Abstract
Conventional biometric identification methods relying on Personally Identifiable Information (PII) pose significant challenges concerning privacy and security. Volatile organic compounds (VOCs) in exhaled breath are unique to individuals and can serve as biomarkers for various diseases, making them a promising tool for both bioidentification and clinical diagnostics. This research investigates the feasibility of utilizing VOCs in exhaled breath as biometric identifiers, employing machine learning algorithms for analysis. Additionally, the research investigates the use of VOCs as non-invasive indicators of health status, specifically for estimating BMI and gender. The study involved 94 participants with an average age of 67 years and an average BMI of 28 kg/m². Exhaled breath samples were collected using a portable electronic nose (e-nose) device, which analyzed the VOCs. Machine learning algorithms were applied to the data to assess the feasibility of identifying individuals and estimating BMI and gender based on VOC patterns. The results indicated that VOC patterns could reliably estimate BMI and gender, and potentially distinguish between individuals, suggesting VOCs as a viable tool for bioidentification. The application of machine learning to VOC data showed promise in non-invasive identification and diagnostics. VOCs in exhaled breath offer a novel, non-invasive method for biometric identification and health monitoring. This approach could overcome the limitations of traditional PII, providing a new avenue for personalized medicine. Further research is needed to enhance the accuracy and applicability of this method in diverse populations.
Similar content being viewed by others
Introduction
Biometrics (from Greek bios = “life”, metron = “measure”) refers to identifying and authenticating individuals using measurable body features, offering a more secure alternative to traditional IDs and passwords. Unlike traditional methods, biometric data cannot be forgotten, stolen, or lost, though they may be subject to alteration. Fingerprints that are widely used in ID cards and passports date back to the 14th century, however, the reliability of fingerprints can be affected by skin disorders and/or moisture making fingerprint recognition imperfect and boosting the development of other features recognition and intensive studies in the evaluation of biometric systems. The biometric features should be universal, durable, unique, and measurable quantitatively, as well as the efficiency of its acquisition, ethically and socially acceptable, and difficult to fake and circumvent1.
Currently, biometrics can be categorized into two main groups:
-
Physiological like fingerprint, face recognition2,3,4,5iris image6,7location of the veins on the hand, etc.
-
Behavioural, such as voice8respiratory profile9,10,11,12,13signature, gait14,15screen touch gestures, screen, or key pressure level as well as typing speed, etc16,17.
The details of each method are provided in the Supplementary Materials.
The most popular are unimodal authentication systems using biometrics, see Fig. 1, but for the sensitivity to noisy data, intraclass variation and similarities, non-universality, and spoofing it needs development and use of several biometric features for identification, where the obtained result are fused and compared with the templates present in the database for proper authentication18,19,20,21,22. Recent reviews also emphasize the importance of integrating sustainable materials and additive manufacturing techniques in biomedical technologies, including the future development of low-waste, eco-friendly diagnostic devices23.
Non-invasive body analysis.
Human breath
Exhaled human breath is a component of the major gases such as nitrogen (78–79%), oxygen (13–16%), and carbon dioxide (4%)24, as well as the volatile organic compounds (VOCs), that can originate endogenously and exogenously, becoming biomarkers of many diseases and becoming a metabolic, physiological, or even pathophysiological source of information.
VOCs can be used as an information source for bioidentification. The research on breath analysis is not new25but nowadays, the exhaled breath pattern has become an interface between medicine and engineering, making it possible to analyse each exhaled molecule.
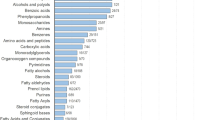
So far, more than 3500 VOCs have been identified in the human breath samples26,27,28however, the total number of VOCs that can be found is not confirmed due to the instrumental limitations. Therefore, depending on the method used for the detection, various numbers could be found in the literature. For example, Phillips et al. have reported 204 VOCs27Smolinska et al. have reported 300–500 VOCs26and Barash et al. have identified more than 500 VOCs in each breath sample29. However, what is interesting from the bioidentification point of view, apart from the group of VOCs that are called biomarkers (which means that their concentrations are highly related to the current condition of the body) and the group of VOCs that are inhaled from the ambient environment, it seems that there is a group of VOCs that are not affected by health conditions either by inhalations and can be associated with each person, similarly to fingerprints. This hypothesis needs to be confirmed on a large scale, although it seems to be a subject of research that is worth investigating. Moreover, the breath can be used for bioauthentication not only as a VOC analysis but also as a breath acoustic analysis11,12,30,31.
Each breath is a good indicator of the state of our health32. Within the technological progress, the breath pattern can be analysed with several tools like gas chromatography (GC) equipped with mass spectrometry (MS)33,34infrared spectroscopy24,35,36chemiluminescence37,38laser ablation spectroscopy39,40and electrochemical tools41,42. These techniques make it possible to detect not only the most common exhaled gases like nitric oxide and carbon dioxide but also a multitude of VOCs, many of which can work as systemic biomarkers or respiratory disease biomarkers43. Systemic biomarkers refer to the body’s functioning, while lung biomarkers are related to processes taking place in the respiratory system44.
Also, carbon nanomaterials such as graphene and carbon nanotubes are increasingly used in advanced sensor platforms for VOC detection due to their high surface area, electrical conductivity, and tunable surface chemistry. These materials enable the development of highly sensitive and selective chemical vapor sensors and biosensors, particularly for non-invasive diagnosis of diseases including cancer, through exhaled breath analysis45.
Recently, a huge effort has been made in the field of fabrication of portable detectors for gas-sensing applications called electronics noses (e-noses), including exhaled human breath analysis. With special emphasis on exhaled breath analysis. In general, the breath sensors can be split into two groups: a targeted approach with a selective chemical sensor, and a unique gas-target pattern recognition with semi-selective/cross-reactive sensors. For example, in46 an e-nose for exhaled breath analysis in the context of silicosis was proposed. The exhaled breath was measured using a matrix, which included 16 sensors made with organic nanofiber. The random forest and extreme gradient boosting-based algorithms as well as k-nearest neighbour (kNN) and support vector machine, were used as breath classifiers. Proposed breath diagnostics methods have good accuracy, in both screening and early detection models. Recently, Thakur et al.47 developed a noninvasive diabetes diagnostics tool. This e-nose used a matrix based on hybrid graphene oxide (GO) field effect transistors (FETs) with a hybrid channel of GO, tungsten trioxide (WO3). As an odour pattern of various breath classifiers, linear discriminant analysis (LDA) and artificial neural networks (ANN), in particular, a multilayer perceptron with an adaptive learning rate of 0.1, were applied. The first one decreases the response vector dimension. To predict the concentration of breath markers, partial least squares (PLS) and multiple linear regression (MLR) were used. The method proposed provides good accuracy, even 99.1%, in the case of artificial breath, and can successfully be applied to the real breath of subjects, who are suffering from diabetes. The exhaled breath analysis in terms of medical applications is well-known, and it has been a subject of research for the last four decades, fractioned exhaled nitric oxide (FeNO)48 and exhaled breath condensate (EBC) analysis49 have been approved by medical societies for clinical use. Others are still under investigation, e.g., diabetes50.
Exhaled breath pattern as a fingerprint
Up until now, several groups of scientists have conducted a study of breath pattern repeatability in humans for breaths taken at different intervals using breath analysis systems such as mass spectrometry, e-nose systems nose containing an array of quartz microbalance sensors (QCM)51.
Martinez-Lozano Sinues et al. examined nine working days of breaths taken from a group of 11 people (on average 18 samples per person) using mass spectrometry. Research showed high inter- and intra-subject variability in circadian levels of exhaled acetone. Principal component analysis (PCA) and the representation of mass spectra in three dimensions showed that for the respondents, there is a combination of samples into separate groups, denoting individual people. Studies have shown that for eleven days (nine working days + weekend), breaths taken from humans contain some of the constant compounds that make it possible to distinguish people, as well as compounds that vary daily and depend on external factors52.
Fasola et al. investigated the repeatability of exhaled breath fingerprints among a group of 15 asthma patients and 30 healthy people. For this purpose, they used Pneumopipe® plus an array of e-nose sensors containing eight QCM oscillating with a resonance frequency of 20 MHz in the thickness-shear mode. For children with asthma, the within-day median intra-class correlation coefficient (ICC) was 0.57, and between-day was 0.66, while for healthy children it was 0.58 and 0.55, respectively. In both study groups, most of the studies were statistically significant in terms of the repeatability of the e-nose system measurements over three visits to the doctor51.
Incalzi et al. examined 25 people aged 65 years and older with chronic obstructive pulmonary disease (COPD) and healthy control samples using an e-nose containing seven quartz microbalance sensors covered with metalloporphyrins. The breaths were collected into Tedlar bags 7 and 15 days after the first measurement. Repeatability of the VOC pattern was observed in subsequent measurements for both sick and healthy people, but the measurements at weekly intervals were more often repeated in Global Initiative for Chronic Obstructive Lung Disease (GOLD) 4 patients than in COPD patients from the GOLD 1–3 group53.
Krilaviciute et al. took breath samples from 1,447 healthy elderly people and then tested them using the GC-MS combined with a thermal desorption system. They analyzed the relationship between the 15 most frequently recurring VOCs in breath and socio-demographic factors and food intake. Studies have shown that it is possible to distinguish the sex of a test subject based on the respiratory profile54.
Wang et al. proposed an e-nose system that can be used to identify people. Breath samples were taken several times from ten people in a 3-L impermeable gas bag (Shanghai Sunrise Instrument Co., China). Three samples were collected from each person during each of the four visits at intervals of at least one week. Samples were analyzed with an electronic nose instrument (eDiagNose) containing 12 individual gas sensors consisting of six types of doped tin dioxide (SnO2) sensors and one type of WO3 sensor. PCA analysis of 10 features selected by mutual information (MI) analysis showed that there is a separation of sample groups belonging to individual people. By training SVM, kNN, Mutual information maximum likelihood (MI ML), and ANN models. The authors have stated that the obtained classification accuracy is 100% on 120 samples with cross-validation (119 training, 1 test) for almost all tested classifiers, regardless of the number of selected features55. However, it seems that the measurement protocol was invalid, i.e., the measurements were conducted 4 times for 3 samples taken from one subject, but from the same bag, and that is why 100% accuracy was obtained. There is a lack of data on how the signal is changing when the same bag is used on various days, for instance.
In turn, Filosa et al. proposed the meta-learning algorithm for the prediction of respiratory flow based on LSTM neural networks56. The data come from the system, which includes a wearable system embedding Fiber Bragg Grating (FBG) sensors and inertial units. It turned out that the algorithm proposed to give a better result, both in static and dynamic conditions, in comparison to machine learning-based methods.
Breath pattern identification can be used to distinguish people using e-nose and AI algorithms, see Fig. 2.
Breath pattern identification.
Stable VOCs as a “known-unknown” problem
Since it is not well established yet that there are sets of stable VOCs that are not affected by a person’s health condition and other environmental factors that are unique to each person due to the small number of people that were used in existing studies, this hypothesis needs to be confirmed on the large scale and can be akin to “known-unknown” problem57.
There are known tools based on machine learning algorithms for predicting unknown chemical compounds because there is a lack of annotations in databases58,59,60. A similar method can be used to discover and classify VOCs in breath to identify them as unique for bioidentification.
This paper discusses the features that are required to recognize a feature as suitable for bioidentification and known biometric features such as recognition of the iris, face, gait, etc. The studies of volatile organic compounds and the verification of their intra and inter-subject repeatability and durability have been reviewed to find out if it is further possible to identify breath patterns as a novel bioidentification method.
Main contributions
This study makes the following key contributions to the field of non-invasive biometric identification and health monitoring through breath analysis:
-
To the best of our knowledge, we demonstrate for the first time the concurrent use of exhaled volatile organic compounds for biometric identification and estimation of physiological traits (specifically, gender and BMI). This integrated approach expands the conventional scope of VOC-based diagnostics, offering a new application domain for breath analysis beyond disease detection.
-
By applying machine learning algorithms to data acquired from a portable electronic nose, we achieved good robust classification performance, which highlights the potential of AI-enhanced e-nose systems for rapid, accessible, and non-invasive biometric screening.
-
The study explores the potential of VOCs for distinguishing individuals, indicating that breath composition contains identifiable, person-specific patterns.
-
Unlike traditional biometric systems that depend on personally identifiable information or external features (e.g., fingerprints, facial recognition), this work introduces a privacy-enhancing alternative by leveraging internal biological signatures.
Related research
Recent advancements have highlighted the potential of exhaled breath analysis, particularly VOCs in non-invasive diagnostics and biometric applications. For instance, a novel pre-clinical method was developed to capture exhaled breath from intubated mice, significantly reducing background contamination and enhancing VOC detection accuracy. This approach, integrating sorbent tubes with respiratory measurement equipment, facilitates high-quality sample collection for downstream GC-MS analysis and supports translational breathomics research61. In the realm of biometric authentication, one study explored fluid dynamics in exhaled breath, achieving over 97.00% confirmation accuracy using an attention U-Net model applied to time-series breath turbulence data. This study demonstrates the potential of breath flow patterns for personal identification. On the other hand, researchers applied AI to VOC profiles for early lung cancer detection, using an agnostic strategy that emphasized compositional breath differences rather than specific biomarkers. This approach shows strong classification performance and the utility of AI-driven breath analysis as a rapid and cost-effective diagnostic tool62. VOC-based diagnostics have also shown promise for colorectal cancer, where 12 Key compounds were statistically identified to differentiate between healthy and affected individuals, with sensitivity and specificity values of 0.80 and 0.85, respectively - highlighting breath analysis as a clinically practical early screening tool63. For chronic respiratory conditions, such as COPD, asthma, and PRISm, comprehensive breathomics approaches using portable micro gas chromatography and machine learning demonstrated high classification performance (AUC up to 0.92), validating their feasibility for real-time, non-invasive disease differentiation62,64.
In breast cancer diagnostics, a large-scale study involving over 1,900 women used high-pressure photon ionization TOF-MS combined with machine learning to distinguish disease presence (AUC = 0.946) and monitor progression, marking a significant advance in breath-based oncology screening65. A multimodal sensor array incorporating metal oxide, electrochemical, and photoionization sensors, paired with a 1D convolutional neural network, also achieved 97.80% accuracy in lung cancer classification, reinforcing the power of deep learning and multi-sensor breath analysis in clinical applications66. Lastly, a standardized VOC identification workflow using Owlstone Medical’s OMNI® platform verified 148 on-breath VOCs across a heterogeneous population. This benchmark dataset supports biomarker discovery and promotes cross-study comparability, addressing a key need for consistency in breathomics research67. In contrast to these studies focused primarily on disease diagnostics, our work uniquely investigates VOCs in exhaled breath as a dual-purpose tool, for both biometric identification and health status estimation (BMI, gender). By leveraging portable e-nose and machine learning models, we show that breath-based VOC signatures can provide a privacy-preserving, scalable alternative to traditional PII-dependent biometrics while enabling non-invasive personal health monitoring. The summary of breath-based biometric and respiratory monitoring studies is presented in Table 1.
Results
In cooperation with the Department of Prosthodontics and Orthodontics, Dental Institute, Faculty of Medicine, Jagiellonian University Medical College, Krakow, Polska (Jagiellonian University ethical committee approval КВЕТ: 1072.6120.40.2023), we have collected exhaled air from people.
Each of the 94 study participants completed a survey containing questions about gender, weight, height, and age. The histogram of the research sample is presented in Figs. 3, 4 and 5. Table 2 provides a summary of key statistics for the patient’s BMI, age, weight, height, and biochemical markers such as glucose, triglycerides, uric acid, and cholesterol. For each variable, the table includes the mean, median, standard deviation, minimum, and maximum values, providing insight into the central tendency and spread of the data. Table 3. summarises the distribution of categorical variables representing the presence or absence of specific diseases. For each condition (e.g., diabetes, hypertension, hypothyroidism, and cancer), the table shows the count of people classified as having the disease (“Yes”) and not having the disease (“No”). We did not ask to follow a special diet or refrain from physical activity before the study to not distorting the results and to simulate the real conditions of the biometric studies. The patients did not smoke cigarettes or drink alcohol immediately before breath collection.
Gender distribution.
BMI distribution.
Age distribution.
To analyse the breath samples collected from participants, we employed several machine learning algorithms designed for regression and classification tasks. Specifically, we used linear regression, Random Forest, Light Gradient Boosting Machine (LGBM), Extreme Gradient Boosting (XGB) for predicting Body Mass Index (BMI), and a set of classification algorithms - logistic regression, Random Forest, Light Gradient Boosting Machine, Extreme Gradient Boosting, and Support Vector Classifier - for gender classification.
These algorithms were selected based on their established performance in biomedical and sensor-based data analysis72,73,74,75,76,77,78,79. The models were trained and evaluated using the dataset of VOC sensor responses obtained from the e-nose system. A brief explanation of each model and its working principles is provided in the Methods section.
The linear regression algorithm was used to predict BMI. The algorithm achieved a mean absolute error (MAE) of 4.09 and a root mean squared error (RMSE) of 6.01. The prediction results on the test set are presented in Fig. 6. Linear regression may be the best model when the simplicity and interpretability of results are important. Although it has a slightly higher MAPE (14.72%) than Random Forest and LGBM, it offers a good balance between other metrics such as MAE, MSE, and Pearson correlation, which is the highest (0.2621). However, it is worth noting that MAE, MSE, and RMSE in linear regression are strongly affected by a single outlier, where the actual value is the lowest and the prediction is the highest. This single point strongly influences these error measures, which can cause these values to appear higher than in other models. Linear regression also has a relatively high standard deviation (4.68), which means that the model is more diverse in its predictions, unlike some other models, such as Random Forest, where the standard deviation is very low (0.34), suggesting that the model predicts similar values for different data. Models such as LGBM or XGB have a standard deviation of 1–3, indicating less variability in the forecasts. Linear Regression is more flexible, which makes it better at dealing with a variety of data while maintaining simplicity and stability. The prediction results on the test set using the Linear Regression model are presented in Fig. 6. Detailed comparison of different regression models’ performance is presented in Table 4.
BMI prediction result.
We compared several classification models to evaluate their performance (Table 5). Researchers employed a logistic regression algorithm for gender classification, achieving an accuracy of 87.5%, precision of 87.5%, recall of 93%, and an F1 score of 90%. SVC also performed very well, achieving 100% sensitivity and an F1 Score of 83.33%. The remaining models, such as Random Forest, LGBMClassifier, and XGBClassifier, achieved similar but significantly lower scores, indicating their less usefulness in the gender prediction based on exhaled breath analysis. The confusion matrix of the Logistic regression model is presented in Fig. 7.
Gender classification confusion matrix.
Discussion
Studies mentioned in this paper report results from breath measurements of individuals and various groups. The findings indicate both intra- and inter-subject variability. However, stable volatile organic compounds have been identified, which, in conjunction with e-nose technology and artificial intelligence, can effectively differentiate individuals52 and their sex54. The unique nature of exhaled breath patterns aligns with existing literature emphasizing their diagnostic and identification potential. For example, Martinez-Lozano Sinues et al.80 demonstrated individual metabolic phenotypes through breath mass spectrometry analysis, supporting the claim that VOCs offer durable and distinguishable profiles. Additionally, studies on e-nose technology have highlighted its feasibility for non-invasive diagnostics, such as its application in the early detection of diseases like diabetes and respiratory conditions47. By leveraging these insights, the current study builds a strong foundation for further exploring VOC-based biometric identification.
Our findings align with previous research demonstrating the potential of e-nose technology for breath analysis in biometric and medical applications. Studies such as those by Fasola et al.. and Incalzi et al. have shown the repeatability and reliability of e-nose measurements across different populations, reinforcing the validity of our approach51,53. Additionally, Krilaviciute et al. provided evidence that breath composition varies by gender, supporting our gender classification results54. The work by15 et al. highlights the importance of rigorous measurement protocols, as overly controlled conditions may lead to inflated accuracy. Finally, Filosa et al.56. demonstrated the advantages of advanced AI methods, suggesting that deep learning techniques could further enhance BMI prediction accuracy in our study. These comparisons emphasize the potential of integrating VOC analysis with AI for robust and practical biometric applications.
Moreover, breath might fulfil another requirement of biometrics, which is durability, because there is proof that breath samples collected from people have statistically significant repeatability even over time points51 and fifteen days53. More specifically, in the study conducted by Fuchs et al.81 to investigate aldehydes as biomarkers of lung cancer, their result showed that aldehydes were detected in all healthy volunteers, smokers, and lung cancer patients. Furthermore, the concentrations of acetaldehyde, propanal, butanal, heptanal, and decanal were stable for cancer patients, smokers, and healthy volunteers. However, all these studies were performed on a small sample of people and for a short period; hence, there is still a need to conduct large-scale research to prove unequivocally that there is a repeatable and unique VOC pattern observed for each person. However, the studies reviewed in this paper suggest that there is great potential for the use of breath as a bioidentification method.
Additionally, replicating or altering an individual’s breath pattern is highly challenging, providing an inherent advantage over methods like face detection, which are vulnerable to spoofing via GAN-generated images5. The development of a comprehensive breath database is essential for advancing this field. Such a database would facilitate the testing of state-of-the-art algorithms, including Siamese neural networks (i.e., a type of deep learning model that uses two or more identical networks to compare and analyse similarities or differences between inputs, such as images or data patterns) and transfer learning models, to improve classification accuracy82. Furthermore, ethical considerations regarding privacy and data security must be addressed, particularly given the sensitive nature of biometric data. Establishing clear regulatory frameworks and adopting secure encryption methods are critical to mitigating risks and ensuring public trust in this emerging technology. Unlike facial recognition, which can be fooled by a photo, video, or 3D model, VOCs are much harder to fake because they rely on the body’s unique metabolic processes. Fingerprints and irises offer high accuracy but can be copied or used without the user’s consent, such as by taking an imprint from a surface. VOCs can increase security where physical contact is a problem, such as in high-sanitation areas where traditional scanners are less practical. Although the technology has yet to be developed, VOCs could complement traditional methods by providing an additional layer of security in specific applications.
Despite its promising findings, this study has notable Limitations. The sample size of 94 participants, primarily white Caucasians, restricts the generalizability of the results. Moreover, the lack of data on participants’ chronic diseases or environmental exposures introduces potential confounding variables. Future research should prioritize recruiting a larger, more diverse cohort, including individuals from different ethnicities, age groups, and health statuses. Additionally, longitudinal studies are necessary to evaluate the stability of VOC patterns over time and across varying conditions. Second, all participants were white Caucasian, so we could not explore differences between races. On the basis of these preliminary findings, future studies should explore several critical avenues. First, the scalability of VOC-based biometrics must be tested with larger, ethnically diverse populations. Second, the integration of multimodal biometrics, combining VOCs with physiological or behavioural traits such as voice or gait, could improve the precision of identification. Third, advancements in sensor technology, such as hybrid e-nose systems with high sensitivity and specificity, would further refine VOC detection. Finally, collaborations with clinical settings to validate the method’s diagnostic utility for comorbid conditions could broaden its applicability in personalized medicine.
Conclusions
Human breath is composed of various volatile organic compounds that can be used as input data to an effective diagnostic algorithm for early detection of a wide range of diseases, even at the onset of the disorder. Exhaled volatile organic compounds are the information source on pathophysiological mechanisms and host response to infections in real-time. The profiling of the volatile organic compounds provides non-invasive monitoring of pathological changes that take place in the body. VOCs can be successively applied in clinical practice, particularly as biomarkers for certain pathologies, including lung disease83.
Our study demonstrates that breath composition varies based on gender and BMI, distinctions that can be detected using an electronic nose (e-nose). For instance, logistic regression achieved an F1 score of 90% for gender classification, while linear regression yielded a mean absolute error (MAE) of 4.06 kg/m² in BMI prediction. These findings highlight the potential of integrating artificial intelligence with VOC analysis for rapid and reliable biometric applications. However, BMI prediction accuracy could be further improved using more advanced algorithms, such as deep neural networks or ensemble methods, which are better suited for capturing complex, non-linear relationships in biometric data.
While the combination of commercially available sensors and machine learning algorithms enables basic biometric differentiation, precise individual identification requires additional sensors and more advanced breath analysis using high-precision instruments like mass spectrometers or gas chromatographs. Reliable bioidentification through VOC analysis necessitates stable e-nose sensors, regular calibration, and careful consideration of environmental and biological factors influencing VOC composition. Long-term data collection is crucial to assess the stability of VOC profiles over time. For effective biometric identification, VOC patterns must be both unique and consistent, which can be validated through high-precision analytical techniques. Preliminary studies confirm the feasibility of e-nose-based bioidentification, with future research focused on expanding the study population and conducting long-term verification.
Challenges and advances in exhaled breath bioidentification
Exhaled breath-based biometric identification, while promising, faces several technical and biological challenges that must be carefully addressed to ensure its reliability, consistency, and scalability in real-world applications. One of the primary challenges is sensor stability and reproducibility. Low-cost e-nose sensors often suffer from signal drift over time and require frequent calibration to maintain accuracy, which limits their long-term reliability in biometric applications. Additionally, environmental and physiological variability, including diet, circadian rhythms, medications, and ambient air quality, can significantly influence VOC profiles, complicating the extraction of consistent individual-specific patterns45,68. Moreover, the lack of standardization in breath sampling protocols, sensor designs, and data preprocessing methods impedes cross-study comparability and broader adoption of breath-based identification systems.
Another key challenge is the lack of standardization in sampling protocols, sensor configurations, and data preprocessing, which impedes reproducibility and hinders cross-study validation84. Most commercially available e-noses cannot yet match the analytical resolution of laboratory-based techniques such as gas chromatography-mass spectrometry (GC-MS) or ion mobility spectrometry (IMS), which are capable of detecting trace-level VOCs with high specificity85. Nevertheless, nanotechnology-enabled sensors, incorporating materials such as graphene, carbon nanotubes, and metal-organic frameworks (MOFs), are rapidly advancing sensor performance by enhancing VOC selectivity and miniaturization potential86,87.
Thus, machine learning and artificial intelligence are being increasingly adopted to address the complexity of VOC datasets. For example, ensemble methods and deep learning architectures are now being explored to capture non-linear relationships in breath data, improving classification and prediction performance beyond traditional models.
To mitigate these challenges, hybrid platforms that combine portable e-nose systems with high-resolution analytical instruments, as well as the application of sustainable and robust materials in sensor fabrication, are emerging as promising solutions88. Furthermore, multi-sensor fusion, longitudinal data acquisition, and the development of standardized protocols will be essential for achieving consistent, individual-specific VOC patterns suitable for reliable bioidentification. As the field progresses, the integration of advanced AI models with next-generation sensors is expected to play a pivotal role in transforming exhaled breath into a practical, privacy-preserving biometric modality.
Materials and methods
Review methodology
The review methodology was based on the PRISMA Statement89 and its extensions: PRISMA-S90,91as well as our personal experience. We considered recent publications, reports, and protocols, and reviewed papers from Scopus databases. The documents used in the presented study were selected based on the procedure presented in Fig. 8. The keywords: “breath pattern” and algorithm” were used; as a result, we obtained: 293 documents, including 196 research papers, 85 conference papers, and 12 reviews. The selection process was done according to the following overall criteria regarding bio identification, biostatistics, and neural networks. Finally, 101 documents were considered. The 85 conference papers, as well as papers without connections to the criteria above, have been excluded from the study.
Literature search flowchart.
Breath collection
We asked participants to draw in air into their lungs and hold it for a few seconds, and then blow a single breath into a Tedlar®Bags92,93,94,95 and then we measured using an e-nose, namely an electronic nose, i.e., a portable device equipped with sensors designed to detect and analyse chemical compounds, such as volatile organic compounds in the air).
E-nose system
Breaths stored in Tedlar® Bags92,93,94,95 were measured using e-nose. E-nose (Fig. 9) consisted of a system pumping air from bags and a set of sensors TGS1820, TGS2620, TGS2600, TGS2660 (Figaro Engineering Inc, Mino, Osaka, Japan), MQ3 (Winsen, ZhengZhou, HeNan, China), 7e4 NO2, 7e4 H2S (SemeaTech, Los Angeles, USA and Shanghai, China), SGX NO2, SGX H2S (SGX SENSORTECH, Switzerland), K33 (Senseair, Delsbo, Sweden), AL-03P, AL-03 S (MGK SENSOR Co., Ltd., Saitama, Japan).
E-nose system.
The breath from each patient’s bag was pumped using a pump in our e-nose system to the sensors in the chamber. The breath was pumped for 25 min, and after each breath dose, the chamber was flushed with air passing through the filter for 15 min. For each bag, the procedure was repeated twice. Data from each sensor was measured and saved every 20 s. Original data was shown in Fig. 10.
Example of original data collected during breath samples measurement (samples no 714, 763, 757, 417). Orange dots mean RG and black dots mean R0.
We used data collected from sensors in the form of relative response RG and sensor response S (Eq. 1) to train machine learning algorithms.
where,
RG – Sensor response to collected breath sample.
R0 – Sensor response to filtered ambient air.
The algorithms’ task was to predict a person’s BMI and gender. The data was divided into training and test sets, and hyperparameter optimization was performed for each algorithm using the Random Search CV method (Scikit-learn)96,97.
Machine learning algorithms
In this study, we applied several widely used machine learning algorithms to predict BMI and classify gender based on VOC sensor data. For BMI regression, we used Linear Regression, which models a linear relationship between features and BMI, and tree-based ensemble methods including Random Forest, LightGBM, and XGBoost, known for their robustness and high accuracy. For gender classification, we tested Logistic Regression - a simple linear classifier using a sigmoid function - alongside Random Forest, LightGBM, XGBoost, and Support Vector Classifier, which maximizes class margins and can model nonlinear patterns via kernels. Random Forest uses bagging to reduce overfitting, LightGBM and XGBoost are fast, scalable gradient boosting methods with built-in regularization, and SVC performs well on smaller, high-dimensional datasets. All models were implemented in Python with Scikit-learn and XGBoost libraries, and hyperparameters were tuned using Randomized Search Cross-Validation to enhance performance while preventing overfitting.
Data availability
The datasets generated during and/or analysed during the current study are not publicly available due to the content of patients’ private medical data but are available from the corresponding author on reasonable request.
References
Jain, A., Bolle, R. & Pankanti, S. Introduction to biometrics. Biometrics 1–41. https://doi.org/10.1007/0-306-47044-6_1 (1996).
Ngo, D. C. L., Teoh, A. B. J. & Goh, A. Biometric hash: High-confidence face recognition. IEEE Trans. Circuits Syst. Video Technol. 16, 771–775 (2006).
Petrescu, R. V. Face recognition as a biometric application. SSRN Electron. J. https://doi.org/10.2139/SSRN.3417325 (2019).
Chen, Y. et al. A face recognition method based on CNN. J. Phys. Conf. Ser. 1395, 012006 (2019).
Wang, X., Guo, H., Hu, S., Chang, M. C. & Lyu, S. GAN-generated Faces Detection: A Survey and New Perspectives (2022).
Strzelczyk, P. Privacy preserving and secure iris-based biometric authentication for computer networks. Journal Telecommunications Inform. Technology 4, 115–118 (2011).
Sandhya, M., Morampudi, M. K., Pruthweraaj, I. & Garepally, P. S. Multi-instance cancelable Iris authentication system using triplet loss for deep learning models. Visual Comput. 1–11 https://doi.org/10.1007/S00371-022-02429-X/TABLES/6 (2022).
Park, H. & Kim, T. User Authentication Method via Speaker Recognition and Speech Synthesis Detection. Security and Communication Networks (2022). (2022).
Ward, M. & Macklem, P. T. The act of breathing and how it fails. Chest 97, 36S–39S (1990).
LoMauro, A., Colli, A., Colombo, L. & Aliverti, A. Breathing patterns recognition: A functional data analysis approach. Comput. Methods Programs Biomed. 217, 106670 (2022).
Chauhan, J. et al. Breathing-based authentication on resource-constrained IoT Breathing-based authentication on resource-constrained IoT devices using recurrent neural networks devices using recurrent neural networks. Aruna SENEVIRATNE Cit. Cit. https://doi.org/10.1109/MC.2018.2381119 (2018).
Bui, M. H. et al. Personalized breath based biometric authentication with wearable multimodality. (2021). https://doi.org/10.48550/arxiv.2110.15941
Kumar, A. K., Ritam, M., Han, L., Guo, S. & Chandra, R. Deep learning for predicting respiratory rate from biosignals. Comput Biol. Med 144, 105338 (2022).
Moon, J. et al. Explainable gait recognition with prototyping encoder–decoder. PLoS One. 17, e0264783 (2022).
Wang, X. & Hu, S. Visual gait recognition based on convolutional block attention network. Multimed Tools Appl. 1–18 https://doi.org/10.1007/S11042-022-12831-1/FIGURES/8 (2022).
Bhattacharyya, D., Ranjan, R., Farkhod, A. A. & Choi, M. Biometric authentication: A review. International J. u-and e-Service 2, 13–28 (2009).
Meng, Y., Wong, D. S., Schlegel, R. & Kwok, L. F. Touch gestures based biometric authentication scheme for touchscreen mobile phones. Lecture Notes Comput. Sci. (including Subser. Lecture Notes Artif. Intell. Lecture Notes Bioinformatics). 7763 LNCS, 331–350 (2013).
Sanjekar, P. S. & Patil, J. B. An overview of multimodal biometrics. Signal. Image Processing: Int. J. (SIPIJ). 4, 57–64 (2013).
Bhateja, A., Pal, S. K. & Kumar, S. A. Comparative study of multimodal biometric authentication system: A review. https://doi.org/10.5958/j.2277-4912.2.2.014
Sadeghi, P., Alshawabkeh, R., Rui, A. & Sun, N. X. A comprehensive review of biomarker sensors for a breathalyzer platform. Sens. 2024. 24, 7263 (2024).
Kim, D., Lee, J., Park, M. K. & Ko, S. H. Recent developments in wearable breath sensors for healthcare monitoring. Communications Materials 2024 5:1 5, 1–14 (2024).
Haick, H. Advances in volatile organic compounds detection: from fundamental research to real-world applications. Appl. Phys. Rev. 11, 40401 (2024).
Pesode, P., Barve, S., Wankhede, S. V. & Ahmad, A. Sustainable Materials and Technologies for Biomedical Applications. Advances in Materials Science and Engineering 6682892 (2023). (2023).
Selvaraj, R., Vasa, N. J., Nagendra, S. M. S. & Mizaikoff, B. Advances in Mid-Infrared Spectroscopy-Based sensing techniques for exhaled breath diagnostics. Molecules 25, 2227 (2020).
DAHLSTROM, H. & ROOS, A. MURPHY, J. P. Cardiogenic Oscillations in Composition of Expired Gas. The ‘Pneumocardiogram’. (1954). https://doi.org/10.1152/jappl.1954.7.3.335 7, 335–339.
Smolinska, A. et al. Profiling of volatile organic compounds in exhaled breath as a strategy to find early predictive signatures of asthma in children. PLoS One 9, e95668 (2014).
Phillips, M. et al. Variation in volatile organic compounds in the breath of normal humans. J. Chromatogr. B Biomed. Sci. Appl. 729, 75–88 (1999).
De Costello, L. A review of the volatiles from the healthy human body. J. Breath. Res. 8, 014001 (2014).
Barash, O. et al. Differentiation between genetic mutations of breast cancer by breath volatolomics. Oncotarget 6, 44864 (2015).
Dai, H., Jiang, J., Ma, J., Huang, H. & Liu, H. Breathing-based continuous non-intrusive user verification leveraging commodity WiFi. J. Commun. Netw. 24, 209–222 (2022).
Lu, L., Liu, L., Hussain, M. J. & Liu, Y. I sense you by breath: speaker recognition via breath biometrics. IEEE Trans. Dependable Secure Comput. 17, 306–319 (2020).
Dweik, R. A. & Amann, A. Exhaled breath analysis: the new frontier in medical testing. J. Breath. Res. 2, 030301 (2008).
Kim, C. et al. Recent trends in exhaled breath diagnosis using an artificial olfactory system. Biosensors (Basel) 11, 337 (2021).
Alkhalifah, Y. et al. VOCCluster: untargeted metabolomics feature clustering approach for clinical breath gas chromatography/mass spectrometry data. Anal. Chem. 92, 2937–2945 (2020).
Karunakaran, K. D. et al. Relationship between age and cerebral hemodynamic response to breath holding: A functional Near-Infrared spectroscopy study. Brain Topogr. 34, 154–166 (2021).
Chen, W. T., Yu, C. H. & Sun, C. W. Altered near-infrared spectroscopy response to breath-holding in patients with fibromyalgia. J Biophotonics 12, e201800142 (2019).
Robinson, J. K., Bollinger, M. J. & Birks, J. W. Luminol/H2O2 chemiluminescence detector for the analysis of nitric oxide in exhaled breath. Anal. Chem. 71, 5131–5136 (1999).
Li, X., Zhang, Z., Tao, L. & Gao, M. Sensitive and selective chemiluminescence assay for hydrogen peroxide in exhaled breath condensate using nanoparticle-based catalysis. Spectrochim Acta Mol. Biomol. Spectrosc. 107, 311–316 (2013).
McCurdy, M. R., Bakhirkin, Y., Wysocki, G., Lewicki, R. & Tittel, F. K. Recent advances of laser-spectroscopy-based techniques for applications in breath analysis. J. Breath. Res. 1, 014001 (2007).
Henderson, B. et al. Laser spectroscopy for breath analysis: towards clinical implementation. Appl Phys. B 124, 161 (2018).
Obermeier, J. et al. Electrochemical sensor system for breath analysis of aldehydes, CO and NO. J. Breath. Res. 9, 016008 (2015).
Gaffney, E. M., Lim, K. & Minteer, S. D. Breath biosensing: using electrochemical enzymatic sensors for detection of biomarkers in human breath. Curr. Opin. Electrochem. 23, 26–30 (2020).
Haddadi, S., Koziel, J. A. & Engelken, T. J. Analytical approaches for detection of breath VOC biomarkers of cattle diseases -A review. Anal. Chim. Acta. 1206, 339565 (2022).
Li, W. et al. VOC biomarkers identification and predictive model construction for lung cancer based on exhaled breath analysis: research protocol for an exploratory study. BMJ Open. 9, e028448 (2019).
Tung, T. T. et al. Carbon nanomaterial sensors for cancer and disease diagnosis. Carbon Nanomaterials Bioimaging Bioanalysis Therapy. 167–202. https://doi.org/10.1002/9781119373476.CH8 (2019).
Xuan, W. et al. Engineering solutions to breath tests based on an e-nose system for silicosis screening and early detection in miners. J Breath. Res 16, 036001 (2022).
Thakur, U. N., Bhardwaj, R., Ajmera, P. K. & Hazra, A. ANN based approach for selective detection of breath acetone by using hybrid GO-FET sensor array. Eng. Res. Express. 4, 25008 (2022).
Ricciardolo, F. L. M., Sorbello, V. & Ciprandi, G. A pathophysiological approach for feno: A biomarker for asthma. Allergol. Immunopathol. (Madr). 43, 609–616 (2015).
Horváth, I. et al. Exhaled breath condensate: methodological recommendations and unresolved questions. Eur. Respir. J. 26, 523–548 (2005).
Rydosz, A. Diabetes without needles: Non-invasive diagnostics and health management. Diabetes Without Needles: Non-invasive Diagnostics Health Manage. 1–302 https://doi.org/10.1016/B978-0-323-99887-1.01001-3 (2022).
Fasola, S. et al. Repeatability of exhaled breath fingerprint collected by a modern sampling system in asthmatic and healthy children. J Breath. Res 13, 036007 (2019).
Martinez-Lozano Sinues, P., Kohler, M. & Zenobi, R. Human breath analysis May support the existence of individual metabolic phenotypes. PLoS One 8, e59909 (2013).
Incalzi, R. A. et al. Reproducibility and respiratory function correlates of exhaled breath fingerprint in chronic obstructive pulmonary disease. PLoS One. 7, e45396 (2012).
Krilaviciute, A. et al. Associations of diet and lifestyle factors with common volatile organic compounds in exhaled breath of average-risk individuals. J Breath. Res 13, 026006 (2019).
Wang, X. R., Lizier, J. T., Berna, A. Z., Bravo, F. G. & Trowell, S. C. Human breath-print identification by E-nose, using information-theoretic feature selection prior to classification. Sens. Actuators B Chem. 217, 165–174 (2015).
Filosa, M. et al. A meta-learning algorithm for respiratory flow prediction from FBG-based wearables in unrestrained conditions. Artif. Intell. Med. 130, 102328 (2022).
Oliveira, R., Tabacof, P. & Valle, E. Known Unknowns: Uncertainty Quality in Bayesian Neural Networks.
Dührkop, K. et al. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nature Biotechnology 2020 39:4 39, 462–471 (2020).
Picache, J. A., May, J. C. & McLean, J. A. Chemical class prediction of unknown biomolecules using ion Mobility-Mass spectrometry and machine learning: supervised inference of feature taxonomy from ensemble randomization. Anal. Chem. 92, 10759–10767 (2020).
Little, J. L., Williams, A. J., Pshenichnov, A. & Tkachenko, V. Identification of ‘known unknowns’ utilizing accurate mass data and chemspider. J. Am. Soc. Mass. Spectrom. 23, 179–185 (2012).
Taylor, A. et al. Development of a new breath collection method for analyzing volatile organic compounds from intubated mouse models. Biol Methods Protoc 9, bpae087 (2024).
Tian, J. et al. Exhaled volatile organic compounds as novel biomarkers for early detection of COPD, asthma, and prism: a cross-sectional study. Respir Res. 26, 1–14 (2025).
Dugheri, S. et al. Breath fingerprint of colorectal cancer patients by gas chromatography-mass spectrometry analysis preparatory to e-nose analyses. Pract. Lab. Med. 45, e00475 (2025).
Mpolokang, A. G., Setlhare, T. C., Bhattacharyya, S. & Chimowa, G. New volatile organic compounds from the exhaled breath of active tuberculosis patients. Scientific Reports 2025 15:1 15, 1–9 (2025).
Zhang, J. et al. Identification potential biomarkers for diagnosis, and progress of breast cancer by using high-pressure photon ionization time-of-flight mass spectrometry. Anal Chim. Acta 1320, 342883 (2024).
Lee, B. et al. Breath analysis system with convolutional neural network (CNN) for early detection of lung cancer. Sens. Actuators B Chem. 409, 135578 (2024).
Vinhas, M. et al. AI applied to volatile organic compound (VOC) profiles from exhaled breath air for early detection of lung cancer. Cancers 2024. 16, 2200 (2024).
Tang, T. et al. Glass based micro total analysis systems: materials, fabrication methods, and applications. Sens. Actuators B Chem. 339, 129859 (2021).
Arulvasan, W., Chou, H., Greenwood, J. et al. High-quality identification of volatile organic compounds (VOCs) originating from breath. Metabolomics 20, 102 (2024). https://doi.org/10.1007/s11306-024-02163-6
Chauhan, J. et al. Performance characterization of deep learning models for Breathing-based authentication on Resource-Constrained devices. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 2, 1–24 (2018).
Li, Y. et al. Electronic nose for the detection and discrimination of volatile organic compounds: application, challenges, and perspectives. TRAC Trends Anal. Chem. 180, 117958 (2024).
Ogunleye, A. & Wang, Q. G. Enhanced XGBoost-Based automatic diagnosis system for chronic kidney disease. IEEE Int. Conf. Control Autom. ICCA 2018-June, 805–810 (2018).
Paleczek, A., Grochala, D. & Rydosz, A. Artificial breath classification using XGBoost algorithm for diabetes detection. Sensors 21, 12 (2021).
Binson, V. A., Subramoniam, M., Sunny, Y. & Mathew, L. Prediction of pulmonary diseases with electronic nose using SVM and XGBoost. IEEE Sens. J. 21, 20886–20895 (2021).
Ke, G. et al. LightGBM: A highly efficient gradient boosting decision tree. Adv Neural Inf. Process. Syst 30, 10159–10183 (2017).
Boubin, M. & Shrestha, S. Microcontroller implementation of support vector machine for detecting blood glucose levels using breath volatile organic compounds. Sens. 2019. 19, 2283 (2019).
Tirzite, M., Bukovskis, M., Strazda, G., Jurka, N. & Taivans, I. Detection of lung cancer in exhaled breath with an electronic nose using support vector machine analysis. J. Breath. Res. 11, 036009 (2017).
Chen, Z., Zheng, Y., Chen, K., Li, H. & Jian, J. Concentration estimator of mixed VOC gases using sensor array with neural networks and decision tree learning. IEEE Sens. J. 17, 1884–1892 (2017).
Sciavicco, G., Manzella, F., Pagliarini, G. & Stan, I. E. The voice of COVID19: breath and cough recording classification with Temporal decision trees and random forests. SSRN Electron. J. https://doi.org/10.2139/SSRN.4102488 (2022).
Martinez-Lozano Sinues, P., Kohler, M. & Zenobi, R. Human breath analysis May support the existence of individual metabolic phenotypes. PLoS One. 8, e59909 (2013).
Fuchs, P., Loeseken, C., Schubert, J. K. & Miekisch, W. Breath gas aldehydes as biomarkers of lung cancer. Int. J. Cancer. 126, 2663–2670 (2010).
Heidari, M., Fouladi, K. & Fouladi-Ghaleh, K. Using Siamese networks with transfer learning for face recognition on Small-Samples datasets. https://doi.org/10.1109/MVIP49855.2020.9116915
Ratiu, I. A., Ligor, T., Bocos-Bintintan, V., Mayhew, C. A. & Buszewski, B. Volatile organic compounds in exhaled breath as fingerprints of lung cancer, asthma and COPD. J. Clin. Med. 2021. 10, Page 32 (10), 32 (2020).
Herbig, J. & Beauchamp, J. Towards standardization in the analysis of breath gas volatiles. J. Breath. Res. 8, 037101 (2014).
Hernández-Vicente, A. et al. Validity of the Polar H7 heart rate sensor for heart rate variability analysis during exercise in different age, body composition and fitness level groups. Sens. 2021. 21, 902 (2021).
Yuan, W. et al. A sensitive and selective mutation detection strategy based on non-canonical DNA structure preference of endonuclease IV. Sens. Actuators B Chem. 359, 131575 (2022).
Yue, F. et al. Highly sensitive polarization rotation measurement through a High-Order vector beam generated by a metasurface. Adv. Mater. Technol. 5, 1901008 (2020).
Tripathi, K. M. et al. Green carbon nanostructured quantum resistive sensors to detect volatile biomarkers. Sustainable Mater. Technol. 16, 1–11 (2018).
Liberati, A. et al. The PRISMA statement for reporting systematic reviews and Meta-Analyses of studies that evaluate health care interventions: explanation and elaboration. PLoS Med. 6, e1000100 (2009).
Rethlefsen, M. L. et al. PRISMA-S: an extension to the PRISMA statement for reporting literature searches in systematic reviews. Syst. Rev. 10, 39 (2021).
Shamseer, L. et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015: elaboration and explanation. BMJ 349 (2015).
Gilchrist, F. J. et al. The suitability of Tedlar bags for breath sampling in medical diagnostic research. Physiol. Meas. 28, 73 (2006).
Mochalski, P., King, J., Unterkofler, K. & Amann, A. Stability of selected volatile breath constituents in tedlar, Kynar and flexfilm sampling bags. Analyst 138, 1405–1418 (2013).
McGarvey, L. J. & Shorten, C. V. The effects of adsorption on the reusability of Tedlar® air sampling bags. AIHAJ - Am. Industrial Hygiene Association. 61, 375–380 (2000).
Beauchamp, J., Herbig, J., Gutmann, R. & Hansel, A. On the use of Tedlar® bags for breath-gas sampling and analysis. J. Breath. Res. 2, 046001 (2008).
Pedregosa, F. A. B. I. A. N. P. E. D. R. E. G. O. S. A. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Buitinck, L. et al. API design for machine learning software: experiences from the scikit-learn project. (2013). https://doi.org/10.48550/arxiv.1309.0238
Acknowledgements
The local ethical committee approved the study protocol (Jagiellonian University КВЕТ: 1072.6120.40.2023). The experiments were conducted following the Declaration of Helsinki, and principles of Good Clinical Practice and written informed consent was obtained from all participants before participation.
Funding
The work was partially supported by the IDUB AGH 4122 grant and the statutory activity at the Institute of Electronics AGH and Polish National Agency for Academic Exchange BPN/BEK/2021/1/00015.
Author information
Authors and Affiliations
Contributions
Anna Paleczek (APA), Justyna Grochala (JG), Dominik Grochala (DG), Agnieszka Pręgowska (APR), Magdalena Osiał (MO), Richard O. Oyeleke (RO), Małgorzata Pihut (MP), Jolanta E. Loster (JL) and Artur Rydosz (AR)Conceptualization: APA, JG, DG, APR, MO, RO, MP, JL, AR; Data curation: APA, JG, DG, AR; Formal analysis: APA, JG, DG, MP, JL, AR; Funding acquisition: AR; Investigation: APA, JG, DG, AR; Methodology: APA, JG, DG, APR, MO, RO, MP, JL, AR; Project administration: MP, AR; Resources: MP, JL, AR; Software: APA; Supervision: MP, JL, AR; Validation: AR; Visualization: APA, APR, MO; Writing – original draft: APA, JG, DG, APR, MO, RO, MP, JL, AR; Writing – review and editing: APA, JG, DG, APR, MO, RO, MP, JL, AR.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Paleczek, A., Grochala, J., Grochala, D. et al. The exhaled breath pattern as a potential method for biometrics identification. Sci Rep 15, 35656 (2025). https://doi.org/10.1038/s41598-025-19463-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19463-z