Abstract
Arboviral diseases transmitted by mosquitoes, including dengue, Chikungunya, Zika, yellow fever, West Nile virus (WNV), and Rift Valley fever (RVF), pose a significant public health challenge globally, particularly impacting populations in low and middle-income countries. Conventional mosquito control methods, which primarily rely on insecticides, face critical challenges, including the development of insecticide resistance and environmental concerns. In this context, Wolbachia, an endosymbiotic bacterium, presents an alternative strategy due to its ability to manipulate mosquito reproduction and impede the transmission of pathogens. This study aimed to detect and assess the genetic diversity of Wolbachia and prophage WO in Ethiopian mosquitoes. Mosquitoes were collected from various ecological niches in the Great Rift Valley. Molecular analyses were performed to identify the presence of Wolbachia using PCR targeting the 16 S rRNA and wsp genes. Additionally, the presence of prophage WO was assessed by detecting the conserved orf7 capsid protein gene. To understand genetic diversity, phylogenetic and genetic diversity analyses were performed. Wolbachia was detected in 44.2% (34/77) of mosquitoes using the 16 S rRNA gene and 46.8% (36/77) using the wsp gene. The highest prevalence was observed in Cx. pipiens complex (100%, 11/11) and Ma. uniformis (92.3%, 12/13). Prophage WO was detected in 46.8% (36/77) of mosquitoes, with evidence of multiple-strain co-infections in Cx. pipiens complex. Phylogenetic analysis classified all isolates within Wolbachia pipientis Supergroup B. This study provides the first preliminary characterization of Wolbachia and prophage WO in Ethiopian mosquitoes, revealing evidence of genetic diversity. These findings lay the conceptual foundation of potential Wolbachia-based vector control strategies in Ethiopia and underscore the need for further studies on strain-specific impacts on vector competence and arboviral transmission dynamics.
Similar content being viewed by others
Introduction
Mosquitoes serve as primary vectors for a wide range of arboviral diseases, including dengue, Chikungunya, Zika, yellow fever, and Rift Valley fever (RVF). These diseases collectively affect millions of people globally, with the burden disproportionately affecting populations in low and middle-income countries1,2. Globally, 146 countries have reported at least one of these arboviral diseases. The number of countries documenting the autochthonous vector-borne transmission of these diseases includes 127 for dengue, 110 for Chikungunya, 92 for Zika, 43 for yellow fever, and 39 for RVF3,4,5,6.
Ethiopia, with its diverse ecological zones and abundant mosquito breeding sites, is at high risk for arboviral outbreaks5,7. Chikungunya, dengue, West Nile virus (WNV), yellow fever, Zika, and RVF8,9,10 has been reported from Ethiopia. Mosquito control efforts remain heavily reliant on chemical insecticides11. However, this approach faces increasing challenges due to insecticide resistance12, unintended impacts on non-target species13, and environmental sustainability concerns14,15. These challenges underscore the urgent need for alternative, environmentally friendly mosquito control strategies.
One promising approach lies in harnessing the potential of Wolbachia, a genus of intracellular bacteria that infects a broad range of arthropods, including mosquitoes16. Wolbachia influences the biology of its arthropod host through various mechanisms, including cytoplasmic incompatibility (CI) and reproductive manipulation17,18. Wolbachia is often regarded as a “master manipulator” of invertebrate biology19, significantly influencing various aspects of mosquito physiology, including reproduction, fitness, and immunity. For instance, Wolbachia induces CI, a reproductive anomaly that prevents uninfected female mosquitoes from producing viable offspring when they mate with Wolbachia-infected males, whereas infected females are compatible with both infected and uninfected males17. This asymmetry gives infected females a strong reproductive advantage, facilitating the spread of Wolbachia in natural populations. CI is primarily driven by the activity of CI factor (cif) gene pairs, which encode toxin–antidote systems, whereby male expression of two genes (cifA and cifB) causes CI20,21,22, while the female expression of cifA rescues the embryo23. This phenomenon, when applied through large-scale releases of infected males, can suppress naïve mosquito populations, making Wolbachia a powerful tool for sustainable vector control. Open-field releases of Wolbachia-infected male mosquitoes have successfully suppressed mosquito populations by over 95%24.
Moreover, Wolbachia infections fundamentally alter the host cell environment, reducing the likelihood of arboviral replication within the vector25,26,27. The primary mechanism is thought to involve disruption of cellular resources and processes, including cholesterol and lipid homeostasis, as well as modulation of host cell cytosine methyltransferase expression, an enzyme essential for viral replication25,27. This intracellular modification severely limits the replication and transmission of a broad range of pathogens, including Dengue virus (DENV)28 and Zika virus (ZIKV)29. This phenomenon, known as pathogen blocking, is significant for arboviral suppression. For example, a Wolbachia release program in Townsville, Australia (2014–2019) achieved a 65% reduction in dengue incidence during the release period and over 99% reduction within two years after the intervention30.
The unique properties of Wolbachia, such as reproductive manipulation and pathogen blocking, are linked to its interactions with prophage WO22, a temperate bacteriophage integrated into the Wolbachia genome. Phage WO is known to be present in the genomes of at least five Wolbachia supergroups, namely A, B, E, F, and S17,19. Prophage WO is known to play a role in the horizontal transfer of Wolbachia genes, affecting replication, host fitness, and spread within mosquito populations31. Prophage WO is believed to play a pivotal role in the induction of CI, thereby influencing host reproductive patterns17,22.
The reproductive manipulation and pathogen-blocking properties of Wolbachia have made Wolbachia an innovative tool in integrated vector management programs, offering an environmentally friendly and effective alternative to traditional chemical methods. Different countries have successfully used Wolbachia to suppress arboviral diseases by releasing Wolbachia-infected mosquitoes into the environment32,33. While Wolbachia-based control strategies have been successfully implemented in some countries, there is limited knowledge on the prevalence and diversity of Wolbachia within Ethiopian mosquito populations. To date, only a single study34 reported the presence of Wolbachia in Ethiopian mosquitoes, specifically in An. stephensi. However, preliminary data on its genetic diversity and association with prophage WO across multiple mosquito species remain lacking. Understanding the genetic variability of Wolbachia in Ethiopian mosquito populations is critical for assessing its feasibility as a vector control strategy and designing effective intervention programs. Therefore, this study aimed to detect and assess the genetic diversity of Wolbachia and prophage WO in Ethiopian mosquitoes.
Materials and methods
Study area and mosquito collection
This study was conducted in Ethiopia’s Great Rift Valley, spanning from Batu (7°55’51"N, 38°42’58"E) to Gawane (10°9’59"N, 40°38’43"E), encompassing urban, peri-urban, and rural habitats (Fig. 1). The selected study areas are thought to support the oviposition and development of mosquito larvae year-round, as they contain large permanent water bodies and experience warmer temperatures. Mosquitoes were collected in August 2023 using CDC light and BG-Sentinel traps placed near water bodies, animal enclosures, and densely populated areas, as well as via hand aspirators from houses. Traps were set between 16:00 and 18:00 and retrieved the following day from 7:00 to 9:00. Captured mosquitoes were frozen at −20 °C and identified morphologically using dichotomous keys from the Walter Reed BioSystemics Unit35,36 and molecularly as described in37.
Map showing the geographic distribution of mosquito collection sites across Ethiopia’s Great Rift Valley. The map was generated using QGIS v3.30.0 (https://qgis.org/).
DNA extraction and molecular identification of mosquitoes
Genomic DNA was extracted from each mosquito using DNeasy Blood & Tissue and MACHEREY-NAGEL NucleoSpin DNA/RNA extraction kits, following the manufacturers’ protocols, with minor modifications. For mosquito identification, amplification of the 710 bp fragment flanking the COI gene was performed using the primer set LCO1490 (5’-GGTCAACAAATCATAAAGATATTGG-3’) and HCO2198 (5’-TAAACTTCAGGGTGACCAAAAAATCA-3’)38. Detailed methodologies for DNA extraction, molecular identification, and mosquito species identification are provided in Leta et al37.
Wolbachia detection
To detect Wolbachia and prophage WO, 77 samples were randomly selected from the original 142 molecularly barcoded mosquito specimens37. This sample size was determined based on logistical and resource constraints. The presence of Wolbachia was determined using PCR assays with primers specific to the 16 S rRNA gene and the Wolbachia surface protein (wsp) gene. For the 16 S rRNA gene, the Wolbachia-specific primers used were WolbF (5′-GAA GAT AAT GAC GGT ACT CAC-3′) and Wspecr (5′-AGC TTC GAG TGA AAC CAA TTC-3′)39. For the wsp gene, the primers utilized were wsp 81 F (5′-TGG TCC AAT AAG TGA TGA AGA AAC-3′) and wsp 691R (5′-AAA AAT TAA ACG CTA CTC CA-3′)40. For both genes, the PCR reaction was performed in a final volume of 25 µl, containing 2.5 µl of 10× buffer, 0.5 µl of 10 mM dNTP mix, 0.5 µl each of 10 µM forward and reverse primers, 0.1 µl of Taq DNA polymerase (5.0 U/µl), 1 µl of the extracted DNA template, and 19.9 µl of ddH2O.
The PCR conditions varied between the two target genes. For the 16 S gene, the protocol included an initial denaturation at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 53 °C for 30 s, and extension at 72 °C for 50 s, with a final extension step at 72 °C for 10 min. For the wsp gene, touchdown PCR was used. Thermal cycling conditions included an initial denaturation at 95 °C for 3 min, followed by a touchdown phase of 13 cycles consisting of denaturation at 95 °C for 45 s, annealing starting at 62 °C and decreasing by 1 °C per cycle until reaching 50 °C for 1 min, and extension at 72 °C for 50 s. This was followed by 30 cycles of denaturation at 95 °C for 45 s, annealing at 50 °C for 45 s, and extension at 72 °C for 50 s, with a final extension step at 72 °C for 10 min.
Prophage WO detection
All samples were then further analyzed for the presence of prophage WO. Prophage WO detection targeted the conserved capsid protein gene orf7, using primers WO-F (5′-CCC ACA TGA GCC AAT GAC GTC TG-3′) and WO-R (5′-CGT TCG CTC TGC AAG TAA CTC CAT TAA AAC-3′) to amplify a 350 bp fragment41,42. The PCR reaction was performed in a final volume of 25 µl, comprising 2.5 µl of 10× buffer, 0.5 µl of 10 mM dNTP mix, 0.5 µl each of 10 µM forward and reverse primers, 0.1 µl of Taq DNA polymerase (5.0 U/µl), 1 µl of the extracted DNA template, and 19.9 µl of ddH₂O.
Thermal cycling conditions included an initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation at 95 °C for 30 s, annealing at 52 °C for 30 s, and extension at 72 °C for 30 s, with a final extension step at 72 °C for 3 min.
Sequencing and analysis
All the PCR products used for the identification of Wolbachia and Prophage WO were visualized on 1.5% agarose gels stained with Midori Green, purified, and sequenced via Sanger sequencing by Eurofins, using the same primers for amplification. Chromatograms were inspected, and forward and reverse sequences were assembled and edited with the BioEdit Sequence Alignment Editor v7.2.5 (https://bioedit.software.informer.com/7.2/)43. For all three genetic markers, sequences were compared with homologous sequences from GenBank using the BLAST (Basic Local Alignment Search Tool) (www.ncbi.nlm.nih.gov/BLAST) and assigned to species level based on over 99% sequence identity to GenBank sequences. Sequence alignments were performed using ClustalW in BioEdit. The phylogenetic analysis was carried out using MEGA v.12.0.9 software (https://www.megasoftware.net/)44. Using the ML model selection in MEGA, the best-fitting maximum likelihood (ML) model was identified for each dataset. Phylogenetic trees were inferred using the Maximum Likelihood (ML) method with appropriate nucleotide substitution models selected for each gene: the Kimura 2-parameter model for the Wolbachia 16 S rRNA gene (16 S), the Kimura 3-parameter model for the Wolbachia surface protein gene (wsp), and the Hasegawa–Kishino–Yano (HKY) model for the prophage WO capsid protein gene (orf7). In all models, node support was evaluated using 1,000 bootstrap replicates to ensure the robustness of the phylogenetic and genetic trees.
Several metrics were calculated for all three genetic markers to assess Wolbachia’s genetic diversity, including the number of polymorphic sites (s), the average pairwise nucleotide differences (k), nucleotide diversity (π), the number of haplotypes (h), haplotype diversity (Hd), and neutrality tests (Fu’s Fs statistic). These analyses were conducted using the DNA Sequence Polymorphism v6.12.03 (http://www.ub.edu/dnasp/)45.
Results
Detection of Wolbachia and prophage WO
A total of 6,601 mosquitoes were collected, of which 4,977 were morphologically identified to the genus level. Of these, 142 mosquitoes were molecularly characterized, and 77, representing multiple genera, including Anopheles, Coquillettidia, Culex, and Mansonia, were subsequently screened for Wolbachia infection. Wolbachia has been detected among all mosquito species examined, although the detection rates differ depending on the specific mosquito species and the genetic marker employed. The detection rates of the Wolbachia 16 S rRNA-encoding gene (16 S), Wolbachia surface protein-encoding gene (wsp), and prophage WO capsid protein gene (orf7) across 12 mosquito species are summarized in Table 1.
The 16 S rRNA gene was detected in 34 out of 77 mosquitoes (44.2%), with the highest detection rates observed in Cx. pipiens complex (100%, 11/11) and Ma. uniformis (92.3%, 12/13). In contrast, the 16 S gene could not be detected in Ae. natronius, An. gambiae, Cx. tenagius, and Cx. univittatus.
The wsp gene was detected in 36 out of 77 mosquitoes (46.8%), with Cx. pipiens complex and Ma. uniformis again showed the highest detection rates (100% and 92.3%, respectively). Notably, Ae. natronius, An. pharoensis, Cq. Microannulata, and Cx. univittatus exhibited 100% wsp gene detection, although with small sample sizes. Lower detection rates were observed in Cx. tritaeniorhynchus (9.5%, 2/21) and Ma. Africana (22.2%, 2/9).
The orf7 gene, associated with the prophage WO, was detected in 36 out of 77 mosquitoes, representing a detection rate of 46.8%. Notably, prophage WO was also detected across all mosquito species examined in this study, although the detection rates varied among species. Similar to the wsp gene, the highest detection rates were observed in the Cx. pipiens complex and Ma. uniformis, with rates of 100% and 92.3%, respectively. In contrast, lower detection rates were recorded for Cx. tritaeniorhynchus (9.5%, 2/21) and Ma. africana (11.1%, 1/9). Most of the orf7 gene sequences could not be submitted to the GenBank database due to multiple chromatogram peaks, particularly in samples from the Cx. pipiens complex, which suggests possible infection by multiple strains of prophage WO.
Genetic diversity of Wolbachia and prophage WO
Wolbachia’s genetic diversity among Ethiopia’s predominant mosquito species was analyzed using sequences from the 16 S rRNA, wsp, and orf7genes. The 16 S rRNA analysis uncovered 83 polymorphic sites within 888 nucleotide positions, with 86 mutations, of which 63 were singleton mutations. In the case of the wsp gene, 33 polymorphic sites were identified among 537 nucleotide positions, with 34 mutations, including 20 singleton mutations. For the orf7 gene, 47 polymorphic sites were identified among 357 nucleotide positions, with 50 mutations, including 15 singleton mutations.
Genetic diversity metrics and neutrality test results varied among mosquito species, although most differences were not statistically significant (Table 2). A significant Fu’s Fs statistic was detected only in the prophage WO capsid protein gene orf7 sequences from Ma. uniformis. According to the 16 S rRNA, the sequences from Cx. pipiens complex showed low levels of haplotype diversity (Hd = 0.56) and nucleotide diversity (π = 0.001), with an average pairwise nucleotide difference (k) of 0.62. In contrast, the sequences from Ma. uniformis exhibited the highest level of genetic variation (Hd = 0.97, π = 0.023), with k = 19.64, and showed the greatest number of polymorphic sites (S = 74).
For the wsp gene, the sequences from both Cx. pipiens complex and Ma. uniformis exhibited moderate haplotype diversity (Hd = 0.82 and 0.68, respectively) and nucleotide diversity (π = 0.006 and 0.008, respectively), with k values of 2.76 and 2.18, respectively. Even if there is a tendency towards a certain pattern, the genetic diversity results for the species analyzed using both the 16 S and wsp genes were not statistically significant, likely due to the small sample size.
For the orf7 gene of prophage WO, sequences from Ma. uniformis exhibited low haplotype diversity (Hd = 0.42) but relatively high nucleotide diversity (π = 0.034) and an average pairwise nucleotide difference (k = 9.78), alongside a strongly positive Fu’s Fs statistic (7.24), suggesting possible balancing selection or population subdivision. These results indicate that the prophage WO orf7 gene in Ethiopian mosquitoes displays evidence of higher sequence divergence and possible historical balancing selection or population structure, reflecting complex evolutionary dynamics between Wolbachia and its associated prophage.
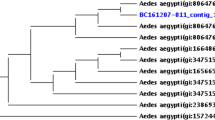
Phylogenetic analysis based on the 16 S rRNA and wsp genes indicates that the Ethiopian Wolbachia isolates cluster within the known Wolbachia supergroups, likely belonging to Supergroup B, which primarily associates with arthropods, including mosquitoes (Figs. 2 and 3). Phylogenetic analysis of the 16 S rRNA gene revealed that sequences from Ma. uniformis and Ma. africana formed a distinct cluster, indicating a stable Wolbachia association within these mosquito species. Similarly, most Wolbachia sequences from members of the Cx. pipiens complex clustered together, with Ethiopian isolates interspersed with reference strains from the United Kingdom, indicating that Wolbachia diversity in these mosquitoes is broadly consistent across geographic regions.
Phylogenetic Analysis of Wolbachia 16 S rRNA Gene Sequences from Ethiopian Mosquitoes. The phylogenetic tree illustrates the evolutionary relationships among Wolbachia isolates based on 16 S rRNA gene sequences. It includes Wolbachia strains obtained from various mosquito species in Ethiopia (marked with **), as well as reference sequences from other geographic regions and hosts. The tree was constructed by the maximum likelihood method using the Kimura 2-parameter nucleotide substitution model, with Ehrlichia ruminantium (NR_074513) serving as the outgroup to root the tree.
Similar to the phylogenetic analysis of the 16 S rRNA gene, the phylogenetic analysis of wsp gene sequences from Ethiopian Wolbachia isolates supports their classification within Supergroup B. The majority of sequences obtained from Ma. uniformis and Cx. pipiens complex mosquitoes grouped together forming strongly supported clades (bootstrap > 95%). In both the 16 S rRNA and wsp gene analyses, although some Ethiopian isolates clustered with strains from other areas, the majority formed a distinct clade, indicating the potential presence of novel strains. This study identified the wPip strain in Cx. neavei and Cx. tritaeniorhynchus, as well as the wNo strain in Cx. tritaeniorhynchus and Cq. microannulata (Fig. 3).
Phylogenetic Analysis of Wolbachia wsp gene sequences from Ethiopian Mosquitoes. The phylogenetic tree illustrates the evolutionary relationships among Wolbachia isolates based on wsp gene sequences. The tree includes Wolbachia strains obtained from various mosquito species in Ethiopia (marked with **) and reference sequences from other geographic regions and hosts. Sequences marked by green stars are wNo strain, and sequences marked by red stars are wPip strains. The tree was constructed by the maximum likelihood method using the Kimura 3-parameter nucleotide substitution model, with Anaplasma marginale (EU676176) as the outgroup to root the tree.
The phylogenetic analysis based on the orf7 gene revealed distinct clustering patterns among mosquito-derived prophage WO sequences from Ethiopia and reference sequences from other countries (Fig. 4). All, except one prophage WO, derived from Wolbachia-infected Ma. uniformis mosquitoes from Ethiopia formed a well-supported clade, indicating a closely related prophage WO lineage circulating within this species. In contrast, the Prophage WO isolate from Wolbachia-infected Ma. africana clustered on a separate branch with moderate bootstrap support, indicating it carries a genetically distinct prophage WO sequence compared to Ma. uniformis. Other isolates, including from C. microannulata, Cx. neavei, and An. coustani also formed their separate branches with moderate to high bootstrap support, highlighting additional host-associated diversity of prophage WO within Ethiopian mosquitoes.
Overall, the phylogenetic analysis complements the genetic diversity analysis by confirming both host-associated and geographically structured patterns of prophage WO diversity in Ethiopian mosquito populations, and it supports the presence of historical diversification and possible balancing selection maintaining prophage variation across mosquito species.
Phylogenetic analysis of prophage WO capsid protein gene (orf7) sequences from Wolbachia-infected mosquitoes. The phylogenetic tree illustrates the evolutionary relationships among Prophage WO isolates based on the capsid protein gene orf7 gene sequences. The tree includes Prophage WO isolates obtained from various Wolbachia-infected mosquito species in Ethiopia (marked with **) and reference sequences from other geographic areas and hosts. The tree was constructed by the maximum likelihood method using Hasegawa–Kishino–Yano (HKY) nucleotide substitution model, with Escherichia phage Lambda (NC_001416) as the outgroup to root the tree.
Discussion
This study reveals the prevalence, genetic diversity, and phylogenetic relationships of Wolbachia strains and their associated prophage WO in mosquito populations from Ethiopia. Using three genetic markers (16 S rRNA, wsp, and orf7), we characterized infection rates and genetic variability across mosquito species, despite sampling limitations and sequencing failures.
Wolbachia and their associated prophage WO were detected across all mosquito species examined. However, their prevalence varied, reflecting the differences in host susceptibility. According to recent analyses, 217 Culicidae species from 22 genera have been screened for Wolbachia, which constitutes only 6% of the total recognized mosquito species46. The Anopheles, Aedes, and Culex genera were particularly notable, collectively accounting for 75% of the species identified as positive for Wolbachia. Of the 22 genera assessed, 17 exhibited at least one species that tested positive for the endosymbiont. The study further illustrated that, among the screened species, 66 were classified as Wolbachia-positive, representing approximately 30% of the total examined. Anopheles spp. were the most frequently detected, while Ae. albopictus and Ae. aegypti have been extensively studied for their associations with Wolbachia46.
In this study, the highest prevalence was observed in the Cx. pipiens complex, which exhibited a 100% infection rate, followed closely by Ma. uniformis at 92.3%. These findings align with previous research, which documented a 97% infection rate in Cx. pipiens47 and a 100% infection rate in Ma. uniformis48, underscoring Wolbachia’s strong affinity for these species. The failure to amplify the 16 S rRNA gene in Ae. natronius, An. gambiae, Cx. tenagius, and Cx. univittatus indicates that these species may harbor strains of Wolbachia that are not detectable with the current primer sets used for 16 S rRNA amplification. Nonetheless, the detection of Wolbachia was successfully achieved in these species using the wsp gene. This highlights the necessity of utilizing multiple genetic markers to accurately evaluate Wolbachia infection rates and strain diversity42. Furthermore, it suggests that whole-genome sequencing may be necessary for a more refined resolution of Wolbachia strains and a better understanding of their evolutionary dynamics49.
The wsp gene, recognized for its high variability and utility in strain differentiation50, was detected in 46.8% of the analyzed mosquito samples. Consistent with findings from the 16 S rRNA marker, a high prevalence of Wolbachia infection was again observed in Cx. pipiens and Ma. uniformis. In contrast, both the 16 S rRNA and wsp markers showed lower detection rates in Cx. tritaeniorhynchus (9.5%) and Ma. africana (22.2%), suggesting either a reduced prevalence of Wolbachia infection or the presence of divergent Wolbachia strains that were not efficiently amplified by the primers used. Notably, a study from Sri Lanka reported a 0% infection rate in Cx. tritaeniorhynchus48, while research from Kenya found a 27% prevalence in Ma. africana51.
While some Ethiopian Wolbachia isolates clustered with previously characterized strains, the majority formed a distinct clade in both the 16 S rRNA and wsp gene phylogenies. This distinct clustering suggests potential genetic divergence and the presence of novel Wolbachia strains unique to Ethiopian mosquito populations. Although confirmation with MLST and whole-genome sequencing is required, our analysis identified previously characterized Wolbachia strains, namely wPip and wNo. The wPip strain was detected in Cx. neavei and Cx. tritaeniorhynchus, both known vectors of arboviruses52,53. wNo strain was found in Cx. tritaeniorhynchus and Cq. microannulata, further expanding the documented host range of these Wolbachia strains. The presence of these strains in Ethiopian mosquito populations highlights their potential role in shaping vector biology and may provide valuable insights for Wolbachia-based biocontrol strategies, as both wPip and wNo strains are known to cause CI54,55.
The clustering of some Ethiopian isolates with reference strains from other regions, such as Cx. pipiens from the UK and USA, suggests a shared evolutionary history and the potential for horizontal transmission19. This finding aligns with previous studies demonstrating the global distribution of Wolbachia strains across geographic regions and host species16,56,57. The close phylogenetic relationship between some Ethiopian isolates and strains from other regions underscores the ability of Wolbachia to infect a wide range of mosquito hosts, facilitating its spread through horizontal transmission19.
The detection of prophage WO across all mosquito species analyzed indicates its extensive distribution among Wolbachia strains in Ethiopia. However, challenges in sequencing some orf7 genes from Cx. pipiens complex, evidenced by multiple peaks in chromatograms, suggests potential co-infections with various prophage strains, a phenomenon that has been documented in previous research42. According to Bourtzis et al.16, an extensive amount of genetic diversity is found within intergenic regions of wPip and is associated with prophage WO. To address the limitations of Sanger sequencing in resolving mixed infections, future studies could utilize PCR cloning or next-generation sequencing (NGS) methods. These approaches would enable a more accurate characterization of multiple prophage WO variants within individual mosquitoes.
While bacteriophages are generally considered rare in obligate intracellular bacteria58, prophage WO demonstrates a broad distribution among diverse Wolbachia-infected insects31,59,60,61,62. Prophage WO has been identified within the genomes of at least five Wolbachia supergroups A, B, E, F, and S. Among these, the majority of known prophage WO elements are found in supergroups A and B, which are predominantly associated with arthropod hosts. Moreover, prophage WO sequences within these supergroups frequently undergo horizontal transfer and recombination, contributing to the genetic diversity and dynamic evolution observed in Wolbachia populations17.
The likely presence of wPip and wNo Wolbachia strains in Ethiopia, alongside the detection of prophage WO among all mosquito species examined, known to contribute to reproductive manipulation and pathogen blocking, is significant. By releasing Wolbachia-infected mosquitoes, vector populations can be sustainably suppressed30, reducing reliance on chemical insecticides and mitigating the environmental impacts of traditional control measures63. Thus, integrating Wolbachia into Ethiopian vector management strategies offers a sustainable alternative to conventional chemical control methods, which face challenges such as insecticide resistance and environmental concerns12,15. By leveraging Wolbachia’s pathogen-blocking and reproductive manipulation capabilities, Ethiopia could enhance its capacity to control the prevalent arboviral diseases. However, further research is necessary to assess the compatibility of novel Wolbachia strains with local mosquito populations and their long-term stability under diverse environmental conditions. Understanding these dynamics is critical for the successful implementation of Wolbachia-based vector control in regions with high disease burdens19,33.
In conclusion, this study provides a preliminary analysis of Wolbachia prevalence, genetic diversity, and phylogenetic relationships in Ethiopian mosquito populations. The high prevalence of Wolbachia in Cx. pipiens complex and Ma. uniformis, along with the potential presence of novel strains, highlights the potential for Wolbachia-based control strategies in Ethiopia. Future studies incorporating MLST genes and whole-genome sequencing could provide deeper insights into the genetic diversity and potential applications in the management of mosquito-borne diseases.
Data availability
All data utilized in the analyses presented in this article are accessible through public databases. The sequences have been deposited in GenBank with accession numbers PV135021-PV135040 and PV187290-PV187301 for the 16 S rRNA encoding gene, PV167821-PV167836 and PV211220-PV211234 for the Wolbachia surface protein encoding gene, and PV167837-PV167849 for the Prophage WO capsid protein gene orf7.
References
Gubler, D. J. The global emergence/resurgence of arboviral diseases as public health problems. Arch. Med. Res. 33, 330–342 (2002).
Madewell, Z. J. Arboviruses and their vectors. South. Med. J. 113, 520–523 (2020).
WHO. Zika epidemiology update -. May (2024). https://www.who.int/publications/m/item/zika-epidemiology-update-may-2024 (2024).
Roo, A. M., De, Vondeling, G. T., Boer, M., Murray, K. & Postma, M. J. The global health and economic burden of Chikungunya from 2011 to 2020 : a driven analysis on the impact of an emerging vector- borne disease. 1–11 (2024). https://doi.org/10.1136/bmjgh-2024-016648
Leta, S. et al. Global risk mapping for major diseases transmitted by Aedes aegypti and Aedes albopictus. Int. J. Infect. Dis. 67, 25–35 (2018).
Tajudeen, Y. A., Oladunjoye, I. O., Mustapha, M. O. & Mustapha, S. T. Ajide-Bamigboye, N. T. Tackling the global health threat of arboviruses: an appraisal of the three holistic approaches to health. Heal Promot Perspect. 11, 371–381 (2021).
Jaleta, M. B. et al. Entomological survey of the potential vectors of rift Valley fever virus and absence of detection of the virus genome from the vectors in various niches in the Southern half of the great rift Valley of Ethiopia. Vet. Med. Sci. 8, 2716–2725 (2022).
Bangoura, S. T. et al. Seroprevalence of seven arboviruses of public health importance in sub- saharan africa: a systematic review and meta- analysis. BMJ Glob Heal. 9, 1–12 (2024).
Ibrahim, M. et al. Sero-prevalence of brucellosis, Q-fever and rift Valley fever in humans and livestock in Somali Region, Ethiopia. PLoS Negl. Trop. Dis. 15, e0008100 (2021).
Endale, A. et al. Sero-prevalence of West nile virus and rift Valley fever virus infections among cattle under extensive production system in South Omo area, Southern Ethiopia. Trop. Anim. Health Prod. https://doi.org/10.1007/s11250-020-02506-0 (2021).
Benelli, G., Jeffries, C. L. & Walker, T. Biological control of mosquito vectors: Past, present, and future. Insects 7, 1–18 (2016).
Hemingway, J. & Ranson, H. Insecticide resistance in insect vectors of human disease. Annu. Rev. Entomol. 45, 371–391 (2000).
Lee, N. S. M., Clements, G. R., Ting, A. S. Y., Wong, Z. H. & Yek, S. H. Persistent mosquito fogging can be detrimental to non-target invertebrates in an urban tropical forest. PeerJ 8, 1–14 (2020).
Ogunlade, S. T. et al. A review: Aedes-Borne arboviral Infections, controls and Wolbachia-Based strategies. VACCINES 9, 1–23 (2021).
Meier, C. J., Rouhier, M. F. & Hillyer, J. F. Chemical Control of Mosquitoes and the Pesticide Treadmill: A Case for Photosensitive Insecticides as Larvicides. Insects 13, (2022).
Bourtzis, K. et al. Harnessing mosquito-Wolbachia symbiosis for vector and disease control. ACTA Trop. 132, S150–S163 (2014).
Kaur, R. et al. Living in the endosymbiotic world of wolbachia: A centennial review. CELL. HOST \& MICROBE. 29, 879–893 (2021).
Chen, H., Ronau, J. A., Beckmann, J. F. & Hochstrasser, M. A wolbachia nuclease and its binding partner provide a distinct mechanism for cytoplasmic incompatibility. Proc. Natl. Acad. Sci. U S A. 116, 22314–22321 (2019).
Werren, J. H., Baldo, L. & Clark, M. E. Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 6, 741–751 (2008).
Beckmann, J. F. et al. The Toxin-Antidote model of cytoplasmic incompatibility: genetics and evolutionary implications. TRENDS Genet. 35, 175–185 (2019).
Beckmann, J. F., Ronau, J. A. & Hochstrasser, M. A Wolbachia deubiquitylating enzyme induces cytoplasmic incompatibility. Nat. Microbiol. 2, 1–7 (2017).
LePage, D. P. et al. Prophage WO genes recapitulate and enhance Wolbachia-induced cytoplasmic incompatibility. Nature 543, 243 (2017).
Shropshire, J. D., On, J., Layton, E. M., Zhou, H. & Bordenstein, S. R. One prophage WO gene rescues cytoplasmic incompatibility in drosophila melanogaster. Proc. Natl. Acad. Sci. U S A. 115, 4987–4991 (2018).
Crawford, J. E. et al. Efficient production of male Wolbachia-infected Aedes aegypti mosquitoes enables large-scale suppression of wild populations (Jun, 10.1038/s41587-020-0471-x, 2020). Nat. Biotechnol. 38, 1000 (2020).
Lindsey, A. R. I., Bhattacharya, T., Newton, I. L. G. & Hardy, R. W. Conflict in the Intracellular Lives of Endosymbionts and Viruses: A Mechanistic Look at Wolbachia-Mediated Pathogen-blocking. VIRUSES-BASEL 10, (2018).
Terradas, G. & McGraw, E. A. Wolbachia-mediated virus blocking in the mosquito vector Aedes aegypti. Curr. Opin. INSECT Sci. 22, 37–44 (2017).
Bhattacharya, T. et al. Differential viral RNA methylation contributes to pathogen blocking in Wolbachia-colonized arthropods. PLOS Pathog 18, 1–25 (2022).
Edenborough, K. M., Flores, H. A., Simmons, C. P. & Fraser, J. E. Using wolbachia to eliminate dengue: will the virus fight back? J. Virol. 95, 1–13 (2021).
Edwards, B., Ghedin, E. & Voronin, D. Wolbachia interferes with Zika virus replication by hijacking cholesterol metabolism in mosquito cells. Microbiol. Spectr. 11, 1–16 (2023).
Ogunlade, S. T., Adekunle, A. I., Meehan, M. T. & McBryde, E. S. Quantifying the impact of wolbachia releases on dengue infection in Townsville, Australia. Sci. Rep. 13, 1–12 (2023).
Kent, B. N. & Bordenstein, S. R. Phage WO of wolbachia: lambda of the endosymbiont world. Trends Microbiol. 18, 173–181 (2010).
Ross, P. A. Designing effective wolbachia release programs for mosquito and arbovirus control. ACTA Trop. 222, 1–13 (2021).
Montenegro, D., Cortés-Cortés, G., Balbuena-Alonso, M. G., Warner, C. & Camps, M. Wolbachia-based emerging strategies for control of vector-transmitted disease. Acta. Trop. 260, 1–22 (2024).
Waymire, E. et al. Wolbachia 16S rRNA haplotypes detected in wild Anopheles stephensi in Eastern Ethiopia. Parasit. Vect. 15, 1–11 (2022).
Potter, M. A. WRBU: keys to the medically important mosquito species: Identification key to the genera of adult mosquitoes for the world. Adapted from Harbach and Sandlant with the addition of Onirion and Verrallina. (2016).
Edwards, F. Mosquitoes of the Ethiopian Region. III. Culicine adults and pupae. British Museum (Nat. Hist.), London.the Trustees of the British Museum, (1941).
Leta, S., Mulatu, T., Yalew, B., Chibssa, T. R. & Paeshuyse, J. D. N. A. Barcoding and blood meal profiling of Ethiopian mosquitoes (Diptera: Culicidae): insights into species identification and host preferences. Parasit. Vect. 18, 1–29 (2025).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Simoes, P. M., Mialdea, G., Reiss, D., Sagot, M. F. & Charlat, S. Wolbachia detection: an assessment of standard PCR protocols. Mol. Ecol. Resour. 11, 567–572 (2011).
Zhou, W., Rousset, F. & O’Neill, S. Phylogeny and PCR-based classification of wolbachia strains using Wsp gene sequences. Proc. R Soc. B Biol. Sci. 265, 509–515 (1998).
Masui, S., Kamoda, S., Sasaki, T. & Ishikawa, H. Distribution and evolution of bacteriophage WO in Wolbachia, the endosymbiont causing sexual alterations in arthropods. J. Mol. Evol. 51, 491–497 (2000).
Su, C. Y., Zhu, D. H. & Yang, X. H. Design and Testing of Effective Primers for Amplification of the orf7 Gene of Phage WO Associated with Andricus hakonensis. Insects 12, (2021).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Kumar, S. et al. MEGA12 : molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol. Biol. Evol. 41, 1–9 (2024).
Librado, P. & Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
da Silva, L. M., Dezordi, F. Z., Paiva, M. H. S. & Wallau, G. L. Systematic Review of Wolbachia Symbiont Detection in Mosquitoes: An Entangled Topic about Methodological Power and True Symbiosis. Pathogens 10, (2021).
Bergman, A. & Hesson, J. C. Wolbachia prevalence in the vector species culex pipiens and culex torrentium in a Sindbis virus-endemic region of Sweden. Parasit. Vect. 14, 1–7 (2021).
Nugapola, N. W. N. P., De Silva, W. A. P. P. & Karunaratne, S. H. P. P. Distribution and phylogeny of Wolbachia strains in wild mosquito populations in Sri Lanka. Parasit. Vect. 10, 1–8 (2017).
Wolfe, T. M. et al. Comparative genome sequencing reveals insights into the dynamics of wolbachia in native and invasive Cherry fruit flies. Mol. Ecol. 30, 6259–6272 (2021).
da Silva, L. M. I. et al. Sequencing and Analysis of Wolbachia Strains from A and B Supergroups Detected in Sylvatic Mosquitoes from Brazil. Microorganisms 12, (2024).
Osei-Poku, J., Han, C., Mbogo, C. M. & Jiggins, F. M. Identification of wolbachia strains in mosquito disease vectors. PLoS One 7, 1–7 (2012).
Fall, G., Diallo, M., Loucoubar, C., Faye, O. & Sall, A. A. Vector competence of culex neavei and culex quinquefasciatus (Diptera: Culicidae) from Senegal for lineages 1, 2, Koutango and a putative new lineage of West nile virus. Am. J. Trop. Med. Hyg. 90, 747–754 (2014).
Faizah, A. N. et al. Evaluating the competence of the primary vector, culex tritaeniorhynchus, and the invasive mosquito species, Aedes japonicus japonicus, in transmitting three Japanese encephalitis virus genotypes. PLoS Negl. Trop. Dis. 14, 1–18 (2020).
Sun, G., Zhang, M., Chen, H. & Hochstrasser, M. The CinB Nuclease from wNo Wolbachia Is Sufficient for Induction of Cytoplasmic Incompatibility in Drosophila. MBio 13, (2022).
Fraser, J. E. et al. Novel phenotype of Wolbachia strain wPip in Aedes aegypti challenges assumptions on mechanisms of Wolbachia-mediated dengue virus inhibition. PLOS Pathog. 16, 1–21 (2020).
Mercant Osuna, A. et al. Diverse novel wolbachia bacteria strains and widespread co-infections with Asaia bacteria in culicine mosquitoes from ecologically diverse regions of Cameroon. Wellcome Open. Res. 8, 267 (2023).
Minwuyelet, A. et al. Symbiotic wolbachia in mosquitoes and its role in reducing the transmission of mosquito-borne diseases: updates and prospects. Front. Microbiol. 14, 1–12 (2023).
Bordenstein, S. R. & Reznikoff, W. S. Mobile DNA in obligate intracellular bacteria. Nat. Rev. Microbiol. 3, 688–699 (2005).
Gavotte, L. et al. A survey of the bacteriophage WO in the endosymbiotic bacteria wolbachia. Mol. Biol. Evol. 24, 427–435 (2007).
Chafee, M. E., Funk, D. J., Harrison, R. G. & Bordenstein, S. R. Lateral phage transfer in obligate intracellular bacteria (Wolbachia): verification from natural populations. Mol. Biol. Evol. 27, 501–505 (2010).
Wang, G. H. et al. Bacteriophage WO can mediate horizontal gene transfer in endosymbiotic Wolbachia genomes. Front Microbiol. 7, 1–16 (2016).
Chauvatcharin, N., Ahantarig, A., Baimai, V. & Kittayapong, P. Bacteriophage WO-B and Wolbachia in natural mosquito hosts: infection incidence, transmission mode and relative density. Mol. Ecol. 15, 2451–2461 (2006).
Garcia, G. A., Hoffmann, A. A., Maciel-de-Freitas, R. & Villela, D. A. M. Aedes aegypti insecticide resistance underlies the success (and failure) of Wolbachia population replacement. Sci. Rep. 10, 1–9 (2020).
Funding
KU Leuven Global Minds Doctoral Scholarship Program (project number 4520196970).
Author information
Authors and Affiliations
Contributions
Conceptualization, field sample collection, laboratory analysis, formal analysis, visualization, and writing the original draft were conducted by SL. Conceptualization, editing, funding acquisition, and supervision were handled by JP & RL. TRC provided supervision. Field data collection was carried out by SL, and laboratory analysis was conducted by SL, BBJ, MMM. All authors have read and agreed to the final published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study received ethical approval from the Research Ethics Review Committee of the College of Veterinary Medicine and Agriculture of Addis Ababa University, certificate reference number VM/ERC/27/05/14/2022. Consent to participate does not apply to this study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Leta, S., Misgana, M.M., Jaleta, B.B. et al. Detection and genetic diversity of Wolbachia and its associated prophage WO in mosquito populations from Ethiopia. Sci Rep 15, 40111 (2025). https://doi.org/10.1038/s41598-025-23896-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23896-x