotide variants (SNVs) in or regulating genes involved in inflammatory pathways6. This was mainly suggested for the association between periodontal disease and ASCVD, with the association with dental caries being less explored. However, early life oral disease (mainly dental caries) was associated with higher burden of atherosclerotic cardiovascular disease in adulthood7. Thus, it is plausible that there is an interplay between dental caries and future risk of ASCVD, which usually begins decades before ASCVD manifests clinically.
Few studies have focused on identifying the overlap in genetic loci associated with dental caries and subclinical ASCVD8, and further research is needed to explore pleiotropic effects between periodontal disease and ASCVD. It is not clear whether periodontal disease and atherosclerosis are outcomes of similar maladaptive inflammatory pathways6. If this is the case, pleiotropic genes may affect the development of both periodontal disease and subclinical ASCVD9. Prior studies have suggested that the association between cardiovascular disease and periodontal disease may have been confounded by genetics6. However, to date, candidate gene association studies have only identified 5 genes that have been associated with both cardiovascular disease and periodontal disease traits (PLG, CAMTA1, VAMP3, VAMP8 and CDKN2B)9,10,11.
Addressing gaps in knowledge by determining whether pleiotropy between oral disease and subclinical ASCVD in fact exists will offer mechanistic insights into disease initiation and progression. We performed independent GWASs to explore potential pleiotropic regions using the combined Dental and Heart SCORE cohorts of 552 subjects. We hypothesize that there will be an intersection of genetic loci associated with oral conditions (i.e., dental caries and periodontal disease) and subclinical ASCVD (i.e., CIMT and CAC). In summary, establishing genetic relationships between these conditions may advance mechanistic understanding in disease susceptibility.
Results
Table 1 presents the study participant demographics and cohort characteristics. Of the 552 subjects, CIMT measurements were obtained for 337 individuals, while 221 subjects had CAC assessment. Notably, data on both measurements were available for 145 participants. In total, 242 participants had periodontitis (44%), and the DMFT mean value was 16 in this cohort (ranging from 0 to 28).
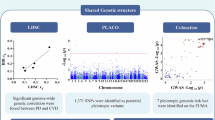
Manhattan plots showing all sequenced SNVs from individuals in the study for (A) DMFT, (B) PSR, (C) CAC, and (D) CIMT. Each dot represents the aggregate significance of an individual SNV. The blue line threshold represents suggestive associations (5 × 10− 5), and the red line threshold represents significant associations (5 × 10− 8). There are approximately 8.5 million SNVs represented and the most significant SNVs associated with each GWAS trait are annotated. Genes marked with an asterisk (*) indicate that the SNV is located near, but not within, the annotated gene.
The Manhattan plots show the SNVs most strongly associated with each phenotype (Fig. 1A–D). Specific associations for each of the top hits are provided in Supplementary Tables 1–4 and Q-Q plots in Suplemmentary Figs. 3, 4, 5 and 6. Genome-wide significant associations were only identified for the CAC phenotype (Fig. 1C, Supplementary Table 3).
The GWAS results for quantitative DMFT scores found suggestive associations with several SNVs (Fig. 1A, supplementary Table 1). Although these results did not reach genome-wide significance of 5 × 10− 8, interesting suggestive associations were identified for the rs79198416 (LOC107985242, near CDC73/KCNT2); p = 7.57 × 10− 7 and for the rs116679466 in IQCA1L, p = 1.15 × 10− 6.
For PSR, there were suggestive associations noted; the rs16854752 (p = 7.44 × 10− 8) is located in the divergent protein kinase domain 2 A (DIPK2A) gene and several other suggestive SNVs are in chromosome 3 (Fig. 1B, supplementary Table 2).
The CAC phenotype showed associations with several SNVs, including rs76676138, rs375047716, rs34729913, rs112265837, rs148981230 (Fig. 1C, supplementary Table 3). The strongest signal was found for SNV rs76676138 which is located near the CCNB3P1 pseudo gene. The other SNVs are located within genes. SNV rs375047716 is in CNTNAP2, and rs34729913 in the uncharacterized area LOC101927281; rs112265837 located in ORAI2. SNV rs148981230 is located in BIN1.
The CIMT phenotype was associated with different SNVs including the rs113152669 in LRBP1; and rs72722175 near ULBP3 (Fig. 1D, supplementary Table 4).
Lastly, there were no regions of shared genetic loci that intersected significantly across phenotypes when a genetic distance of a maximum of 1 Mb was considered for shared genetic associations between traits (p < 5 × 10− 8). The closest significant overlap was identified between the rs375047716 and rs11767967 with CAC levels and the rs116679466 with higher DMFT scores (p = 7.57 × 10− 07, Table 2). The SNV rs375047716 is located in chromosome 7, CNTNAP2 gene. It is approximately 4 million bp away from rs116679466. None of these SNVs are in linkage disequilibrium (Supplementary Fig. 2). However, there were several genetic loci that overlapped between significant CAC results (p < 5 × 10− 8) with suggestive (p < 5 × 10− 5) DMFT and PSR results (Table 2).
Discussion
Assessments of genetic loci associated with both oral disease and ASCVD reported four genetic loci associated with both traits (reviewed by Loos6. Additionally, PLG variants have been reported as associated with both chronic and aggressive periodontitis12 and this gene has been suggested to also play a role in atherosclerosis in clinical studies13,14. Nevertheless, most of these studies were conducted using separate datasets for oral health and ASCVD10,15. In contrast, the present study utilizes a cohort with both oral and ASCVD data collected from the same individuals.
There were no SNVs that significantly intersected across the phenotypes. However, there were several associations with each phenotype individually that are of note. For the DMFT phenotype, SNVs rs79198416 and rs116679466 showed suggestive associations approaching genome-wide significance. The rs79198416 variant lies within an uncharacterized locus LOC107985242 which requires further investigation. The nearest gene is the EEF1A1P14 which is in a genetic locus that has been associated with childhood obesity in a Hispanic population16. The rs116679466 is located within the IQCA1L gene which has been associated with ATP binding17. The PSR phenotype had several SNVs located in the DIPK2A gene that has been shown to have expression in cardiomyocyte proliferation18, which is a novel finding. DIPK2A is involved with other processes including negative regulation of smooth muscle cell apoptotic process, positive regulation of protein kinase C activity, and autophagy19. It has also been demonstrated to inhibit apoptosis and enhance cell survival19. Interestingly, the products of DIPK2A and VAMP8 have been shown to interact19 which is a gene previously identified to overlap in coronary artery disease and periodontitis15. The CAC phenotype showed significant association with SNVs rs76676138, rs373047716, rs34729913, rs112265837, and rs148981230, to name a few. SNV rs76676138 is located near the CCNB3P1 pseudogene. SNV rs373047716 is located in the CNTNAP2 gene which is the largest gene on chromosome 7 and has been associated with multiple neurodevelopmental disorders20. BIN1 is highly relevant in age-related cognitive decline21 but the identified SNV (rs148981230) has never been specifically reported as associated with disease in the literature. The SNV rs112265837 is located in the ORAI2 gene which has been previously associated with calcium channel activity in the plasma membrane and oral cancer cell migration22. The CIMT phenotype was associated with rs113152669 in LRP1B gene which belongs to the LDL receptor family, and several SNVs in or near ULBP3 gene.
Previously identified associations between dental caries and ASCVD may have been confounded by diet and the microbiome, while an involvement of a common SNV seems unlikely in this population according to our results. Dental caries occurs when oral bacteria, especially Streptococcus mutans, metabolize sugars, producing acids that demineralize enamel23,24. This bacterium is not only a key factor for caries but has also been found in cardiovascular specimens, such as heart valves and atheromatous plaques25.
On the other hand, periodontal disease is characterized by inflammation of the gingival tissue and the bone that supports the teeth. This chronic, low-grade inflammation is thought to contribute to a systemic inflammatory response. The inflammation is triggered when bacteria infiltrate periodontal pockets, prompting an immune response from the host. Without appropriate treatment, such bacterial infections can lead to the loss of alveolar bone, loss of teeth, and other complications related to oral health. The two main proposed mechanisms involved with the connection between periodontal disease and ASCVD suggest that subgingival pathogens may directly invade endothelial cells, or an indirect mechanism created by elevated levels of inflammatory cytokines due to periodontitis stimulating a chronic inflammatory response. The first mechanism is supported by the presence of periodontal disease bacteria formed in atherosclerotic lesions in the coronary arteries and the second by the presence of increased levels of IL-1b, IL-6, IL-8 and TNF-a which are cytokines also related to ASCVD26. Interestingly, we recently identified a shared sphingomyelin metabolite in gingivitis and CIMT in this same cohort, which may explain some of these mechanisms27.
In summary, we did not identify significantly associated genetic loci contributing to both oral and atherosclerotic cardiovascular diseases. We found some suggestive intersections that included 1q43, 2q33.1, 3p24.2, 3p25.3, 11q14.2, and 17p13.3 loci that may require further investigation using larger sample sizes. The lack of significant results indicates that the associations between oral health and ASCVD observed in previous studies may be less attributable to shared genetic susceptibility and more influenced by environmental, behavioral, and microbiome-related factors, potentially pointing to a more direct causal relationship. However, this present exploratory study had noted limitations. Given the cross-sectional design, we cannot determine causality between oral conditions and ASCVD to confirm a causal relationship. Additionally, as mentioned before, our sample size could have been a limiting factor in detecting significant associations and a potential overlap. We have also established a liberal significance threshold of 5 × 10− 8 in our GWASs analyses which may have led to finding potential false positive results for CAC trait. Nevertheless, strengths of the study include the presence of precise oral and cardiovascular phenotypes in the Dental/Heart SCORE cohort and the presence of whole genome sequencing data that allowed us to conduct comprehensive GWASs in this population. In conclusion, future studies should explore additional shared multi-omics influences to elucidate mechanistic insights affecting oral and atherosclerotic cardiovascular diseases.
Methods
Participants
The research participants were recruited through a community-based study called Heart Strategies Concentrated on Risk Evaluation (Heart SCORE) that began in 2003 and enrolled 2,000 participants. The study’s primary goal was to investigate racial disparities in cardiovascular disease28. Participants were age 45 to 75, lived in the greater Pittsburgh metropolitan area, and free of known health conditions likely to restrict life expectancy to less than five years. Dental assessments and related data collection were carried out between 2007 and 2010 on a subgroup of 552 participants, known as the Dental Strategies Concentrating on Risk Evaluation (Dental SCORE) study.
All participants in the Heart/Dental SCORE studies provided written informed consent, as approved by the University of Pittsburgh Institutional Review Board. The procedures strictly adhere to regulations and protocols set forth by the Center for Oral Health Research in Appalachia, cohort 1 (COHRA1). The COHRA1/Dental SCORE protocols and genetic data are accessible on dbGaP (accession number phs001591.v1. p1), and extended phenotypic data are available on FaceBase (FaceBase.org, DentalSCORE Record ID: 3P4VSA, Accession: FB00001361, DOI: https://doi.org/10.25550/3P-4VSA).
The cohort data contain comprehensive information on subclinical markers of ASCVD, clinical variables such as blood pressure (i.e., normal, prehypertensive, hypertensive stage 1, and hypertensive stage 2), diabetes, smoking status, HDL cholesterol, and total cholesterol, and self-reported demographic details (including race and ethnicity) for all participants28,29. Participants were categorized based on their smoking habits as current, former, or never smoker.
Phenotypes assessed
Oral health status phenotypes
Oral examinations were carried out by experienced dental professionals, including dentists, dental hygienists, and dental assistants. Inter- and intra-examiner concordances were high as reported elsewhere29,30. The assessment included (i) dental caries, assessed using the decayed, missing (due to caries), and filled teeth index (DMFT) as a continuous trait, and (ii) periodontal health, evaluated with the Periodontal Scoring and Recording index (PSR). For the PSR trait, we assigned a pocket depth score to each sextant of an individual’s mouth based on the most severe finding. Scores were 1 (less than 3.5 mm of pocket depth), 2 (3.5 to 5.5 mm of pocket depth), and 3 (more than 5.5 mm of pocket depth). We used the highest PSR score among the sextants (numbered 1–3) in the analysis.
ASCVD phenotypes
Two subclinical markers of ASCVD were assessed in this study, carotid artery intima-media thickness (CIMT) and coronary artery calcification (CAC). CIMT was measured in millimeters using ultrasound and codified as a continuous trait with the highest value between right and left being used for analysis. Electron-beam computed tomography (EBCT) was used to assess CAC (total volume in millimeters) values and measure the severity of calcification31 as a continuous trait. The CAC score was determined using the Agatston score, which is the sum of the attenuation (in Hounsfield units) and area of CAC lesions in the coronary arteries32.
Genome-wide association analyses and intersection between traits
GWASs were carried out separately on each of the four phenotypes. Study participants were genotyped on the Illumina Infinium Multi-Ethnic Global-8 v1.0 array and subsequently imputed using the Michigan Imputation Server based on the HRC (Version r1.1 2016) reference panel, genome build GRCh37/hg19. Approximately 8.5 million SNVs that met Hardy Weinberg equilibrium (HWE), and imputation quality filtering thresholds33 were initially considered for the GWASs, and further filtered for minor allele frequency as appropriate based on the sample size available for each GWAS. Sex and age were included as covariates in the regression in each GWAS as well as principal components of ancestry to account for population admixture.
Genome wide p-value thresholds of 5 × 10− 8 and 10− 5 were used to identify significant and suggestive associations respectively. SNVs were annotated to genes including or adjacent to the SNV. We also examined whether the associated loci from any of our GWASs overlapped between the four (oral and ASCVD) phenotypes. For the purpose of this comparison, search intervals of 1 megabase length were drawn around suggestive and significant loci from each GWAS and p-values were examined across all four GWASs within each search interval.
The analysis was completed using PLINK genetic analysis software (version 2.050)34 including the 3 most significant principal components of ancestry to adjust for population substructure (Supplementary Fig. 1). Linear regression was used to analyze DMFT, CIMT, and CAC; and PSR was analyzed using a logistic regression framework. The Fastman R package35 was used to generate Manhattan and Q-Q plots.
Data availability
The dataset generated and/or analyzed during the current study are available from the corresponding author on reasonable request. The COHRA1/DentalSCORE protocols and genetic data are accessible on dbGaP (https://www.ncbi.nlm.nih.gov/gap; accession number phs001591.v1. p1), and extended phenotypic data are available on FaceBase (FaceBase.org; DentalSCORE Record ID:3P4VSA Accession: FB00001361 DOI:10.25550/3P-4VSA).
References
Sabharwal, A., Stellrecht, E. & Scannapieco, F. A. Associations between dental caries and systemic diseases: a scoping review. BMC Oral Health. 21, 472. https://doi.org/10.1186/s12903-021-01803-w (2021).
Pitts, N. B., Twetman, S., Fisher, J. & Marsh, P. D. Understanding dental caries as a non-communicable disease. Br. Dent. J. 231, 749–753. https://doi.org/10.1038/s41415-021-3775-4 (2021).
Bashir, N. Z. Update on the prevalence of untreated caries in the US adult population, 2017–2020. J. Am. Dent. Assoc. 153, 300–308. https://doi.org/10.1016/j.adaj.2021.09.004 (2022).
Lee, H. et al. Association between dental health and obstructive coronary artery disease: an observational study. BMC Cardiovasc. Disord. 19, 98. https://doi.org/10.1186/s12872-019-1080-9 (2019).
McEvoy, J. W. et al. Coronary artery calcium progression: an important clinical measurement? A review of published reports. J. Am. Coll. Cardiol. 56, 1613–1622. https://doi.org/10.1016/j.jacc.2010.06.038 (2010).
Loos, B. G. & Van Dyke, T. E. The role of inflammation and genetics in periodontal disease. Periodontol 2000. 83, 26–39. https://doi.org/10.1111/prd.12297 (2020).
Pussinen, P. J. et al. Association of childhood oral infections with cardiovascular risk factors and subclinical atherosclerosis in adulthood. JAMA Netw. Open. 2, e192523. https://doi.org/10.1001/jamanetworkopen.2019.2523 (2019).
Priyamvara, A. et al. Periodontal inflammation and the risk of cardiovascular disease. Curr. Atheroscler Rep. 22, 28. https://doi.org/10.1007/s11883-020-00848-6 (2020).
Aarabi, G. et al. Genetic susceptibility contributing to periodontal and cardiovascular disease. J. Dent. Res. 96, 610–617. https://doi.org/10.1177/0022034517699786 (2017).
Schaefer, A. S. et al. Genetic evidence for PLASMINOGEN as a shared genetic risk factor of coronary artery disease and periodontitis. Circ. Cardiovasc. Genet. 8, 159–167. https://doi.org/10.1161/CIRCGENETICS.114.000554 (2015).
Schaefer, A. S. et al. Identification of a shared genetic susceptibility locus for coronary heart disease and periodontitis. PLoS Genet. 5, e1000378. https://doi.org/10.1371/journal.pgen.1000378 (2009).
Munz, M. et al. A haplotype block downstream of plasminogen is associated with chronic and aggressive periodontitis. J. Clin. Periodontol. 44, 962–970. https://doi.org/10.1111/jcpe.12749 (2017).
Wang, H. et al. Effect of two lipoprotein (a)-Associated genetic variants on plasminogen levels and fibrinolysis. G3 (Bethesda). 6, 3525–3532. https://doi.org/10.1534/g3.116.034702 (2016).
Bonomi, A. et al. Analysis of the genetic variants associated with Circulating levels of sgp130. Results from the IMPROVE study. Genes Immun. 21, 100–108. https://doi.org/10.1038/s41435-019-0090-z (2020).
Munz, M. et al. Genome-wide association meta-analysis of coronary artery disease and periodontitis reveals a novel shared risk locus. Sci. Rep. 8, 13678. https://doi.org/10.1038/s41598-018-31980-8 (2018).
Comuzzie, A. G. et al. Novel genetic loci identified for the pathophysiology of childhood obesity in the Hispanic population. PLoS One. 7, e51954. https://doi.org/10.1371/journal.pone.0051954 (2012).
Alliance of Genome Resources. Harmonizing model organism data in the alliance of genome resources. Genetics 220 https://doi.org/10.1093/genetics/iyac022 (2022).
Beigi, F. et al. C3orf58, a novel paracrine protein, stimulates cardiomyocyte cell-cycle progression through the PI3K-AKT-CDK7 pathway. Circ. Res. 113, 372–380. https://doi.org/10.1161/CIRCRESAHA.113.301075 (2013).
Tian, X. et al. DIPK2A promotes STX17- and VAMP7-mediated autophagosome-lysosome fusion by binding to VAMP7B. Autophagy 16, 797–810. https://doi.org/10.1080/15548627.2019.1637199 (2020).
Rodenas-Cuadrado, P., Ho, J. & Vernes, S. C. Shining a light on CNTNAP2: complex functions to complex disorders. Eur. J. Hum. Genet. 22, 171–178. https://doi.org/10.1038/ejhg.2013.100 (2014).
Seshadri, S. et al. Genome-wide analysis of genetic loci associated with alzheimer disease. JAMA 303, 1832–1840. https://doi.org/10.1001/jama.2010.574 (2010).
Singh, A. K. et al. Orai-1 and Orai-2 regulate oral cancer cell migration and colonisation by suppressing Akt/mTOR/NF-kappaB signalling. Life Sci. 261, 118372. https://doi.org/10.1016/j.lfs.2020.118372 (2020).
Machiulskiene, V. et al. Terminology of dental caries and dental caries management: consensus report of a workshop organized by ORCA and cariology research group of IADR. Caries Res. 54, 7–14. https://doi.org/10.1159/000503309 (2020).
Lemos, J. A. et al. The biology of Streptococcus mutans. Microbiol. Spectr. 7 https://doi.org/10.1128/microbiolspec.GPP3-0051-2018 (2019).
Soto-Barreras, U. et al. Peripheral arterial disease associated with caries and periodontal disease. J. Periodontol. 84, 486–494. https://doi.org/10.1902/jop.2012.120051 (2013).
Zardawi, F., Gul, S., Abdulkareem, A., Sha, A. & Yates, J. Association between periodontal disease and atherosclerotic cardiovascular diseases: revisited. Front. Cardiovasc. Med. 7, 625579. https://doi.org/10.3389/fcvm.2020.625579 (2020).
Bezamat, M. et al. Oral disease and atherosclerosis May be associated with overlapping metabolic pathways. JDR Clin. Trans. Res. 10, 315–323. https://doi.org/10.1177/23800844241280383 (2025).
Aiyer, A. N. et al. Racial differences in coronary artery calcification are not attributed to differences in lipoprotein particle sizes: the heart strategies concentrating on risk evaluation (Heart SCORE) study. Am. Heart J. 153, 328–334. https://doi.org/10.1016/j.ahj.2006.11.002 (2007).
Polk, D. E. et al. Study protocol of the center for oral health research in appalachia (COHRA) etiology study. BMC Oral Health. 8, 18. https://doi.org/10.1186/1472-6831-8-18 (2008).
Wendell, S. et al. Taste genes associated with dental caries. J. Dent. Res. 89, 1198–1202. https://doi.org/10.1177/0022034510381502 (2010).
Arad, Y., Spadaro, L. A., Goodman, K., Newstein, D. & Guerci, A. D. Prediction of coronary events with electron beam computed tomography. J. Am. Coll. Cardiol. 36, 1253–1260. https://doi.org/10.1016/s0735-1097(00)00872-x (2000).
Obisesan, O. H., Osei, A. D., Uddin, S. M. I., Dzaye, O. & Blaha, M. J. An update on coronary artery calcium interpretation at chest and cardiac CT. Radiol. Cardiothorac. Imaging. 3, e200484. https://doi.org/10.1148/ryct.2021200484 (2021).
Orlova, E. et al. Association of early childhood caries with bitter taste receptors: A meta-analysis of genome-wide association studies and transcriptome-wide association study. Genes (Basel) 14. https://doi.org/10.3390/genes14010059 (2022).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575. https://doi.org/10.1086/519795 (2007).
Paria, S. S., Rahman, S. R., Adhikari, K. & fastman A fast algorithm for visualizing GWAS results using Manhattan and Q-Q plots. htttps://arXiv.org/abs/2022.2004.2019.488738. https://doi.org/10.1101/2022.04.19.488738 (2022).
Acknowledgements
We would like to thank the participants of the Heart and Dental SCORE projects. Mariana Bezamat was partially supported by the National Heart, Lung, and Blood Institute under Award number R25HL105400 to Víctor G. Dávila-Román, Dabeeru C. Rao and Lisa de las Fuentes. Dental SCORE was supported by NIH grants R01-DE014899, U01-DE018903, X01-HG009878 to Drs. Marazita, McNeil, Shaffer, and Foxman. The Heart SCORE study was funded by the Pennsylvania Department of Health (ME-02-384). The department specifically disclaims responsibility for any analyses, interpretations, or conclusions. Additional funding was provided by National Institutes of Health grant R01HL089292. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceptualization: Mariana Bezamat, Anum Saeed, Lisa de las Fuentes, Lei Liu, Steven Reis, Mary Marazita.Investigation: Lei Liu, Mariana Bezamat, Nandita Mukhopadhyay, Dylan Baxter, Lisa de las Fuentes, Steven Reis, Mary Marazita, Daniel McNeil, John Shaffer, Betsy Foxman.Methodology: Nandita Mukhopadhyay, Lei Liu, Anum Saeed, Dylan Baxter, Mariana Bezamat, Steven Reis, Mary Marazita.Software: Dylan Baxter, Nandita Mukhopadhyay, Mariana Bezamat.Writing original draft: Mariana BezamatReview and editing: Anum Saeed, Steven Reis, Mary Marazita, Lei Liu, Dylan Baxter, Lisa de las Fuentes, Daniel W. McNeil, John Shaffer, Betsy Foxman.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bezamat, M., Baxter, D.J., Mukhopadhyay, N. et al. Genetic susceptibility to oral and atherosclerotic cardiovascular diseases based on dental and heart SCORE studies. Sci Rep 15, 33257 (2025). https://doi.org/10.1038/s41598-025-18651-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-18651-1