Abstract
Individuals differ in when and how quickly they experience puberty, and these differences in pubertal development are associated with lifelong health and mortality. We conducted sex-specific epigenome-wide association analyses of salivary DNA-methylation (DNAm) samples from ~ 3500 adolescents and identified 373 DNAm sites significantly associated with pubertal age, pace of pubertal development, and/or onset of early puberty by age 9. These DNAm signals converged with results from previous genomic and transcriptomic studies of puberty and with pan-mammalian epigenomic studies of age. Genomic annotations and trait enrichment results implicate child maltreatment and toxicant exposures as relevant for puberty. We developed a novel DNAm biomarker of pubertal age that was consistently associated with earlier age at menarche across three adolescent cohorts. Saliva DNAm is sensitive to signatures of pubertal development and suggests molecular links between reproductive maturation and both embryonic development and biological aging across species.
Similar content being viewed by others
Introduction
Puberty is a complex, coordinated set of neurobiological, endocrine, and morphological changes that produce reproductive maturity1. People differ in their age at pubertal onset, which typically occurs between ages 8 and 13 in girls and ages 9 and 14 in boys2. Individual differences in puberty are highly heritable and respond to multiple endogenous and environmental cues, including nutrition and early life adversity3,4. In recent birth cohorts, puberty has onset at progressively younger ages, occurring about three months earlier per decade. This historical change has intensified interest in better understanding the mechanisms responsible for regulating the timing of puberty5.
Pubertal development is intertwined with the aging process6,7. Children who experience puberty earlier tend to have poorer adult health, including higher risks of obesity, type 2 diabetes, cardiovascular disease, breast, endometrial, and prostate cancers, and earlier mortality8,9,10. Within and across species, earlier reproductive maturity is associated with shorter lifespan11,12,13,14, and both earlier puberty and earlier mortality, in humans, are linked to a common set of adverse early environments6,15. The connections between puberty and aging can be understood in terms of evolutionary theories7. Life history models posit that accelerated sexual maturity is adaptive within a threatening or impoverished environment because it increases the likelihood of reproduction before an early death16, while programmatic theories of aging posit that genetically-programmed processes necessary for early development become futile or detrimental in late life17. Integrating theories and methods from research on development and research on aging, therefore, has the potential to address critical questions in both fields.
From animal studies, DNA-methylation (DNAm) is known to be a critical biological mechanism for the onset and pace of mammalian puberty4,18,19,20. However, relatively few epigenetic studies have examined differences in human pubertal development21,22,23,24. Even fewer studies have focused on DNAm measured in saliva, which is more easily obtained from pediatric samples than other tissues25. In contrast, copious epigenetic research in adults has identified DNAm biomarkers of aging and aging-related disease in blood and other tissues, aka “epigenetic clocks”26,27. Epigenetic aging clocks, when calculated in saliva samples statistically corrected for cell composition, have been associated with individual differences in puberty, although results are mixed and differ across clocks28,29,30. This preliminary evidence suggests that saliva DNAm might be sensitive to some of the biological signals that connect pubertal development to later-life biological aging, but much variation in pubertal development remains uncaptured by existing epigenetic aging clocks.
Here, we conduct sex-specific epigenome wide association studies (EWAS) to identify DNAm markers of pubertal development in adolescents. We operationalize puberty in three different ways: (1) pubertal age, (2) pubertal pace, and (3) early puberty. Our EWAS analyses use saliva DNAm samples, accounting for cell composition, from two adolescent cohorts: (1) the Texas Twin Project (TTP, N = 1,185, spanning ages 8 to 18), which contributed data on pubertal age and pubertal pace, and (2) the Future of Families and Child Well-Being Study (FFCW, N = 1,956, measured at age 9), which contributed data on early puberty. We then use elastic net regression and data from the TTP to develop a novel DNAm biomarker of pubertal age (the phenotype measured in the largest number of participants) and test its relationships with pubertal development, age at menarche, and epigenetic clocks in a hold-out sample of the TTP, the FFCW cohort, as well as a third cohort: (3) the German Twin Family Panel (TwinLife, N = 395, spanning ages 14 to 29).
Results
Complementary research designs identify multiple dimensions of pubertal development
We distinguished three dimensions of pubertal development (Fig. 1). First, using the TTP cohort (Table 1, Methods), we measured participants’ pubertal age, which reflects the maturity of an adolescent’s physical development and is scaled in years. For example, a girl with a chronological age of 14 but a pubertal age of 15 shows physical changes in skin, height, body hair, breasts, and menstruation that are typical of 15-year-old girls; she is “older” in her physical development than her same-aged peers. Pubertal age was highly correlated, but not synonymous, with chronological age in girls (r = 0.83, 95% CI = 0.81 to 0.86) and in boys (r = 0.79, 95% CI = 0.76 to 0.82).
Schematic representations of pubertal age, pubertal pace, and early puberty. (A) Pubertal age is calculated by comparing an adolescent’s self-reported physical development (changes in secondary sex characteristics and, for girls, the onset of menarche) to sex-specific age norms. An adolescent’s pubertal age is the age at which their level of physical maturity is typical for their sex. Pubertal age is a continuous variable that varies both within- and between-persons, increases with chronological age, and can be measured cross-sectionally in age-heterogeneous cohorts. (B) Pubertal pace is a measure of the rapidity of change in an adolescent’s self-reported physical development that is calculated by comparing an adolescent with themselves across time. Here, we take the difference in pubertal age across two longitudinal timepoints divided by the lag (in years) between timepoints. Pubertal pace is a continuous variable that varies both within- and between-persons and has a non-linear relationship with chronological age, as the changes of puberty initially accelerate and then decelerate. (C) Early puberty is measured by assessing whether an adolescent has already experienced key milestones of physical development (menarche or voice break) by a young age (in this study, 9 years old). Unlike pubertal age and pubertal pace, early puberty is a categorical variable that differs between-persons, but does not change within-persons, and is measured in an age-homogenous cohort.
Second, pubertal pace was measured in a two-wave longitudinal subset of TTP participants who were ages 10–13 at wave I (N = 62 girls and 61 boys; Table 1, Methods). Pubertal pace reflects how rapidly an adolescent is maturing in their physical development over time and was not significantly correlated with chronological age in either girls (r = 0.06 [−0.19, 0.31] at wave I; r = −0.10 [−0.34, 0.16] at wave II) or boys (r = 0.20 [−0.05, 0.43] at wave I; r = 0.14 [−0.11, 0.38] at wave II). Youth with faster pubertal pace had lower pubertal ages at wave I (r = −0.52 [−0.68, −0.31] in boys; r = −0.28 [−0.49, − 0.03] in girls) and higher pubertal ages at wave II (r = 0.45 [0.22, 0.63] in boys; r = 0.49 [0.27, 0.66] in girls).
Third, early puberty, which we measured in 9-year-olds from the FFCW (Table 1, Methods), is a binary variable that reflects whether an adolescent has already reached key milestones in reproductive development (menarche or voice break) by a young age, here by age 9.
Sex-specific EWASs identify DNAm markers unique to each dimension of pubertal development
Pubertal age
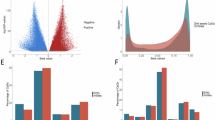
We controlled for immune and epithelial cell composition in all EWASs (see Methods). In girls, 106 CpG sites were significantly associated with pubertal age at epigenome-wide significance (p < 3.909 e-07) and an additional 320 (426 total) at suggestive significance level (p < 2.703 e-05; see Methods for calculation of significance levels) (Fig. 2; Supplementary Table S1). In boys, 155 CpG sites were epigenome-wide significant and an additional 739 (894 total) were suggestive (Supplementary Table S2). The correlation between Z statistics across all CpG sites between boys and girls was 0.84 (SE = 0.0006) indicating moderately high overlap of DNAm patterns characteristic of pubertal age across sexes.
Manhattan plots of results from EWASs of pubertal development. Red line denotes the epigenome-wide significance level (3.9e-07), and blue line denotes the suggestive significance level (2.7e-05). See Supplementary Methods for additional information on calculation of significance levels.
Pubertal pace
In girls, 148 CpG sites were significantly associated with pubertal pace at epigenome-wide significance and an additional 743 (891 total) at suggestive significance level (Fig. 2; Supplementary Table S3). In boys, 264 CpG sites were epigenome-wide significant for pubertal pace, and an additional 1,181 (1,455 total) were suggestive (Supplementary Table S4). Within each sex, the CpG sites associated with pubertal pace were largely distinct from those associated with pubertal age; however, one CpG site for boys pubertal age (cg23564243) was suggestively associated with boys’ pubertal pace. The correlation between Z statistics in boys and girls was − 0.11 (SE = 0.00, 95% CI = −0.11, −0.11), indicating that some CpG sites associated with pubertal pace have opposite directions of effect in boys vs. girls.
Early puberty
In girls, 58 CpG sites were significantly associated with early puberty at epigenome-wide significance and an additional 173 (231 total) at suggestive significance level (Fig. 2, Supplementary Table S5). In boys, 19 CpG sites were epigenome-wide significant and an additional 73 (92 total) were suggestive (Supplementary Table S6). The correlation between Z statistics between boys and girls was 0.21 (SE = 0.0016). No epigenome-wide significant CpG sites for early puberty showed epigenome-wide significance for pubertal age or pace.
Results from EWASs of puberty converge with insights from genomic, transcriptomic, and pan-mammalian analyses of puberty and aging
To identify candidate mechanisms of interest for the regulation of pubertal development and its contribution to aging, we cross-referenced our results with previous genomic, transcriptomic, and epigenomic studies of puberty and aging. Specifically, we mapped each suggestive hit from the six EWASs to the nearest gene (3,659 genes total). We then compared this gene set to (1) genes found to be significantly associated with earlier age at menarche or voice break in previous large-scale GWASs (887 genes)31, (2) genes differentially expressed with advancing female pubertal development in a longitudinal study of leucocyte gene expression (108 genes)32, and (3) genes that were differentially methylated with age in a pan-mammalian study of 185 species (1188 genes)33. Out of the 1998 genes identified in these previous studies, 402 (20%) mapped to CpG sites associated with at least one puberty phenotype in our EWAS analyses, which is higher than would be expected by chance based on Fisher’s exact test assuming 20,000 genes (OR = 1.14, p = 0.015) (Supplementary Table S7).
Table 2 summarizes the 26 genes that were consistently implicated across our EWASs and previous research (at least 1 significant association in prior studies and at least 3 significant associations across the 6 EWASs and 3 prior studies). This set includes several genes (ZIC1, ZIC4, FOXG1, GATA4) coding for transcription factors critical to embryonic development of neural, cardiac, and testicular tissues. It also includes multiple genes with established relevance to aging and aging-related disease, including DCAKD, PRKCB, and BDNF (associated with Parkinson’s disease, Alzheimer’s disease and neurocognitive decline34,35,36 and FOXO3 (robustly associated with longevity37. Some aging-related genes that are differentially methylated with pubertal development are known to have opposing effects in early development versus adulthood: NMT1 codes for an enzyme that is essential in early development but is inhibited by senolytic agents38, and EIF4G3 codes for a translation initiation complex that is necessary for embryonic development but prolongs longevity when suppressed in adulthood39.
Methylation in known aging-related genes was significantly associated with puberty even when analyzing pubertal phenotypes that control for age-related variation (i.e., for pubertal pace and early puberty). Considering aging-related genes through the lens of puberty also suggests new hypotheses regarding their mechanism. For instance, methylation of LHFPL4 was strongly implicated in the pan-mammalian EWAS of age, but “the known function of LHFPL4 in synaptic clustering of γ-aminobutyric acid (GABA) receptors does not [on its own] provide a clear connection to aging across tissues.”33 In contrast, GABA receptors do have a clear connection to the onset of puberty, as tonic inhibition by GABA neurotransmission suppresses secretion of gonadotropin-releasing hormone, and the removal of central inhibition triggers the onset of puberty40.
Several consistently implicated genes have connections to psychosocial function, including PPM1F, the expression of which is regulated by psychosocial stress in animal models41; SDK1, which is differentially methylated in children who have been maltreated42; and CSMD2, which is associated with schizophrenia, a severe psychiatric disorder that commonly onsets in adolescence43.
We tested the 403 genes with converging evidence across our EWASs and previous research for over-representation in Gene Ontology biological processes and identified significant enrichment in 81 biological processes after FDR correction (Supplementary Table S8). We used affinity propagation to reduce redundancy, resulting in 17 biological processes identified as the most representative (Fig. 3). The most enriched process was endocrine system development. Many processes pertain specifically to embryonic development (mesenchyme development, mesoderm development, central nervous system differentiation, cardiac chamber development, sensory organ morphogenesis).
Overrepresented biological processes in set of genes with converging evidence from current EWASs and previous research for involvement in puberty.
Trait enrichment analyses reveal epigenetic connections with aging, diseases, child abuse, and toxicant exposures
We conducted trait enrichment analyses that compared CpG sites associated with pubertal phenotypes to previously reported results from the EWAS Atlas (Methods, Supplementary Table S9). Across our 6 EWASs, there was significant enrichment for 90 traits; of these, 28 replicated across at least 2 EWASs (selected traits highlighted in Fig. 4). One of the strongest enrichment results was for tooth mobility, which increases during puberty, pregnancy, and menopause. Our EWAS results were also strongly enriched for CpG sites previously identified in studies of vitamin B12 and folic acid (vitamin B9) supplementation, and in studies of birth defects (orofacial cleft) that supplementation with these B vitamins can prevent. Dietary B9 and B12 are essential for DNA synthesis, repair, and methylation, and are particularly critical during periods of rapid cell growth44.
Key trait enrichment results. Trait enrichments that replicate across two or more of the six EWASs are in bold.
Results from our EWASs were enriched for SETD1B-syndrome and Kabuki syndrome, rare autosomal dominant disorders characterized by developmental delay (no participants in our study are known to have either disorder). Pathogenic variants causing Kabuki syndrome have been identified in KMT2D45, which, like SETD1B, codes for a methyltransferase enzyme critical for histone modifications characteristic of transcriptionally active genes. Inducing expression of KMT2D and SETD1B in vitro blocks cellular senescence46. Other genetic disorders that were significant in trait enrichment analyses of pubertal age were Werner syndrome, which involves the rapid onset of premature aging at puberty, and Klinefelter syndrome, a sex chromosome aneuploidy (XXY) that involves lower testosterone levels, reduced sperm production, and slower testicular growth.
One of the most consistent trait enrichment findings was for child maltreatment, and specifically sexual abuse. Finally, trait enrichment results implicated several environmental exposures, including prenatal mercury exposure, air pollution exposure, smoking and maternal smoking, maternal lead exposure, and exposure to polychlorinated biphenyls (PCBs), a class of synthetic organic chemicals that accumulates in fatty tissue, biomagnifies up the food chain, and disrupts endocrine function.
Elastic net regression generates a DNAm-based biomarker of pubertal age in boys and girls that is correlated with but distinct from epigenetic aging clocks
For our elastic net analyses, we focused on pubertal age, the phenotype for which we had the most available data. We divided the TTP dataset into a training set (75% of families) and test set (25% of families) and used elastic net regression to generate DNAm-based predictors of pubertal age in boys and girls. Probes that had low reliability or were unavailable across both arrays (Illumina 450 K and EPIC) were not included. Given the strong correlation between pubertal age and chronological age in the TTP sample, and the well-established chronological age signal in the human epigenome, we also excluded probes significantly associated at the suggestive level with chronological age in TTP sample (Supplementary Table S10). The final lambda value selected by the analysis was 0.27 for girls and 0.28 for boys (Supplementary Methods). The resulting algorithms included 195 CpG sites for girls (Supplementary Table S11) and 166 sites in boys (Supplementary Table S12). Three CpG sites overlapped across sexes (cg24250902, cg27180636, and cg04165099).
To calculate DNAmPubertalAge, we then applied the resulting algorithms, separately in each sex, to the TTP test set, to the FFCW cohort, and to the TwinLife cohort. Within these cohorts, we also calculated DNAmPubertalAgeAccel, which residualized DNAmPubertalAge for chronological age, and estimated associations between these DNAm biomarkers and measures of pubertal development and body size (height and BMI). In the FFCW study, pubertal development was measured at age 9, and age at menarche and body size was measured at age 15. In the TwinLife study, 14- to 16-year-old participants reported on pubertal development and body size, and all post-menarcheal females ages 15 to 29 reported age at menarche.
Analysis of 8 technical replicates in girls and 11 technical replicates in boys in TTP suggested good measurement reliability of DNAmPubertalAge (intraclass correlation coefficients of 0.88 [95% CI 0.55 to 0.97] in girls and 0.87 [95% CI 0.59 to 0.96] in boys).
We report results with DNAmPubertalAge and DNAmPubertalAgeAccel. Males and females in the TTP, females in the FFCW, and females in TwinLife who had greater DNAmPubertalAge reported more advanced pubertal development (Table 3). After controlling for chronological age, however, DNAmPubertalAgeAccel was associated with more advanced pubertal development only among females from TwinLife. DNAmPubertalAgeAccel was also consistently and significantly associated with greater BMI and with earlier age at menarche in females from all three cohorts (rs range from − 0.23 to −0.13; Fig. 5). Associations for males were less consistent, with DNAmPubertalAgeAccel significantly associated only with height among 15-year-old boys in the FFCW.
Associations between DNAmPubertalAgeAccel and age at menarche across three adolescent to young adult cohorts
Finally, using the TTP test set, 15-year-olds from FFCW, and 14- to 16-year-olds from TwinLife, we estimated the relationships between DNAmPubertalAgeAccel and five previously developed epigenetic aging clocks, including Horvath Multi-Tissue, Horvath Skin-and-Blood, GrimAge, PhenoAge, and DunedinPACE. All epigenetic clocks were residualized for cell composition, and all were residualized for age except for DunedinPACE, which was trained on longitudinal data reflecting change over time among same-aged participants. Very few individual CpG sites included in the calculation of DNAmPubertalAge overlapped with the CpG sites included in the epigenetic aging clocks (1–7 CpG sites in girls and 0–5 sites in boys; Supplementary Table S13). Nonetheless, in girls, there was moderate convergence between DNAmPubertalAgeAccel and all epigenetic clocks tested (rs ranging from 0.15 to 0.60 across clocks and cohorts, median r = 0.58; Supplementary Table S14). In boys, DNAmPubertalAgeAccel was consistently significantly correlated with accelerations on the Horvath and PhenoAge clocks (rs ranging from 0.18 to 0.54, median r = 0.46) but not GrimAge or DunedinPACE clocks. The correlations between DNAmPubertalAgeAccel with the epigenetic aging clocks were comparable to the correlations among the different epigenetic aging clocks, which are presented in Supplementary Table S15-S17.
Discussion
We used data from three adolescent cohorts with complementary research designs to distinguish three facets of pubertal development (pubertal age, pace of pubertal development, and early puberty) and conducted sex-specific EWASs that identified 373 DNAm markers significantly associated with puberty phenotypes. Individual CpG sites associated with puberty were largely distinct across puberty phenotypes and, for pubertal age and early puberty, across sexes. However, downstream analyses that mapped associated DNAm markers to genes found that multiple genes replicated across sexes, puberty phenotypes, and cohorts. Moreover, our EWAS results converged with results from previous genomic and transcriptomic studies of puberty (conducted using blood samples), as well as with a previous pan-mammalian DNAm study of aging (conducted using multiple tissues). These results support the conclusion that saliva DNAm reflects signals of biological processes involved in the regulation of human puberty and links reproductive maturation to biological aging across species.
Our analyses of salivary DNAm from adolescents reveal molecular connections between puberty and both the early and late stages of the human lifespan. Genes differentially methylated with earlier, more advanced, or faster puberty are enriched for biological processes linked to embryonic development. For instance, trait enrichment analyses highlight overlap between DNAm markers identified in our EWASs and those found in previous studies of neural tube defects, including conditions like orofacial cleft. At the same time, our EWASs of puberty uncovered DNAm differences in genes that are well-established in aging, aging-related disease, and longevity, including FOXO3, BDNF, NMT1, and LHFPL4. Notably, LHFPL4, which regulates inhibitory GABA synapses, was robustly associated with aging in a previous pan-mammalian EWAS. The current evidence linking LHFPL4 to puberty suggests a new hypothesis regarding the mechanism of its relationship with aging, as removal of GABA inhibition is a critical trigger for the onset of mammalian puberty.
In addition, our novel DNAm biomarker of pubertal age, DNAmPubertalAge was moderately correlated with epigenetic aging clocks. This result is more striking because our elastic net models specifically filtered out probes significantly associated with chronological age and the resulting algorithm included very few CpG sites included in the calculation of any epigenetic clock. These results align with the epigenetic clock theory of aging, which suggests that DNAm serves as a molecular link connecting developmental processes from conception through reproductive maturity, to post-maturity maintenance and ultimately, death26. Furthermore, they are broadly consistent with the pseudo-programmatic theory of aging, which posits that aging is an evolved byproduct of the same genetically-determined developmental program of cellular and molecular events that is optimized to produce a reproductively mature adult7,17,33,47,48. Yet, we acknowledge that there is substantial disagreement on the definition of aging, which affects its hypothesized relationship to reproductive development49.
Our results also suggested pathways by which human pubertal development is influenced by psychosocial adversity, as predicted by life-history evolutionary theories6. Consistent with epidemiological research50, DNAm markers associated with pubertal age in both sexes, and with early puberty in girls, showed significant trait enrichment for child maltreatment and sexual abuse. Additionally, DNAm markers in or near SDK1, which has been previously shown to be differentially methylated in maltreated children, were significantly associated with pubertal pace in both sexes and early puberty in boys.
The DNAmPubertalAge biomarker showed more promising validation results in females than in males. It was modestly but consistently associated with earlier age at menarche among post-menarcheal females across three cohorts that differed in age span and demographic composition. In contrast, our efforts to develop a DNAm-based biomarker for male pubertal development were less successful, likely due to limited statistical power and the inherent challenges of separating puberty-specific signals from those related to chronological age in adolescents. Although this is the largest DNAm study of puberty to-date, our discovery sample sizes remain relatively small compared to DNAm research in adults.
Future studies aiming to develop DNAm biomarkers for puberty would greatly benefit from larger-scale data collection in adolescents, with more detailed, sex-specific, multi-modal, and longitudinal measurement of pubertal development. These studies would also benefit from probing how specific our results are to saliva DNAm compared to other peripheral tissue types. Saliva is easily accessible in pediatric cohorts and, similar to blood, contains a substantial proportion of immune cells and thus holds great promise for child-focused DNAm research51. Other directions for future research are epigenetic studies of youth taking gonadotropin-releasing hormone analogues to suppress puberty52 and youth with constitutional delay of puberty53, as chronological age and pubertal development are uniquely decoupled in these adolescents. Finally, it is unclear how current and future DNAm biomarkers of puberty will associate with psychological and physical health outcomes beyond the adolescent period.
In conclusion, our analyses identify molecular connections between pubertal development and both ends of the human lifespan, from embryonic development to late-life aging, and implicate early life adversities, including child maltreatment and toxicant exposures, as risk factors for accelerated pubertal development, a growing public health concern. We anticipate that the development of sex-specific DNAm biomarkers for reproductive development will continue to improve as more data becomes available on peripubertal children and adolescents, and that research that integrates tools and perspectives from research on development and aging will advance both fields.
Methods
Participants
Participants were drawn from three cohorts. The Texas Twin Project (TTP) is a study of school-aged twins and other multiples recruited from public school rosters and public outreach in the Austin- and Houston- metropolitan areas54. Participants ranged in age from 8 to 18. Ethical approval for TTP was granted by the University of Texas at Austin Institutional Review Board. The Future of Families and Child Wellbeing Study (FFCW) is a representative 20-city longitudinal birth cohort that recruited children born between 1998 and 2000 in the United States55. Families were randomly selected from 72 hospitals at the birth of the target child, and unmarried parents were oversampled. Epigenetic data was collected at ages 9 and 15. Ethical approval for FFCW was granted by the Princeton University Institutional Review Board. The German Twin Family Panel (TwinLife) is a representative, longitudinal study of twins that was conducted annually between 2014 and 2024. Ethical approval for TwinLife was granted by the German Psychological Association (protocol numbers: RR 11.2009 and RR 09.2013). We focus here on a subsample of 14- to 16-year-olds, who reported on the physical changes of puberty, and a second subsample of 15- to 29-year-old females who reported on age at menarche. All research was performed in accordance with named guidelines and regulations and informed consent was obtained from all participants and/or their legal guardians. Table 1 summarizes the demographic characteristics of all included participants. See Supplementary Methods for additional details on participant sampling, recruitment, consent, and exclusion criteria.
Measures
Pubertal development
Participants in the TTP completed the Pubertal Development Scale (PDS)56, which asked them to rate physical changes in skin, height, body hair, facial hair (boys), voice (boys), and breasts (girls) on a 1 (“has not yet begun to change”) to 4 (“has finished changing”) scale. Girls also reported on the onset of menstruation (1 = “no”, 4 = “yes”) and the age (in years) at which it occurred. For both sexes, ratings were averaged across non-missing items to create an overall PDS score (Petersen et al., 1988). Scores for participants missing 20% or more of the individual items were omitted.
Using longitudinal data on self-reported puberty in N = 3,516 participants from the full TTP sample, including participants who contributed no epigenetic data, we estimated the linear and quadratic effects of age on average PDS score [Girls: age-centered \(\:\beta\:\) = 0.27, p < 0.001, age-squared \(\:\beta\:\) = −0.016, p < 0.001; Boys: age-centered \(\:\beta\:\) = 0.22, p < 0.001, age-squared \(\:\beta\:\) = 0.0068, p < 0.001]. These coefficients were used to rescale self-reported pubertal development into pubertal age, i.e., the age in years at which their level of pubertal development is the norm.
Pubertal pace was then calculated in a longitudinal subsample of TTP participants as the difference in participants’ pubertal age between waves I and II, divided by the lag (in years) between waves.
In FFCWS at year 9, the PDS was completed by the primary caregiver rather than the child. The mean of the items for cases with complete data was calculated and then rescaled into pubertal age using the coefficients estimated in the TTP cohort. At age 15, female participants reported their age at menarche. For girls in the FFCWS cohort, early puberty was defined as having experienced menarche at the age 9 assessment or reporting age of menarche less than 9 at the year 15 assessment. For boys, early puberty was defined as ratings greater than or equal to “2” (beginning to change) on the voice change and facial hair items of the PDS at age 9.
In TwinLife, pubertal development was measured using a one-item adaptation of the German translation of the PDS57. Girls were asked, “In puberty you get hair under your armpits and in the pubic area. For girls, the breasts begin to grow and they get their period (menstrual bleeding). Have you noticed whether any of these changes have already started for you?”. Boys were asked, “In puberty you get hair under your armpits and in the pubic area. For boys, facial hair starts to grow and the voice changes. Have you noticed whether any of these changes have already started for you?”. The item was rated on a 4-point scale (1= “Not started yet” to 4 = “I already have the body of an adult.”). Age at menarche (in years) was assessed by self-report.
Body size
In TTP, BMI was calculated using height and weight, transformed to sex- and age-normed z scores according to the method published by the US Centers for Disease Control and Prevention. Prior to the onset of the COVID-19 pandemic in March 2020, height and weight were measured in-laboratory using a stadiometer and research-grade scale. After data collection resumed in June 2021, height and weight were collected via self-report during virtual assessments.
In FFCW, at age 9, height and weight were collected during the in-home assessment by trained interviewers with uniform scales and measuring tapes. At age 15, 80% of the sample had their height and weight collected by an interviewer as at age 9, and the remaining 20% self-reported.
In TwinLife, height and weight were measured from self-report.
DNA-methylation
See Supplementary Methods for more information on acquisition and quality control procedures. In TTP and TwinLife, the Infinium MethylationEPIC BeadChip kit version 1(Illumina, Inc., San Diego, CA) was used to assess methylation levels in saliva at 850,000 methylation sites. Probes on sex chromosomes were included. Surrogate variable analysis was used to correct methylation values for batch effects using the “combat” function in the SVA package58. In TTP, sex was included as a covariate for batch correction. Elastic-net regression analyses were conducted using combat-corrected data. In the FFCW, due to the timing of assay completion, 40% of the saliva samples were completed using the Illumina 450K chip (n = 1811) and the remaining 60% used the Illumina EPIC chip (nsamples =2558). All analyses of FFCW data were run separately for 450K and EPIC and then meta-analyzed.
Analyses
Epigenome-wide association studies
We used data from the TTP to perform EWASs of pubertal age and pubertal pace. In analyses of pubertal age, analyses were restricted to the first timepoint for individuals who contributed more than one epigenetic sample. In analyses of pubertal pace, analyses were limited to individuals who contributed epigenetic samples at more than one timepoint and who were between the ages of 10 and 13 at the first time point (wave I). Pubertal pace was analyzed in relation to DNA-methylation at the second time point (wave II).
For each probe, we fit a linear model using the lmFit function in the ‘limma’ package with methylation M-value as the outcome and pubertal development as the exposure of interest. The lmFit algorithm uses an empirical Bayes approach that computes a moderated t-statistic for each probe, for which the standard error is smoothed across all probes in the array for a more efficient standard error estimate59. Immune and epithelial cell composition were included as covariates in each lmFit model. To address potential confounding bias by shared DNAm between Twin-pairs, generalized estimating equations (GEE) were used from the ‘geepack’ R package (v1.3.10) (see Supplementary Methods for more information)60. We then used the Bacon package in Bioconductor to correct for inflation in models with lambda values > 1.0. This method constructs an empirical null distribution for the data using a Gibbs Sampling algorithm61.
We used FFCW data to perform EWAS of early puberty. For each probe, we fit a Linear model using the lmFit function in the Limma package with methylation M-value as the outcome and early puberty as the exposure of interest. EWAS were performed separately in the 450k and EPIC samples; results were then meta-analyzed using the ‘DExMA’ R Package (v1.10.7)62. Meta-analysis was performed using Fisher’s combined probability test. Standard errors were calculated using inverse variance weighting.
Cell composition
For all cohorts, cell composition of immune and epithelial cells were estimated using the R package “EpiDISH”63,64. In all EWAS, we statistically controlled for immune and epithelial cell composition.
Significance thresholds
Array-based methylation data is known to contain a significant portion of invariant probes, violating the assumption of independence in Bonferroni correction. To find the number of independent tests in each dataset, we employed two methods: (1) a more conservative method for calculating correlated regions of probes, CoMeBack65, and (2) a method for finding the number of independent linear combinations of probes, Matrix Spectral Decomposition (MSD)66. Because the CoMeBack and MSD thresholds were similar across datasets, we used thresholds calculated from the FFCW EPIC sample (the most stringent thresholds), resulting in an epigenome-wide threshold of 3.9e-07 (calculated from CoMeBack) and a suggestive threshold of 2.7e-05 (calculated from MSD).
Enrichment and annotation analyses
EWAS hits were annotated using Illumina provided support files for both the EPIC and 450k arrays. Enrichment analyses were performed using the EWAS Open Platform toolkit67. Genome assembly GRCh38 (hg38) was used to identify the nearest gene, chromosome, position, and probe strand for suggestive hits from puberty EWAS. Trait enrichment analysis runs weighted Fisher’s exact test to calculate the co-occurrence probability between inputted CpGs and trait-related probes on EWAS Atlas.
We also tested genes with convergent evidence for involvement in puberty and/or aging against a non-redundant representative set of Gene Ontology biological processes using WebGestalt68 and identified overrepresented gene sets. We ran WebGestalt from the web interface at http://www.webgestalt.org/ and used the default parameters (minimum number of IDs in the category: 5; maximum number of IDs in the category: 2000; FDR Method: BH; and a reference gene set of all mapped NCBI Entrez Gene IDs).
Elastic net regressions of DNA-methylation and pubertal age
We restricted elastic net regression analyses to probes that (1) passed quality control filters described in the Supplementary Methods and are present on both the 450k and EPIC arrays, (2) had an intraclass correlation coefficient (ICC) > 0.4 in 19 TTP technical replicates, (3) were nominally significant probes from the EWAS of pubertal age in each sex in TTP. We further filtered out probes that were suggestively associated with chronological age in a sex-pooled EWAS (Supplementary Methods) in the TTP sample. This resulted in 13,166 input probes for girls and 12,284 input probes for boys.
In a 75% discovery set of the TTP data, we used elastic-net-regression to develop a DNAm algorithm of pubertal age, separately for each sex. The TTP discovery set was a random sample of families stratified by zygosity and sex, resulting in six family types: MZ female, MZ male, DZ female-female, DZ male-male, DZ female-male, and triplets. Repeated measures were included in the discovery test set, resulting in a slightly different sample than the EWAS analytic sample (N = 581 samples from 450 girls and N = 586 samples from 448 boys; Supplementary Methods). Following the method established by Horvath et al., we fixed the alpha parameter of each elastic net at 0.5, while allowing the lambda value to vary69. The lambda value for each elastic net regression was chosen by applying 10-fold cross-validation within the training sample using the cv.glmnet function from the R package glmnet70. Participants from the same family were assigned to the same fold.
We then used the resulting algorithm to calculate DNAmPubertalAge scores across cohorts, and residualized this score for chronological age to calculate DNAmPubertalAgeAccel. Given that the input probes for the elastic net analyses (n = 13,166 for girls and 12,284 for boys) were filtered based on results from the TTP pubertal age EWAS analyses, the FFCW and TwinLife cohorts may be considered independent validation samples for the elastic net analyses. We also calculated, in all three cohorts, five epigenetic aging clocks (GrimAge, PhenoAge, DunedinPACE, Horvath Multi-Tissue, and Horvath Skin-and-Blood; see Supplementary Methods). We residualized all DNAm measures for cell composition and standardized them prior to analyses. To test associations with phenotypes in FFCW, we performed fixed-effects regression models using the lme4 package in R.71 In TTP and TwinLife, we performed linear mixed models that included a random intercept to correct for non-independence of twins within families, using the lmerTest package in R72. Regression models did not include covariates. A p-value of less than 0.05 indicated a statistically significant association. Marginal R2 for linear mixed models were calculated using the MuMIn package in R73.
Data availability
To ensure the privacy of participants and minimize deductive disclosure risks, data from the Texas Twin Project Access are available to academic researchers via a restricted data use contract with the University of Texas at Austin. Researchers may apply for restricted-use data at https://redcap.prc.utexas.edu/redcap/surveys/?s=NJJFAXT4JN7A9AR7. Data from the Future of Families and Child Well-Being study are free to download and can be accessed through two data repositories: ICPSR’s Data Sharing for Demographic Research (DSDR) archive (https://www.icpsr.umich.edu/web/pages/DSDR/index.html) and Princeton University’s Office of Population Research (OPR) data archive (https://oprdata.princeton.edu/archive/restricted/Default.aspx). For more detailed instructions, please visit the study’s FAQ page (https://ffcws.princeton.edu/faq). To protect the confidentiality of survey respondents, some FFCWS data is available only through a Restricted Use Contract (https://ffcws.princeton.edu/restricted). Data from the Twin Life study can be accessed with a data use contract and are released only for academic research and teaching after the data depositor’s written authorization. Additional information and applications for phenotypic data access can be found at the GESIS Community Data repository (https://search.gesis.org/research_data/ZA6701). In order to protect the confidentiality of the respondents, molecular genetic and epigenetic data, also in combination with phenotypic data, are only available via a restricted data use agreement with the TwinLife Data Center at Bielefeld University.
References
Dorn, L. D. & Biro, F. M. Puberty and its measurement: A decade in review. J. Res. Adolesc. 21, 180–195 (2011).
Parent, A-S. et al. The timing of normal puberty and the age limits of sexual precocity: variations around the world, secular trends, and changes after migration. Endocr. Rev. 24, 668–693 (2003).
Banerjee, I. & Clayton, P. The genetic basis for the timing of human puberty. J. Neuroendocrinol. 19, 831–838 (2007).
Aylwin, C. F., Toro, C. A., Shirtcliff, E. & Lomniczi, A. Emerging genetic and epigenetic mechanisms underlying pubertal maturation in adolescence. J. Res. Adolesc. 29, 54–79 (2019).
Eckert-Lind, C. et al. Worldwide secular trends in age at pubertal onset assessed by breast development among girls: A systematic review and Meta-analysis. JAMA Pediatr. 174, e195881 (2020).
Belsky, J. Early-life adversity accelerates child and adolescent development. Curr. Dir. Psychol. Sci. 28, 241–246 (2019).
Raffington, L. Utilizing epigenetics to study the shared nature of development and biological aging across the lifespan. Npj Sci. Learn. 9, 1–8 (2024).
Elks, C. E. et al. Age at menarche and type 2 diabetes risk: the EPIC-InterAct study. Diabetes Care. 36, 3526–3534 (2013).
Lakshman, R. et al. Early age at menarche associated with cardiovascular disease and mortality. J. Clin. Endocrinol. Metab. 94, 4953–4960 (2009).
Day, F. R. et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat. Genet. 49, 834–841 (2017).
Xiang, Y. et al. Early menarche and childbirth accelerate aging-related outcomes and age-related diseases: Evidence for antagonistic pleiotropy in humans. eLife 13, RP102447 (2024).
Li, C. Z. et al. Epigenetic predictors of species maximum life span and other life-history traits in mammals. Sci. Adv. 10, eadm7273 (2024).
Yuan, R. et al. Genetic coregulation of age of female sexual maturation and lifespan through Circulating IGF1 among inbred mouse strains. Proc. Natl. Acad. Sci. 109, 8224–8229 (2012).
Harvey, P. H. & Zammuto, R. M. Patterns of mortality and age at first reproduction in natural populations of mammals. Nature 315, 319–320 (1985).
Hayward, M. D. & Gorman, B. K. The long arm of childhood: the influence of early-life social conditions on men’s mortality. Demography 41, 87–107 (2004).
Belsky, J. The development of human reproductive strategies: progress and prospects. Curr. Dir. Psychol. Sci. 21, 310–316 (2012).
Gems, D., Virk, R. S. & de Magalhães, J. P. Epigenetic clocks and programmatic aging. Ageing Res. Rev. 101, 102546 (2024).
Argente, J. et al. Molecular basis of normal and pathological puberty: from basic mechanisms to clinical implications. Lancet Diabetes Endocrinol. 11, 203–216 (2023).
Lomniczi, A. et al. Epigenetic control of female puberty. Nat. Neurosci. 16, 281–289 (2013).
Rzeczkowska, P. A., Hou, H., Wilson, M. D. & Palmert, M. R. Epigenetics: A new player in the regulation of mammalian puberty. Neuroendocrinology 99, 139–155 (2014).
Bessa, D. S. et al. Methylome profiling of healthy and central precocious puberty girls. Clin. Epigenetics. 10, 146 (2018).
Almstrup, K. et al. Pubertal development in healthy children is mirrored by DNA methylation patterns in peripheral blood. Sci Rep [Internet]. [cited 2018 Sep 20];6. (2016). Available from: http://www.nature.com/articles/srep28657
Moore, S. R. et al. Distinctions between sex and time in patterns of DNA methylation across puberty. BMC Genom. 21, 389 (2020).
Thompson, E. E. et al. Global DNA methylation changes spanning puberty are near predicted estrogen-responsive genes and enriched for genes involved in endocrine and immune processes. Clin Epigenetics [Internet]. [cited 2018 Sep 20];10. Available from: https://clinicalepigeneticsjournal.biomedcentral.com/articles/ (2018). https://doi.org/10.1186/s13148-018-0491-2
Nishitani, S., Parets, S. E., Haas, B. W. & Smith, A. K. DNA methylation analysis from saliva samples for epidemiological studies. Epigenetics 13, 352–362 (2018).
Horvath, S. & Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 19, 371–384 (2018).
Crimmins, E. M., Klopack, E. T. & Kim, J. K. Generations of epigenetic clocks and their links to socioeconomic status in the health and retirement study. Epigenomics 16, 1031–1042 (2024).
Raffington, L. et al. Socioeconomic disadvantage and the Pace of biological aging in children. Pediatrics 147, e2020024406 (2021).
Hamlat, E. J. et al. Early life adversity predicts an accelerated cellular aging phenotype through early timing of puberty. Psychol. Med. 53, 7720–7728 (2023).
Goering, M. et al. Examining Health Behaviors as Mechanisms Linking Earlier Pubertal Timing with Accelerated Epigenetic Aging in Late Adolescence. J Youth Adolesc [Internet]. [cited 2024 Dec 17]; (2024). Available from: https://doi.org/10.1007/s10964-024-02096-2
Kentistou, K. A. et al. Understanding the genetic complexity of puberty timing across the allele frequency spectrum. Nat. Genet. 56, 1397–1411 (2024).
Resztak, J. A. et al. Analysis of transcriptional changes in the immune system associated with pubertal development in a longitudinal cohort of children with asthma. Nat. Commun. 14, 230 (2023).
Lu, A. T. et al. Universal DNA methylation age across mammalian tissues. Nat. Aging. 3, 1144–1166 (2023).
Tapia-Arancibia, L., Aliaga, E., Silhol, M. & Arancibia, S. New insights into brain BDNF function in normal aging and alzheimer disease. Brain Res. Rev. 59, 201–220 (2008).
Barbu, M. C. et al. Expression quantitative trait loci-derived scores and white matter microstructure in UK biobank: a novel approach to integrating genetics and neuroimaging. Transl Psychiatry. 10, 1–12 (2020).
Zhou, Z. et al. Molecular identification of protein kinase C beta in alzheimer’s disease. Aging 12, 21798–21808 (2020).
Torigoe, T. H. et al. Novel protective effect of the FOXO3 longevity genotype on mechanisms of cellular aging in Okinawans. Npj Aging. 10, 1–9 (2024).
McHugh, D. et al. COPI vesicle formation and N-myristoylation are targetable vulnerabilities of senescent cells. Nat. Cell. Biol. 25, 1804–1820 (2023).
Howard, A. & Rogers, A. N. Role of translation initiation factor 4G in lifespan regulation and age-related health. Ageing Res. Rev. 13, 115–124 (2014).
Watanabe, M., Fukuda, A. & Nabekura, J. The role of GABA in the regulation of GnRH neurons. Front Neurosci [Internet]. [cited 2024 Nov 13];8. Available from: https://www.frontiersin.org/journals/neuroscience/articles/ (2014). https://doi.org/10.3389/fnins.2014.00387/full
Wingo, A. P. et al. Expression of the PPM1F gene is regulated by stress and associated with anxiety and depression. Biol. Psychiatry. 83, 284–295 (2018).
Roberts, A. L. et al. Exposure to childhood abuse is associated with human sperm DNA methylation. Transl Psychiatry. 8, 1–11 (2018).
Håvik, B. et al. The complement control-related genes CSMD1 and CSMD2 associate to schizophrenia. Biol. Psychiatry. 70, 35–42 (2011).
Crider, K. S. et al. A review of molecular mechanisms and the evidence for folate’s Role2. Adv. Nutr. 3, 21–38 (2012).
Van Laarhoven, P. M. et al. Kabuki syndrome genes KMT2D and KDM6A: functional analyses demonstrate critical roles in craniofacial, heart and brain development. Hum. Mol. Genet. 24, 4443–4453 (2015).
Nacarelli, T. et al. The methyltransferase enzymes KMT2D, SETD1B, and ASH1L are key mediators of both metabolic and epigenetic changes during cellular senescence. Mol. Biol. Cell. 33, ar36 (2022).
de Magalhães, J. P. & Church, G. M. Genomes optimize reproduction: aging as a consequence of the developmental program. Physiology 20, 252–259 (2005).
Zwaan, B. J. Linking development and aging. Sci. Aging Knowl. Environ. 2003, pe32–pe32 (2003).
Gladyshev, V. N. et al. Disagreement on foundational principles of biological aging. PNAS Nexus. 3, pgae499 (2024).
Mendle, J., Ryan, R. M. & McKone, K. M. Early childhood maltreatment and pubertal development: replication in a Population-Based sample. J. Res. Adolesc. Off J. Soc. Res. Adolesc. 26, 595–602 (2016).
Middleton, L. Y. M. et al. Saliva cell type DNA methylation reference panel for epidemiological studies in children. Epigenetics 17, 161–177 (2022).
Puberty blockers for transgender and gender-diverse youth [Internet]. Mayo Clin. [cited 2024 Dec 18]. Available from: https://www.mayoclinic.org/diseases-conditions/gender-dysphoria/in-depth/pubertal-blockers/art-20459075
Zhu, J. & Chan, Y-M. Adult consequences of Self-Limited delayed puberty. Pediatrics 139, e20163177 (2017).
Harden, K. P., Tucker-Drob, E. M. & Tackett, J. L. The Texas twin project. Twin Res. Hum. Genet. 16, 385–390 (2013).
Reichman, N. E., Teitler, J. O., Garfinkel, I. & McLanahan, S. S. Fragile families: sample and design. Child. Youth Serv. Rev. 23, 303–326 (2001).
Petersen, A. C., Crockett, L., Richards, M. & Boxer, A. A self-report measure of pubertal status: reliability, validity, and initial norms. J. Youth Adolesc. 17, 117–133 (1988).
Watzlawik, M. The perception of similarities and differences among adolescent siblings: identification and deidentification of twins and nontwins. J. Adolesc. Res. 24, 561–578 (2009).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Halekoh, U., Højsgaard, S. & Yan, J. The R Package geepack for Generalized Estimating Equations. J Stat Softw [Internet]. [cited 2024 Aug 23];15. (2006). Available from: http://www.jstatsoft.org/v15/i02/
Van Iterson, M., Van Zwet, E. W., the BIOS Consortium & Heijmans, B. T. Controlling bias and inflation in epigenome- and transcriptome-wide association studies using the empirical null distribution. Genome Biol. 18, 19 (2017).
Villatoro-García, J. A. et al. DExMA: an R package for performing gene expression Meta-Analysis with missing genes. Mathematics 10, 3376 (2022).
Middleton, L. Y. M. et al. Saliva cell type DNA methylation reference panel for epidemiological studies in children. Epigenetics 17, 161–177 (2022).
Zheng, S. C., Breeze, C. E., Beck, S. & Teschendorff, A. E. Identification of differentially methylated cell types in epigenome-wide association studies. Nat. Methods. 15, 1059–1066 (2018).
Gatev, E., Gladish, N., Mostafavi, S. & Kobor, M. S. CoMeBack: DNA methylation array data analysis for co-methylated regions. Bioinformatics 36, 2675–2683 (2020).
Hagenbeek, F. A. et al. Intergenerational transmission of complex traits and the offspring methylome [Internet]. medRxiv; 2024 [cited 2024 Dec 17]. p. 2024.04.15.24305824. Available from: https://www.medrxiv.org/content/https://doi.org/10.1101/2024.04.15.24305824v2
Xiong, Z. et al. EWAS open platform: integrated data, knowledge and toolkit for epigenome-wide association study. Nucleic Acids Res. 50, D1004–D1009 (2022).
Elizarraras, J. M. et al. WebGestalt 2024: faster gene set analysis and new support for metabolomics and multi-omics. Nucleic Acids Res. 52, W415–W421 (2024).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Friedman, J. H., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting Linear Mixed-Effects Models Using lme4. J Stat Softw [Internet]. [cited 2023 Feb 4];67. (2015). Available from: http://www.jstatsoft.org/v67/i01/
Kuznetsova, A., Brockhoff, P. B., Christensen, R. H. B., Jensen, S. P. & lmerTest Tests in Linear Mixed Effects Models [Internet]. 2020 [cited 2024 Dec 20]. Available from: https://cran.r-project.org/web/packages/lmerTest/index.html
Bartoń, K. & MuMIn Multi-Model Inference. R package version 1.48.11 [Internet]. 2025 [cited 2025 Aug 19]. Available from: https://CRAN.R-project.org/package=MuMIn
Acknowledgements
None
Funding
Collection and analysis of data for the Texas Twin Project were supported by grant R01HD092548 from the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) to KPH and grant R01HD114724 from NICHD to KPH and LR. KPH and ETD are Faculty Research Associates of the Population Research Center at the University of Texas at Austin, which is supported by a National Institutes of Health grant (P2CHD042849). ETD is also a member of the Center on Aging and Population Sciences at the University of Texas at Austin, which is supported by the National Institutes of Health (P30AG066614). Collection and analysis of data for the Future of Families and Child Well-Being Study was funded by grants 2R01HD036916, R01HD039135, and R01HD040421. CM and DN were further supported with funds from the National Institutes of Health (R01MD011716, R01AG071071, R01HD076592, R01MH103761). KPH, ETD, LR, and CM have been supported by the Jacobs Foundation. YW, AMS, DC, EB, LR are supported by the Max Planck Society. LR is further supported by European Union grant #101073237. YW is further supported by the European Union’s Horizon Europe research and innovation program under the Marie Skłodowska-Curie grant agreement (No. 101150809). Collection and analysis of data for the German Twin Family Panel (TwinLife) was funded by the grant 220286500 from the German Research Foundation (DFG) awarded to Martin Diewald, Rainer Riemann, Frank M. Spinath, Christian Kandler, and Bastian Mönkediek. The molecular genetic extension project of TwinLife was funded by the DFG grant 428902522 awarded to Martin Diewald, Peter Krawitz, Markus M. Noethen, Rainer Riemann, and Frank M. Spinath. The epigenetic change satellite project of TwinLife was funded by the DFG grant 458609264 awarded to Elisabeth Binder, Martin Diewald, Andreas J. Forstner, Christian Kandler, Markus M. Noethen, and Frank M. Spinath. AMS, BM, and EB have been supported by the German Research Foundation. JAG was supported in part by an NICHD training grant to the Population Studies Center at the University of Michigan (T32HD007339).
Author information
Authors and Affiliations
Contributions
AJD, TS, JAG, YEW, and KPH conducted statistical analyses. KPH, LR, and CM conceptualized the study and supervised statistical analyses. AMS, DC, BM, CKLP, AJF, EBB, LS, DAN, CM, ETD, and KPH contributed to participant recruitment, data collection, and data pre-processing. AJD, LR, and KPH drafted the manuscript, and all authors provided critical feedback and revisions on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
deSteiguer, A.J., Smith, T., Goode, J.A. et al. Salivary DNA methylation and pubertal development in adolescents. Sci Rep 15, 35970 (2025). https://doi.org/10.1038/s41598-025-19588-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19588-1