Abstract
In this study, Shiga toxin-producing E. coli (STEC) were detected in 64% of all tested buffalo meat samples, distributed as 62% of buffalo meat chops and 66% of ground buffalo meat. Six serotypes (O55:H7, O111:H8, O145, O157:H7, O26:H11, and O103:H2) were identified, of which, O55:H7 (37.26%) and O111:H8 (21.29%) were the most predominant. The eaeA, stx1, stx2, and EHEC-hlyA were detected by multiplex PCR in all E. coli isolates at different percentages, and multiple virulence patterns with stx2/stx1/eae were the most prevalent pattern indicating that the serotypes will be more virulent in pathogenesis. The isolates showed high resistance rates against Erythromycin (100%), Clindamycin (86.7%), Nalidixic acid (84.4%), Cefepime (73.4%), and Ampicillin (62.7%), Cefazolin (53.2%) and Tetracycline (51.7%) and most of the isolates (73%) were classified as multi-drug resistant (MDR). For the first time, we identified a high frequency of Cefotaxime-resistant (28.5%) and Colistin-resistant (16.3%) E. coli from buffalo meat and this becomes a huge problem because Cefotaxime and colistin are typically used as a last resort to treat complex infections caused by multidrug-resistant Gram-negative bacilli. The emergence of virulent and multidrug-resistant E. coli strains in food seriously threatens human health. This highlights the need for antimicrobial stewardship programs in developing countries like Egypt to minimize the spread and emergence of these strains and prevent their dissemination to humans.
Similar content being viewed by others
Introduction
Buffalo has several physiological and genetic characteristics including the ability to withstand disease better than other animals, strong musculature, and tolerance to a wide range of environmental and nutritional changes, such as water scarcity, poor green fodders, high temperatures, and rough topography. All these features enable them to be one of the most important meat-producing animals in Egypt because they produce 381,000 tons of buffalo meat annually, which accounts for 45.3% of the native red meat produced in the country1. Although buffalo meat is highly nutritious food as they supply humans with protein of high biological value, minerals, fat, and polyunsaturated fatty acid with low cholesterol content, they are considered a good medium for the growth and multiplication of most spoilage and pathogenic bacteria contaminating carcasses during different stages of slaughtering and dressing from polluted water, air, dirty skin, intestinal contents, cutting tools, infected personnel, faulty slaughtering procedures, processing machines, handling, and storage.
Escherichia coli (E. coli) frequently inhabit the gastrointestinal tracts of both humans and animals as a normal component of their gut flora. Despite the fact that the majority of E. coli is harmless, some strains that have acquired specific virulence characteristics can cause extra-intestinal and diarrheal diseases in both humans and animals2. Diarrheagenic E. coli (DEC) is a major contributor to diarrhea outbreaks worldwide, accounting for 30 to 40% of acute diarrhea cases in children under five in developing nations3. The DEC is divided into pathotypes, each of which has distinct host preferences, worldwide occurrence, disease effects, and modes of transmission4. Shiga toxin-producing E. coli (STEC), Enterotoxigenic E. coli (ETEC), Enteropathogenic E. coli (EPEC), Enteroinvasive E. coli (EIEC), Enteroaggregative E. coli (EAEC), and Diffusely adherent E. coli (DAEC) are the six main pathotypes of DEC. Each E. coli pathotype has unique virulence factors encoded by particular gene clusters and unique pathogenic mechanisms5. These pathogenicity-linked genes may control characteristics such as motility, iron acquisition, adhesion, invasion, attachment, and toxin activity2.
According to reports, the most common cause of human gastroenteritis worldwide associated with the consumption of contaminated bovine meat is Escherichia coli, specifically Shiga toxin-producing ones (STEC)5. According to the roles that various serotypes play in human outbreaks, Shiga toxin-producing E. coli (STECs) can be divided into O157 and non-O157 serogroups. The number of human infections associated with non-O157 STEC has increased recently, and researchers in the US and Europe have discovered that non-O157 serogroups such as O26, O45, O103, O111, O113, O121, and O145 are responsible for roughly one-third of STEC infections. The reports indicated that STEC causes 2,801,000 acute illnesses worldwide yearly and leads to 3,890 cases of HUS, 270 cases of end-stage renal disease, and 230 deaths in the US, more than $1 billion each year6.
The presence of virulence genes such as Stx1, Stx2, Intimin (eaeA gene), and hemolysin (hlyA) is responsible for the pathogenicity and severity of Shiga toxin Escherichia coli strains (STEC) and is the primary cause of the clinical manifestations of STEC-induced infections in humans beings like bloody diarrhea that progress to life threaten hemolytic uremic syndrome7. The symptoms of STEC-induced human gastroenteritis may include diarrhea (roughly half of the cases are bloody diarrhea), hemorrhagic colitis (HC), hemolytic-uremic syndrome (HUS), which can cause kidney failure in children, thrombocytopenic purpura (TTP) in adults, mild fever, and cramping in the abdomen with an incubation period of three to four days. Most infected cases recover without medication, and the illness typically lasts 7 to 9 days. The severity of symptoms varies according to the host’s health, the implicated serogroups, and the age of the person. Serious symptoms are more likely to occur in older people, young children, and those with compromised immune systems8.
In recent decades, the occurrence of multidrug-resistant (MDR) E. coli strains, particularly STEC ones isolated from different foods of animal origin has increased worldwide, with more difficulty in the treatment of such infections. The widespread overuse or abuse of primarily common antibiotics as preventative, therapeutic, or growth promoters in meat-producing animals is the cause of the elevated rates of multidrug-resistant (MDR) microorganisms during the food production process9. Annually, more than 2.8 million Americans contract antibiotic-resistant infections, and more than 35,000 of them die as a result10. One of the most important multidrug-resistant (MDR) bacteria is Colistin and Cefotaxime-resistant E. coli which become a huge problem because Cefotaxime and colistin are typically used as a last resort to treat complex infections caused by multidrug-resistant Gram-negative bacilli. Contamination of raw meat with MDR E. coli may carry great public health hazards on human health due to the probable transmission of antibiotic-resistance genes to pathogenic bacteria11.
To the best of our knowledge, the information regarding the isolation of E. coli strains that produce Shiga toxin (STEC), particularly O157 and non-O157 STEC strains from Egyptian buffalo meat is lacking and the incidence of MDR Escherichia coli isolates against most used antibiotics are globally increased, thus the purpose of this work is to investigate the isolations of E. coli that produce Shiga toxin, including their prevalence, serotyping, virulence genes, and phenotypic antimicrobial resistance profile, in addition to public health hazards of Cefotaxime-resistant and Colistin-resistant E. coli recovered from buffalo meat sold in Mansoura city, Egypt.
Materials and methods
Collection of samples
Two hundred buffalo meat samples (100 samples each of ground buffalo meat and buffalo meat chops) were gathered from various retail butcher shops located throughout Mansoura, Egypt, between October 2021 and February 2022. After being aseptically placed in a polyethylene bag and numbered, each 200 g sample was quickly delivered to the Food Hygiene, Safety, and Technology Laboratory, Faculty of Veterinary Medicine, Mansoura University where the microbiological analyses were carried out. An overview of the study design is shown in Fig. 1.
A graphical abstract of the study design, workflow, and results.
Isolation and identification of Escherichia coli strains
After homogenizing 25 g of each buffalo meat sample with 225 milliliters of sterile modified tryptone soy broth (Oxoid, CMO989) that contained 40 milligrams of vancomycin per liter, the samples were incubated for 24 h at 37 °C. A loopful of the enriched culture was streaked onto sorbitol MacConkey agar (Oxoid, CMO813) supplemented with cefixime (0.05 mg/L) (Oxoid, SR0172E) and potassium tellurite (2.5 mg/L). The plates were then incubated for 24 h at 37 °C to check for typical colonies of Escherichia coli O157 and non-O157 strains. For additional biochemical identification and confirmation, five pink colonies (sorbitol-fermenting; SF) and five colorless colonies (non-sorbitol-fermenting; NSF) were chosen and subcultured onto nutrient agar (Oxoid, CM0003s) slopes. Along with other biochemical tests like indole, methyl red, Voges-Proskauer, and citrate utilization, 263 colonies of the NSF (colorless) and SF (pink) strains were examined for sorbitol fermentation, glucuronidase activity, and enterohemolysin production12.
Serological identification of E. coli strains
Rapid diagnostic testing was used to perform serological identification on E. coli isolates that had been biochemically identified. According to the instruction manuals, the E. coli polyvalent and monovalent antisera kit sets for O and H antigens (Denka Seiken Co., Ltd., Tokyo, Japan) were prepared to identify enteropathogenic serotypes.
Molecular characterization of E. coli isolates
Isolation of genomic DNA
Using QIAamp® genomic DNA extraction kits (QIAGEN, Germantown, MD, USA), the genomic DNA of the E. coli strains, the positive (E. coli O157:H7 reference strains), and a nonpathogenic negative control strain (E. coli K12 DH5α) that lacks any virulence gene were extracted following the manufacturer’s instructions and used as a template for PCR assays.
PCR for the identification of virulence genes and the rfbEO157 gene specific to E. coli O157.
The presence of rfbEO157 was examined in all of the 32 isolates that were serologically recognized as E. coli O157 strains. All of the 263 serologically identified E. coli strains were examined to discover specific virulence genes, such as EHEC-hlyA, eaeA, stx1, and stx2. The target genes’ primer sets for PCR amplification are listed in (Table 1).
The PCR amplification of the rfbE-O157 gene specific for E. coli O157:H7 was applied using a SimpliAmp thermal cycler (Thermo Fisher Scientific Inc, UK). GoTaq® Green Master Mix (Promega Corporation, Madison, USA). A 25-µl reaction mixture that contained 12.5 µl GoTaq® Green Master Mix, 2 µl of E. coli DNA template, 1 µl from each of the forward and reverse primers (0.4 µM each), 8.5 µl nuclease-free water. The protocol of PCR cycling was done as an initial denaturation at 95 °C for 4 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 56 °C for 30 s, and extension at 72 °C for 30 s, followed by a final extension at 72 °C for 5 min.
Multiplex PCR assays were carried out for the detection of stx1, stx2, eaeA, and hylA genes using a 50-µl reaction mixture that contained 25 µl GoTaq® Green Master Mix, 4 µl of E. coli DNA template, 0.5 µl from each of the forward and reverse primers (0.2 µM each), 19 µl nuclease-free water. The protocol of PCR cycling was done as an initial denaturation at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 58 °C for 40 s, and extension at 72 °C for 60 s, followed by a final extension at 72 °C for 7 min.
To separate the PCR products from each reaction mixture, ten microliters of each amplified PCR product were electrophoresed in 1.5% agarose gel (Puregene™, India) for 50 min at 95 V and then visualized under an ultraviolet transilluminator (Acculab, Montréal, Québec, Canada).
Antimicrobial resistance profile and multiple antibiotic resistance (MAR) index of E. coli strains
The disk diffusion susceptibility technique, as outlined by the Clinical and Laboratory Standards Institute, was used to determine the antimicrobial susceptibility profile of the 263 identified E. coli strains17. At varying concentrations, the method was used on sixteen antimicrobial discs from ten different antimicrobial classes (Oxoid Limited, Basingstoke, Hampshire, UK) including Polymyxin: Colistin (25 µg); Cephalosporins: Cefotaxime (30 µg), Cefazolin (30 µg); Amikacin (30 µg), Gentamicin (10 µg); Carbapenemes: Meropenem (10 µg), Cefepime (30 µg); Fluoroquinolones: 5 µg of ciprofloxacin and 30 µg of nalidixic acid; Erythromycin (15 µg) and azithromycin (15 µg) are macrolides. Penicillins: Ampicillin (10 µg), Amoxycillin-Clavulanic acid (30 µg); Lincomycins: Clindamycin (10 µg); Sulfonamides: Sulfamethoxazole (25 µg); and Tetracyclines: Tetracycline (30 µg).
Isolates were classified as susceptible, intermediate, or resistant following the National Committee for Clinical Laboratory Standards guidelines17. Isolates with intermediate susceptibility were regarded as resistant. The reference strain for antibiotic disc control was E. coliATCC 25 922. According to Singh et al18., the ratio of antimicrobial resistances to the total number of antimicrobials analyzed was used to calculate the multiple antibiotic resistances (MAR) index for each resistance pattern. A high danger of contamination is indicated when a MAR value is greater than 0.2. The isolate is considered MAR when they are resistant to at least three antimicrobials from two or more classes.
Results and discussion
Prevalence of STEC serotypes among examined Buffalo meat samples
Shiga toxin-producing Escherichia coli (STEC) are the causative agent of hemolytic uremic syndrome (HUS) and hemorrhagic colitis (HC) in humans, and outbreaks are mostly associated with bovine food sources. The majority of HC and HUS outbreaks in several nations, mostly in Europe and Latin America, have been attributed to the STEC O157:H7 serotype. But in the last fifteen years, non-O157 serotypes have emerged as important enteric pathogens and are commonly associated with HC and HUS in several countries, including the US, Ireland, Japan, Argentina, Chile, Germany, and Australia19,20. The HUS is characterized by acute renal failure in children, microangiopathic hemolytic anemia, and thrombocytopenia. Typically, STEC affects children under the age of five, the elderly, and patients with immuno-compromised systems. The ease of transmission and extremely low infectious dose (less than 10 cells) of this bacterium emphasize its significance as a foodborne pathogen21.
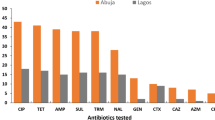
Of all buffalo meat samples tested, 64% (128/200) had STEC, which was distributed as 62% (62/100) of buffalo meat chops and 66% (66/100) of ground buffalo meat. By serological examination, six types of STEC were recovered from buffalo meat samples. Interestingly, O55:H7 was the most predominant serotype as it was present in 98 (37.26%) buffalo meat samples consisting of 45 and 53 meat chops and ground meat, respectively. The second most common serotypes, however, were O111:H7, O145, and O157:H7, which were recovered from 56 (21.29%), 46 (17.49%), and 32 (12.16%) buffalo meat samples distributed as 25, 21, and 14 of meat chops and 31, 25 and 18 of ground meat, respectively. On the other hand, O26:H11 and O103:H2 were the least recovered strains as they were present in 17 (6.46%) and 14 (5.32%) of buffalo meat samples distributed as 7 and 4 of meat chops and 10 and 10 of ground buffalo meat, respectively (Fig. 2a and b). The prevalence of E. coli O157:H7 in this study was 12.16%, while that of other E. coli serotypes was 87.83%. Comparable to our findings, the percentages of E. coli O157 and non-O157 in beef samples from the Hamadan industrial slaughterhouse were 12.5% and 87.5%, respectively22. In Nigerian meat products, the incidence of E. coli O157 versus non-O157 was likewise comparable at 28.8% and 79.2%23. In contrast, E. coli O157 and non-O157 serotypes were found in beef carcasses at 30% and 41%, respectively24. The STEC serogroups O26, O111, O103, and O145 found in this investigation are known to be extremely harmful to humans and have been implicated in food poisoning outbreaks with public health problems worldwide.
Serological identification of the 263 Escherichia coli isolates recovered from Egyptian buffalo meat (a). Distribution frequency of the different Escherichia coli serotypes identified in the buffalo meat chops and buffalo ground meat (b).
Numerous studies conducted globally have identified a broad range of distinct STEC serotypes from food samples and food-producing animals by varying rates of isolation. A similar finding in the same country was that the O55:H7 serotype was reported as being the most prevalent and O157:H7 serotypes come in the second order25. Another similar interesting study was recorded in Nigeria, in which O55:H7, O111:H8, and O145 were the most common serotypes isolated from buffalo meat cuts, meanwhile, O111 and O26 were the most predominant serotypes recovered from meat products23. Correspondingly, Fernández et al.26 reported that the most commonly recovered serotypes were O26:H11, O91:H21/H–, O103:H2, O111:H–, and O145:H28. In the same context, Nés tcheverría and Lía Padola27 found that O26:H11, O145: H–, and O157:H7 serotypes were among the most prevalent serotypes isolated from bovine samples. Furthermore, the strains of EHEC O26:H11 and O157:H7 were found to be the most frequently isolated from human patients infected with STEC in Germany and other countries28,29,30. In contrast to our findings, typical O26, O103, and O157 strains were the least recovered serotypes which represented only 1.8% of STEC isolates from food31. The cross-contamination of organisms that typically reside in the gut of food animals, including buffalo, and are expelled with the animal feces may be the cause of the comparatively higher incidence of E. coli O55:H7, O111:H7, O145, and O157:H7 among the tested buffalo meat samples in this study. Therefore, it is highly probable that E. coli will contaminate the meat during the slaughtering process, as well as, from the contaminated hide, particularly if poor evisceration occurs.
Virulence gene distribution and molecular characterization of E. coli strains isolated from buffalo meat
Several virulence factors are responsible for the pathogenicity of Shiga toxin-producing Escherichia coli (STEC) strains. These elements include the synthesis of at least one of the two Shiga toxins (stx1 and/or stx2), intimin (eaeA), and enterohemolysin (EHEC-hlyA) which cause human bloody diarrhea that progresses to life-threatening Hemolytic Uremic Syndrome. Shiga toxins can also bind to cellular receptors and prevent the synthesis of proteins in many organs, including the kidney, brain, and liver, leading to serious illnesses32.
The results of the serological identification of E. coli isolates showed that only 32 isolates (12.16%) were serotyped as O157:H7, whereas 231 isolates (87.83%) were serotyped as non-O157 strains, such as O55:H7, O111:H8, O145, O26:H11, and O103:H2. The presence of the rfbEO157 gene, which is specific for E. coli O157 genetic identification, was further verified by PCR in 32 E. coli O157:H7 isolates (Fig. 3). Multiplex PCR was used to determine whether E. coli O157:H7 (n = 32) and non-O157 isolates (n = 231) had eaeA, stx1, stx2, and EHEC-hlyA, which were found at the expected molecular sizes of 890 bp, 614 bp, 779 bp, and 165 bp, respectively (Fig. 4).
Representative Agarose gel electrophoresis for PCR product of the amplified rfbEO157 gene for confirmation of the Enterohaemorrhagic E. coli O157:H7. Six microliters from the amplified DNA at the expected molecular size of 129 bp for the rfbEO157 gene were separated by electrophoresis on 1.5% agarose gel and visualized under UV light after being stained with ethidium bromide. Lane M: 100 bp ladder as molecular size DNA marker. Lane C+: amplified product using E. coli O157:H7 Sakai (EHEC) genome template as a positive control reference strain. Lane C–: E. amplified product using coli K12 DH5α genome template as a negative control strain; Lanes with the key numbers from 1 to 10 represented the positive strains for target genes.
Representative agarose gel electrophoresis for the multiplex PCR products of the amplified virulence genes (stx1, stx2, eaeA, and hlyA) showing their distribution pattern among the six different serotypes of the Enterohaemorrhagic E. coli isolates. Ten microliters from the amplified DNA were separated by electrophoresis on 1.5% agarose gel and visualized under UV light after being stained with ethidium bromide. Amplicons from positive strains exhibited the expected molecular size of 614, 779, 890, and 165 bp for stx1, stx2, eaeA, and hlyA genes, respectively. Lane M: 100 bp ladder as molecular size DNA marker. Lane C+: amplified product using a template from E. coli O157:H7 Sakai (EHEC) genome as a positive control reference strain. Lane C–: E. amplified product using coli K12 DH5α genome template as a negative control strain; Lanes with the key numbers 1, 2, 3, 4, 5, and 6 denote representative isolates from E. coli Serotypes O55:H7, O111:H8, O145, O157:H7, O26:H11, and O103:H2, respectively. The letter “A” denotes a positive pattern for stx1 and stx2 genes only. The letter “B” denotes a positive pattern for stx1, stx2, and eaeA genes. The letter “C” denotes a positive pattern for stx1, stx2, eaeA, and hlyA genes. The letter “D” denotes a positive pattern for stx2, eaeA genes only. The letter “E” denotes a positive pattern for stx2, eaeA, and hlyA genes. The letter “F” denotes a positive pattern for stx1, eaeA, and hlyA genes. The letter G denotes a positive pattern for stx1, stx2, and hlyA genes.
Shiga toxin genes (stx1 and stx2)
Shigatoxins, which are encoded by the stx1 and stx2 genes and carried by lysogenic phages, are the primary virulence factor of STEC. While there are variations in both Shiga toxin genes, stx2 is the most heterogeneous group. Compared to strains that only harbor stx1 or even both, those that are stx2 positive may be more virulent and are more commonly associated with HUS30.
All the 263 E. coli isolates found in buffalo meat during the current investigation were Shiga toxin-producing (STEC), as shown in Table (2). The most common virulence gene found in STEC strains was Shiga toxin 2 (stx2), which was found in 88.21% (232/263) of the isolates distributed into 98 (O55:H7), 56 (O111:H8), 32 (O157:H7), 27 (O145), 14 (O103:H2) and 5 (O26:H11). However, only 69.58% (183/263) of the isolates had only stx1 consisting of 87 (O55:H7), 40 (O111:H8), 19 (O145), 16 (O157:H7), 12 (O26:H11) and 9 (O103:H2). Furthermore, 57.79% (152/263) of total isolates carried both stx1 and stx2 (Table 2). As was previously mentioned, strains carrying the stx2 gene might be more severe than those carrying the stx1 gene, or even strains carrying both stx1 and stx2. Amazingly, stx2 had been reported to be 1000 times more cytotoxic to the human kidney’s microvascular endothelial cells than stx133,34.
In STEC strains from animal-based food, the frequency of occurrence and the correlation between the stx1 and stx2genes differed from study to study. Sallam et al35. found that of the E. coli O157 isolates from retail beef in Egypt, stx2 was found in 86.66% of cases and stx1 in 46.7%. Sallam et al.25, in another study in Egypt, discovered that stx2, stx1, and both stx1 and stx2 were present in 97.6%, 27.8%, and 25.4% of STEC O157:H7 and O55:H7 isolated from camel meat, respectively. In Germany, Slanec et al.36 and Beutin et al.31 found that stx2 was present in 70.7% and 81% of STEC strains recovered from fresh meat samples, respectively. According to Fayemi et al.23, 18.75% of the isolated E. coli O157:H7 from Nigerian meat samples contained stx2. In Ethiopia, stx2 was found in 75% and 56% of recovered E. coli O157 from beef samples, respectively37,38. Lee et al.39 found that stx2 and both stx1 and stx2 were present in 64% and 14% of the STEC strains obtained from fresh beef in Korea, respectively. The stx1 gene, stx2 gene, and both stx1 and stx2 genes were present in 29%, 51%, and 20% of the STEC isolates discovered in minced beef tested in Spain, respectively40. Furthermore, Hessain et al.41, Samad et al.42, and Manage et al.24 found that both stx1 and stx2 genes were present in 45.45%, 10%, and 22.9% of isolated E. coli O157:H7 in Saudi Arabia, Pakistan, and Canada, respectively. In contrast to our findings, the only E. coli O157 isolates found in Iranian camel meat didn’t contain the virulent genes stx1 and stx243.
Intimin (eaeA) and enterohemolysin (hlyA) genes
Another prevalent virulence factor is Intimin, which is necessary for the bacteria to adhere closely to epithelial cells and produce the characteristic histopathological lesion known as “attaching and effacing” (A/E). Regarding stx genes, eae was found in stx1-positive STEC strains more often than stx2-positive strains, and calves had higher levels of both eae and stx1 than adult cattle. Intimin and Shiga toxins are well-established virulence factors that have been thoroughly studied. Though several studies are required to fully understand the role that EHEC hlyA may play in the pathogenicity of EHEC infection, other research has found that EHEC hlyA, which is present in all E. coli O157:H7 strains, the majority of non-O157 EHEC strains, and most of eaeA-positive STEC isolates from animals, may serve as both a virulence marker and a virulence factor. Perhaps during an EHEC infection, Shiga toxins, Intimin, and EHEC hlyA work together to cause illness. Combinations of eaeA and EHEC- hlyA might be helpful in distinguishing virulent STEC from less harmful or harmless STEC44,45.
After stx2, the Intimin (eaeA) gene was the second most common virulence gene in the current investigation, found in 77.94% (205/263) of the total isolates. However, hlyA gene was present in 45.24% (119/263) of the isolates. The eaeA gene was present in all (100%) O157:H7, (100%) O145, and (100%) O26:H11 isolates, 75% of O111:H8 isolates, 62.28% of O55:H7 isolates and 35.71% of O103:H2 isolates recovered from the buffalo meat in the current investigation (Table 2). The STEC isolates carrying 77.94% of eaeA gene in the present study was lower than that reported in Egypt (93.3%), Ireland (95.3%), and in the UK (100%) by Sallam et al.35; Cagney et al.46, and Chapman et al.47, respectively, but higher than that reported in beef meat from Spain (10%) by Mora et al.48, in Germany (5%) and by Beutin et al.31, who were unable to identify the eaeA gene in any of the STEC strains that recovered from meat products. However, the prevalence of eaeA gene observed from buffalo meat in the present study was comparable to that recorded in Egypt by Salam et al.25 who found that 82.4% of E. coli O55:H7 isolates recovered from camel meat harbored eaeA gene.
An additional virulence-associated marker, among STEC serotypes, is enterohemolysin which is encoded by hlyA gene and implicated in E. coli’s virulence. It was indicated from our results that 71.87% of E. coli O157:H7, 76.08% of E. coli O145, 70.58% of E. coli O26:H11, 28.57% of E. coli O111:H8, 24.48% of E. coli O55:H7 and 64.28% of E. coli O103:H2 isolates were positive for hlyA gene with an overall prevalence of 45.24% (119/263) among the STEC isolates tested (Table 2).
Combination patterns of stx1, stx2, EaeA and HlyA genes in STEC strains
It was indicated from the results of the multiplex PCR assay showed that there was no isolate possessed a single virulence pattern (stx1 only, stx2 only, eaeA only, and hlyA only), and all recovered isolates in this study showed multiple virulence patterns which may indicate that the serotypes will be more virulent in pathogenesis. A total of 7 diverse virulence profiles of STEC isolates were recovered throughout the present study which arranged in the following decreasing order: stx2/stx1/eae (54 isolates), stx2/stx1 (49 isolates), stx2/eae (41 isolates), stx2/stx1/eae/hlyA (40 isolates), stx2/eae/hlyA (39 isolates), stx1/eae/hlyA (31 isolates), and stx2/stx1/hlyA (9 isolates). The most predominant virulence profile was stx2/stx1/eae, followed by stx2/stx1 profile which may indicate a serious zoonotic threat in this geographic region. Several diverse virulence profiles of STEC isolates were recorded by several publications which have clarified a strong correlation between these virulence genes and the capacity of STEC isolates to infect humans with serious illnesses, especially HUS and bloody diarrhea39,44,49,50,51.
Antibiotic resistance profiles and multiple antibiotic resistances (MAR) index of STEC serotypes
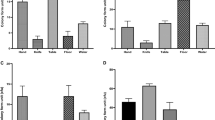
All E. coli isolates recovered from buffalo meat samples were tested for antimicrobial resistance against 16 antimicrobial agents, each with different modes of action, and widely used in human and veterinary medicine. Very high resistance rates were detected against Erythromycin (100%), Clindamycin (86.7%), Nalidixic acid (84.4%), Cefepime (73.4%), and Ampicillin (62.7%), followed by Cefazolin (53.2%), Tetracycline (51.7%). On the other hand, low resistance rates of 16.3%, 12.5%, 7.6%, 5.3%, 2.7%, and 1.5% were noticed against Colistin, Amikacin, Ciprofloxacin, Azithromycin, Gentamicin and Meropenem (Table 3). According to the current findings, every isolate exhibited resistance to at least one or more of the antibiotics under investigation, with 100% demonstrating resistance to erythromycin (E). At the same time, the most effective antibiotics were Meropenem (98.5%), Gentamicin (94.7%), and Azithromycin (93.5%), followed by Ciprofloxacin (92.4%), Amikacin (85.6%), and Colistin (78.3%), with other susceptibility patterns shown in Table (3). The number and % of E. coli strains resistant to the different antimicrobial agents (n = 16) tested are shown in Table (4). Furthermore, the results of the study revealed that 71.43%, 62.5%, 60.87%, 68.75%, 58.82% and 64.29% of E. coli O55:H7, O111:H8, O157:H7, O26:H11, O103:H2 and O103:H2 isolates, respectively were classified as multi-drug resistant (MDR), and different resistance profiles were showed in Table (5). Moreover, 192 isolates (73%) showed a multiple antibiotic resistance index (MAR) exceeding 0.2, whereas 71 isolates (27%) displayed a MAR index below 0.2. With a MAR index of 1.0, it is noteworthy that 1.52% (4/263) of E. coli isolates (all of which are from the O55:H7 serovars) showed resistance to all 16 tested antibiotics (Table 5).
Regarding the results of Amikacin and Gentamicin which appeared to be as two of the most effective antibiotics in treatment of the pathogenic E. coli, nearly similar susceptibility rates were recorded in a previous Egyptian study by Sallam et al.25 who found that 60.3% and 83.3% of the recovered E. coli O157:H7 and E. coli O55:H7 isolates from camel meat were sensitive toward Amikacin and Gentamicin, respectively. Additionally, 100% of the E. coli O157:H7 isolated from raw cattle meat in Ethiopia were susceptible to amikacin and gentamicin, according to Hiko et al.52. Other effective antibiotics in the treatment of E. coli recorded in the current study were Meropenem and Ciprofloxacin and these findings were supported by Debbarma et al.53 who found that 100% and 94.67% of E. coli isolates were susceptible to Meropenem and Ciprofloxacin, respectively. Also, Sallam et al.25 and El-Ghareeb et al.54 recorded nearly similar susceptibility rates (77.8% and 82.4%) against Ciprofloxacin. Because of their limited application in animal production, amikacin, gentamicin, meropenem, ciprofloxacin, and colistin have comparatively lower rates of resistance.
Our findings on erythromycin and clindamycin were comparable to those of a prior study by Sallam et al.25, which discovered that all of the E. coli isolates from Egyptian camel meat showed resistance to erythromycin and Clindamycin and this was supported by the findings of Snodgrass and Motaparthi (2021)55, who mentioned that Clindamycin was effective against Gram-positive and anaerobic bacteria, but it wasn’t effective against E. coli. Very high resistance rates of 92.9%, 71.4%, and 48.4% were noted for Tetracycline, Ampicillin, and Sulfamethoxazole-trimethoprim, respectively25. Other higher resistance rates (77.7%), (62.5%) and (60%) against tetracycline were recorded by Shekh et al.56; Babolhavaeji et al.22; Debbarma et al.53, respectively. Concerning Nalidixic acid and Ampicillin, lower resistance rates (20% and 40%) were recorded in Bangladesh57. Regarding the resistance against Amoxycillin-Clavulanic acid, a very low resistance rate (4%) was recorded in Ethiopia by Haile et al.58, which give an indication that Amoxycillin-Clavulanic acid is still effective in the treatment of E. coli as it contained B- lactamase inhibitor. Geographical location, study design, testing methodologies, guidelines for interpreting the results, locally approved medications, farm-level management, and antibiotic misuse or overuse may all be contributing factors to the variations in resistance patterns observed in various studies59.
Colistin- and Cefotaxime-resistant E. coli isolated from Buffalo meat
The emergence of multidrug-resistant bacteria and the sharp rise in antibiotic resistance have further reduced the number of available treatments for bacterial illnesses by various antibiotics60. As a result, colistin, an older and less accessible antibiotic, has been reintroduced globally as a last-resort medication for the treatment of serious infections caused by MDR Gram-negative bacteria61. Cefotaxime, a third-generation cephalosporin, is another preferred antibiotic that is typically used as a last resort to treat complex infections. However, the extensive application of antibiotics, especially colistin, and cefotaxime, for veterinary treatment, disease prevention, and growth promotion has led to the emergence and the increase of colistin-resistant and cefotaxime-resistant bacteria, compromising the effectiveness of colistin and cefotaxime15. Plasmid-mediated colistin resistance genes, such as mcr-1 to mcr-10 genes, maybe the cause of the colistin resistance62. Colistin- and cefotaxime-resistant Escherichia coli have been documented in food, animals, humans, and the environment worldwide61,63,64,65,66,67. However, in Egypt, a few studies have isolated colistin or cefotaxime-resistant E. coli from food samples, especially from buffalo meat.
Data regarding Colistin and Cefotaxime resistance in bacteria from animals and food of animal origin are relatively scarce. It was not until 2014, that colistin and Cefotaxime were incorporated into the mandatory antimicrobial panel for Enterobacteriaceae as part of the European surveillance of bacteria originating from animals. To the best of our knowledge, these antibiotics had not been included in the antimicrobial resistance monitoring programs of numerous other countries until quite recently. In this investigation, isolated E. coli strains from buffalo meat showed a high frequency (28.5%) of Cefotaxime resistance. Concerningly, these strains exhibited simultaneous resistance to quinolones, aminoglycosides, sulfonamides, and cephalosporins. More worryingly, 16.3% of E. coli strains exhibited resistance to Colistin. Cefotaxime and colistin are frequently used as a last resort to treat infections caused by multidrug-resistant Gram-negative bacilli62. The spread of Colistin-resistant and Cefotaxime-resistant pathogens has attracted global attention. Here, we report the first cases of Colistin-resistant and Cefotaxime-resistant E. coli from buffalo meat worldwide. These strains were multidrug-resistant and were resistant to third-generation cephalosporins Cefotaxime, Colistin, Erythromycin, Clindamycin, Nalidixic acid, Tetracycline, Amoxycillin-Clavulanic acid and Sulfamethoxazole. The emergence of such strains further complicates the treatment of E. coli infections. Nearly similar results of Colistin resistance (20.0%) and a high frequency (82.9%) of cefotaxime resistance were observed in E. coli O157 non-H7 strains isolated from retail food in China68.
Our results support the generally accepted theory that buffalo is a significant source of multidrug-resistant E. coli and highlight the importance of antibiotic susceptibility surveys in determining the best course of treatment for E. coli strains originating from cows. The information also emphasizes the necessity of establishing antimicrobial stewardship initiatives in developing countries, such as Egypt, in order to maximize their therapeutic use and minimize the spread and emergence of strains that are resistant to antibiotics.
Conclusions
The high frequency of Cefotaxime-resistant (28.5%) and Colistin-resistant (16.3%) E. coli from buffalo meat and this becomes a huge problem because Cefotaxime and Colistin are typically used as a last resort to treat complex infections caused by multidrug-resistant Gram-negative bacilli. The great majority of STEC (73%) isolates were classified as multi-drug resistant (MDR) which indicated overuse and/or misuse of the antibiotics. Additionally, the virulence of E. coli isolates was determined by the existence of the eaeA, stx1, stx2, and EHEC-hlyA genes which were detected in all E. coli isolates at different percentages and multiple virulence patterns with stx2/stx1/eae were the most prevalent pattern indicating that the serotypes will be more virulent in pathogenesis. Therefore, the emergence of virulent and multidrug-resistant E. coli strains in buffalo meat can constitute a significant public health concern, emphasizing the necessity of the implementation of a better antimicrobial stewardship program in developing countries like Egypt to minimize the spread and emergence of these strains and prevent their dissemination to humans. In addition, applying control measures based on the prevention of contamination of meat at slaughter and butcher shops level through designing a multi-hurdle approach, establishing a comprehensive HACCP plan tailored to food plant, and creating detailed SOPs, as well as proper cooking of meat at private households.
Data availability
All data supporting the findings of this study are included within the article. No datasets were generated or analysed during the current study.
References
FAO. FAOSTAT Database. food and agriculture organization of the united nations, Rome, Italy. (2019). Available at: https://www.fao.org/faostat/en
Kaper, J. B., Nataro, J. P. & Mobley, H. L. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2, 123–140. https://doi.org/10.1038/nrmicro818 (2004).
Miliwebsky, E. et al. Human diarrheal infections: diagnosis of diarrheagenic Escherichia coli Pathotypes in Escherichia coli in the Americas. ed. A.G. Torres (Switzerland: Springer). ;343–369. (2016).
Croxen, M. A. & Finlay, B. B. Molecular mechanisms of Escherichia coli pathogenicity. Nat. Rev. Microbiol. 8, 26–38. https://doi.org/10.1038/nrmicro2265 (2010).
Pakbin, B., Brück, W. M. & Rossen, J. W. Virulence factors of enteric pathogenic Escherichia coli: a review. Int. J. Mol. Sci. 22, 9922. https://doi.org/10.3390/ijms22189922 (2021).
Majowicz, S. E. et al. Global incidence of human Shiga toxin producing Escherichia coli infections and deaths: a systematic review and knowledge synthesis. Food Pathog Dis. 6, 447e55. https://doi.org/10.1089/fpd.2013.1704 (2014).
Gonzalez-Escalona, N. & Kase, J. A. Virulence gene profiles and phylogeny of Shiga toxin-positive Escherichia coli strains isolated from FDA regulated foods during 2010–2017. PLoS One. 14, e0214620. https://doi.org/10.1371/journal.pone.0214620 (2019).
Joseph, A. et al. Shiga toxin-associated hemolytic uremic syndrome. A Narrative Review Toxins 20201;2:67. https://doi.org/10.3390/toxins12020067
Capita, R. & Alonso-Calleja, C. Antibiotic-resistant bacteria: A challenge for the food industry. Crit. Rev. Food Sci. Nutr. 53, 11–48. https://doi.org/10.1080/10408398.2010.519837 (2013).
CDC. Antibiotic resistance threats in the United States, 2019, Center for disease control and prevention, U.S. department of health and human services, Atlanta, GA. (2019). https://doi.org/10.15620/cdc:82532
Chabou, S. et al. Real-time quantitative PCR assay with Taqman1 probe for rapid detection of MCR-1 plasmid-mediated colistin resistance. New. Microbes New. Infect. 13, 71–74. https://doi.org/10.1016/j.nmni.2016.06.017 (2016).
ISO 16654. (R 2018). Microbiology of Food and Animal Feeding Stuffs e Horizontal Method for the Detection of Escherichia coli O157. International Organization for Standardization: Geneva, Switzerland. Available online: (2001). https://www.iso.org/standard/29821.html
Dhanashree, B. & Mallya, S. Detection of Shiga-toxigenic Escherichia coli (STEC) in diarrhoeagenic stool and meat samples in Mangalore, India. Indian J. Med. Res. 128, 271–277 (2008).
Mazaheri, S., Ahrabi, S. & Aslani, M. Shiga toxin-producing Escherichia coli isolated from lettuce samples in Tehran, Iran. Jundishapur J. Microbiol. 7 (11), 1–6. https://doi.org/10.5812/jjm.12346 (2014).
Hessain, A. M. et al. Molecular characterization of Escherichia coli O157:H7 recovered from meat and meat products relevant to human health in Riyadh, Saudi Arabia. Saudi J. Biol. Sci. 22 (6), 725–729. https://doi.org/10.1016/j.sjbs.2015.06.009 (2015).
Sharma, V. K. Real-time reverse transcription-multiplex PCR for simultaneous and specific detection of RfbE and EaeA genes of Escherichia coli O157:H7. Molecul Cell. Prob. 20 (5), 298–306. https://doi.org/10.1016/j.mcp.2006.03.001 (2006).
CLSI., (Clinical and Laboratory Standards Institute). Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Wayne, PA: M07USA., 2018. (2018). https://clsi.org/media/1928/m07ed11_sample.pdf
Singh, S., Yadav, A. S., Singh, S. M. & Bharti, P. Prevalence of Salmonella in chicken eggs collected from poultry farms and marketing channels and their antimicrobial resistance. Food Res. Int. 43, 2027–2030. https://doi.org/10.1016/j.foodres.2010.06.001 (2010).
Bettelheim, K. A. The non-O157 shiga-toxigenic (verocytotoxigenic) Escherichia coli; under-rated pathogens. Crit. Rev. Microbiol. 33, 67–87. https://doi.org/10.1080/10408410601172172 (2007).
Essendoubi, S. et al. Prevalence of Shiga toxin-producing Escherichia coli (STEC) O157:H7, six non-O157 STECs, and Salmonella on beef carcasses in provincially licensed abattoirs in alberta, Canada. Food Cont. 105, 226–232. https://doi.org/10.1016/j.foodcont.2019.05.032 (2019).
Travert, B. et al. Shiga toxin-associated haemolytic uremic syndrome: specificities of adult patients and implications for critical care management. Toxins 13 (5), 306. https://doi.org/10.3390/toxins13050306 (2021).
Babolhavaeji, K. et al. Prevalence of Shiga Toxin-producing Escherichia coli O157 and Non-O157 serogroups isolated from fresh Raw beef meat samples in an industrial slaughterhouse. Int. J. Microbiol. 1978952. https://doi.org/10.1155/2021/1978952 (2021).
Fayemi, O. E. et al. Prevalence, characterization and antibiotic resistance of Shiga toxigenic Escherichia coli serogroups isolated from fresh beef and locally processed ready-to-eat meat products in Lagos, Nigeria. Int. J. Food Microbiol. 347, 109191. https://doi.org/10.1016/j.ijfoodmicro.2021.109191 (2021).
Manage, D. P. et al. Detection of pathogenic Escherichia coli on potentially contaminated beef carcasses using cassette PCR and conventional PCR. BMC Microbiol. 19 (1), 175. https://doi.org/10.1186/s12866-019-1541-4 (2019).
Sallam, K. I. et al. cefotaxime-, ciprofloxacin-, and extensively drug-resistant Escherichia coli O157:H7 and O55:H7 in camel meat. Foods 12, 1443. https://doi.org/10.3390/foods12071443 (2023).
Fernández, D., Sanz, M. E., Parma, A. E. & Padola, N. L. Short communication: characterization of Shiga toxin-producing Escherichia coli isolated from newborn, milk-fed and growing dairy calves. J. Dairy. Sci. 95, 5340–5343. https://doi.org/10.3168/jds.2011-5140 (2012).
Nés tcheverría, A. & Lía Padola, N. Shiga toxin-producing Escherichia coli factors involved in virulence and cattle colonization. Virulence 4, 366–372. https://doi.org/10.4161/viru.24642 (2013).
Nuesch-Inderbinen, M., Treier, A., Stevens, M. J. & Stephan, R. Whole genome sequence-based characterization of Shiga toxin-producing Escherichia coli isolated from game meat originating from several European countries. Sci. Rep. 13, 3247. https://doi.org/10.1038/s41598-023-30333-4 (2023).
Rodwell, E. V. et al. The epidemiology of Shiga toxin-producing Escherichia coli O26:H11 (clonal complex 29) in England, 2014–2021. J. Infect. 86, 552–562. https://doi.org/10.1016/j.jinf.2023.04.006 (2023).
Minary, K. et al. Outbreak of hemolytic uremic syndrome with unusually severe clinical presentation caused by Shiga toxin-producing Escherichia coli O26:H11 in France. Arch. Pedia. 29, 448–452. https://doi.org/10.1016/j.arcped.2022.05.011 (2022).
Beutin, L. et al. Identification of human-pathogenic strains of Shiga toxin-producing Escherichia coli from food by a combination of serotyping and molecular typing of Shiga toxin genes. Appl. Environ. Microbiol. 73, 4769–4775. https://doi.org/10.1128/AEM.00873-07 (2007).
Smith, J. L., Fratamico, P. M. & Gunther, N. W. Shiga toxin-producing Escherichia coli. Adv. Appl. Microbiol. 86, 145–197. https://doi.org/10.1016/B978-0-12-800262-9.00003-2 (2014).
Karama, M. et al. Molecular profiling and antimicrobial resistance of Shiga toxin-producing Escherichia coli O26, O45, O103, O121, O145 and O157 isolates from cattle on cowcalf operations in South Africa. Sci. Rep. 9 (1), 11930. https://doi.org/10.1038/s41598-019-47948-1 (2019).
Bauwens, A. et al. Differential cytotoxic actions of Shiga toxin 1 and Shiga toxin 2 on microvascular and macrovascular endothelial cells. Thromb. Haemost. 105, 515–528. https://doi.org/10.1160/TH10-02-0140 (2011).
Sallam, K. I., Mohammed, M. A., Ahdy, A. M. & Tamura, T. Prevalence, genetic characterization and virulence genes of sorbitol-fermenting Escherichia coli O157: H-and E. coli O157: H7 isolated from retail beef. Int. J. Food Microbiol. 165 (3), 295–301. https://doi.org/10.1016/j.ijfoodmicro.2013.05.024 (2013).
Slanec, T., Fruth, A., Creuzburg, K. & Schmidt, H. Molecular analysis of virulence profiles and Shiga toxin genes in food-borne Shiga toxin-producing Escherichia coli. Appl. Environ. Microbiol. 75, 6187–6197. https://doi.org/10.1128/AEM.00874-09 (2009).
Gutema, F. D. et al. Occurrence, molecular characteristics, and antimicrobial resistance of Escherichia coli O157 in cattle, beef, and humans in Bishoftu Town, central Ethiopia. Foodborne Pathog Dis. 18 (1). https://doi.org/10.1089/fpd.2020.2830 (2021).
Ayenew, H. Y., Mitiku, B. A. & Tesema, T. S. Occurrence of virulence genes and antimicrobial resistance of e. coli o157:h7 isolated from the beef carcass of Bahir Dar city, Ethiopia. Vet. Med. Int. 8046680. https://doi.org/10.1155/2021/8046680 (2021).
Lee, G. Y., Jang, H. I., Hwang, I. G. & Rhee, M. S. Prevalence and classification of pathogenic Escherichia coli isolated from fresh beef, poultry, and pork in Korea. Int. J. Food Microbiol. 134, 196–200. https://doi.org/10.1016/j.ijfoodmicro.2009.06.013 (2009).
Mora, A. et al. Serotypes, virulence genes and intimin types of Shiga toxin (Verocytotoxin)–producing Escherichia coli isolate from minced beef in Lugo (Spain) from 1995 through 2003. BMC Microbiol. 7, 13. https://doi.org/10.1186/1471-2180-7-13 (2007).
Hessain, A. M. et al. Molecular characterization of Escherichia coli O157:H7 recovered from meat and meat products relevant to human health in Riyadh, Saudi Arabia. Saudi J. Biol. Sci. 22, 725–729. https://doi.org/10.1016/j.sjbs.2015.06.009 (2015).
Samad, A. et al. Multiplex polymerase chain reaction detection of Shiga toxin genes and antibiotic sensitivity of Escherichia coli O157:H7 isolated from beef meat in Quetta, Pakistan. J. Food Saf. 38, 1–8. https://doi.org/10.1111/jfs.12540 (2018).
Rahimi, E., Kazemeini, H. R. & Salajegheh, M. Escherichia coli O157:H7/NM prevalence in Raw beef, camel, sheep, goat, and water Buffalo meat in Fars and Khuzestan provinces, Iran. Vet. Res. Forum. 3, 13–17 (2012).
Blanco, M. et al. Serotypes, virulence genes and intimin types of Shiga toxin (verotoxin)-producing Escherichia coli isolates from cattle in Spain and identification of a new intimin variant gene (eae). J. Clin. Microbiol. 42, 645–651. https://doi.org/10.1128/JCM.42.2.645-651.200435 (2004).
Guth, B. E., Prado, V. & Rivas, M. Shiga toxin-producing Escherichia coli. In: Torres AG, ed. Pathogenic Escherichia coli in Latin America. Bentham Science Publishers Ltd: United States, ;65–83. (2010).
Cagney, C. et al. Prevalence and numbers of Escherichia coli O157:H7 in minced beef and beef burgers from butcher shops and supermarkets in the Republic of Ireland. Food Microbiol. 21, 203–212. https://doi.org/10.1016/S0740-0020(03)00052-2 (2004).
Chapman, P. A., Cornell, J. & Green, C. Infection with verocytotoxin-producing Escherichia coli O157 during a visit to an inner City open farm. Epidemiol. Infect. 125, 531–536. https://doi.org/10.1017/s0950268800004775 (2000).
Mora, A. et al. Antimicrobial resistance of Shiga toxin (verotoxin)-producing Escherichia coli O157 and non-O157 strains isolated from humans, cattle, sheep and food in Spain. Res. Microbiol. 156, 793–806. https://doi.org/10.1016/j.resmic.2005.03.006 (2005).
Binandeh, F., Pajohi-Alamoti, M., Mahmoodi, P. & Ahangari, A. Evaluation of stx1, stx2, hlyA, and EaeA virulence genes in Escherichia coli O157:H7 isolated from meat (beef and mutton) in Hamedan, Iran, during 2015–2016. Int. J. Enteric Pathog. 8 (2), 55–59. https://doi.org/10.34172/ijep.2020.12 (2020).
Aidar-Ugrinovich, L. et al. Serotypes, virulence genes, and intimin types of Shiga toxin-producing Escherichia coli (STEC) and enteropathogenic E. coli (EPEC) isolated from calves in Sao Paulo, Brazil. Int. J. Food Microbiol. 115, 297–306. https://doi.org/10.1016/j.ijfoodmicro.2006.10.046 (2007).
Srivani, M. et al. Prevalence and antimicrobial resistance pattern of Shiga toxigenic Escherichia coli in diarrheic Buffalo calves. Vet. World. 10 (7), 774. https://doi.org/10.14202/vetworld.2017.774-778 (2017).
Hiko, A., Asrat, D. & Zewde, G. Occurrence of Escherichia coli O157: H7 in retail Raw meat products in Ethiopia. J. Infect. Dev. Count. 2 (5), 389–393. https://doi.org/10.3855/jidc.203 (2008).
Debbarma, M., Deka, D., Tolenkhomba, C. T. & Rajesh, J. B. Microbiological contamination of retail meat from Mizoram (India) with special reference to molecular detection and multi-drug resistance of Escherichia coli. Indian J. Vet. Sci. Anim. Biotechnol. 18, 32–35 (2022). https://www.researchgate.net/publication/376486167
El-Ghareeb, W. R. et al. Isolation and identification of extended spectrum-lactamases (ESBLs) Escherichia coli from minced camel meat in Eastern Province, Saudi Arabia. Thai J. Vet. Med. 50, 155–161 (2020). https://he01.tcithaijo.org/index.php/tjvm/article/view/243726
Snodgrass, A. & Motaparthi, K. Systemic antibacterial agents. In: (eds Wolverton, S. E. & Wu, J. J.) Comprehensive Dermatologic Drug Therapy. 4th ed. Elsevier; :95–96. (2021).
Shekh, C. S. et al. Isolation of pathogenic Escherichia coli from Buffalo meat sold in Parbhani city, Maharashtra, India. Vet. World. 6 (5), 277–279. https://doi.org/10.5455/vetworld.2013.277-279 (2013).
Parvej, M. S. et al. Prevalence and characteristics of Shiga-toxin-producing Escherichia coli (STEC) isolated from beef slaughterhouse. J. Adv. Vet. Anim. Res. 5, 218–225. https://doi.org/10.5455/javar.2018.e271 (2018).
Haile, W. Prevalence and sources of contamination of cattle meat at municipal abattoir and butcheries as well as its public health importance in Addis Ababa, Ethiopia [dissertation]. AAU Institutional Repository, ; 25–56. (2017). http://etd.aau.edu.et/handle/123456789/21782
Sallam, K. I., Abd-Elghany, S. M., Elhadidy, M. & Tamura, T. Molecular characterization and antimicrobial resistance profile of methicillin-resistant Staphylococcus aureus in retail chicken. J. Food prot. 78 (10), 1879–1884. https://doi.org/10.4315/0362-028X.JFP-15-150 (2015).
Bush, K. et al. Tackling antibiotic resistance. Nat. Rev. Microbiol. 9, 894–896. https://doi.org/10.1038/nrmicro2693 (2011).
Baron, S., Hadjadj, L., Rolain, J. M. & Olaitan, A. O. Molecular mechanisms of polymyxin resistance: known and unknown. Int. J. Antimicrob. Agents. 48 (6), 583–591. https://doi.org/10.1016/j.ijantimicag.2016.06.023 (2016).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in china: a Microbiological and molecular biological study. Lancet infect. Dis. 16, 161–168. https://doi.org/10.1016/S1473-3099(15)00424-7 (2016).
Maamar, E. et al. Emergence of plasmid-mediated colistin resistance in CMY-2-producing Escherichia coli of lineage ST2197 in a Tunisian poultry farm. Int. J. Food Microbiol. 269, 60–63. https://doi.org/10.1016/j.ijfoodmicro.2018.01.017 (2018).
Clemente, L. et al. Revealing mcr-1-positive ESBL-producing Escherichia coli strains among Enterobacteriaceae from food-producing animals (bovine, swine and poultry) and meat (bovine and swine), Portugal, 2010–2015. Int. J. Food Microbiol. 296, 37–42. https://doi.org/10.1016/j.ijfoodmicro.2019.02.006 (2019).
Hassen, B. et al. High prevalence of mcr-1 encoding colistin resistance and first identification of blaCTX-M55 in ESBL/CMY-2-producing Escherichia coli isolated from chicken faces and retail meat in Tunisia. Int. J. Food Microbiol. 318, 108478. https://doi.org/10.1016/j.ijfoodmicro.2019.108478 (2020).
Le, P. Q. et al. Prevalence of mobile colistin resistance (mcr) genes in extended-spectrum β-lactamase-producing Escherichia coli isolated from retail Raw foods in Nha Trang, Vietnam. Int. J. Food Microbiol. 346, 109164. https://doi.org/10.1016/j.ijfoodmicro.2021.109164 (2021).
Zhang, X. et al. Colistin resistance prevalence in Escherichia coli from domestic animals in intensive breeding farms of Jiangsu Province. Int. J. Food Microbiol. 291, 87–90. https://doi.org/10.1016/j.ijfoodmicro.2018.11.013 (2019).
Zhang, W. et al. Prevalence of colistin resistance gene mcr-1 in Escherichia coli isolated from chickens in central China, 2014 to 2019. J. Glob Antimicrob. Resist. 29, 241–246. https://doi.org/10.1016/j.jgar.2022.03.024 (2022).
Acknowledgements
The authors gratefully acknowledge the funding of the Deanship of Graduate Studies and Scientific Research, Jazan University, Saudi Arabia, through Project number: (JU- 20250272 -DGSSR- RP -2025).
Funding
The authors gratefully acknowledge the funding of the Deanship of Graduate Studies and Scientific Research, Jazan University, Saudi Arabia, through Project number: (JU- 20250272 -DGSSR- RP -2025).
Author information
Authors and Affiliations
Contributions
Khalid Ibrahim Sallam Writing – review & editing, Writing original draft, Data curation. Mohamed Z. Sayed‑Ahmed Writing – review & editing, resources, funding acquisition. Hanan Ahmed Zaher Methodology, Formal analysis. Amira ElSayeh Investigation, Formal analysis, Conceptualization. Hilal A. Thaibah Formal analysis, Conceptualization. Moaddey Alfarhan Formal analysis, Conceptualization. Sarfaraz Ahmad Validation, Software, Conceptualization. Nawazish Alam Formal analysis, Conceptualization. Samir Mohammed Abd-Elghany Writing - original draft, Writing – review & editing, Formal analysis, Data curation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sallam, K.I., Sayed-Ahmed, M.Z., Zaher, H.A. et al. Colistin- and cefotaxime-resistant Shiga toxin-producing Escherichia coli (STEC) in buffalo meat. Sci Rep 15, 36485 (2025). https://doi.org/10.1038/s41598-025-22013-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-22013-2